Details of the Target
General Information of Target
| Target ID | LDTP12608 | |||||
|---|---|---|---|---|---|---|
| Target Name | Phosphatidylinositol-3-phosphatase SAC1 (SACM1L) | |||||
| Gene Name | SACM1L | |||||
| Gene ID | 22908 | |||||
| Synonyms |
KIAA0851; SAC1; Phosphatidylinositol-3-phosphatase SAC1; EC 3.1.3.64; Phosphatidylinositol-4-phosphate phosphatase; Suppressor of actin mutations 1-like protein |
|||||
| 3D Structure | ||||||
| Sequence |
MAAAPALKHWRTTLERVEKFVSPLYFTDCNLRGRLFGASCPVAVLSSFLTPERLPYQEAV
QRDFRPAQVGDSFGPTWWTCWFRVELTIPEAWVGQEVHLCWESDGEGLVWRDGEPVQGLT KEGEKTSYVLTDRLGERDPRSLTLYVEVACNGLLGAGKGSMIAAPDPEKMFQLSRAELAV FHRDVHMLLVDLELLLGIAKGLGKDNQRSFQALYTANQMVNVCDPAQPETFPVAQALASR FFGQHGGESQHTIHATGHCHIDTAWLWPFKETVRKCARSWVTALQLMERNPEFIFACSQA QQLEWVKSRYPGLYSRIQEFACRGQFVPVGGTWVEMDGNLPSGEAMVRQFLQGQNFFLQE FGKMCSEFWLPDTFGYSAQLPQIMHGCGIRRFLTQKLSWNLVNSFPHHTFFWEGLDGSRV LVHFPPGDSYGMQGSVEEVLKTVANNRDKGRANHSAFLFGFGDGGGGPTQTMLDRLKRLS NTDGLPRVQLSSPRQLFSALESDSEQLCTWVGELFLELHNGTYTTHAQIKKGNRECERIL HDVELLSSLALARSAQFLYPAAQLQHLWRLLLLNQFHDVVTGSCIQMVAEEAMCHYEDIR SHGNTLLSAAAAALCAGEPGPEGLLIVNTLPWKRIEVMALPKPGGAHSLALVTVPSMGYA PVPPPTSLQPLLPQQPVFVVQETDGSVTLDNGIIRVKLDPTGRLTSLVLVASGREAIAEG AVGNQFVLFDDVPLYWDAWDVMDYHLETRKPVLGQAGTLAVGTEGGLRGSAWFLLQISPN SRLSQEVVLDVGCPYVRFHTEVHWHEAHKFLKVEFPARVRSSQATYEIQFGHLQRPTHYN TSWDWARFEVWAHRWMDLSEHGFGLALLNDCKYGASVRGSILSLSLLRAPKAPDATADTG RHEFTYALMPHKGSFQDAGVIQAAYSLNFPLLALPAPSPAPATSWSAFSVSSPAVVLETV KQAESSPQRRSLVLRLYEAHGSHVDCWLHLSLPVQEAILCDLLERPDPAGHLTLRDNRLK LTFSPFQVLSLLLVLQPPPH |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Phosphoinositide phosphatase which catalyzes the hydrolysis of phosphatidylinositol 4-phosphate (PtdIns(4)P). Can also catalyze the hydrolysis of phosphatidylinositol 3-phosphate (PtdIns(3)P) and has low activity towards phosphatidylinositol-3,5-bisphosphate (PtdIns(3,5)P2). Shows a very robust PtdIns(4)P phosphatase activity when it binds PtdIns(4)P in a 'cis' configuration in the cellular environment, with much less activity seen when it binds PtdIns(4)P in 'trans' configuration. PtdIns(4)P phosphatase activity (when it binds PtdIns(4)P in 'trans' configuration) is enhanced in the presence of PLEKHA3.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AGS | SNV: p.Q302H | . | |||
| RL952 | SNV: p.G562R | DBIA Probe Info | |||
| U118MG | SNV: p.A52T | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
13.62 | LDD0402 | [1] | |
|
A-EBA Probe Info |
 |
3.32 | LDD0215 | [2] | |
|
TH211 Probe Info |
 |
Y272(9.53) | LDD0257 | [3] | |
|
TH216 Probe Info |
 |
Y272(20.00) | LDD0259 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K483(4.06) | LDD3494 | [4] | |
|
Probe 1 Probe Info |
 |
Y449(11.40) | LDD3495 | [5] | |
|
BTD Probe Info |
 |
C445(0.75) | LDD2109 | [6] | |
|
AHL-Pu-1 Probe Info |
 |
C445(2.10) | LDD0173 | [7] | |
|
HPAP Probe Info |
 |
3.26 | LDD0064 | [8] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [9] | |
|
DBIA Probe Info |
 |
C389(3.56); C392(3.56) | LDD0080 | [10] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [11] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [12] | |
|
Lodoacetamide azide Probe Info |
 |
C445(0.00); C389(0.00); C392(0.00) | LDD0037 | [11] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [13] | |
|
WYneN Probe Info |
 |
C445(0.00); C392(0.00); C389(0.00) | LDD0021 | [14] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [15] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [14] | |
|
IPM Probe Info |
 |
C445(0.00); C392(0.00); C389(0.00) | LDD0005 | [14] | |
|
NHS Probe Info |
 |
K483(0.00); K10(0.00) | LDD0010 | [14] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [14] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [16] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [17] | |
|
Acrolein Probe Info |
 |
C23(0.00); C392(0.00); C445(0.00); C389(0.00) | LDD0217 | [18] | |
|
Crotonaldehyde Probe Info |
 |
C445(0.00); C23(0.00) | LDD0219 | [18] | |
|
Methacrolein Probe Info |
 |
C445(0.00); C23(0.00); C389(0.00); C392(0.00) | LDD0218 | [18] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [19] | |
|
AOyne Probe Info |
 |
11.90 | LDD0443 | [20] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
8.11 | LDD1740 | [21] | |
|
C091 Probe Info |
 |
10.13 | LDD1782 | [21] | |
|
C092 Probe Info |
 |
16.45 | LDD1783 | [21] | |
|
C094 Probe Info |
 |
31.12 | LDD1785 | [21] | |
|
C134 Probe Info |
 |
20.97 | LDD1816 | [21] | |
|
C159 Probe Info |
 |
7.67 | LDD1839 | [21] | |
|
C218 Probe Info |
 |
11.96 | LDD1892 | [21] | |
|
C231 Probe Info |
 |
14.32 | LDD1904 | [21] | |
|
C235 Probe Info |
 |
19.03 | LDD1908 | [21] | |
|
C289 Probe Info |
 |
31.34 | LDD1959 | [21] | |
|
C349 Probe Info |
 |
9.71 | LDD2010 | [21] | |
|
C350 Probe Info |
 |
22.63 | LDD2011 | [21] | |
|
C362 Probe Info |
 |
25.46 | LDD2023 | [21] | |
|
C388 Probe Info |
 |
47.18 | LDD2047 | [21] | |
|
C390 Probe Info |
 |
20.68 | LDD2049 | [21] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [22] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [22] | |
|
FFF probe14 Probe Info |
 |
12.58 | LDD0477 | [22] | |
|
FFF probe2 Probe Info |
 |
19.80 | LDD0463 | [22] | |
|
FFF probe3 Probe Info |
 |
10.35 | LDD0464 | [22] | |
|
JN0003 Probe Info |
 |
20.00 | LDD0469 | [22] | |
|
Photoindomethacin Probe Info |
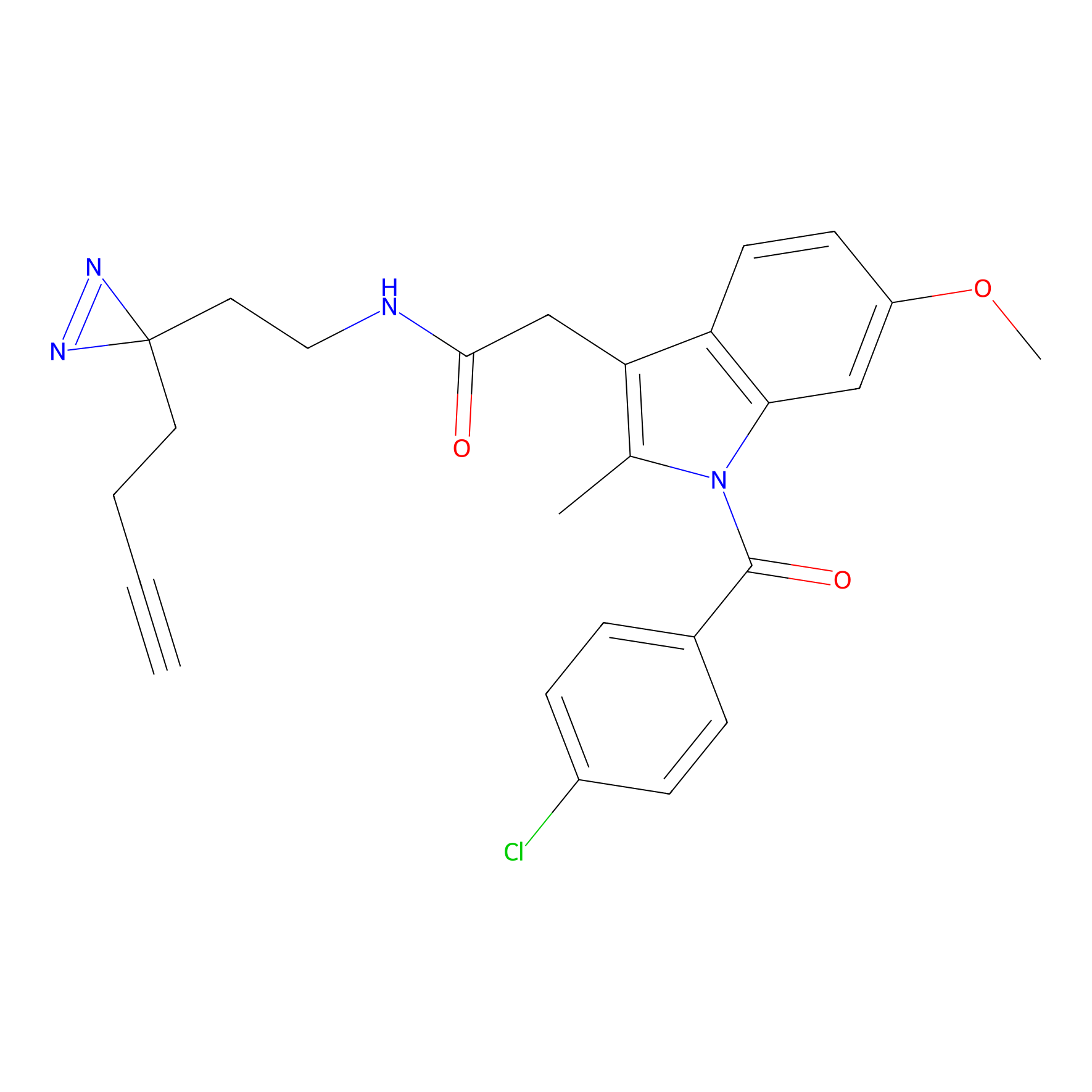 |
N.A. | LDD0154 | [23] | |
|
OEA-DA Probe Info |
 |
7.43 | LDD0046 | [24] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C445(0.90) | LDD2117 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C445(2.10) | LDD0173 | [7] |
| LDCM0214 | AC1 | HCT 116 | C389(1.05); C392(1.00); C445(0.95) | LDD0531 | [10] |
| LDCM0215 | AC10 | HCT 116 | C389(1.13); C392(1.26); C445(0.84) | LDD0532 | [10] |
| LDCM0216 | AC100 | HCT 116 | C389(1.04); C392(1.04); C445(0.80) | LDD0533 | [10] |
| LDCM0217 | AC101 | HCT 116 | C389(0.92); C392(1.03); C445(0.87) | LDD0534 | [10] |
| LDCM0218 | AC102 | HCT 116 | C389(1.02); C392(0.99); C445(0.73) | LDD0535 | [10] |
| LDCM0219 | AC103 | HCT 116 | C389(0.93); C392(0.97); C445(0.75) | LDD0536 | [10] |
| LDCM0220 | AC104 | HCT 116 | C389(1.04); C392(1.05); C445(0.79) | LDD0537 | [10] |
| LDCM0221 | AC105 | HCT 116 | C389(0.91); C392(0.93); C445(0.76) | LDD0538 | [10] |
| LDCM0222 | AC106 | HCT 116 | C389(1.05); C392(1.14); C445(0.92) | LDD0539 | [10] |
| LDCM0223 | AC107 | HCT 116 | C389(0.91); C392(0.93); C445(0.76) | LDD0540 | [10] |
| LDCM0224 | AC108 | HCT 116 | C389(0.99); C392(0.96); C445(0.90) | LDD0541 | [10] |
| LDCM0225 | AC109 | HCT 116 | C389(1.03); C392(1.00); C445(0.98) | LDD0542 | [10] |
| LDCM0226 | AC11 | HCT 116 | C389(1.03); C392(1.05); C445(0.77) | LDD0543 | [10] |
| LDCM0227 | AC110 | HCT 116 | C389(1.02); C392(0.98); C445(0.82) | LDD0544 | [10] |
| LDCM0228 | AC111 | HCT 116 | C389(0.94); C392(1.01); C445(0.84) | LDD0545 | [10] |
| LDCM0229 | AC112 | HCT 116 | C389(0.94); C392(0.96); C445(0.63) | LDD0546 | [10] |
| LDCM0230 | AC113 | HCT 116 | C389(0.79); C392(0.92); C445(1.06) | LDD0547 | [10] |
| LDCM0231 | AC114 | HCT 116 | C389(1.00); C392(0.95); C445(0.76) | LDD0548 | [10] |
| LDCM0232 | AC115 | HCT 116 | C389(0.84); C392(0.95); C445(0.66) | LDD0549 | [10] |
| LDCM0233 | AC116 | HCT 116 | C389(0.93); C392(1.04); C445(0.70) | LDD0550 | [10] |
| LDCM0234 | AC117 | HCT 116 | C389(0.96); C392(1.00); C445(0.82) | LDD0551 | [10] |
| LDCM0235 | AC118 | HCT 116 | C389(1.03); C392(1.09); C445(0.94) | LDD0552 | [10] |
| LDCM0236 | AC119 | HCT 116 | C389(1.02); C392(1.03); C445(0.86) | LDD0553 | [10] |
| LDCM0237 | AC12 | HCT 116 | C389(1.13); C392(1.34); C445(1.02) | LDD0554 | [10] |
| LDCM0238 | AC120 | HCT 116 | C389(0.99); C392(1.09); C445(0.74) | LDD0555 | [10] |
| LDCM0239 | AC121 | HCT 116 | C389(1.41); C392(1.19); C445(1.09) | LDD0556 | [10] |
| LDCM0240 | AC122 | HCT 116 | C389(1.06); C392(1.13); C445(0.87) | LDD0557 | [10] |
| LDCM0241 | AC123 | HCT 116 | C389(1.06); C392(0.96); C445(1.02) | LDD0558 | [10] |
| LDCM0242 | AC124 | HCT 116 | C389(1.14); C392(1.07); C445(0.98) | LDD0559 | [10] |
| LDCM0243 | AC125 | HCT 116 | C389(1.03); C392(1.08); C445(0.89) | LDD0560 | [10] |
| LDCM0244 | AC126 | HCT 116 | C389(1.24); C392(1.19); C445(0.76) | LDD0561 | [10] |
| LDCM0245 | AC127 | HCT 116 | C389(1.24); C392(1.22); C445(0.78) | LDD0562 | [10] |
| LDCM0246 | AC128 | HCT 116 | C389(1.00); C392(1.09); C445(1.11) | LDD0563 | [10] |
| LDCM0247 | AC129 | HCT 116 | C389(1.14); C392(1.21); C445(1.06) | LDD0564 | [10] |
| LDCM0249 | AC130 | HCT 116 | C389(0.99); C392(1.09); C445(0.98) | LDD0566 | [10] |
| LDCM0250 | AC131 | HCT 116 | C389(1.18); C392(1.21); C445(0.96) | LDD0567 | [10] |
| LDCM0251 | AC132 | HCT 116 | C389(1.16); C392(1.18); C445(0.91) | LDD0568 | [10] |
| LDCM0252 | AC133 | HCT 116 | C389(1.04); C392(1.16); C445(0.92) | LDD0569 | [10] |
| LDCM0253 | AC134 | HCT 116 | C389(0.92); C392(1.10); C445(0.86) | LDD0570 | [10] |
| LDCM0254 | AC135 | HCT 116 | C389(0.98); C392(1.11); C445(0.90) | LDD0571 | [10] |
| LDCM0255 | AC136 | HCT 116 | C389(1.04); C392(1.11); C445(0.80) | LDD0572 | [10] |
| LDCM0256 | AC137 | HCT 116 | C389(1.03); C392(1.09); C445(0.79) | LDD0573 | [10] |
| LDCM0257 | AC138 | HCT 116 | C389(1.01); C392(1.07); C445(0.71) | LDD0574 | [10] |
| LDCM0258 | AC139 | HCT 116 | C389(1.10); C392(1.19); C445(0.76) | LDD0575 | [10] |
| LDCM0259 | AC14 | HCT 116 | C389(0.99); C392(1.01); C445(0.80) | LDD0576 | [10] |
| LDCM0260 | AC140 | HCT 116 | C389(1.09); C392(1.15); C445(0.73) | LDD0577 | [10] |
| LDCM0261 | AC141 | HCT 116 | C389(1.03); C392(1.14); C445(0.69) | LDD0578 | [10] |
| LDCM0262 | AC142 | HCT 116 | C389(1.24); C392(1.26); C445(0.76) | LDD0579 | [10] |
| LDCM0263 | AC143 | HCT 116 | C389(0.81); C445(0.85); C392(0.96) | LDD0580 | [10] |
| LDCM0264 | AC144 | HCT 116 | C445(0.54); C389(0.78); C392(0.94) | LDD0581 | [10] |
| LDCM0265 | AC145 | HCT 116 | C445(0.70); C389(0.87); C392(0.93) | LDD0582 | [10] |
| LDCM0266 | AC146 | HCT 116 | C445(0.60); C389(0.83); C392(0.94) | LDD0583 | [10] |
| LDCM0267 | AC147 | HCT 116 | C445(0.56); C389(0.71); C392(1.03) | LDD0584 | [10] |
| LDCM0268 | AC148 | HCT 116 | C445(0.43); C389(0.89); C392(1.07) | LDD0585 | [10] |
| LDCM0269 | AC149 | HCT 116 | C445(0.55); C389(0.92); C392(0.98) | LDD0586 | [10] |
| LDCM0270 | AC15 | HCT 116 | C445(0.79); C392(1.09); C389(1.10) | LDD0587 | [10] |
| LDCM0271 | AC150 | HCT 116 | C445(0.74); C389(0.89); C392(1.09) | LDD0588 | [10] |
| LDCM0272 | AC151 | HCT 116 | C445(0.68); C389(0.79); C392(1.16) | LDD0589 | [10] |
| LDCM0273 | AC152 | HCT 116 | C445(0.51); C389(0.69); C392(0.82) | LDD0590 | [10] |
| LDCM0274 | AC153 | HCT 116 | C445(0.53); C389(0.89); C392(1.08) | LDD0591 | [10] |
| LDCM0621 | AC154 | HCT 116 | C389(0.80); C392(0.92); C445(0.71) | LDD2158 | [10] |
| LDCM0622 | AC155 | HCT 116 | C389(0.83); C392(0.92); C445(0.55) | LDD2159 | [10] |
| LDCM0623 | AC156 | HCT 116 | C389(0.97); C392(1.27); C445(0.73) | LDD2160 | [10] |
| LDCM0624 | AC157 | HCT 116 | C389(1.04); C392(1.21); C445(0.80) | LDD2161 | [10] |
| LDCM0276 | AC17 | HCT 116 | C392(0.96); C445(1.03); C389(1.05) | LDD0593 | [10] |
| LDCM0277 | AC18 | HCT 116 | C445(0.81); C389(0.90); C392(0.91) | LDD0594 | [10] |
| LDCM0278 | AC19 | HCT 116 | C445(0.93); C389(0.94); C392(0.99) | LDD0595 | [10] |
| LDCM0279 | AC2 | HCT 116 | C445(0.92); C392(1.04); C389(1.09) | LDD0596 | [10] |
| LDCM0280 | AC20 | HCT 116 | C392(0.91); C389(0.97); C445(1.03) | LDD0597 | [10] |
| LDCM0281 | AC21 | HCT 116 | C389(0.87); C445(0.95); C392(0.97) | LDD0598 | [10] |
| LDCM0282 | AC22 | HCT 116 | C389(1.07); C445(1.09); C392(1.16) | LDD0599 | [10] |
| LDCM0283 | AC23 | HCT 116 | C392(0.86); C389(0.93); C445(1.02) | LDD0600 | [10] |
| LDCM0284 | AC24 | HCT 116 | C392(1.08); C389(1.10); C445(1.11) | LDD0601 | [10] |
| LDCM0285 | AC25 | HCT 116 | C392(0.93); C445(0.99); C389(1.13) | LDD0602 | [10] |
| LDCM0286 | AC26 | HCT 116 | C445(0.81); C392(0.99); C389(0.99) | LDD0603 | [10] |
| LDCM0287 | AC27 | HCT 116 | C445(0.82); C392(0.82); C389(0.90) | LDD0604 | [10] |
| LDCM0288 | AC28 | HCT 116 | C392(0.78); C445(0.84); C389(0.88) | LDD0605 | [10] |
| LDCM0289 | AC29 | HCT 116 | C445(0.77); C392(0.81); C389(0.87) | LDD0606 | [10] |
| LDCM0290 | AC3 | HCT 116 | C445(0.99); C392(1.07); C389(1.10) | LDD0607 | [10] |
| LDCM0291 | AC30 | HCT 116 | C445(0.74); C392(0.84); C389(0.86) | LDD0608 | [10] |
| LDCM0292 | AC31 | HCT 116 | C445(0.83); C392(0.94); C389(0.97) | LDD0609 | [10] |
| LDCM0293 | AC32 | HCT 116 | C445(0.62); C389(0.96); C392(0.97) | LDD0610 | [10] |
| LDCM0294 | AC33 | HCT 116 | C445(0.71); C392(0.90); C389(1.00) | LDD0611 | [10] |
| LDCM0295 | AC34 | HCT 116 | C445(0.64); C392(0.89); C389(0.97) | LDD0612 | [10] |
| LDCM0296 | AC35 | HCT 116 | C392(0.93); C389(0.94); C445(1.19) | LDD0613 | [10] |
| LDCM0297 | AC36 | HCT 116 | C392(1.02); C389(1.05); C445(1.33) | LDD0614 | [10] |
| LDCM0298 | AC37 | HCT 116 | C392(1.14); C389(1.17); C445(1.25) | LDD0615 | [10] |
| LDCM0299 | AC38 | HCT 116 | C392(1.07); C389(1.08); C445(1.25) | LDD0616 | [10] |
| LDCM0300 | AC39 | HCT 116 | C392(1.05); C389(1.06); C445(1.08) | LDD0617 | [10] |
| LDCM0301 | AC4 | HCT 116 | C445(0.97); C392(1.05); C389(1.09) | LDD0618 | [10] |
| LDCM0302 | AC40 | HCT 116 | C445(0.96); C389(0.97); C392(0.98) | LDD0619 | [10] |
| LDCM0303 | AC41 | HCT 116 | C389(1.07); C392(1.09); C445(1.18) | LDD0620 | [10] |
| LDCM0304 | AC42 | HCT 116 | C445(0.98); C389(0.99); C392(1.01) | LDD0621 | [10] |
| LDCM0305 | AC43 | HCT 116 | C392(0.99); C389(1.01); C445(1.06) | LDD0622 | [10] |
| LDCM0306 | AC44 | HCT 116 | C392(0.97); C389(0.98); C445(1.11) | LDD0623 | [10] |
| LDCM0307 | AC45 | HCT 116 | C445(0.88); C392(0.99); C389(1.01) | LDD0624 | [10] |
| LDCM0308 | AC46 | HCT 116 | C392(0.89); C389(0.96); C445(0.97) | LDD0625 | [10] |
| LDCM0309 | AC47 | HCT 116 | C389(0.98); C445(1.03); C392(1.05) | LDD0626 | [10] |
| LDCM0310 | AC48 | HCT 116 | C389(0.98); C445(0.99); C392(1.01) | LDD0627 | [10] |
| LDCM0311 | AC49 | HCT 116 | C445(0.70); C389(0.79); C392(0.92) | LDD0628 | [10] |
| LDCM0312 | AC5 | HCT 116 | C445(0.95); C392(1.05); C389(1.11) | LDD0629 | [10] |
| LDCM0313 | AC50 | HCT 116 | C392(0.80); C445(0.80); C389(0.84) | LDD0630 | [10] |
| LDCM0314 | AC51 | HCT 116 | C392(0.86); C389(0.99); C445(1.34) | LDD0631 | [10] |
| LDCM0315 | AC52 | HCT 116 | C389(0.91); C392(0.93); C445(1.02) | LDD0632 | [10] |
| LDCM0316 | AC53 | HCT 116 | C445(0.78); C392(0.83); C389(0.88) | LDD0633 | [10] |
| LDCM0317 | AC54 | HCT 116 | C389(0.83); C445(0.85); C392(0.87) | LDD0634 | [10] |
| LDCM0318 | AC55 | HCT 116 | C445(0.80); C392(0.84); C389(0.91) | LDD0635 | [10] |
| LDCM0319 | AC56 | HCT 116 | C445(0.72); C389(0.92); C392(1.00) | LDD0636 | [10] |
| LDCM0320 | AC57 | HCT 116 | C445(0.82); C389(0.92); C392(1.11) | LDD0637 | [10] |
| LDCM0321 | AC58 | HCT 116 | C445(0.69); C389(0.85); C392(0.98) | LDD0638 | [10] |
| LDCM0322 | AC59 | HCT 116 | C445(0.73); C389(0.80); C392(1.05) | LDD0639 | [10] |
| LDCM0323 | AC6 | HCT 116 | C445(0.69); C392(1.02); C389(1.07) | LDD0640 | [10] |
| LDCM0324 | AC60 | HCT 116 | C445(0.79); C389(0.83); C392(1.08) | LDD0641 | [10] |
| LDCM0325 | AC61 | HCT 116 | C445(0.71); C389(0.80); C392(1.11) | LDD0642 | [10] |
| LDCM0326 | AC62 | HCT 116 | C389(0.77); C445(0.85); C392(0.94) | LDD0643 | [10] |
| LDCM0327 | AC63 | HCT 116 | C389(0.87); C445(0.90); C392(1.00) | LDD0644 | [10] |
| LDCM0328 | AC64 | HCT 116 | C445(0.86); C389(0.88); C392(1.02) | LDD0645 | [10] |
| LDCM0329 | AC65 | HCT 116 | C445(0.73); C389(0.96); C392(1.17) | LDD0646 | [10] |
| LDCM0330 | AC66 | HCT 116 | C445(0.75); C389(0.80); C392(1.07) | LDD0647 | [10] |
| LDCM0331 | AC67 | HCT 116 | C445(0.62); C389(0.79); C392(1.07) | LDD0648 | [10] |
| LDCM0332 | AC68 | HCT 116 | C445(0.82); C392(0.89); C389(0.94) | LDD0649 | [10] |
| LDCM0333 | AC69 | HCT 116 | C445(0.74); C392(0.87); C389(1.00) | LDD0650 | [10] |
| LDCM0334 | AC7 | HCT 116 | C445(0.87); C389(1.10); C392(1.16) | LDD0651 | [10] |
| LDCM0335 | AC70 | HCT 116 | C445(0.63); C389(0.86); C392(0.88) | LDD0652 | [10] |
| LDCM0336 | AC71 | HCT 116 | C445(0.93); C389(1.11); C392(1.13) | LDD0653 | [10] |
| LDCM0337 | AC72 | HCT 116 | C445(0.76); C389(1.00); C392(1.04) | LDD0654 | [10] |
| LDCM0338 | AC73 | HCT 116 | C445(0.62); C389(0.97); C392(1.00) | LDD0655 | [10] |
| LDCM0339 | AC74 | HCT 116 | C445(0.68); C389(1.10); C392(1.18) | LDD0656 | [10] |
| LDCM0340 | AC75 | HCT 116 | C445(0.63); C389(0.95); C392(1.11) | LDD0657 | [10] |
| LDCM0341 | AC76 | HCT 116 | C445(0.78); C392(0.89); C389(0.98) | LDD0658 | [10] |
| LDCM0342 | AC77 | HCT 116 | C445(0.66); C389(0.84); C392(0.85) | LDD0659 | [10] |
| LDCM0343 | AC78 | HCT 116 | C445(0.84); C389(0.92); C392(0.98) | LDD0660 | [10] |
| LDCM0344 | AC79 | HCT 116 | C445(0.83); C389(0.96); C392(1.03) | LDD0661 | [10] |
| LDCM0345 | AC8 | HCT 116 | C445(0.79); C392(1.02); C389(1.06) | LDD0662 | [10] |
| LDCM0346 | AC80 | HCT 116 | C445(0.78); C392(0.88); C389(1.01) | LDD0663 | [10] |
| LDCM0347 | AC81 | HCT 116 | C392(1.06); C389(1.07); C445(1.09) | LDD0664 | [10] |
| LDCM0348 | AC82 | HCT 116 | C445(0.71); C389(0.95); C392(1.06) | LDD0665 | [10] |
| LDCM0349 | AC83 | HCT 116 | C445(0.75); C392(0.86); C389(0.96) | LDD0666 | [10] |
| LDCM0350 | AC84 | HCT 116 | C445(0.77); C392(0.83); C389(0.99) | LDD0667 | [10] |
| LDCM0351 | AC85 | HCT 116 | C445(0.81); C392(0.93); C389(1.09) | LDD0668 | [10] |
| LDCM0352 | AC86 | HCT 116 | C445(0.89); C389(0.97); C392(1.00) | LDD0669 | [10] |
| LDCM0353 | AC87 | HCT 116 | C392(1.02); C445(1.07); C389(1.19) | LDD0670 | [10] |
| LDCM0354 | AC88 | HCT 116 | C392(0.82); C445(0.98); C389(1.12) | LDD0671 | [10] |
| LDCM0355 | AC89 | HCT 116 | C445(0.76); C392(0.81); C389(1.06) | LDD0672 | [10] |
| LDCM0357 | AC90 | HCT 116 | C389(1.07); C392(1.31); C445(1.40) | LDD0674 | [10] |
| LDCM0358 | AC91 | HCT 116 | C445(0.74); C392(0.81); C389(1.00) | LDD0675 | [10] |
| LDCM0359 | AC92 | HCT 116 | C445(0.76); C392(0.81); C389(0.92) | LDD0676 | [10] |
| LDCM0360 | AC93 | HCT 116 | C392(0.78); C389(0.85); C445(0.95) | LDD0677 | [10] |
| LDCM0361 | AC94 | HCT 116 | C392(0.83); C389(1.00); C445(1.02) | LDD0678 | [10] |
| LDCM0362 | AC95 | HCT 116 | C389(0.92); C392(0.95); C445(0.99) | LDD0679 | [10] |
| LDCM0363 | AC96 | HCT 116 | C392(0.86); C445(0.92); C389(0.97) | LDD0680 | [10] |
| LDCM0364 | AC97 | HCT 116 | C445(0.74); C392(0.80); C389(0.89) | LDD0681 | [10] |
| LDCM0365 | AC98 | HCT 116 | C445(0.56); C389(0.93); C392(0.99) | LDD0682 | [10] |
| LDCM0366 | AC99 | HCT 116 | C445(0.94); C389(1.01); C392(1.03) | LDD0683 | [10] |
| LDCM0248 | AKOS034007472 | HCT 116 | C389(1.20); C392(1.14); C445(1.00) | LDD0565 | [10] |
| LDCM0356 | AKOS034007680 | HCT 116 | C445(0.90); C389(0.99); C392(1.00) | LDD0673 | [10] |
| LDCM0275 | AKOS034007705 | HCT 116 | C445(0.60); C389(1.15); C392(1.23) | LDD0592 | [10] |
| LDCM0156 | Aniline | NCI-H1299 | 11.42 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | C23(0.00); C445(0.00); H467(0.00); C392(0.00) | LDD0222 | [18] |
| LDCM0367 | CL1 | HCT 116 | C392(0.93); C389(0.97); C445(1.28) | LDD0684 | [10] |
| LDCM0368 | CL10 | HCT 116 | C445(0.70); C392(0.95); C389(1.08) | LDD0685 | [10] |
| LDCM0369 | CL100 | HCT 116 | C445(0.86); C392(1.00); C389(1.00) | LDD0686 | [10] |
| LDCM0370 | CL101 | HCT 116 | C445(0.68); C392(0.93); C389(0.97) | LDD0687 | [10] |
| LDCM0371 | CL102 | HCT 116 | C392(0.93); C389(0.94); C445(0.98) | LDD0688 | [10] |
| LDCM0372 | CL103 | HCT 116 | C445(1.00); C392(1.15); C389(1.16) | LDD0689 | [10] |
| LDCM0373 | CL104 | HCT 116 | C445(0.91); C392(0.94); C389(0.94) | LDD0690 | [10] |
| LDCM0374 | CL105 | HCT 116 | C392(0.90); C445(0.91); C389(0.92) | LDD0691 | [10] |
| LDCM0375 | CL106 | HCT 116 | C445(0.78); C392(0.79); C389(0.90) | LDD0692 | [10] |
| LDCM0376 | CL107 | HCT 116 | C445(0.79); C389(0.92); C392(1.01) | LDD0693 | [10] |
| LDCM0377 | CL108 | HCT 116 | C445(0.76); C392(0.93); C389(0.95) | LDD0694 | [10] |
| LDCM0378 | CL109 | HCT 116 | C392(0.79); C389(0.81); C445(0.83) | LDD0695 | [10] |
| LDCM0379 | CL11 | HCT 116 | C445(0.71); C392(0.88); C389(1.01) | LDD0696 | [10] |
| LDCM0380 | CL110 | HCT 116 | C445(0.72); C392(0.89); C389(1.01) | LDD0697 | [10] |
| LDCM0381 | CL111 | HCT 116 | C389(0.83); C392(0.87); C445(0.88) | LDD0698 | [10] |
| LDCM0382 | CL112 | HCT 116 | C392(0.91); C389(0.98); C445(1.11) | LDD0699 | [10] |
| LDCM0383 | CL113 | HCT 116 | C445(0.67); C392(0.92); C389(1.01) | LDD0700 | [10] |
| LDCM0384 | CL114 | HCT 116 | C392(0.82); C445(0.82); C389(0.87) | LDD0701 | [10] |
| LDCM0385 | CL115 | HCT 116 | C445(0.80); C392(0.91); C389(0.92) | LDD0702 | [10] |
| LDCM0386 | CL116 | HCT 116 | C445(0.81); C389(0.97); C392(0.97) | LDD0703 | [10] |
| LDCM0387 | CL117 | HCT 116 | C445(0.85); C392(1.04); C389(1.05) | LDD0704 | [10] |
| LDCM0388 | CL118 | HCT 116 | C445(1.01); C389(1.02); C392(1.04) | LDD0705 | [10] |
| LDCM0389 | CL119 | HCT 116 | C392(0.90); C389(0.92); C445(1.00) | LDD0706 | [10] |
| LDCM0390 | CL12 | HCT 116 | C445(0.71); C392(0.94); C389(0.95) | LDD0707 | [10] |
| LDCM0391 | CL120 | HCT 116 | C389(1.08); C392(1.09); C445(1.13) | LDD0708 | [10] |
| LDCM0392 | CL121 | HCT 116 | C392(0.52); C445(0.68); C389(0.81) | LDD0709 | [10] |
| LDCM0393 | CL122 | HCT 116 | C389(0.85); C445(0.94); C392(0.98) | LDD0710 | [10] |
| LDCM0394 | CL123 | HCT 116 | C445(0.77); C389(0.96); C392(1.02) | LDD0711 | [10] |
| LDCM0395 | CL124 | HCT 116 | C445(0.75); C389(0.86); C392(1.00) | LDD0712 | [10] |
| LDCM0396 | CL125 | HCT 116 | C445(1.11); C389(1.29); C392(1.43) | LDD0713 | [10] |
| LDCM0397 | CL126 | HCT 116 | C389(0.81); C445(0.87); C392(0.94) | LDD0714 | [10] |
| LDCM0398 | CL127 | HCT 116 | C445(0.95); C389(1.10); C392(1.19) | LDD0715 | [10] |
| LDCM0399 | CL128 | HCT 116 | C445(0.71); C389(0.79); C392(0.96) | LDD0716 | [10] |
| LDCM0400 | CL13 | HCT 116 | C445(0.79); C389(0.99); C392(1.02) | LDD0717 | [10] |
| LDCM0401 | CL14 | HCT 116 | C392(0.92); C389(0.94); C445(0.96) | LDD0718 | [10] |
| LDCM0402 | CL15 | HCT 116 | C445(0.80); C389(0.90); C392(0.90) | LDD0719 | [10] |
| LDCM0403 | CL16 | HCT 116 | C392(0.73); C445(0.85); C389(0.93) | LDD0720 | [10] |
| LDCM0404 | CL17 | HCT 116 | C445(0.78); C392(0.84); C389(0.97) | LDD0721 | [10] |
| LDCM0405 | CL18 | HCT 116 | C445(0.68); C392(0.84); C389(1.00) | LDD0722 | [10] |
| LDCM0406 | CL19 | HCT 116 | C445(0.66); C392(0.83); C389(0.97) | LDD0723 | [10] |
| LDCM0407 | CL2 | HCT 116 | C392(0.92); C389(1.00); C445(1.23) | LDD0724 | [10] |
| LDCM0408 | CL20 | HCT 116 | C445(0.55); C392(0.70); C389(0.88) | LDD0725 | [10] |
| LDCM0409 | CL21 | HCT 116 | C445(0.55); C392(0.75); C389(0.96) | LDD0726 | [10] |
| LDCM0410 | CL22 | HCT 116 | C445(0.57); C392(0.82); C389(1.03) | LDD0727 | [10] |
| LDCM0411 | CL23 | HCT 116 | C445(0.65); C392(0.82); C389(0.87) | LDD0728 | [10] |
| LDCM0412 | CL24 | HCT 116 | C389(0.93); C392(0.70); C445(0.59) | LDD0729 | [10] |
| LDCM0413 | CL25 | HCT 116 | C389(0.83); C392(0.70); C445(0.63) | LDD0730 | [10] |
| LDCM0414 | CL26 | HCT 116 | C389(0.92); C392(0.71); C445(0.70) | LDD0731 | [10] |
| LDCM0415 | CL27 | HCT 116 | C389(0.93); C392(0.74); C445(0.72) | LDD0732 | [10] |
| LDCM0416 | CL28 | HCT 116 | C389(0.88); C392(0.69); C445(0.55) | LDD0733 | [10] |
| LDCM0417 | CL29 | HCT 116 | C389(0.98); C392(0.79); C445(0.59) | LDD0734 | [10] |
| LDCM0418 | CL3 | HCT 116 | C389(1.03); C392(0.95); C445(1.09) | LDD0735 | [10] |
| LDCM0419 | CL30 | HCT 116 | C389(0.92); C392(0.83); C445(0.74) | LDD0736 | [10] |
| LDCM0420 | CL31 | HCT 116 | C389(0.87); C392(0.79); C445(0.94) | LDD0737 | [10] |
| LDCM0421 | CL32 | HCT 116 | C389(0.73); C392(0.71); C445(0.77) | LDD0738 | [10] |
| LDCM0422 | CL33 | HCT 116 | C389(0.72); C392(0.74); C445(0.86) | LDD0739 | [10] |
| LDCM0423 | CL34 | HCT 116 | C389(0.74); C392(0.71); C445(0.78) | LDD0740 | [10] |
| LDCM0424 | CL35 | HCT 116 | C389(0.71); C392(0.73); C445(0.78) | LDD0741 | [10] |
| LDCM0425 | CL36 | HCT 116 | C389(0.72); C392(0.73); C445(0.97) | LDD0742 | [10] |
| LDCM0426 | CL37 | HCT 116 | C389(0.79); C392(0.80); C445(0.85) | LDD0743 | [10] |
| LDCM0428 | CL39 | HCT 116 | C389(0.77); C392(0.77); C445(0.81) | LDD0745 | [10] |
| LDCM0429 | CL4 | HCT 116 | C389(0.98); C392(0.83); C445(1.00) | LDD0746 | [10] |
| LDCM0430 | CL40 | HCT 116 | C389(0.71); C392(0.72); C445(0.78) | LDD0747 | [10] |
| LDCM0431 | CL41 | HCT 116 | C389(0.76); C392(0.78); C445(0.86) | LDD0748 | [10] |
| LDCM0432 | CL42 | HCT 116 | C389(0.72); C392(0.77); C445(0.80) | LDD0749 | [10] |
| LDCM0433 | CL43 | HCT 116 | C389(0.78); C392(0.77); C445(0.80) | LDD0750 | [10] |
| LDCM0434 | CL44 | HCT 116 | C389(0.76); C392(0.78); C445(0.79) | LDD0751 | [10] |
| LDCM0435 | CL45 | HCT 116 | C389(0.78); C392(0.75); C445(0.77) | LDD0752 | [10] |
| LDCM0436 | CL46 | HCT 116 | C389(1.26); C392(1.21); C445(0.90) | LDD0753 | [10] |
| LDCM0437 | CL47 | HCT 116 | C389(1.28); C392(1.17); C445(0.82) | LDD0754 | [10] |
| LDCM0438 | CL48 | HCT 116 | C389(1.21); C392(1.12); C445(0.86) | LDD0755 | [10] |
| LDCM0439 | CL49 | HCT 116 | C389(1.04); C392(0.96); C445(0.95) | LDD0756 | [10] |
| LDCM0440 | CL5 | HCT 116 | C389(1.06); C392(0.95); C445(1.01) | LDD0757 | [10] |
| LDCM0441 | CL50 | HCT 116 | C389(1.13); C392(1.08); C445(0.92) | LDD0758 | [10] |
| LDCM0442 | CL51 | HCT 116 | C389(1.25); C392(1.32); C445(1.01) | LDD0759 | [10] |
| LDCM0443 | CL52 | HCT 116 | C389(1.29); C392(1.26); C445(0.86) | LDD0760 | [10] |
| LDCM0444 | CL53 | HCT 116 | C389(1.11); C392(1.18); C445(0.84) | LDD0761 | [10] |
| LDCM0445 | CL54 | HCT 116 | C389(1.22); C392(1.06); C445(0.79) | LDD0762 | [10] |
| LDCM0446 | CL55 | HCT 116 | C389(1.23); C392(1.07); C445(1.02) | LDD0763 | [10] |
| LDCM0447 | CL56 | HCT 116 | C389(1.06); C392(0.94); C445(0.90) | LDD0764 | [10] |
| LDCM0448 | CL57 | HCT 116 | C389(1.19); C392(1.05); C445(0.91) | LDD0765 | [10] |
| LDCM0449 | CL58 | HCT 116 | C389(1.18); C392(1.11); C445(0.81) | LDD0766 | [10] |
| LDCM0450 | CL59 | HCT 116 | C389(1.22); C392(1.15); C445(1.04) | LDD0767 | [10] |
| LDCM0451 | CL6 | HCT 116 | C389(0.96); C392(0.86); C445(0.83) | LDD0768 | [10] |
| LDCM0452 | CL60 | HCT 116 | C389(1.04); C392(1.01); C445(0.82) | LDD0769 | [10] |
| LDCM0453 | CL61 | HCT 116 | C389(1.06); C392(1.07); C445(0.92) | LDD0770 | [10] |
| LDCM0454 | CL62 | HCT 116 | C389(1.00); C392(1.01); C445(0.87) | LDD0771 | [10] |
| LDCM0455 | CL63 | HCT 116 | C389(0.79); C392(0.92); C445(0.79) | LDD0772 | [10] |
| LDCM0456 | CL64 | HCT 116 | C389(0.84); C392(0.95); C445(0.65) | LDD0773 | [10] |
| LDCM0457 | CL65 | HCT 116 | C389(1.10); C392(1.18); C445(0.81) | LDD0774 | [10] |
| LDCM0458 | CL66 | HCT 116 | C389(1.05); C392(1.07); C445(0.60) | LDD0775 | [10] |
| LDCM0459 | CL67 | HCT 116 | C389(0.97); C392(0.95); C445(0.66) | LDD0776 | [10] |
| LDCM0460 | CL68 | HCT 116 | C389(1.04); C392(1.03); C445(0.66) | LDD0777 | [10] |
| LDCM0461 | CL69 | HCT 116 | C389(0.84); C392(0.86); C445(0.66) | LDD0778 | [10] |
| LDCM0462 | CL7 | HCT 116 | C389(1.06); C392(1.01); C445(0.83) | LDD0779 | [10] |
| LDCM0463 | CL70 | HCT 116 | C389(0.87); C392(0.87); C445(0.68) | LDD0780 | [10] |
| LDCM0464 | CL71 | HCT 116 | C389(0.92); C392(0.92); C445(0.73) | LDD0781 | [10] |
| LDCM0465 | CL72 | HCT 116 | C389(1.08); C392(1.04); C445(0.93) | LDD0782 | [10] |
| LDCM0466 | CL73 | HCT 116 | C389(0.98); C392(1.01); C445(0.67) | LDD0783 | [10] |
| LDCM0467 | CL74 | HCT 116 | C389(1.01); C392(1.00); C445(0.67) | LDD0784 | [10] |
| LDCM0469 | CL76 | HCT 116 | C389(0.88); C392(0.89); C445(0.71) | LDD0786 | [10] |
| LDCM0470 | CL77 | HCT 116 | C389(0.89); C392(0.90); C445(0.93) | LDD0787 | [10] |
| LDCM0471 | CL78 | HCT 116 | C389(0.91); C392(0.91); C445(0.78) | LDD0788 | [10] |
| LDCM0472 | CL79 | HCT 116 | C389(0.84); C392(0.84); C445(0.65) | LDD0789 | [10] |
| LDCM0473 | CL8 | HCT 116 | C389(0.95); C392(0.90); C445(0.69) | LDD0790 | [10] |
| LDCM0474 | CL80 | HCT 116 | C389(0.83); C392(0.89); C445(0.73) | LDD0791 | [10] |
| LDCM0475 | CL81 | HCT 116 | C389(0.83); C392(0.87); C445(0.74) | LDD0792 | [10] |
| LDCM0476 | CL82 | HCT 116 | C389(0.94); C392(0.89); C445(0.62) | LDD0793 | [10] |
| LDCM0477 | CL83 | HCT 116 | C389(0.86); C392(0.84); C445(0.83) | LDD0794 | [10] |
| LDCM0478 | CL84 | HCT 116 | C389(0.81); C392(0.79); C445(0.53) | LDD0795 | [10] |
| LDCM0479 | CL85 | HCT 116 | C389(0.80); C392(0.79); C445(0.67) | LDD0796 | [10] |
| LDCM0480 | CL86 | HCT 116 | C389(0.83); C392(0.83); C445(0.86) | LDD0797 | [10] |
| LDCM0481 | CL87 | HCT 116 | C389(0.93); C392(0.91); C445(0.77) | LDD0798 | [10] |
| LDCM0482 | CL88 | HCT 116 | C389(0.89); C392(0.92); C445(0.72) | LDD0799 | [10] |
| LDCM0483 | CL89 | HCT 116 | C389(0.91); C392(0.91); C445(0.57) | LDD0800 | [10] |
| LDCM0484 | CL9 | HCT 116 | C389(0.95); C392(0.89); C445(0.88) | LDD0801 | [10] |
| LDCM0485 | CL90 | HCT 116 | C389(0.98); C392(0.96); C445(0.97) | LDD0802 | [10] |
| LDCM0486 | CL91 | HCT 116 | C389(1.04); C392(0.98); C445(0.93) | LDD0803 | [10] |
| LDCM0487 | CL92 | HCT 116 | C389(1.03); C392(1.01); C445(0.87) | LDD0804 | [10] |
| LDCM0488 | CL93 | HCT 116 | C389(1.13); C392(1.07); C445(1.02) | LDD0805 | [10] |
| LDCM0489 | CL94 | HCT 116 | C389(0.97); C392(0.90); C445(0.78) | LDD0806 | [10] |
| LDCM0490 | CL95 | HCT 116 | C389(0.95); C392(0.95); C445(0.79) | LDD0807 | [10] |
| LDCM0491 | CL96 | HCT 116 | C389(0.94); C392(0.97); C445(0.84) | LDD0808 | [10] |
| LDCM0492 | CL97 | HCT 116 | C389(0.96); C392(0.97); C445(0.80) | LDD0809 | [10] |
| LDCM0493 | CL98 | HCT 116 | C389(1.00); C392(0.98); C445(0.78) | LDD0810 | [10] |
| LDCM0494 | CL99 | HCT 116 | C389(1.02); C392(1.04); C445(0.90) | LDD0811 | [10] |
| LDCM0495 | E2913 | HEK-293T | C445(1.10); C199(1.16); C23(1.05); C392(1.03) | LDD1698 | [25] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C445(1.49) | LDD1702 | [6] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [9] |
| LDCM0625 | F8 | Ramos | C445(1.48); 1.61; C344(3.06) | LDD2187 | [26] |
| LDCM0572 | Fragment10 | Ramos | C445(1.29) | LDD2189 | [26] |
| LDCM0573 | Fragment11 | Ramos | C445(0.31); 2.93 | LDD2190 | [26] |
| LDCM0574 | Fragment12 | Ramos | C445(1.31) | LDD2191 | [26] |
| LDCM0575 | Fragment13 | Ramos | C445(0.89); 1.26; C344(1.54) | LDD2192 | [26] |
| LDCM0576 | Fragment14 | Ramos | C445(1.38); 2.77; C344(1.81) | LDD2193 | [26] |
| LDCM0579 | Fragment20 | Ramos | C445(3.93); C344(0.04) | LDD2194 | [26] |
| LDCM0580 | Fragment21 | Ramos | C445(0.94); 1.65; C344(0.86) | LDD2195 | [26] |
| LDCM0582 | Fragment23 | Ramos | C445(0.74); 1.13 | LDD2196 | [26] |
| LDCM0578 | Fragment27 | Ramos | C445(0.93); 0.94 | LDD2197 | [26] |
| LDCM0586 | Fragment28 | Ramos | C445(0.59); 1.70; C344(1.22) | LDD2198 | [26] |
| LDCM0588 | Fragment30 | Ramos | C445(1.03); 1.71; C344(1.49) | LDD2199 | [26] |
| LDCM0589 | Fragment31 | Ramos | C445(0.92); 1.31 | LDD2200 | [26] |
| LDCM0590 | Fragment32 | Ramos | C445(1.31) | LDD2201 | [26] |
| LDCM0468 | Fragment33 | HCT 116 | C389(1.20); C392(1.20); C445(0.74) | LDD0785 | [10] |
| LDCM0596 | Fragment38 | Ramos | C445(1.10); 1.30 | LDD2203 | [26] |
| LDCM0566 | Fragment4 | Ramos | C445(1.56) | LDD2184 | [26] |
| LDCM0427 | Fragment51 | HCT 116 | C389(0.79); C392(0.69); C445(0.70) | LDD0744 | [10] |
| LDCM0610 | Fragment52 | Ramos | C445(1.35); 1.57; C344(2.84) | LDD2204 | [26] |
| LDCM0614 | Fragment56 | Ramos | C445(1.16); 1.04; C344(0.35) | LDD2205 | [26] |
| LDCM0569 | Fragment7 | Ramos | C445(1.42); 4.84; C344(0.09) | LDD2186 | [26] |
| LDCM0571 | Fragment9 | Ramos | C445(2.95) | LDD2188 | [26] |
| LDCM0107 | IAA | HeLa | C23(0.00); H467(0.00); C392(0.00) | LDD0221 | [18] |
| LDCM0022 | KB02 | HCT 116 | C389(3.56); C392(3.56) | LDD0080 | [10] |
| LDCM0023 | KB03 | HCT 116 | C389(4.34); C392(4.34) | LDD0081 | [10] |
| LDCM0024 | KB05 | HCT 116 | C389(2.11); C392(2.11) | LDD0082 | [10] |
| LDCM0109 | NEM | HeLa | H341(0.00); H467(0.00) | LDD0224 | [18] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C445(0.75) | LDD2109 | [6] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C445(1.00) | LDD2123 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C445(0.57) | LDD2125 | [6] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C445(0.98) | LDD2127 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C445(1.10) | LDD2136 | [6] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C445(0.96) | LDD2137 | [6] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C389(1.00); C389(0.25); C392(0.25) | LDD2206 | [27] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C344(0.76); C445(0.65); C389(0.27); C392(0.27) | LDD2207 | [27] |
| LDCM0014 | Panhematin | hPBMC | 3.26 | LDD0064 | [8] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Cyclic AMP-responsive element-binding protein 3-like protein 1 (CREB3L1) | BZIP family | Q96BA8 | |||
GPCR
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| B-cell antigen receptor complex-associated protein alpha chain (CD79A) | . | P11912 | |||
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Atypical chemokine receptor 2 (ACKR2) | G-protein coupled receptor 1 family | O00590 | |||
Other
References
