Details of the Target
General Information of Target
| Target ID | LDTP06499 | |||||
|---|---|---|---|---|---|---|
| Target Name | V-type proton ATPase subunit S1 (ATP6AP1) | |||||
| Gene Name | ATP6AP1 | |||||
| Gene ID | 537 | |||||
| Synonyms |
ATP6IP1; ATP6S1; VATPS1; XAP3; V-type proton ATPase subunit S1; V-ATPase subunit S1; Protein XAP-3; V-ATPase Ac45 subunit; V-ATPase S1 accessory protein; Vacuolar proton pump subunit S1 |
|||||
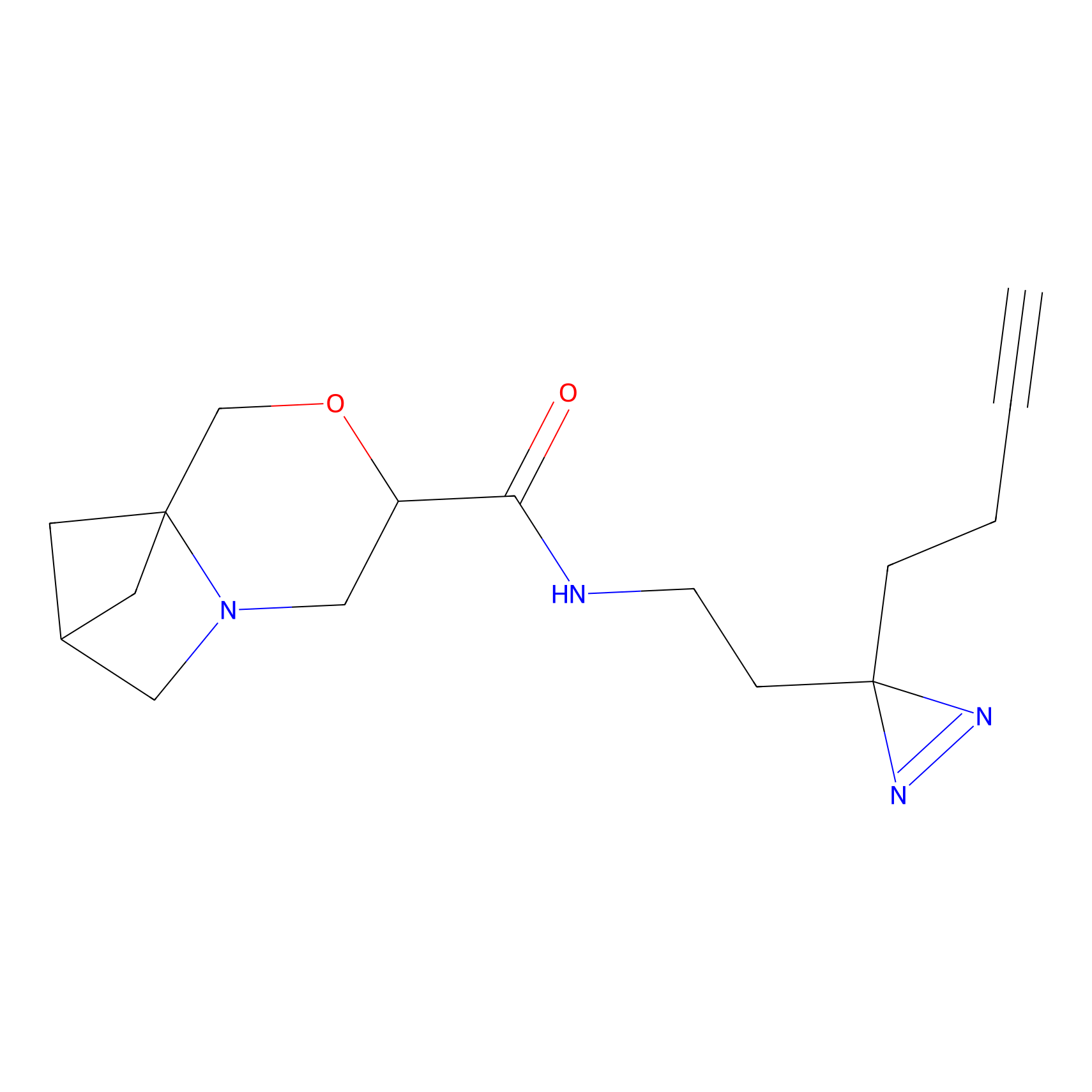
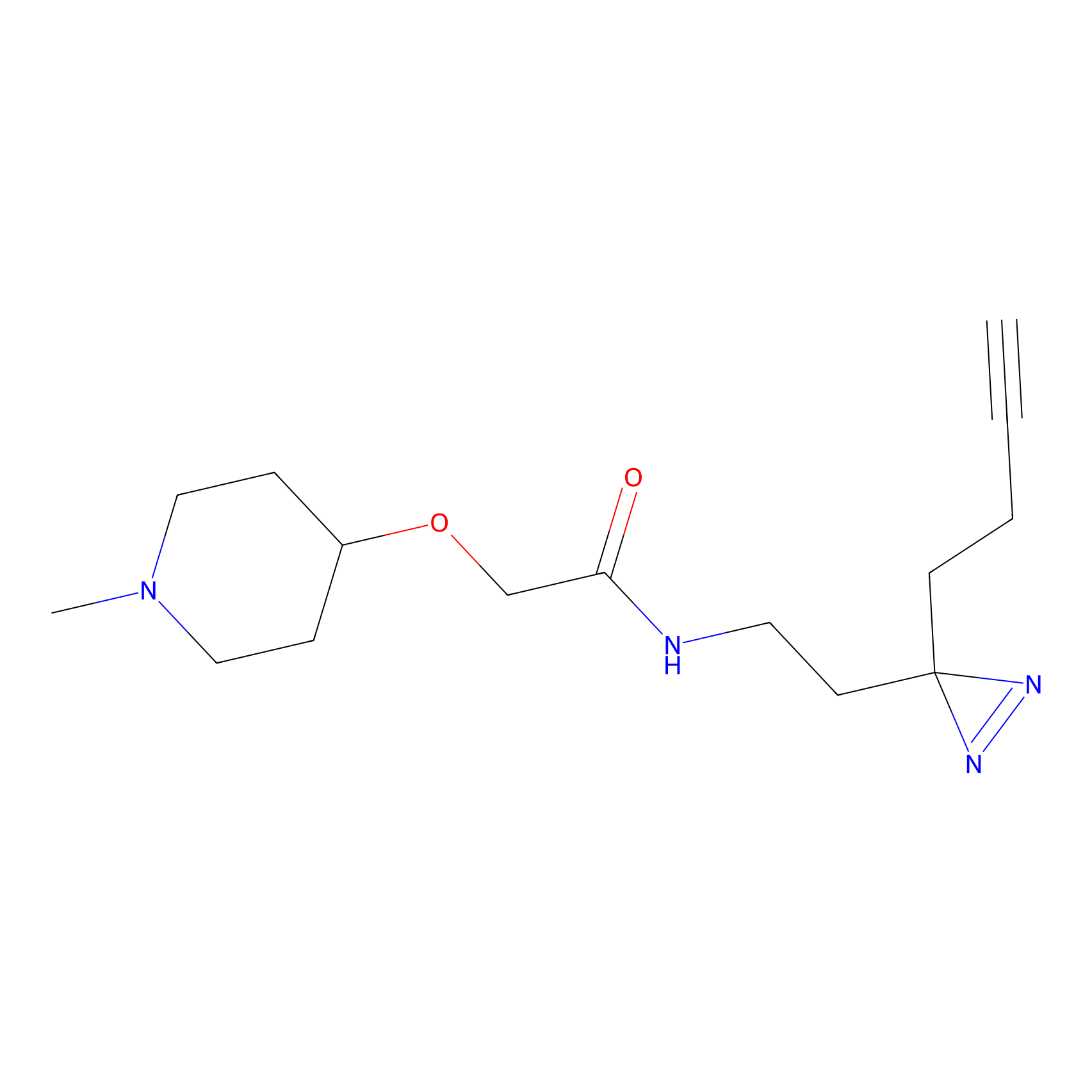
| 3D Structure | ||||||
| Sequence |
MMAAMATARVRMGPRCAQALWRMPWLPVFLSLAAAAAAAAAEQQVPLVLWSSDRDLWAPA
ADTHEGHITSDLQLSTYLDPALELGPRNVLLFLQDKLSIEDFTAYGGVFGNKQDSAFSNL ENALDLAPSSLVLPAVDWYAVSTLTTYLQEKLGASPLHVDLATLRELKLNASLPALLLIR LPYTASSGLMAPREVLTGNDEVIGQVLSTLKSEDVPYTAALTAVRPSRVARDVAVVAGGL GRQLLQKQPVSPVIHPPVSYNDTAPRILFWAQNFSVAYKDQWEDLTPLTFGVQELNLTGS FWNDSFARLSLTYERLFGTTVTFKFILANRLYPVSARHWFTMERLEVHSNGSVAYFNASQ VTGPSIYSFHCEYVSSLSKKGSLLVARTQPSPWQMMLQDFQIQAFNVMGEQFSYASDCAS FFSPGIWMGLLTSLFMLFIFTYGLHMILSLKTMDRFDDHKGPTISLTQIV |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Vacuolar ATPase subunit S1 family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Accessory subunit of the proton-transporting vacuolar (V)-ATPase protein pump, which is required for luminal acidification of secretory vesicles. Guides the V-type ATPase into specialized subcellular compartments, such as neuroendocrine regulated secretory vesicles or the ruffled border of the osteoclast, thereby regulating its activity. Involved in membrane trafficking and Ca(2+)-dependent membrane fusion. May play a role in the assembly of the V-type ATPase complex (Probable). In aerobic conditions, involved in intracellular iron homeostasis, thus triggering the activity of Fe(2+) prolyl hydroxylase (PHD) enzymes, and leading to HIF1A hydroxylation and subsequent proteasomal degradation. In islets of Langerhans cells, may regulate the acidification of dense-core secretory granules.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
5.17 | LDD0318 | [1] | |
|
OPA-S-S-alkyne Probe Info |
 |
K380(1.79) | LDD3494 | [2] | |
|
Probe 1 Probe Info |
 |
Y332(10.83) | LDD3495 | [3] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0222 | [4] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [5] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [5] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [6] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References