Details of the Target
General Information of Target
| Target ID | LDTP07504 | |||||
|---|---|---|---|---|---|---|
| Target Name | Lysophospholipid acyltransferase 5 (LPCAT3) | |||||
| Gene Name | LPCAT3 | |||||
| Gene ID | 10162 | |||||
| Synonyms |
MBOAT5; OACT5; Lysophospholipid acyltransferase 5; LPLAT 5; EC 2.3.1.-; 1-acylglycerophosphocholine O-acyltransferase; EC 2.3.1.23; 1-acylglycerophosphoethanolamine O-acyltransferase; EC 2.3.1.n7; 1-acylglycerophosphoserine O-acyltransferase; EC 2.3.1.n6; Lysophosphatidylcholine acyltransferase; LPCAT; Lyso-PC acyltransferase; Lysophosphatidylcholine acyltransferase 3; Lyso-PC acyltransferase 3; Lysophosphatidylserine acyltransferase; LPSAT; Lyso-PS acyltransferase; Membrane-bound O-acyltransferase domain-containing protein 5; O-acyltransferase domain-containing protein 5
|
|||||
| 3D Structure | ||||||
| Sequence |
MASSAEGDEGTVVALAGVLQSGFQELSLNKLATSLGASEQALRLIISIFLGYPFALFYRH
YLFYKETYLIHLFHTFTGLSIAYFNFGNQLYHSLLCIVLQFLILRLMGRTITAVLTTFCF QMAYLLAGYYYTATGNYDIKWTMPHCVLTLKLIGLAVDYFDGGKDQNSLSSEQQKYAIRG VPSLLEVAGFSYFYGAFLVGPQFSMNHYMKLVQGELIDIPGKIPNSIIPALKRLSLGLFY LVGYTLLSPHITEDYLLTEDYDNHPFWFRCMYMLIWGKFVLYKYVTCWLVTEGVCILTGL GFNGFEEKGKAKWDACANMKVWLFETNPRFTGTIASFNINTNAWVARYIFKRLKFLGNKE LSQGLSLLFLALWHGLHSGYLVCFQMEFLIVIVERQAARLIQESPTLSKLAAITVLQPFY YLVQQTIHWLFMGYSMTAFCLFTWDKWLKVYKSIYFLGHIFFLSLLFILPYIHKAMVPRK EKLKKME |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Membrane-bound acyltransferase family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Lysophospholipid O-acyltransferase (LPLAT) that catalyzes the reacylation step of the phospholipid remodeling process also known as the Lands cycle. Catalyzes transfer of the fatty acyl chain from fatty acyl-CoA to 1-acyl lysophospholipid to form various classes of phospholipids. Converts 1-acyl lysophosphatidylcholine (LPC) into phosphatidylcholine (PC) (LPCAT activity), 1-acyl lysophosphatidylserine (LPS) into phosphatidylserine (PS) (LPSAT activity) and 1-acyl lysophosphatidylethanolamine (LPE) into phosphatidylethanolamine (PE) (LPEAT activity). Favors polyunsaturated fatty acyl-CoAs as acyl donors compared to saturated fatty acyl-CoAs. Has higher activity for LPC acyl acceptors compared to LPEs and LPSs. Can also transfer the fatty acyl chain from fatty acyl-CoA to 1-O-alkyl lysophospholipid or 1-O-alkenyl lysophospholipid with lower efficiency. Acts as a major LPC O-acyltransferase in liver and intestine. As a component of the liver X receptor/NR1H3 or NR1H2 signaling pathway, mainly catalyzes the incorporation of arachidonate into PCs of endoplasmic reticulum (ER) membranes, increasing membrane dynamics and enabling triacylglycerols transfer to nascent very low-density lipoprotein (VLDL) particles. Promotes processing of sterol regulatory protein SREBF1 in hepatocytes, likely by facilitating the translocation of SREBF1-SCAP complex from ER to the Golgi apparatus. Participates in mechanisms by which the liver X receptor/NR1H3 or NR1H2 signaling pathway counteracts lipid-induced ER stress response and inflammation. Down-regulates hepatic inflammation by limiting arachidonic acid availability for synthesis of inflammatory eicosanoids, such as prostaglandins. In enterocytes, acts as a component of a gut-brain feedback loop that coordinates dietary lipid absorption and food intake. Regulates the abundance of PCs containing linoleate and arachidonate in enterocyte membranes, enabling passive diffusion of fatty acids and cholesterol across the membrane for efficient chylomicron assembly. In the intestinal crypt, acts as a component of dietary-responsive phospholipid-cholesterol axis, regulating the biosynthesis of cholesterol and its mitogenic effects on intestinal stem cells.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [1] | |
|
AF-1 Probe Info |
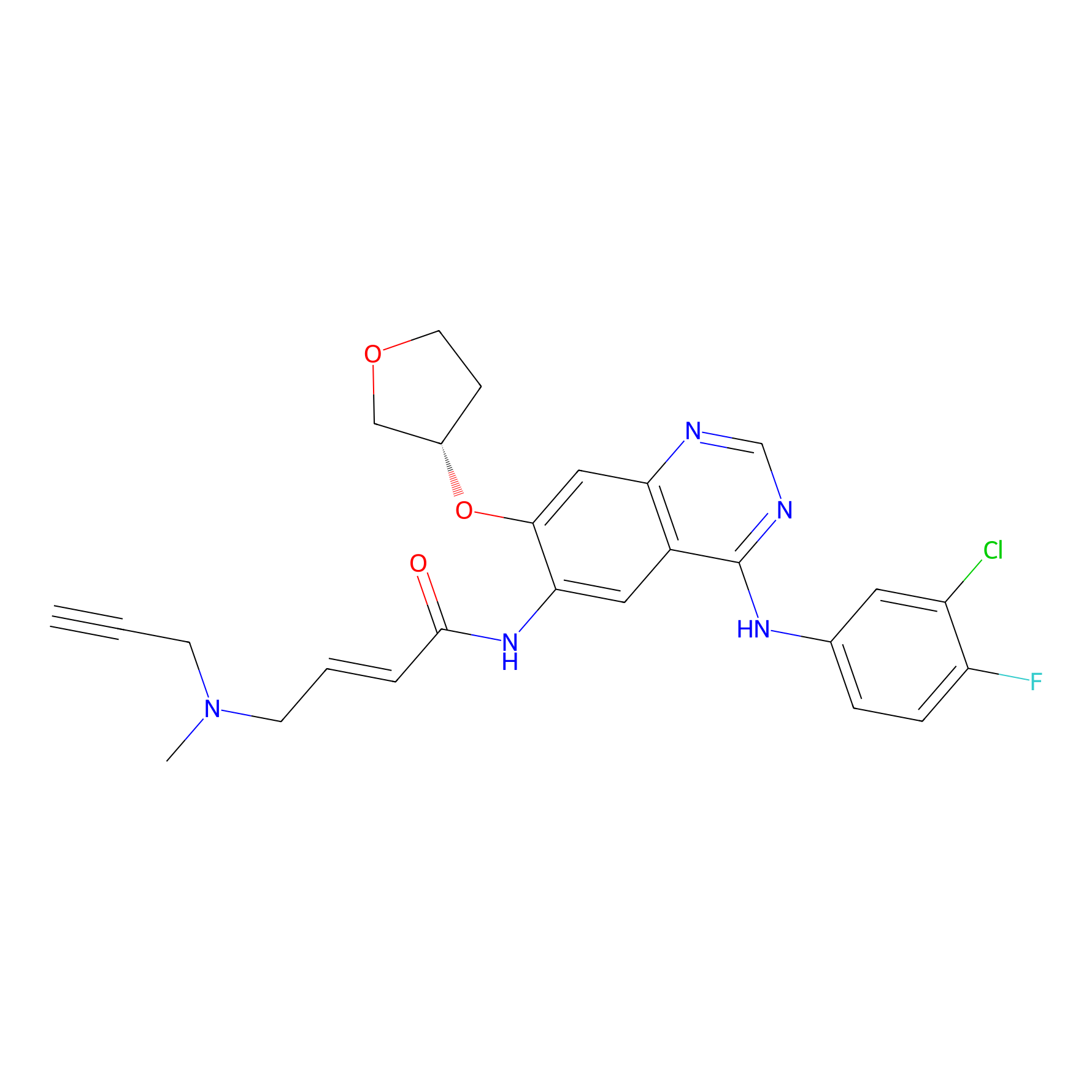 |
2.33 | LDD0421 | [2] | |
|
DBIA Probe Info |
 |
C316(2.07) | LDD0080 | [3] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [4] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0164 | [5] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [6] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C141 Probe Info |
 |
10.27 | LDD1823 | [8] | |
|
C145 Probe Info |
 |
11.39 | LDD1827 | [8] | |
|
C147 Probe Info |
 |
7.84 | LDD1829 | [8] | |
|
C225 Probe Info |
 |
5.90 | LDD1898 | [8] | |
|
C228 Probe Info |
 |
17.15 | LDD1901 | [8] | |
|
C231 Probe Info |
 |
11.71 | LDD1904 | [8] | |
|
C233 Probe Info |
 |
8.94 | LDD1906 | [8] | |
|
C280 Probe Info |
 |
9.65 | LDD1950 | [8] | |
|
C285 Probe Info |
 |
28.25 | LDD1955 | [8] | |
|
C290 Probe Info |
 |
6.06 | LDD1960 | [8] | |
|
FFF probe11 Probe Info |
 |
7.28 | LDD0471 | [9] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [9] | |
|
FFF probe15 Probe Info |
 |
5.78 | LDD0478 | [9] | |
|
FFF probe3 Probe Info |
 |
6.46 | LDD0465 | [9] | |
|
FFF probe6 Probe Info |
 |
12.29 | LDD0467 | [9] | |
|
FFF probe7 Probe Info |
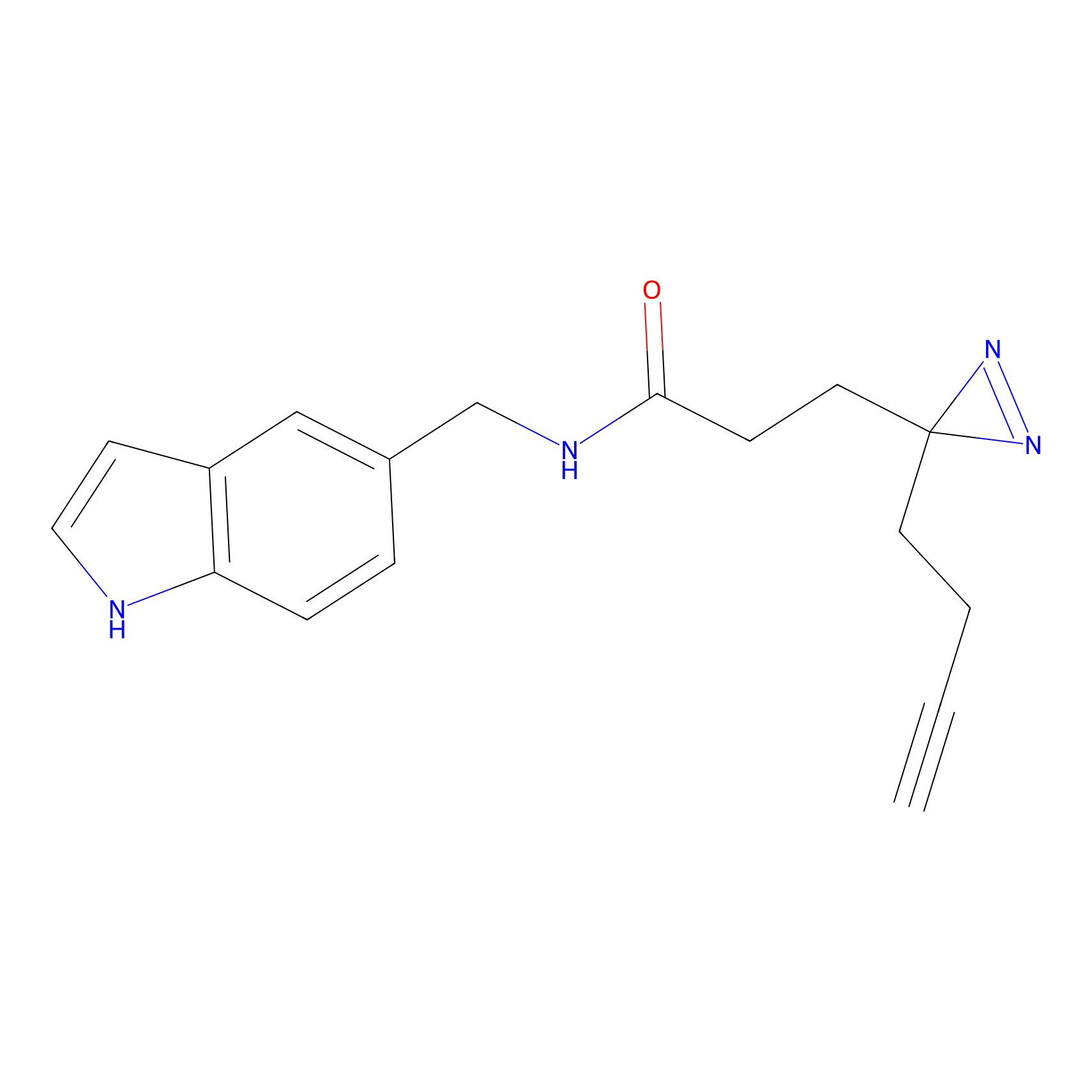 |
19.13 | LDD0483 | [9] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [10] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [10] | |
|
AX-1 Probe Info |
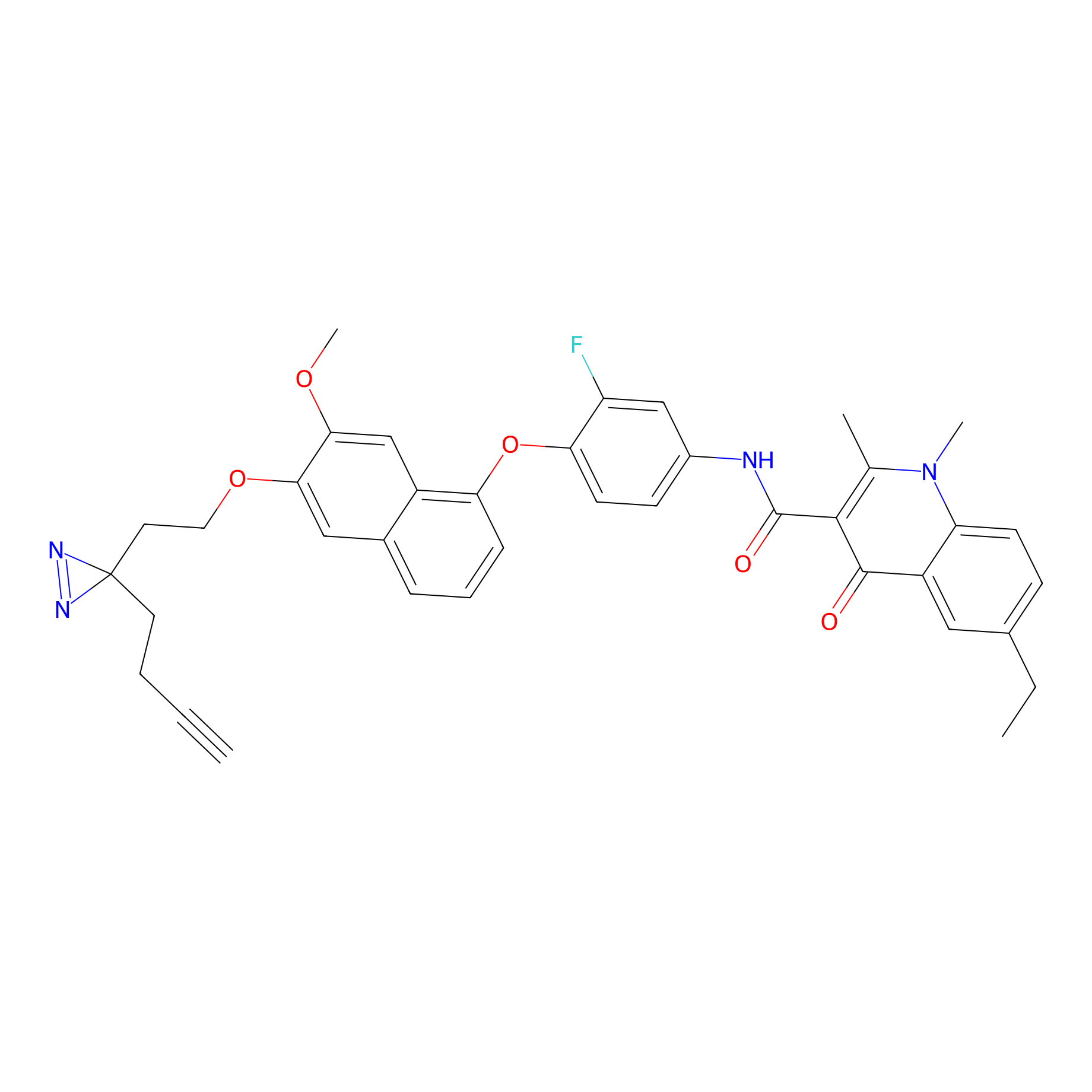 |
3.19 | LDD0441 | [11] | |
|
A-DA Probe Info |
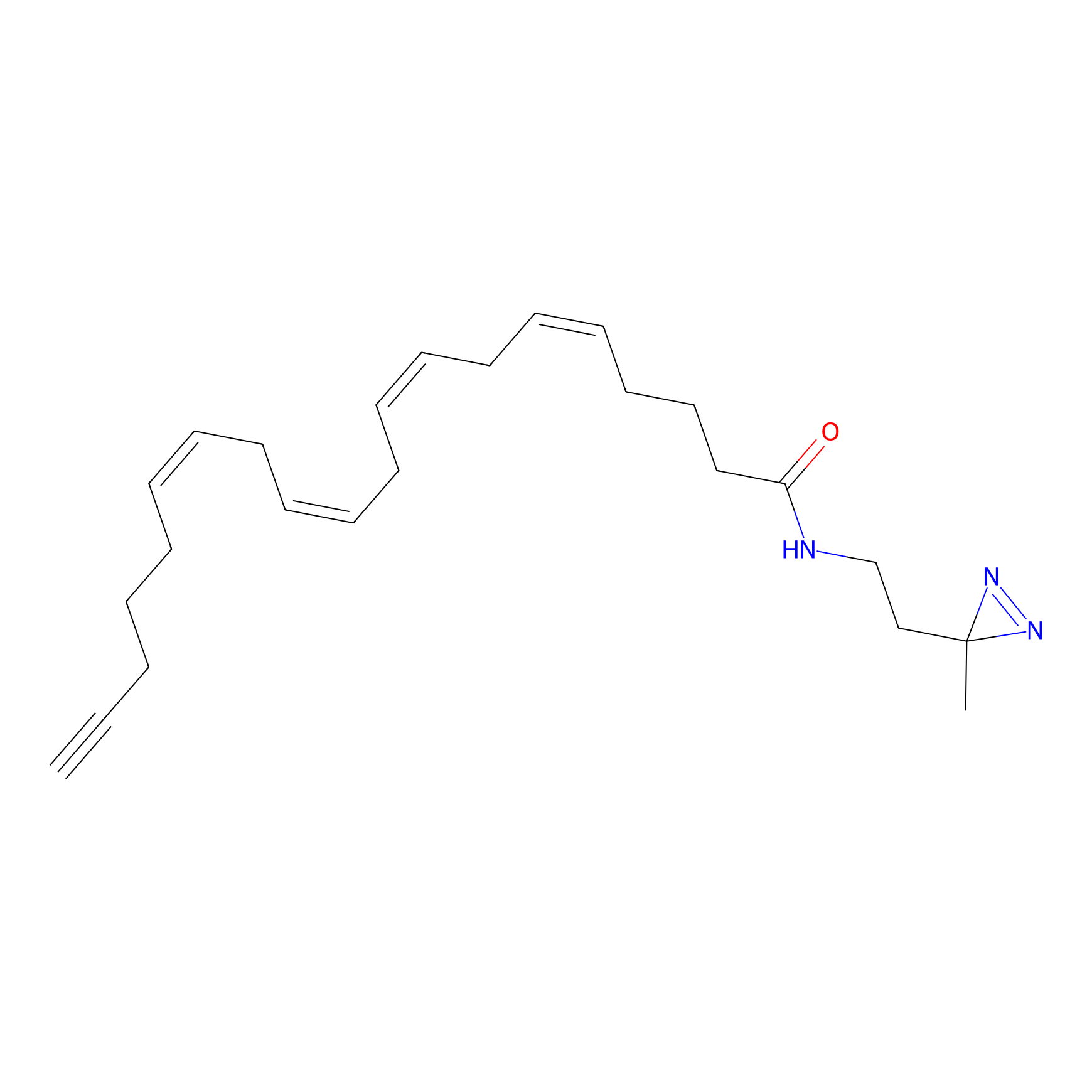 |
3.29 | LDD0142 | [12] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0176 | 9im | MDA-MB-231 | 3.19 | LDD0441 | [11] |
| LDCM0214 | AC1 | HCT 116 | C316(0.98) | LDD0531 | [3] |
| LDCM0215 | AC10 | PaTu 8988t | C316(0.77) | LDD1094 | [3] |
| LDCM0216 | AC100 | HCT 116 | C316(0.75) | LDD0533 | [3] |
| LDCM0217 | AC101 | HCT 116 | C316(0.72) | LDD0534 | [3] |
| LDCM0218 | AC102 | HCT 116 | C316(0.65) | LDD0535 | [3] |
| LDCM0219 | AC103 | HCT 116 | C316(0.61) | LDD0536 | [3] |
| LDCM0220 | AC104 | HCT 116 | C316(0.71) | LDD0537 | [3] |
| LDCM0221 | AC105 | HCT 116 | C316(0.60) | LDD0538 | [3] |
| LDCM0222 | AC106 | HCT 116 | C316(0.62) | LDD0539 | [3] |
| LDCM0223 | AC107 | HCT 116 | C316(1.09) | LDD0540 | [3] |
| LDCM0224 | AC108 | HCT 116 | C316(1.02) | LDD0541 | [3] |
| LDCM0225 | AC109 | HCT 116 | C316(0.94) | LDD0542 | [3] |
| LDCM0226 | AC11 | PaTu 8988t | C316(0.80) | LDD1105 | [3] |
| LDCM0227 | AC110 | HCT 116 | C316(0.89) | LDD0544 | [3] |
| LDCM0228 | AC111 | HCT 116 | C316(0.91) | LDD0545 | [3] |
| LDCM0229 | AC112 | HCT 116 | C316(1.02) | LDD0546 | [3] |
| LDCM0230 | AC113 | HCT 116 | C316(0.90) | LDD0547 | [3] |
| LDCM0231 | AC114 | HCT 116 | C316(0.81) | LDD0548 | [3] |
| LDCM0232 | AC115 | HCT 116 | C316(0.85) | LDD0549 | [3] |
| LDCM0233 | AC116 | HCT 116 | C316(0.99) | LDD0550 | [3] |
| LDCM0234 | AC117 | HCT 116 | C316(1.16) | LDD0551 | [3] |
| LDCM0235 | AC118 | HCT 116 | C316(1.06) | LDD0552 | [3] |
| LDCM0236 | AC119 | HCT 116 | C316(1.00) | LDD0553 | [3] |
| LDCM0237 | AC12 | PaTu 8988t | C316(0.79) | LDD1116 | [3] |
| LDCM0238 | AC120 | HCT 116 | C316(0.77) | LDD0555 | [3] |
| LDCM0239 | AC121 | HCT 116 | C316(1.09) | LDD0556 | [3] |
| LDCM0240 | AC122 | HCT 116 | C316(0.81) | LDD0557 | [3] |
| LDCM0241 | AC123 | HCT 116 | C316(0.95) | LDD0558 | [3] |
| LDCM0242 | AC124 | HCT 116 | C316(1.07) | LDD0559 | [3] |
| LDCM0243 | AC125 | HCT 116 | C316(0.79) | LDD0560 | [3] |
| LDCM0244 | AC126 | HCT 116 | C316(0.90) | LDD0561 | [3] |
| LDCM0245 | AC127 | HCT 116 | C316(1.12) | LDD0562 | [3] |
| LDCM0246 | AC128 | HCT 116 | C316(1.09) | LDD0563 | [3] |
| LDCM0247 | AC129 | HCT 116 | C316(0.89) | LDD0564 | [3] |
| LDCM0249 | AC130 | HCT 116 | C316(0.92) | LDD0566 | [3] |
| LDCM0250 | AC131 | HCT 116 | C316(0.87) | LDD0567 | [3] |
| LDCM0251 | AC132 | HCT 116 | C316(1.10) | LDD0568 | [3] |
| LDCM0252 | AC133 | HCT 116 | C316(0.98) | LDD0569 | [3] |
| LDCM0253 | AC134 | HCT 116 | C316(0.96) | LDD0570 | [3] |
| LDCM0254 | AC135 | HCT 116 | C316(1.09) | LDD0571 | [3] |
| LDCM0255 | AC136 | HCT 116 | C316(1.10) | LDD0572 | [3] |
| LDCM0256 | AC137 | HCT 116 | C316(1.16) | LDD0573 | [3] |
| LDCM0257 | AC138 | HCT 116 | C316(1.13) | LDD0574 | [3] |
| LDCM0258 | AC139 | HCT 116 | C316(1.04) | LDD0575 | [3] |
| LDCM0259 | AC14 | PaTu 8988t | C316(0.90) | LDD1138 | [3] |
| LDCM0260 | AC140 | HCT 116 | C316(1.04) | LDD0577 | [3] |
| LDCM0261 | AC141 | HCT 116 | C316(1.06) | LDD0578 | [3] |
| LDCM0262 | AC142 | HCT 116 | C316(1.12) | LDD0579 | [3] |
| LDCM0263 | AC143 | HCT 116 | C316(0.96) | LDD0580 | [3] |
| LDCM0264 | AC144 | HCT 116 | C316(0.66) | LDD0581 | [3] |
| LDCM0265 | AC145 | HCT 116 | C316(0.73) | LDD0582 | [3] |
| LDCM0266 | AC146 | HCT 116 | C316(0.78) | LDD0583 | [3] |
| LDCM0267 | AC147 | HCT 116 | C316(0.76) | LDD0584 | [3] |
| LDCM0268 | AC148 | HCT 116 | C316(0.86) | LDD0585 | [3] |
| LDCM0269 | AC149 | HCT 116 | C316(0.70) | LDD0586 | [3] |
| LDCM0270 | AC15 | PaTu 8988t | C316(0.80) | LDD1149 | [3] |
| LDCM0271 | AC150 | HCT 116 | C316(0.78) | LDD0588 | [3] |
| LDCM0272 | AC151 | HCT 116 | C316(0.69) | LDD0589 | [3] |
| LDCM0273 | AC152 | HCT 116 | C316(0.71) | LDD0590 | [3] |
| LDCM0274 | AC153 | HCT 116 | C316(0.79) | LDD0591 | [3] |
| LDCM0621 | AC154 | HCT 116 | C316(0.72) | LDD2158 | [3] |
| LDCM0622 | AC155 | HCT 116 | C316(0.65) | LDD2159 | [3] |
| LDCM0623 | AC156 | HCT 116 | C316(0.77) | LDD2160 | [3] |
| LDCM0624 | AC157 | HCT 116 | C316(0.81) | LDD2161 | [3] |
| LDCM0276 | AC17 | HCT 116 | C316(1.05) | LDD0593 | [3] |
| LDCM0277 | AC18 | HCT 116 | C316(0.73) | LDD0594 | [3] |
| LDCM0278 | AC19 | HCT 116 | C316(0.80) | LDD0595 | [3] |
| LDCM0279 | AC2 | HCT 116 | C316(0.95) | LDD0596 | [3] |
| LDCM0280 | AC20 | HCT 116 | C316(0.86) | LDD0597 | [3] |
| LDCM0281 | AC21 | HCT 116 | C316(0.85) | LDD0598 | [3] |
| LDCM0282 | AC22 | HCT 116 | C316(1.10) | LDD0599 | [3] |
| LDCM0283 | AC23 | HCT 116 | C316(0.94) | LDD0600 | [3] |
| LDCM0284 | AC24 | HCT 116 | C316(0.93) | LDD0601 | [3] |
| LDCM0285 | AC25 | HCT 116 | C316(0.92) | LDD0602 | [3] |
| LDCM0286 | AC26 | HCT 116 | C316(0.90) | LDD0603 | [3] |
| LDCM0287 | AC27 | HCT 116 | C316(0.67) | LDD0604 | [3] |
| LDCM0288 | AC28 | HCT 116 | C316(0.94) | LDD0605 | [3] |
| LDCM0289 | AC29 | HCT 116 | C316(0.74) | LDD0606 | [3] |
| LDCM0290 | AC3 | HCT 116 | C316(1.03) | LDD0607 | [3] |
| LDCM0291 | AC30 | HCT 116 | C316(0.72) | LDD0608 | [3] |
| LDCM0292 | AC31 | HCT 116 | C316(0.75) | LDD0609 | [3] |
| LDCM0293 | AC32 | HCT 116 | C316(0.73) | LDD0610 | [3] |
| LDCM0294 | AC33 | HCT 116 | C316(0.71) | LDD0611 | [3] |
| LDCM0295 | AC34 | HCT 116 | C316(0.68) | LDD0612 | [3] |
| LDCM0296 | AC35 | HCT 116 | C316(1.30) | LDD0613 | [3] |
| LDCM0297 | AC36 | HCT 116 | C316(1.26) | LDD0614 | [3] |
| LDCM0298 | AC37 | HCT 116 | C316(1.18) | LDD0615 | [3] |
| LDCM0299 | AC38 | HCT 116 | C316(1.10) | LDD0616 | [3] |
| LDCM0300 | AC39 | HCT 116 | C316(1.04) | LDD0617 | [3] |
| LDCM0301 | AC4 | HCT 116 | C316(0.97) | LDD0618 | [3] |
| LDCM0302 | AC40 | HCT 116 | C316(1.03) | LDD0619 | [3] |
| LDCM0303 | AC41 | HCT 116 | C316(1.14) | LDD0620 | [3] |
| LDCM0304 | AC42 | HCT 116 | C316(1.05) | LDD0621 | [3] |
| LDCM0305 | AC43 | HCT 116 | C316(1.10) | LDD0622 | [3] |
| LDCM0306 | AC44 | HCT 116 | C316(1.03) | LDD0623 | [3] |
| LDCM0307 | AC45 | HCT 116 | C316(1.03) | LDD0624 | [3] |
| LDCM0308 | AC46 | HCT 116 | C316(1.06) | LDD0625 | [3] |
| LDCM0309 | AC47 | HCT 116 | C316(0.93) | LDD0626 | [3] |
| LDCM0310 | AC48 | HCT 116 | C316(0.89) | LDD0627 | [3] |
| LDCM0311 | AC49 | HCT 116 | C316(0.99) | LDD0628 | [3] |
| LDCM0312 | AC5 | HCT 116 | C316(1.05) | LDD0629 | [3] |
| LDCM0313 | AC50 | HCT 116 | C316(0.92) | LDD0630 | [3] |
| LDCM0314 | AC51 | HCT 116 | C316(0.92) | LDD0631 | [3] |
| LDCM0315 | AC52 | HCT 116 | C316(0.86) | LDD0632 | [3] |
| LDCM0316 | AC53 | HCT 116 | C316(0.91) | LDD0633 | [3] |
| LDCM0317 | AC54 | HCT 116 | C316(1.03) | LDD0634 | [3] |
| LDCM0318 | AC55 | HCT 116 | C316(1.02) | LDD0635 | [3] |
| LDCM0319 | AC56 | HCT 116 | C316(1.06) | LDD0636 | [3] |
| LDCM0320 | AC57 | HCT 116 | C316(1.11) | LDD0637 | [3] |
| LDCM0321 | AC58 | HCT 116 | C316(1.05) | LDD0638 | [3] |
| LDCM0322 | AC59 | HCT 116 | C316(0.90) | LDD0639 | [3] |
| LDCM0323 | AC6 | PaTu 8988t | C316(0.79) | LDD1202 | [3] |
| LDCM0324 | AC60 | HCT 116 | C316(0.92) | LDD0641 | [3] |
| LDCM0325 | AC61 | HCT 116 | C316(1.02) | LDD0642 | [3] |
| LDCM0326 | AC62 | HCT 116 | C316(0.97) | LDD0643 | [3] |
| LDCM0327 | AC63 | HCT 116 | C316(0.98) | LDD0644 | [3] |
| LDCM0328 | AC64 | HCT 116 | C316(0.97) | LDD0645 | [3] |
| LDCM0329 | AC65 | HCT 116 | C316(0.95) | LDD0646 | [3] |
| LDCM0330 | AC66 | HCT 116 | C316(1.01) | LDD0647 | [3] |
| LDCM0331 | AC67 | HCT 116 | C316(1.04) | LDD0648 | [3] |
| LDCM0332 | AC68 | HCT 116 | C316(0.97) | LDD0649 | [3] |
| LDCM0333 | AC69 | HCT 116 | C316(0.95) | LDD0650 | [3] |
| LDCM0334 | AC7 | PaTu 8988t | C316(1.00) | LDD1213 | [3] |
| LDCM0335 | AC70 | HCT 116 | C316(0.91) | LDD0652 | [3] |
| LDCM0336 | AC71 | HCT 116 | C316(0.98) | LDD0653 | [3] |
| LDCM0337 | AC72 | HCT 116 | C316(0.87) | LDD0654 | [3] |
| LDCM0338 | AC73 | HCT 116 | C316(0.98) | LDD0655 | [3] |
| LDCM0339 | AC74 | HCT 116 | C316(1.08) | LDD0656 | [3] |
| LDCM0340 | AC75 | HCT 116 | C316(1.14) | LDD0657 | [3] |
| LDCM0341 | AC76 | HCT 116 | C316(1.00) | LDD0658 | [3] |
| LDCM0342 | AC77 | HCT 116 | C316(0.86) | LDD0659 | [3] |
| LDCM0343 | AC78 | HCT 116 | C316(0.89) | LDD0660 | [3] |
| LDCM0344 | AC79 | HCT 116 | C316(1.14) | LDD0661 | [3] |
| LDCM0345 | AC8 | PaTu 8988t | C316(0.92) | LDD1224 | [3] |
| LDCM0346 | AC80 | HCT 116 | C316(0.97) | LDD0663 | [3] |
| LDCM0347 | AC81 | HCT 116 | C316(1.28) | LDD0664 | [3] |
| LDCM0348 | AC82 | HCT 116 | C316(1.29) | LDD0665 | [3] |
| LDCM0349 | AC83 | HCT 116 | C316(1.09) | LDD0666 | [3] |
| LDCM0350 | AC84 | HCT 116 | C316(0.99) | LDD0667 | [3] |
| LDCM0351 | AC85 | HCT 116 | C316(1.07) | LDD0668 | [3] |
| LDCM0352 | AC86 | HCT 116 | C316(1.04) | LDD0669 | [3] |
| LDCM0353 | AC87 | HCT 116 | C316(1.07) | LDD0670 | [3] |
| LDCM0354 | AC88 | HCT 116 | C316(1.04) | LDD0671 | [3] |
| LDCM0355 | AC89 | HCT 116 | C316(0.98) | LDD0672 | [3] |
| LDCM0357 | AC90 | HCT 116 | C316(1.11) | LDD0674 | [3] |
| LDCM0358 | AC91 | HCT 116 | C316(1.06) | LDD0675 | [3] |
| LDCM0359 | AC92 | HCT 116 | C316(1.10) | LDD0676 | [3] |
| LDCM0360 | AC93 | HCT 116 | C316(0.96) | LDD0677 | [3] |
| LDCM0361 | AC94 | HCT 116 | C316(1.45) | LDD0678 | [3] |
| LDCM0362 | AC95 | HCT 116 | C316(1.05) | LDD0679 | [3] |
| LDCM0363 | AC96 | HCT 116 | C316(1.20) | LDD0680 | [3] |
| LDCM0364 | AC97 | HCT 116 | C316(1.05) | LDD0681 | [3] |
| LDCM0365 | AC98 | HCT 116 | C316(0.68) | LDD0682 | [3] |
| LDCM0366 | AC99 | HCT 116 | C316(0.70) | LDD0683 | [3] |
| LDCM0166 | Afatinib | A431 | 2.33 | LDD0421 | [2] |
| LDCM0248 | AKOS034007472 | PaTu 8988t | C316(0.77) | LDD1127 | [3] |
| LDCM0356 | AKOS034007680 | PaTu 8988t | C316(0.79) | LDD1235 | [3] |
| LDCM0275 | AKOS034007705 | PaTu 8988t | C316(0.76) | LDD1154 | [3] |
| LDCM0083 | Avasimibe | A-549 | 2.05 | LDD0143 | [12] |
| LDCM0367 | CL1 | HCT 116 | C316(1.34) | LDD0684 | [3] |
| LDCM0368 | CL10 | HCT 116 | C316(1.05) | LDD0685 | [3] |
| LDCM0369 | CL100 | HCT 116 | C316(0.96) | LDD0686 | [3] |
| LDCM0370 | CL101 | PaTu 8988t | C316(0.79) | LDD1249 | [3] |
| LDCM0371 | CL102 | PaTu 8988t | C316(0.76) | LDD1250 | [3] |
| LDCM0372 | CL103 | PaTu 8988t | C316(0.79) | LDD1251 | [3] |
| LDCM0373 | CL104 | PaTu 8988t | C316(0.75) | LDD1252 | [3] |
| LDCM0374 | CL105 | HCT 116 | C316(0.81) | LDD0691 | [3] |
| LDCM0375 | CL106 | HCT 116 | C316(0.81) | LDD0692 | [3] |
| LDCM0376 | CL107 | HCT 116 | C316(0.80) | LDD0693 | [3] |
| LDCM0377 | CL108 | HCT 116 | C316(0.95) | LDD0694 | [3] |
| LDCM0378 | CL109 | HCT 116 | C316(0.82) | LDD0695 | [3] |
| LDCM0379 | CL11 | HCT 116 | C316(1.02) | LDD0696 | [3] |
| LDCM0380 | CL110 | HCT 116 | C316(0.77) | LDD0697 | [3] |
| LDCM0381 | CL111 | HCT 116 | C316(0.80) | LDD0698 | [3] |
| LDCM0382 | CL112 | HCT 116 | C316(0.94) | LDD0699 | [3] |
| LDCM0383 | CL113 | HCT 116 | C316(0.68) | LDD0700 | [3] |
| LDCM0384 | CL114 | HCT 116 | C316(0.88) | LDD0701 | [3] |
| LDCM0385 | CL115 | HCT 116 | C316(0.72) | LDD0702 | [3] |
| LDCM0386 | CL116 | HCT 116 | C316(0.81) | LDD0703 | [3] |
| LDCM0387 | CL117 | HCT 116 | C316(1.25) | LDD0704 | [3] |
| LDCM0388 | CL118 | HCT 116 | C316(1.07) | LDD0705 | [3] |
| LDCM0389 | CL119 | HCT 116 | C316(1.00) | LDD0706 | [3] |
| LDCM0390 | CL12 | HCT 116 | C316(1.00) | LDD0707 | [3] |
| LDCM0391 | CL120 | HCT 116 | C316(1.11) | LDD0708 | [3] |
| LDCM0392 | CL121 | HCT 116 | C316(0.93) | LDD0709 | [3] |
| LDCM0393 | CL122 | HCT 116 | C316(1.10) | LDD0710 | [3] |
| LDCM0394 | CL123 | HCT 116 | C316(0.87) | LDD0711 | [3] |
| LDCM0395 | CL124 | HCT 116 | C316(1.01) | LDD0712 | [3] |
| LDCM0396 | CL125 | HCT 116 | C316(1.06) | LDD0713 | [3] |
| LDCM0397 | CL126 | HCT 116 | C316(0.85) | LDD0714 | [3] |
| LDCM0398 | CL127 | HCT 116 | C316(0.91) | LDD0715 | [3] |
| LDCM0399 | CL128 | HCT 116 | C316(1.05) | LDD0716 | [3] |
| LDCM0400 | CL13 | HCT 116 | C316(0.98) | LDD0717 | [3] |
| LDCM0401 | CL14 | HCT 116 | C316(0.96) | LDD0718 | [3] |
| LDCM0402 | CL15 | HCT 116 | C316(1.09) | LDD0719 | [3] |
| LDCM0403 | CL16 | HCT 116 | C316(0.91) | LDD0720 | [3] |
| LDCM0404 | CL17 | HCT 116 | C316(0.93) | LDD0721 | [3] |
| LDCM0405 | CL18 | HCT 116 | C316(0.85) | LDD0722 | [3] |
| LDCM0406 | CL19 | HCT 116 | C316(0.86) | LDD0723 | [3] |
| LDCM0407 | CL2 | HCT 116 | C316(1.19) | LDD0724 | [3] |
| LDCM0408 | CL20 | HCT 116 | C316(0.74) | LDD0725 | [3] |
| LDCM0409 | CL21 | HCT 116 | C316(0.75) | LDD0726 | [3] |
| LDCM0410 | CL22 | HCT 116 | C316(0.87) | LDD0727 | [3] |
| LDCM0411 | CL23 | HCT 116 | C316(0.74) | LDD0728 | [3] |
| LDCM0412 | CL24 | HCT 116 | C316(0.83) | LDD0729 | [3] |
| LDCM0413 | CL25 | HCT 116 | C316(0.83) | LDD0730 | [3] |
| LDCM0414 | CL26 | HCT 116 | C316(0.92) | LDD0731 | [3] |
| LDCM0415 | CL27 | HCT 116 | C316(0.76) | LDD0732 | [3] |
| LDCM0416 | CL28 | HCT 116 | C316(0.74) | LDD0733 | [3] |
| LDCM0417 | CL29 | HCT 116 | C316(0.87) | LDD0734 | [3] |
| LDCM0418 | CL3 | HCT 116 | C316(1.19) | LDD0735 | [3] |
| LDCM0419 | CL30 | HCT 116 | C316(1.02) | LDD0736 | [3] |
| LDCM0420 | CL31 | HCT 116 | C316(1.08) | LDD0737 | [3] |
| LDCM0421 | CL32 | HCT 116 | C316(1.09) | LDD0738 | [3] |
| LDCM0422 | CL33 | HCT 116 | C316(1.01) | LDD0739 | [3] |
| LDCM0423 | CL34 | HCT 116 | C316(1.08) | LDD0740 | [3] |
| LDCM0424 | CL35 | HCT 116 | C316(0.89) | LDD0741 | [3] |
| LDCM0425 | CL36 | HCT 116 | C316(1.11) | LDD0742 | [3] |
| LDCM0426 | CL37 | HCT 116 | C316(1.00) | LDD0743 | [3] |
| LDCM0428 | CL39 | HCT 116 | C316(1.11) | LDD0745 | [3] |
| LDCM0429 | CL4 | HCT 116 | C316(0.93) | LDD0746 | [3] |
| LDCM0430 | CL40 | HCT 116 | C316(1.21) | LDD0747 | [3] |
| LDCM0431 | CL41 | HCT 116 | C316(1.10) | LDD0748 | [3] |
| LDCM0432 | CL42 | HCT 116 | C316(1.02) | LDD0749 | [3] |
| LDCM0433 | CL43 | HCT 116 | C316(1.14) | LDD0750 | [3] |
| LDCM0434 | CL44 | HCT 116 | C316(1.04) | LDD0751 | [3] |
| LDCM0435 | CL45 | HCT 116 | C316(1.15) | LDD0752 | [3] |
| LDCM0436 | CL46 | HCT 116 | C316(0.90) | LDD0753 | [3] |
| LDCM0437 | CL47 | HCT 116 | C316(1.04) | LDD0754 | [3] |
| LDCM0438 | CL48 | HCT 116 | C316(0.87) | LDD0755 | [3] |
| LDCM0439 | CL49 | HCT 116 | C316(1.12) | LDD0756 | [3] |
| LDCM0440 | CL5 | HCT 116 | C316(0.98) | LDD0757 | [3] |
| LDCM0441 | CL50 | HCT 116 | C316(0.90) | LDD0758 | [3] |
| LDCM0442 | CL51 | HCT 116 | C316(1.18) | LDD0759 | [3] |
| LDCM0443 | CL52 | HCT 116 | C316(0.80) | LDD0760 | [3] |
| LDCM0444 | CL53 | HCT 116 | C316(0.79) | LDD0761 | [3] |
| LDCM0445 | CL54 | HCT 116 | C316(0.87) | LDD0762 | [3] |
| LDCM0446 | CL55 | HCT 116 | C316(1.00) | LDD0763 | [3] |
| LDCM0447 | CL56 | HCT 116 | C316(1.13) | LDD0764 | [3] |
| LDCM0448 | CL57 | HCT 116 | C316(0.96) | LDD0765 | [3] |
| LDCM0449 | CL58 | HCT 116 | C316(1.03) | LDD0766 | [3] |
| LDCM0450 | CL59 | HCT 116 | C316(1.09) | LDD0767 | [3] |
| LDCM0451 | CL6 | HCT 116 | C316(1.06) | LDD0768 | [3] |
| LDCM0452 | CL60 | HCT 116 | C316(1.09) | LDD0769 | [3] |
| LDCM0453 | CL61 | HCT 116 | C316(0.91) | LDD0770 | [3] |
| LDCM0454 | CL62 | HCT 116 | C316(0.92) | LDD0771 | [3] |
| LDCM0455 | CL63 | HCT 116 | C316(0.75) | LDD0772 | [3] |
| LDCM0456 | CL64 | HCT 116 | C316(0.81) | LDD0773 | [3] |
| LDCM0457 | CL65 | HCT 116 | C316(0.91) | LDD0774 | [3] |
| LDCM0458 | CL66 | HCT 116 | C316(0.75) | LDD0775 | [3] |
| LDCM0459 | CL67 | HCT 116 | C316(0.74) | LDD0776 | [3] |
| LDCM0460 | CL68 | HCT 116 | C316(0.70) | LDD0777 | [3] |
| LDCM0461 | CL69 | HCT 116 | C316(0.72) | LDD0778 | [3] |
| LDCM0462 | CL7 | HCT 116 | C316(1.05) | LDD0779 | [3] |
| LDCM0463 | CL70 | HCT 116 | C316(0.79) | LDD0780 | [3] |
| LDCM0464 | CL71 | HCT 116 | C316(0.70) | LDD0781 | [3] |
| LDCM0465 | CL72 | HCT 116 | C316(0.77) | LDD0782 | [3] |
| LDCM0466 | CL73 | HCT 116 | C316(0.71) | LDD0783 | [3] |
| LDCM0467 | CL74 | HCT 116 | C316(0.70) | LDD0784 | [3] |
| LDCM0469 | CL76 | HCT 116 | C316(0.91) | LDD0786 | [3] |
| LDCM0470 | CL77 | HCT 116 | C316(0.85) | LDD0787 | [3] |
| LDCM0471 | CL78 | HCT 116 | C316(0.93) | LDD0788 | [3] |
| LDCM0472 | CL79 | HCT 116 | C316(0.89) | LDD0789 | [3] |
| LDCM0473 | CL8 | HCT 116 | C316(0.88) | LDD0790 | [3] |
| LDCM0474 | CL80 | HCT 116 | C316(1.07) | LDD0791 | [3] |
| LDCM0475 | CL81 | HCT 116 | C316(0.91) | LDD0792 | [3] |
| LDCM0476 | CL82 | HCT 116 | C316(1.10) | LDD0793 | [3] |
| LDCM0477 | CL83 | HCT 116 | C316(0.90) | LDD0794 | [3] |
| LDCM0478 | CL84 | HCT 116 | C316(0.98) | LDD0795 | [3] |
| LDCM0479 | CL85 | HCT 116 | C316(0.86) | LDD0796 | [3] |
| LDCM0480 | CL86 | HCT 116 | C316(0.83) | LDD0797 | [3] |
| LDCM0481 | CL87 | HCT 116 | C316(0.82) | LDD0798 | [3] |
| LDCM0482 | CL88 | HCT 116 | C316(0.81) | LDD0799 | [3] |
| LDCM0483 | CL89 | HCT 116 | C316(0.94) | LDD0800 | [3] |
| LDCM0484 | CL9 | HCT 116 | C316(0.92) | LDD0801 | [3] |
| LDCM0485 | CL90 | HCT 116 | C316(1.01) | LDD0802 | [3] |
| LDCM0486 | CL91 | HCT 116 | C316(0.90) | LDD0803 | [3] |
| LDCM0487 | CL92 | HCT 116 | C316(1.09) | LDD0804 | [3] |
| LDCM0488 | CL93 | HCT 116 | C316(1.14) | LDD0805 | [3] |
| LDCM0489 | CL94 | HCT 116 | C316(0.91) | LDD0806 | [3] |
| LDCM0490 | CL95 | HCT 116 | C316(0.91) | LDD0807 | [3] |
| LDCM0491 | CL96 | HCT 116 | C316(0.95) | LDD0808 | [3] |
| LDCM0492 | CL97 | HCT 116 | C316(0.91) | LDD0809 | [3] |
| LDCM0493 | CL98 | HCT 116 | C316(0.91) | LDD0810 | [3] |
| LDCM0494 | CL99 | HCT 116 | C316(0.90) | LDD0811 | [3] |
| LDCM0182 | Compound 18 | HEK-293T | 10.15 | LDD0501 | [9] |
| LDCM0191 | Compound 21 | HEK-293T | 8.73 | LDD0493 | [9] |
| LDCM0190 | Compound 34 | HEK-293T | 4.91 | LDD0497 | [9] |
| LDCM0192 | Compound 35 | HEK-293T | 4.55 | LDD0491 | [9] |
| LDCM0193 | Compound 36 | HEK-293T | 9.80 | LDD0494 | [9] |
| LDCM0195 | Compound 38 | HEK-293T | 4.28 | LDD0499 | [9] |
| LDCM0196 | Compound 39 | HEK-293T | 8.70 | LDD0496 | [9] |
| LDCM0197 | Compound 40 | HEK-293T | 12.99 | LDD0495 | [9] |
| LDCM0495 | E2913 | HEK-293T | C316(0.85); C270(0.99) | LDD1698 | [14] |
| LDCM0625 | F8 | Ramos | C316(1.09) | LDD2187 | [15] |
| LDCM0082 | FK866 | A-549 | 3.29 | LDD0142 | [12] |
| LDCM0573 | Fragment11 | Ramos | C316(2.21) | LDD2190 | [15] |
| LDCM0574 | Fragment12 | Ramos | C316(8.46) | LDD2191 | [15] |
| LDCM0575 | Fragment13 | Ramos | C316(1.27) | LDD2192 | [15] |
| LDCM0582 | Fragment23 | Ramos | C316(1.09) | LDD2196 | [15] |
| LDCM0586 | Fragment28 | Ramos | C316(0.53) | LDD2198 | [15] |
| LDCM0588 | Fragment30 | Ramos | C316(2.83) | LDD2199 | [15] |
| LDCM0468 | Fragment33 | HCT 116 | C316(0.74) | LDD0785 | [3] |
| LDCM0427 | Fragment51 | HCT 116 | C316(1.62) | LDD0744 | [3] |
| LDCM0022 | KB02 | HCT 116 | C316(2.07) | LDD0080 | [3] |
| LDCM0023 | KB03 | HCT 116 | C316(0.59) | LDD0081 | [3] |
| LDCM0024 | KB05 | HCT 116 | C316(1.04) | LDD0082 | [3] |
| LDCM0084 | Ro 48-8071 | A-549 | 2.71 | LDD0145 | [12] |
References
