Details of the Target
General Information of Target
| Target ID | LDTP03610 | |||||
|---|---|---|---|---|---|---|
| Target Name | Cytochrome b-c1 complex subunit 1, mitochondrial (UQCRC1) | |||||
| Gene Name | UQCRC1 | |||||
| Gene ID | 7384 | |||||
| Synonyms |
Cytochrome b-c1 complex subunit 1, mitochondrial; Complex III subunit 1; Core protein I; Ubiquinol-cytochrome-c reductase complex core protein 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MAASVVCRAATAGAQVLLRARRSPALLRTPALRSTATFAQALQFVPETQVSLLDNGLRVA
SEQSSQPTCTVGVWIDVGSRFETEKNNGAGYFLEHLAFKGTKNRPGSALEKEVESMGAHL NAYSTREHTAYYIKALSKDLPKAVELLGDIVQNCSLEDSQIEKERDVILREMQENDASMR DVVFNYLHATAFQGTPLAQAVEGPSENVRKLSRADLTEYLSTHYKAPRMVLAAAGGVEHQ QLLDLAQKHLGGIPWTYAEDAVPTLTPCRFTGSEIRHRDDALPFAHVAIAVEGPGWASPD NVALQVANAIIGHYDCTYGGGVHLSSPLASGAVANKLCQSFQTFSICYAETGLLGAHFVC DRMKIDDMMFVLQGQWMRLCTSATESEVARGKNILRNALVSHLDGTTPVCEDIGRSLLTY GRRIPLAEWESRIAEVDASVVREICSKYIYDQCPAVAGYGPIEQLPDYNRIRSGMFWLRF |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Peptidase M16 family, UQCRC1/QCR1 subfamily
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Component of the ubiquinol-cytochrome c oxidoreductase, a multisubunit transmembrane complex that is part of the mitochondrial electron transport chain which drives oxidative phosphorylation. The respiratory chain contains 3 multisubunit complexes succinate dehydrogenase (complex II, CII), ubiquinol-cytochrome c oxidoreductase (cytochrome b-c1 complex, complex III, CIII) and cytochrome c oxidase (complex IV, CIV), that cooperate to transfer electrons derived from NADH and succinate to molecular oxygen, creating an electrochemical gradient over the inner membrane that drives transmembrane transport and the ATP synthase. The cytochrome b-c1 complex catalyzes electron transfer from ubiquinol to cytochrome c, linking this redox reaction to translocation of protons across the mitochondrial inner membrane, with protons being carried across the membrane as hydrogens on the quinol. In the process called Q cycle, 2 protons are consumed from the matrix, 4 protons are released into the intermembrane space and 2 electrons are passed to cytochrome c. The 2 core subunits UQCRC1/QCR1 and UQCRC2/QCR2 are homologous to the 2 mitochondrial-processing peptidase (MPP) subunits beta-MPP and alpha-MPP respectively, and they seem to have preserved their MPP processing properties. May be involved in the in situ processing of UQCRFS1 into the mature Rieske protein and its mitochondrial targeting sequence (MTS)/subunit 9 when incorporated into complex III (Probable). Seems to play an important role in the maintenance of proper mitochondrial function in nigral dopaminergic neurons.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
6.09 | LDD0402 | [1] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [2] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
W1 Probe Info |
 |
17.50 | LDD0235 | [3] | |
|
TH211 Probe Info |
 |
Y132(20.00) | LDD0260 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [5] | |
|
ONAyne Probe Info |
 |
K225(8.33) | LDD0274 | [6] | |
|
STPyne Probe Info |
 |
K134(2.04); K138(8.13); K225(10.00) | LDD0277 | [6] | |
|
Probe 1 Probe Info |
 |
Y123(17.38); Y420(25.02) | LDD3495 | [7] | |
|
JZ128-DTB Probe Info |
 |
C453(0.00); C410(0.00) | LDD0462 | [8] | |
|
EA-probe Probe Info |
 |
C453(0.78) | LDD2210 | [9] | |
|
DBIA Probe Info |
 |
C453(0.87) | LDD0078 | [10] | |
|
5E-2FA Probe Info |
 |
H119(0.00); H402(0.00) | LDD2235 | [11] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C154(0.00); C268(0.00); C380(0.00); C410(0.00) | LDD0038 | [12] | |
|
IA-alkyne Probe Info |
 |
C268(0.00); C380(0.00); C154(0.00); C410(0.00) | LDD0032 | [13] | |
|
IPIAA_L Probe Info |
 |
C380(0.00); C154(0.00) | LDD0031 | [14] | |
|
Lodoacetamide azide Probe Info |
 |
C154(0.00); C268(0.00); C380(0.00); C410(0.00) | LDD0037 | [12] | |
|
Alkyne tyramide Probe Info |
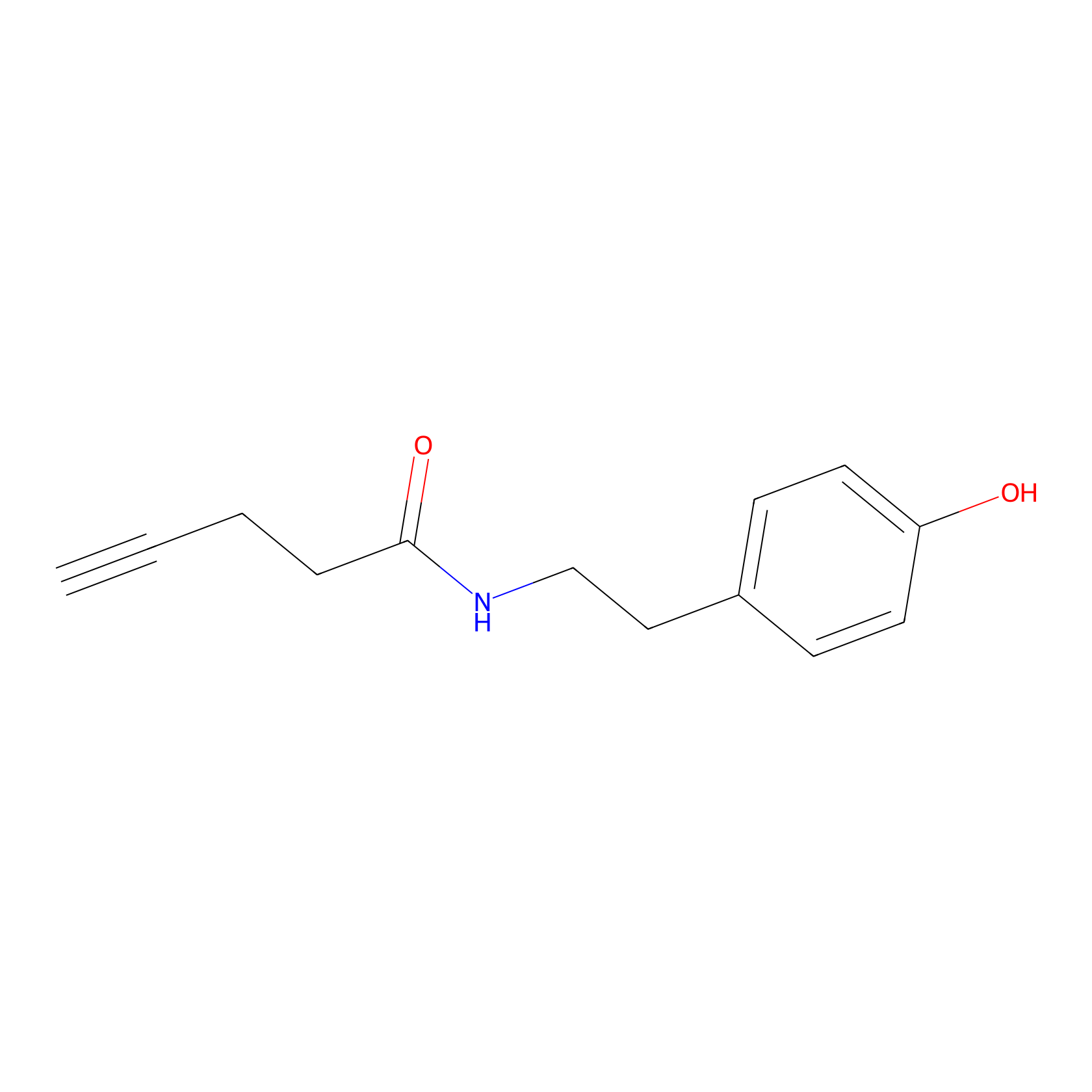 |
Y123(0.00); Y420(0.00) | LDD0003 | [15] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [15] | |
|
IPM Probe Info |
 |
C154(0.00); C380(0.00); C453(0.00); C410(0.00) | LDD0025 | [16] | |
|
JW-RF-010 Probe Info |
 |
C410(0.00); C154(0.00); C453(0.00); C380(0.00) | LDD0026 | [16] | |
|
NAIA_4 Probe Info |
 |
C154(0.00); C268(0.00); C380(0.00); C410(0.00) | LDD2226 | [17] | |
|
TFBX Probe Info |
 |
C380(0.00); C7(0.00); C445(0.00) | LDD0027 | [16] | |
|
WYneN Probe Info |
 |
C154(0.00); C410(0.00); C380(0.00) | LDD0021 | [15] | |
|
WYneO Probe Info |
 |
C380(0.00); C410(0.00) | LDD0022 | [15] | |
|
Compound 10 Probe Info |
 |
C268(0.00); C380(0.00); C410(0.00); C453(0.00) | LDD2216 | [18] | |
|
Compound 11 Probe Info |
 |
C268(0.00); C380(0.00); C410(0.00) | LDD2213 | [18] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0152 | [19] | |
|
SF Probe Info |
 |
Y219(0.00); Y420(0.00) | LDD0028 | [20] | |
|
Phosphinate-6 Probe Info |
 |
C410(0.00); C154(0.00); C453(0.00); C268(0.00) | LDD0018 | [21] | |
|
Ox-W18 Probe Info |
 |
W429(0.00); W255(0.00) | LDD2175 | [22] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [23] | |
|
Acrolein Probe Info |
 |
C410(0.00); C154(0.00); C453(0.00); C380(0.00) | LDD0217 | [24] | |
|
Cinnamaldehyde Probe Info |
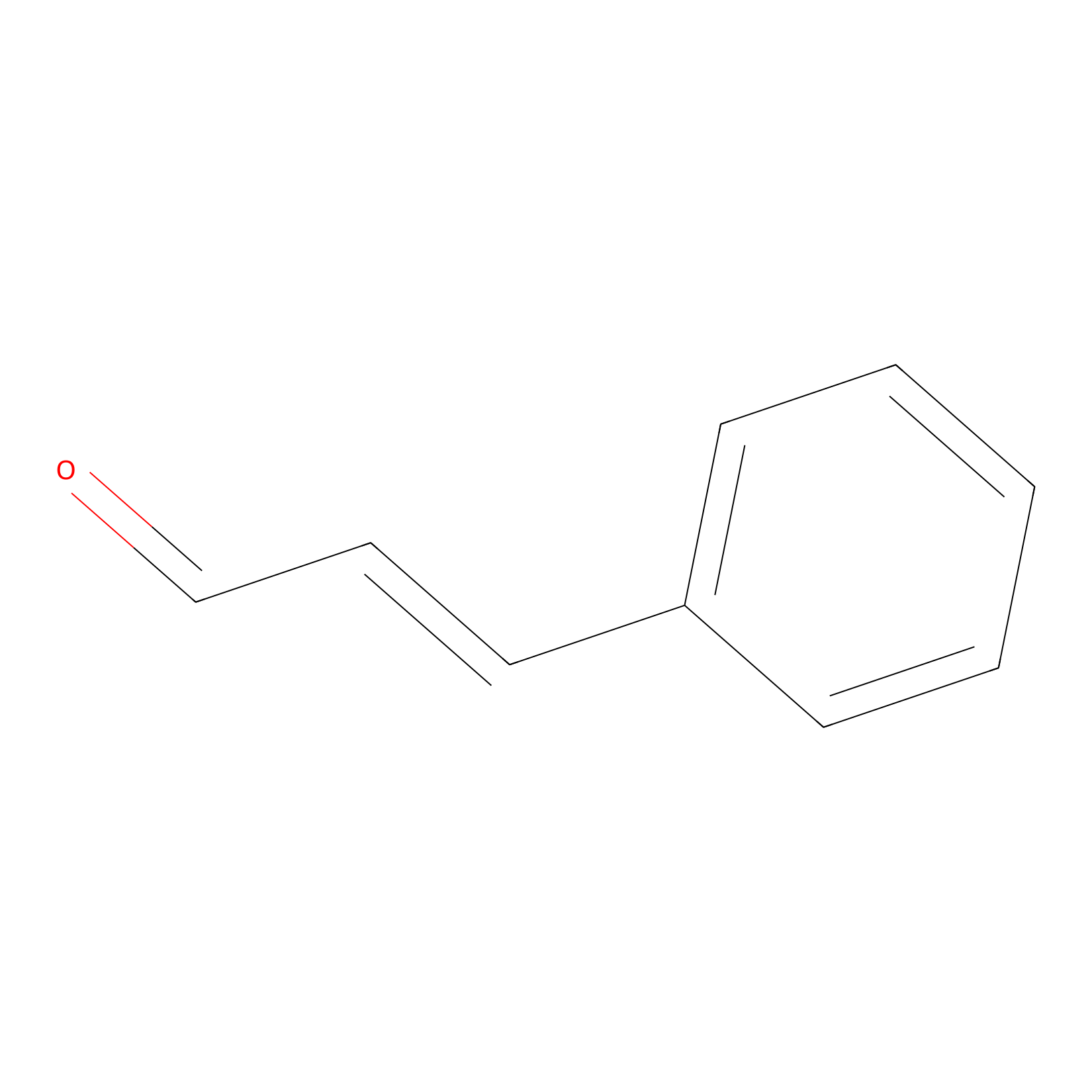 |
C380(0.00); C410(0.00) | LDD0220 | [24] | |
|
Crotonaldehyde Probe Info |
 |
C410(0.00); C380(0.00); C453(0.00) | LDD0219 | [24] | |
|
Methacrolein Probe Info |
 |
C410(0.00); C380(0.00) | LDD0218 | [24] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [25] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [26] | |
|
NAIA_5 Probe Info |
 |
C154(0.00); C268(0.00); C380(0.00); C410(0.00) | LDD2223 | [17] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [26] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [26] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C139 Probe Info |
 |
10.56 | LDD1821 | [27] | |
|
C217 Probe Info |
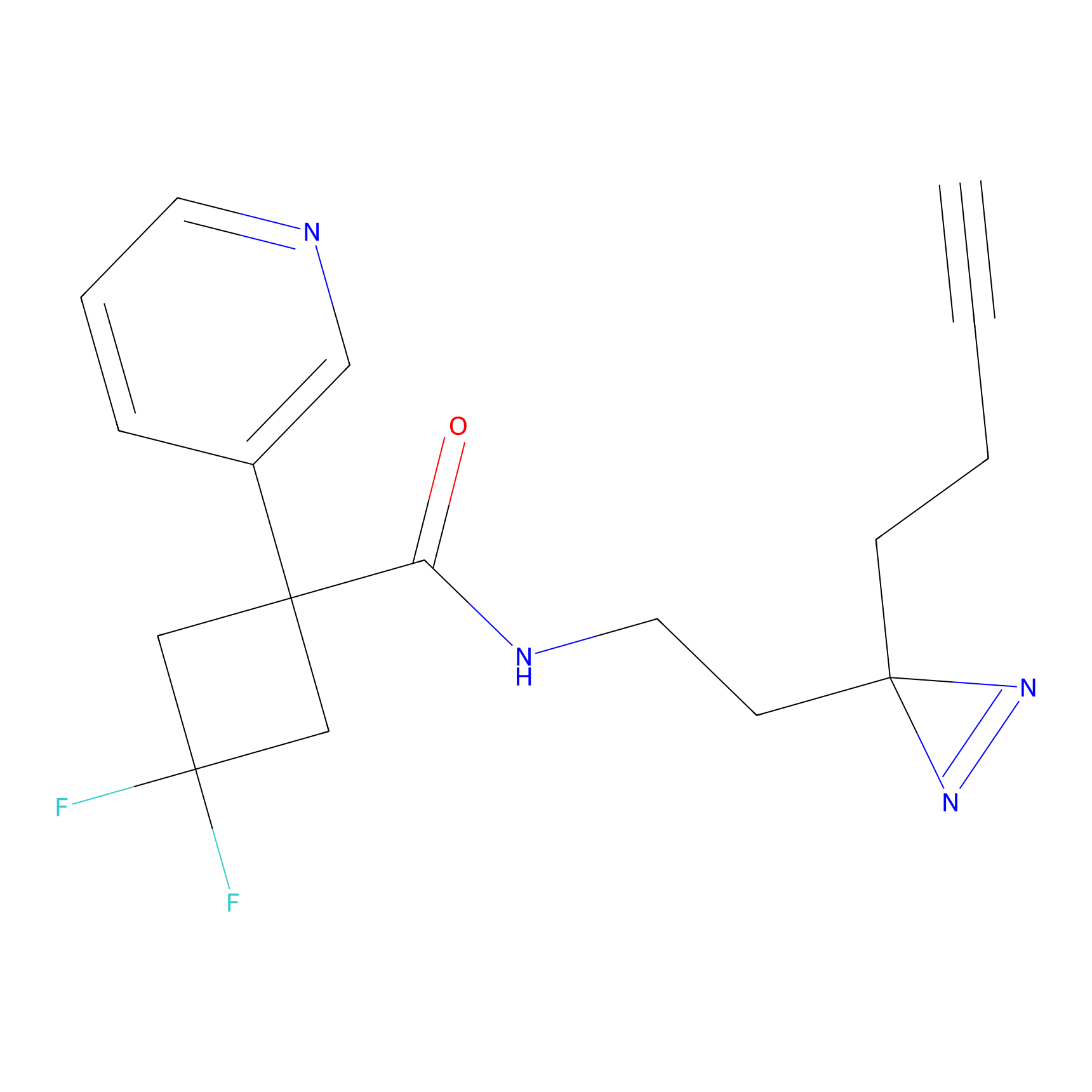 |
5.31 | LDD1891 | [27] | |
|
C234 Probe Info |
 |
5.58 | LDD1907 | [27] | |
|
C366 Probe Info |
 |
6.28 | LDD2027 | [27] | |
|
C423 Probe Info |
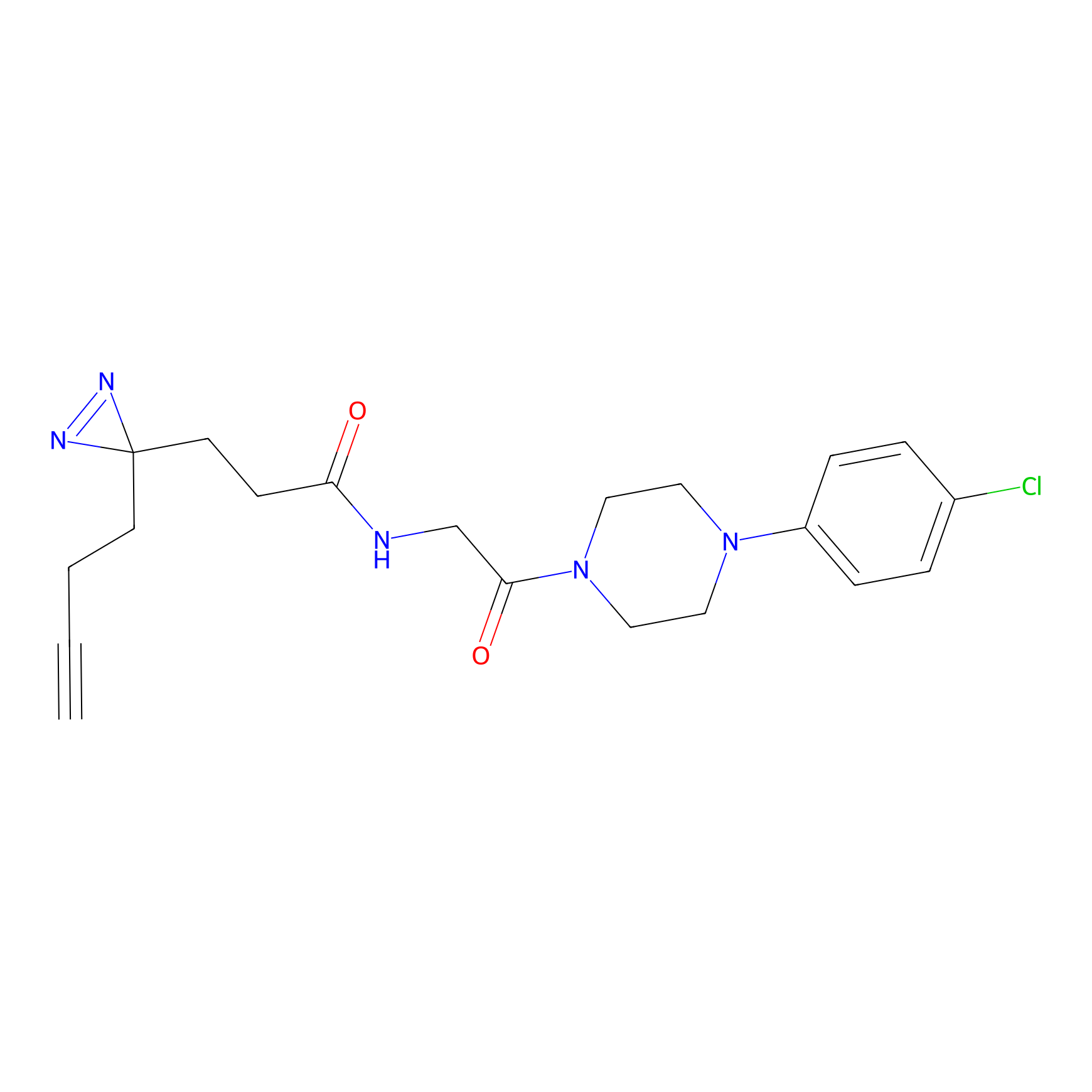 |
7.46 | LDD2078 | [27] | |
|
FFF probe11 Probe Info |
 |
12.73 | LDD0471 | [28] | |
|
FFF probe13 Probe Info |
 |
15.96 | LDD0475 | [28] | |
|
FFF probe14 Probe Info |
 |
14.59 | LDD0477 | [28] | |
|
FFF probe2 Probe Info |
 |
12.29 | LDD0463 | [28] | |
|
FFF probe3 Probe Info |
 |
11.41 | LDD0464 | [28] | |
|
FFF probe4 Probe Info |
 |
12.39 | LDD0466 | [28] | |
|
FFF probe7 Probe Info |
 |
12.33 | LDD0483 | [28] | |
|
JN0003 Probe Info |
 |
13.11 | LDD0469 | [28] | |
|
BD-F Probe Info |
 |
G405(0.00); A457(0.00); P454(0.00); E411(0.00) | LDD0024 | [29] | |
|
LD-F Probe Info |
 |
E411(0.00); V409(0.00); P408(0.00) | LDD0015 | [29] | |
|
OEA-DA Probe Info |
 |
6.66 | LDD0046 | [30] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0069 | [31] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C380(0.98) | LDD2142 | [32] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C380(1.10); C410(0.99) | LDD2112 | [32] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C380(0.57); C410(0.51) | LDD2095 | [32] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C380(0.59); C410(0.80) | LDD2130 | [32] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C380(1.00); C410(0.88) | LDD2117 | [32] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C410(1.44) | LDD2152 | [32] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C380(0.52); C410(0.64) | LDD2132 | [32] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C380(0.45) | LDD2131 | [32] |
| LDCM0214 | AC1 | HCT 116 | C453(1.02) | LDD0531 | [10] |
| LDCM0215 | AC10 | HCT 116 | C453(0.41); C268(0.72); C410(0.96) | LDD0532 | [10] |
| LDCM0216 | AC100 | HCT 116 | C453(0.93); C380(0.95); C410(1.54) | LDD0533 | [10] |
| LDCM0217 | AC101 | HCT 116 | C453(1.04); C380(0.94); C410(1.22) | LDD0534 | [10] |
| LDCM0218 | AC102 | HCT 116 | C453(1.24); C380(0.91); C410(1.08) | LDD0535 | [10] |
| LDCM0219 | AC103 | HCT 116 | C453(1.27); C380(0.64); C410(1.22) | LDD0536 | [10] |
| LDCM0220 | AC104 | HCT 116 | C453(1.11); C380(0.60); C410(1.39) | LDD0537 | [10] |
| LDCM0221 | AC105 | HCT 116 | C453(1.13); C380(0.84); C410(1.23) | LDD0538 | [10] |
| LDCM0222 | AC106 | HCT 116 | C453(1.25); C380(0.79); C410(1.40) | LDD0539 | [10] |
| LDCM0223 | AC107 | HCT 116 | C453(1.05); C380(0.76); C410(1.33) | LDD0540 | [10] |
| LDCM0224 | AC108 | HCT 116 | C453(1.13); C380(0.86); C410(1.15) | LDD0541 | [10] |
| LDCM0225 | AC109 | HCT 116 | C453(1.09); C380(0.70); C410(1.06) | LDD0542 | [10] |
| LDCM0226 | AC11 | HCT 116 | C453(0.47); C268(0.68); C410(0.63) | LDD0543 | [10] |
| LDCM0227 | AC110 | HCT 116 | C453(1.36); C380(0.79); C410(1.45) | LDD0544 | [10] |
| LDCM0228 | AC111 | HCT 116 | C453(1.31); C380(0.71); C410(1.05) | LDD0545 | [10] |
| LDCM0229 | AC112 | HCT 116 | C453(1.32); C380(0.92); C410(0.82) | LDD0546 | [10] |
| LDCM0230 | AC113 | HCT 116 | C453(1.13); C268(1.15); C380(0.93) | LDD0547 | [10] |
| LDCM0231 | AC114 | HCT 116 | C453(0.90); C268(0.87); C380(0.82) | LDD0548 | [10] |
| LDCM0232 | AC115 | HCT 116 | C453(0.99); C268(1.00); C380(0.89) | LDD0549 | [10] |
| LDCM0233 | AC116 | HCT 116 | C453(0.90); C268(1.08); C380(0.94) | LDD0550 | [10] |
| LDCM0234 | AC117 | HCT 116 | C453(0.94); C268(1.16); C380(0.87) | LDD0551 | [10] |
| LDCM0235 | AC118 | HCT 116 | C453(0.89); C268(1.12); C380(1.03) | LDD0552 | [10] |
| LDCM0236 | AC119 | HCT 116 | C453(0.88); C268(1.38); C380(1.00) | LDD0553 | [10] |
| LDCM0237 | AC12 | HCT 116 | C453(0.63); C268(0.75); C410(0.78) | LDD0554 | [10] |
| LDCM0238 | AC120 | HCT 116 | C453(2.07); C268(0.81); C380(1.13) | LDD0555 | [10] |
| LDCM0239 | AC121 | HCT 116 | C453(1.29); C268(1.39); C380(0.89) | LDD0556 | [10] |
| LDCM0240 | AC122 | HCT 116 | C453(1.08); C268(0.93); C380(0.97) | LDD0557 | [10] |
| LDCM0241 | AC123 | HCT 116 | C453(2.04); C268(1.09); C380(1.14) | LDD0558 | [10] |
| LDCM0242 | AC124 | HCT 116 | C453(0.95); C268(1.32); C380(0.89) | LDD0559 | [10] |
| LDCM0243 | AC125 | HCT 116 | C453(1.31); C268(1.02); C380(1.00) | LDD0560 | [10] |
| LDCM0244 | AC126 | HCT 116 | C453(1.10); C268(1.04); C380(0.98) | LDD0561 | [10] |
| LDCM0245 | AC127 | HCT 116 | C453(1.16); C268(1.01); C380(0.84) | LDD0562 | [10] |
| LDCM0246 | AC128 | HCT 116 | C453(3.12); C410(0.86) | LDD0563 | [10] |
| LDCM0247 | AC129 | HCT 116 | C453(1.19); C410(1.78) | LDD0564 | [10] |
| LDCM0249 | AC130 | HCT 116 | C453(1.13); C410(1.19) | LDD0566 | [10] |
| LDCM0250 | AC131 | HCT 116 | C453(2.71); C410(2.02) | LDD0567 | [10] |
| LDCM0251 | AC132 | HCT 116 | C453(1.03); C410(2.20) | LDD0568 | [10] |
| LDCM0252 | AC133 | HCT 116 | C453(2.55); C410(1.46) | LDD0569 | [10] |
| LDCM0253 | AC134 | HCT 116 | C453(3.70); C410(1.48) | LDD0570 | [10] |
| LDCM0254 | AC135 | HCT 116 | C453(1.02); C410(1.44) | LDD0571 | [10] |
| LDCM0255 | AC136 | HCT 116 | C453(1.48); C410(1.12) | LDD0572 | [10] |
| LDCM0256 | AC137 | HCT 116 | C453(1.18); C410(1.37) | LDD0573 | [10] |
| LDCM0257 | AC138 | HCT 116 | C453(1.51); C410(1.34) | LDD0574 | [10] |
| LDCM0258 | AC139 | HCT 116 | C453(0.98); C410(1.78) | LDD0575 | [10] |
| LDCM0259 | AC14 | HCT 116 | C453(0.87); C268(0.86); C410(0.92) | LDD0576 | [10] |
| LDCM0260 | AC140 | HCT 116 | C453(1.04); C410(1.53) | LDD0577 | [10] |
| LDCM0261 | AC141 | HCT 116 | C453(1.15); C410(1.50) | LDD0578 | [10] |
| LDCM0262 | AC142 | HCT 116 | C453(0.96); C410(2.15) | LDD0579 | [10] |
| LDCM0263 | AC143 | HCT 116 | C453(0.82); C410(0.75) | LDD0580 | [10] |
| LDCM0264 | AC144 | HCT 116 | C410(0.57); C453(1.38) | LDD0581 | [10] |
| LDCM0265 | AC145 | HCT 116 | C410(0.66); C453(0.96) | LDD0582 | [10] |
| LDCM0266 | AC146 | HCT 116 | C410(0.65); C453(1.13) | LDD0583 | [10] |
| LDCM0267 | AC147 | HCT 116 | C410(0.49); C453(1.24) | LDD0584 | [10] |
| LDCM0268 | AC148 | HCT 116 | C410(0.41); C453(1.00) | LDD0585 | [10] |
| LDCM0269 | AC149 | HCT 116 | C410(0.55); C453(1.05) | LDD0586 | [10] |
| LDCM0270 | AC15 | HCT 116 | C410(0.72); C453(0.82); C268(0.98) | LDD0587 | [10] |
| LDCM0271 | AC150 | HCT 116 | C410(0.77); C453(0.95) | LDD0588 | [10] |
| LDCM0272 | AC151 | HCT 116 | C410(0.87); C453(0.92) | LDD0589 | [10] |
| LDCM0273 | AC152 | HCT 116 | C410(0.43); C453(1.24) | LDD0590 | [10] |
| LDCM0274 | AC153 | HCT 116 | C410(0.39); C453(0.76) | LDD0591 | [10] |
| LDCM0621 | AC154 | HCT 116 | C453(0.86); C410(0.56) | LDD2158 | [10] |
| LDCM0622 | AC155 | HCT 116 | C453(0.63); C410(0.60) | LDD2159 | [10] |
| LDCM0623 | AC156 | HCT 116 | C453(0.93); C410(1.07) | LDD2160 | [10] |
| LDCM0624 | AC157 | HCT 116 | C453(0.93); C410(1.17) | LDD2161 | [10] |
| LDCM0276 | AC17 | HCT 116 | C410(0.77); C380(1.29); C453(1.71) | LDD0593 | [10] |
| LDCM0277 | AC18 | HCT 116 | C410(0.37); C453(0.91); C380(1.14) | LDD0594 | [10] |
| LDCM0278 | AC19 | HCT 116 | C410(0.68); C380(1.36); C453(1.73) | LDD0595 | [10] |
| LDCM0279 | AC2 | HCT 116 | C453(0.71) | LDD0596 | [10] |
| LDCM0280 | AC20 | HCT 116 | C410(0.89); C453(1.52); C380(1.76) | LDD0597 | [10] |
| LDCM0281 | AC21 | HCT 116 | C410(0.77); C380(1.23); C453(1.31) | LDD0598 | [10] |
| LDCM0282 | AC22 | HCT 116 | C410(0.70); C453(1.30); C380(1.33) | LDD0599 | [10] |
| LDCM0283 | AC23 | HCT 116 | C410(0.84); C453(1.56); C380(1.72) | LDD0600 | [10] |
| LDCM0284 | AC24 | HCT 116 | C380(1.32); C410(1.37); C453(1.80) | LDD0601 | [10] |
| LDCM0285 | AC25 | HCT 116 | C453(1.01); C380(1.49) | LDD0602 | [10] |
| LDCM0286 | AC26 | HCT 116 | C380(0.54); C453(0.87) | LDD0603 | [10] |
| LDCM0287 | AC27 | HCT 116 | C380(0.64); C453(1.38) | LDD0604 | [10] |
| LDCM0288 | AC28 | HCT 116 | C380(1.00); C453(1.12) | LDD0605 | [10] |
| LDCM0289 | AC29 | HCT 116 | C380(0.66); C453(0.96) | LDD0606 | [10] |
| LDCM0290 | AC3 | HCT 116 | C453(0.78) | LDD0607 | [10] |
| LDCM0291 | AC30 | HCT 116 | C380(0.56); C453(0.95) | LDD0608 | [10] |
| LDCM0292 | AC31 | HCT 116 | C380(0.72); C453(0.99) | LDD0609 | [10] |
| LDCM0293 | AC32 | HCT 116 | C380(0.61); C453(1.00) | LDD0610 | [10] |
| LDCM0294 | AC33 | HCT 116 | C380(0.92); C453(1.28) | LDD0611 | [10] |
| LDCM0295 | AC34 | HCT 116 | C380(0.73); C453(1.36) | LDD0612 | [10] |
| LDCM0296 | AC35 | HCT 116 | C453(0.84); C380(0.86); C410(1.01) | LDD0613 | [10] |
| LDCM0297 | AC36 | HCT 116 | C380(0.76); C453(0.84); C410(1.61) | LDD0614 | [10] |
| LDCM0298 | AC37 | HCT 116 | C453(0.85); C380(1.04); C410(1.41) | LDD0615 | [10] |
| LDCM0299 | AC38 | HCT 116 | C453(0.73); C380(0.74); C410(1.64) | LDD0616 | [10] |
| LDCM0300 | AC39 | HCT 116 | C453(0.68); C380(0.73); C410(0.93) | LDD0617 | [10] |
| LDCM0301 | AC4 | HCT 116 | C453(0.64) | LDD0618 | [10] |
| LDCM0302 | AC40 | HCT 116 | C410(0.69); C453(0.78); C380(0.81) | LDD0619 | [10] |
| LDCM0303 | AC41 | HCT 116 | C453(0.94); C380(1.05); C410(1.09) | LDD0620 | [10] |
| LDCM0304 | AC42 | HCT 116 | C453(0.87); C410(0.91); C380(0.92) | LDD0621 | [10] |
| LDCM0305 | AC43 | HCT 116 | C453(0.69); C380(0.73); C410(1.07) | LDD0622 | [10] |
| LDCM0306 | AC44 | HCT 116 | C410(0.85); C453(1.03); C380(1.54) | LDD0623 | [10] |
| LDCM0307 | AC45 | HCT 116 | C410(0.67); C453(0.84); C380(0.92) | LDD0624 | [10] |
| LDCM0308 | AC46 | HCT 116 | C380(0.40); C453(0.83); C268(0.99) | LDD0625 | [10] |
| LDCM0309 | AC47 | HCT 116 | C380(0.38); C268(0.66); C453(0.81) | LDD0626 | [10] |
| LDCM0310 | AC48 | HCT 116 | C380(0.51); C268(0.66); C453(0.67) | LDD0627 | [10] |
| LDCM0311 | AC49 | HCT 116 | C380(0.33); C268(0.40); C453(1.20) | LDD0628 | [10] |
| LDCM0312 | AC5 | HCT 116 | C453(0.53) | LDD0629 | [10] |
| LDCM0313 | AC50 | HCT 116 | C380(0.31); C268(0.36); C453(1.43) | LDD0630 | [10] |
| LDCM0314 | AC51 | HCT 116 | C380(0.30); C453(0.98); C268(1.59) | LDD0631 | [10] |
| LDCM0315 | AC52 | HCT 116 | C380(0.40); C268(0.65); C453(1.13) | LDD0632 | [10] |
| LDCM0316 | AC53 | HCT 116 | C268(0.55); C453(1.25); C380(1.62) | LDD0633 | [10] |
| LDCM0317 | AC54 | HCT 116 | C268(0.49); C380(0.50); C453(1.01) | LDD0634 | [10] |
| LDCM0318 | AC55 | HCT 116 | C268(0.40); C380(0.83); C453(1.00) | LDD0635 | [10] |
| LDCM0319 | AC56 | HCT 116 | C268(0.27); C380(0.33); C453(1.04) | LDD0636 | [10] |
| LDCM0320 | AC57 | HCT 116 | C453(0.99) | LDD0637 | [10] |
| LDCM0321 | AC58 | HCT 116 | C453(1.26) | LDD0638 | [10] |
| LDCM0322 | AC59 | HCT 116 | C453(1.22) | LDD0639 | [10] |
| LDCM0323 | AC6 | HCT 116 | C453(0.78); C268(0.93); C410(0.96) | LDD0640 | [10] |
| LDCM0324 | AC60 | HCT 116 | C453(1.42) | LDD0641 | [10] |
| LDCM0325 | AC61 | HCT 116 | C453(0.96) | LDD0642 | [10] |
| LDCM0326 | AC62 | HCT 116 | C453(1.07) | LDD0643 | [10] |
| LDCM0327 | AC63 | HCT 116 | C453(1.29) | LDD0644 | [10] |
| LDCM0328 | AC64 | HCT 116 | C453(1.38) | LDD0645 | [10] |
| LDCM0329 | AC65 | HCT 116 | C453(1.22) | LDD0646 | [10] |
| LDCM0330 | AC66 | HCT 116 | C453(1.33) | LDD0647 | [10] |
| LDCM0331 | AC67 | HCT 116 | C453(1.43) | LDD0648 | [10] |
| LDCM0332 | AC68 | HCT 116 | C268(0.60); C380(0.66); C453(0.97) | LDD0649 | [10] |
| LDCM0333 | AC69 | HCT 116 | C268(0.52); C453(0.90); C380(1.15) | LDD0650 | [10] |
| LDCM0334 | AC7 | HCT 116 | C453(0.50); C410(0.84); C268(0.92) | LDD0651 | [10] |
| LDCM0335 | AC70 | HCT 116 | C268(0.32); C453(0.75); C380(1.22) | LDD0652 | [10] |
| LDCM0336 | AC71 | HCT 116 | C268(0.75); C453(0.81); C380(1.53) | LDD0653 | [10] |
| LDCM0337 | AC72 | HCT 116 | C268(0.59); C380(0.70); C453(0.76) | LDD0654 | [10] |
| LDCM0338 | AC73 | HCT 116 | C453(0.78); C268(0.39); C380(0.72) | LDD0655 | [10] |
| LDCM0339 | AC74 | HCT 116 | C268(0.37); C453(0.87); C380(1.14) | LDD0656 | [10] |
| LDCM0340 | AC75 | HCT 116 | C268(0.33); C380(0.75); C453(0.91) | LDD0657 | [10] |
| LDCM0341 | AC76 | HCT 116 | C268(0.42); C453(0.83) | LDD0658 | [10] |
| LDCM0342 | AC77 | HCT 116 | C268(0.45); C380(0.70); C453(1.32) | LDD0659 | [10] |
| LDCM0343 | AC78 | HCT 116 | C268(0.83); C453(0.91); C380(1.12) | LDD0660 | [10] |
| LDCM0344 | AC79 | HCT 116 | C268(0.77); C453(0.86); C380(0.95) | LDD0661 | [10] |
| LDCM0345 | AC8 | HCT 116 | C410(0.64); C453(0.76); C268(0.91) | LDD0662 | [10] |
| LDCM0346 | AC80 | HCT 116 | C268(0.43); C453(0.85); C380(3.31) | LDD0663 | [10] |
| LDCM0347 | AC81 | HCT 116 | C453(0.77); C268(0.93); C380(1.08) | LDD0664 | [10] |
| LDCM0348 | AC82 | HCT 116 | C268(0.16); C453(0.81); C380(0.88) | LDD0665 | [10] |
| LDCM0349 | AC83 | HCT 116 | C380(0.35); C410(0.50); C453(1.01) | LDD0666 | [10] |
| LDCM0350 | AC84 | HCT 116 | C380(0.36); C410(0.46); C453(1.01) | LDD0667 | [10] |
| LDCM0351 | AC85 | HCT 116 | C410(0.66); C380(0.77); C453(0.82) | LDD0668 | [10] |
| LDCM0352 | AC86 | HCT 116 | C380(0.75); C453(0.85); C410(1.02) | LDD0669 | [10] |
| LDCM0353 | AC87 | HCT 116 | C453(0.74); C410(0.90); C380(1.05) | LDD0670 | [10] |
| LDCM0354 | AC88 | HCT 116 | C380(0.62); C453(0.82); C410(0.90) | LDD0671 | [10] |
| LDCM0355 | AC89 | HCT 116 | C380(0.45); C410(0.77); C453(0.88) | LDD0672 | [10] |
| LDCM0357 | AC90 | HCT 116 | C453(0.65); C380(1.08); C410(1.51) | LDD0674 | [10] |
| LDCM0358 | AC91 | HCT 116 | C380(0.35); C410(0.53); C453(1.06) | LDD0675 | [10] |
| LDCM0359 | AC92 | HCT 116 | C380(0.41); C410(0.61); C453(0.63) | LDD0676 | [10] |
| LDCM0360 | AC93 | HCT 116 | C410(0.69); C380(0.69); C453(0.82) | LDD0677 | [10] |
| LDCM0361 | AC94 | HCT 116 | C410(0.66); C453(0.68); C380(0.70) | LDD0678 | [10] |
| LDCM0362 | AC95 | HCT 116 | C380(0.68); C453(0.81); C410(0.99) | LDD0679 | [10] |
| LDCM0363 | AC96 | HCT 116 | C380(0.61); C410(0.81); C453(0.82) | LDD0680 | [10] |
| LDCM0364 | AC97 | HCT 116 | C380(0.42); C410(0.54); C453(0.83) | LDD0681 | [10] |
| LDCM0365 | AC98 | HCT 116 | C380(0.42); C410(0.86); C453(1.15) | LDD0682 | [10] |
| LDCM0366 | AC99 | HCT 116 | C410(0.86); C453(1.01); C380(1.02) | LDD0683 | [10] |
| LDCM0545 | Acetamide | MDA-MB-231 | C380(0.49) | LDD2138 | [32] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C380(0.84); C410(0.63) | LDD2113 | [32] |
| LDCM0248 | AKOS034007472 | HCT 116 | C453(0.42); C268(0.83); C410(0.95) | LDD0565 | [10] |
| LDCM0356 | AKOS034007680 | HCT 116 | C453(0.45); C268(0.77); C410(0.87) | LDD0673 | [10] |
| LDCM0275 | AKOS034007705 | HCT 116 | C410(0.38); C453(0.74); C268(1.03) | LDD0592 | [10] |
| LDCM0156 | Aniline | NCI-H1299 | 9.73 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C453(0.87) | LDD0078 | [10] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C410(0.78) | LDD2091 | [32] |
| LDCM0630 | CCW28-3 | 231MFP | C453(2.11); C268(1.17) | LDD2214 | [33] |
| LDCM0108 | Chloroacetamide | HeLa | C154(0.00); C410(0.00); C453(0.00); C268(0.00) | LDD0222 | [24] |
| LDCM0367 | CL1 | HCT 116 | C380(0.76); C453(1.31) | LDD0684 | [10] |
| LDCM0368 | CL10 | HCT 116 | C453(0.93); C380(1.00) | LDD0685 | [10] |
| LDCM0369 | CL100 | HCT 116 | C453(0.74) | LDD0686 | [10] |
| LDCM0370 | CL101 | HCT 116 | C453(0.70); C410(0.80); C268(0.96) | LDD0687 | [10] |
| LDCM0371 | CL102 | HCT 116 | C453(0.46); C268(0.85); C410(1.22) | LDD0688 | [10] |
| LDCM0372 | CL103 | HCT 116 | C453(0.57); C410(0.83); C268(0.85) | LDD0689 | [10] |
| LDCM0373 | CL104 | HCT 116 | C453(0.69); C268(0.71); C410(0.88) | LDD0690 | [10] |
| LDCM0374 | CL105 | HCT 116 | C410(0.70); C453(0.82); C380(1.20) | LDD0691 | [10] |
| LDCM0375 | CL106 | HCT 116 | C410(0.30); C453(0.77); C380(1.13) | LDD0692 | [10] |
| LDCM0376 | CL107 | HCT 116 | C410(0.41); C453(0.75); C380(1.35) | LDD0693 | [10] |
| LDCM0377 | CL108 | HCT 116 | C410(0.30); C453(0.86); C380(1.20) | LDD0694 | [10] |
| LDCM0378 | CL109 | HCT 116 | C410(0.73); C380(1.33); C453(1.57) | LDD0695 | [10] |
| LDCM0379 | CL11 | HCT 116 | C380(0.97); C453(1.48) | LDD0696 | [10] |
| LDCM0380 | CL110 | HCT 116 | C410(0.43); C453(1.07); C380(1.56) | LDD0697 | [10] |
| LDCM0381 | CL111 | HCT 116 | C410(0.76); C380(1.27); C453(1.49) | LDD0698 | [10] |
| LDCM0382 | CL112 | HCT 116 | C453(0.71); C380(0.88) | LDD0699 | [10] |
| LDCM0383 | CL113 | HCT 116 | C380(0.81); C453(1.25) | LDD0700 | [10] |
| LDCM0384 | CL114 | HCT 116 | C380(0.40); C453(1.17) | LDD0701 | [10] |
| LDCM0385 | CL115 | HCT 116 | C380(0.88); C453(1.09) | LDD0702 | [10] |
| LDCM0386 | CL116 | HCT 116 | C453(0.74); C380(0.97) | LDD0703 | [10] |
| LDCM0387 | CL117 | HCT 116 | C410(0.48); C380(0.55); C453(0.64) | LDD0704 | [10] |
| LDCM0388 | CL118 | HCT 116 | C380(0.92); C453(0.96); C410(0.97) | LDD0705 | [10] |
| LDCM0389 | CL119 | HCT 116 | C410(0.59); C453(0.85); C380(1.32) | LDD0706 | [10] |
| LDCM0390 | CL12 | HCT 116 | C380(0.68); C453(1.77) | LDD0707 | [10] |
| LDCM0391 | CL120 | HCT 116 | C380(0.88); C453(0.91); C410(1.14) | LDD0708 | [10] |
| LDCM0392 | CL121 | HCT 116 | C380(0.59); C268(0.67); C453(1.09) | LDD0709 | [10] |
| LDCM0393 | CL122 | HCT 116 | C268(0.55); C380(0.62); C453(0.76) | LDD0710 | [10] |
| LDCM0394 | CL123 | HCT 116 | C380(0.33); C268(0.51); C453(1.13) | LDD0711 | [10] |
| LDCM0395 | CL124 | HCT 116 | C380(0.44); C268(0.50); C453(0.99) | LDD0712 | [10] |
| LDCM0396 | CL125 | HCT 116 | C453(1.07) | LDD0713 | [10] |
| LDCM0397 | CL126 | HCT 116 | C453(1.22) | LDD0714 | [10] |
| LDCM0398 | CL127 | HCT 116 | C453(1.01) | LDD0715 | [10] |
| LDCM0399 | CL128 | HCT 116 | C453(1.29) | LDD0716 | [10] |
| LDCM0400 | CL13 | HCT 116 | C380(0.69); C453(1.31) | LDD0717 | [10] |
| LDCM0401 | CL14 | HCT 116 | C380(1.01); C453(1.57) | LDD0718 | [10] |
| LDCM0402 | CL15 | HCT 116 | C380(1.47); C453(1.55) | LDD0719 | [10] |
| LDCM0403 | CL16 | HEK-293T | C268(1.00); C410(1.12); C445(1.21); C453(1.05) | LDD1001 | [10] |
| LDCM0404 | CL17 | HCT 116 | C453(1.24) | LDD0721 | [10] |
| LDCM0405 | CL18 | HCT 116 | C453(0.83) | LDD0722 | [10] |
| LDCM0406 | CL19 | HCT 116 | C453(1.02) | LDD0723 | [10] |
| LDCM0407 | CL2 | HCT 116 | C380(1.09); C453(1.20) | LDD0724 | [10] |
| LDCM0408 | CL20 | HCT 116 | C453(1.67) | LDD0725 | [10] |
| LDCM0409 | CL21 | HCT 116 | C453(1.08) | LDD0726 | [10] |
| LDCM0410 | CL22 | HCT 116 | C453(1.42) | LDD0727 | [10] |
| LDCM0411 | CL23 | HCT 116 | C453(0.92) | LDD0728 | [10] |
| LDCM0412 | CL24 | HCT 116 | C453(0.92) | LDD0729 | [10] |
| LDCM0413 | CL25 | HCT 116 | C453(0.86) | LDD0730 | [10] |
| LDCM0414 | CL26 | HCT 116 | C453(0.92) | LDD0731 | [10] |
| LDCM0415 | CL27 | HCT 116 | C453(0.82) | LDD0732 | [10] |
| LDCM0416 | CL28 | HCT 116 | C453(0.87) | LDD0733 | [10] |
| LDCM0417 | CL29 | HCT 116 | C453(1.97) | LDD0734 | [10] |
| LDCM0418 | CL3 | HCT 116 | C453(1.29); C380(1.19) | LDD0735 | [10] |
| LDCM0419 | CL30 | HCT 116 | C453(0.84) | LDD0736 | [10] |
| LDCM0420 | CL31 | HCT 116 | C453(1.08); C268(0.79); C380(1.10) | LDD0737 | [10] |
| LDCM0421 | CL32 | HCT 116 | C453(1.05); C268(0.50); C380(1.03) | LDD0738 | [10] |
| LDCM0422 | CL33 | HCT 116 | C453(1.90); C268(0.72); C380(0.86) | LDD0739 | [10] |
| LDCM0423 | CL34 | HCT 116 | C453(1.24); C268(0.34); C380(0.63) | LDD0740 | [10] |
| LDCM0424 | CL35 | HCT 116 | C453(1.60); C268(0.39); C380(0.99) | LDD0741 | [10] |
| LDCM0425 | CL36 | HCT 116 | C453(1.57); C268(0.48); C380(0.56) | LDD0742 | [10] |
| LDCM0426 | CL37 | HCT 116 | C453(4.46); C268(0.38); C380(0.62) | LDD0743 | [10] |
| LDCM0428 | CL39 | HCT 116 | C453(1.87); C268(0.53); C380(1.48) | LDD0745 | [10] |
| LDCM0429 | CL4 | HCT 116 | C453(1.48); C380(0.85) | LDD0746 | [10] |
| LDCM0430 | CL40 | HCT 116 | C453(1.79); C268(0.41); C380(1.22) | LDD0747 | [10] |
| LDCM0431 | CL41 | HCT 116 | C453(1.41); C268(0.58); C380(1.03) | LDD0748 | [10] |
| LDCM0432 | CL42 | HCT 116 | C453(1.21); C268(0.22); C380(0.84) | LDD0749 | [10] |
| LDCM0433 | CL43 | HCT 116 | C453(2.33); C268(0.43); C380(1.54) | LDD0750 | [10] |
| LDCM0434 | CL44 | HCT 116 | C453(1.73); C268(0.45); C380(1.30) | LDD0751 | [10] |
| LDCM0435 | CL45 | HCT 116 | C453(1.17); C268(0.32); C380(1.46) | LDD0752 | [10] |
| LDCM0436 | CL46 | HCT 116 | C453(1.86); C380(0.63) | LDD0753 | [10] |
| LDCM0437 | CL47 | HCT 116 | C453(2.15); C380(0.87) | LDD0754 | [10] |
| LDCM0438 | CL48 | HCT 116 | C453(1.99); C380(0.67) | LDD0755 | [10] |
| LDCM0439 | CL49 | HCT 116 | C453(2.80); C380(0.75) | LDD0756 | [10] |
| LDCM0440 | CL5 | HCT 116 | C453(1.41); C380(0.88) | LDD0757 | [10] |
| LDCM0441 | CL50 | HCT 116 | C453(1.66); C380(1.02) | LDD0758 | [10] |
| LDCM0442 | CL51 | HCT 116 | C453(1.60); C380(1.34) | LDD0759 | [10] |
| LDCM0443 | CL52 | HCT 116 | C453(1.27); C380(1.01) | LDD0760 | [10] |
| LDCM0444 | CL53 | HCT 116 | C453(3.82); C380(0.75) | LDD0761 | [10] |
| LDCM0445 | CL54 | HCT 116 | C453(3.70); C380(0.60) | LDD0762 | [10] |
| LDCM0446 | CL55 | HCT 116 | C453(2.46); C380(0.87) | LDD0763 | [10] |
| LDCM0447 | CL56 | HCT 116 | C453(1.52); C380(1.15) | LDD0764 | [10] |
| LDCM0448 | CL57 | HCT 116 | C453(1.62); C380(0.65) | LDD0765 | [10] |
| LDCM0449 | CL58 | HCT 116 | C453(2.10); C380(1.00) | LDD0766 | [10] |
| LDCM0450 | CL59 | HCT 116 | C453(1.47); C380(0.88) | LDD0767 | [10] |
| LDCM0451 | CL6 | HCT 116 | C453(1.43); C380(1.33) | LDD0768 | [10] |
| LDCM0452 | CL60 | HCT 116 | C453(1.82); C380(0.67) | LDD0769 | [10] |
| LDCM0453 | CL61 | HCT 116 | C453(0.97) | LDD0770 | [10] |
| LDCM0454 | CL62 | HCT 116 | C453(0.95) | LDD0771 | [10] |
| LDCM0455 | CL63 | HCT 116 | C453(1.08) | LDD0772 | [10] |
| LDCM0456 | CL64 | HCT 116 | C453(1.02) | LDD0773 | [10] |
| LDCM0457 | CL65 | HCT 116 | C453(1.10) | LDD0774 | [10] |
| LDCM0458 | CL66 | HCT 116 | C453(0.92) | LDD0775 | [10] |
| LDCM0459 | CL67 | HCT 116 | C453(1.27) | LDD0776 | [10] |
| LDCM0460 | CL68 | HCT 116 | C453(2.16) | LDD0777 | [10] |
| LDCM0461 | CL69 | HCT 116 | C453(1.35) | LDD0778 | [10] |
| LDCM0462 | CL7 | HCT 116 | C453(0.88); C380(0.79) | LDD0779 | [10] |
| LDCM0463 | CL70 | HCT 116 | C453(1.51) | LDD0780 | [10] |
| LDCM0464 | CL71 | HCT 116 | C453(1.40) | LDD0781 | [10] |
| LDCM0465 | CL72 | HCT 116 | C453(1.66) | LDD0782 | [10] |
| LDCM0466 | CL73 | HCT 116 | C453(2.13) | LDD0783 | [10] |
| LDCM0467 | CL74 | HCT 116 | C453(1.85) | LDD0784 | [10] |
| LDCM0469 | CL76 | HCT 116 | C453(0.85) | LDD0786 | [10] |
| LDCM0470 | CL77 | HCT 116 | C453(1.39) | LDD0787 | [10] |
| LDCM0471 | CL78 | HCT 116 | C453(0.85) | LDD0788 | [10] |
| LDCM0472 | CL79 | HCT 116 | C453(1.06) | LDD0789 | [10] |
| LDCM0473 | CL8 | HCT 116 | C453(1.28); C380(0.90) | LDD0790 | [10] |
| LDCM0474 | CL80 | HCT 116 | C453(0.80) | LDD0791 | [10] |
| LDCM0475 | CL81 | HCT 116 | C453(0.85) | LDD0792 | [10] |
| LDCM0476 | CL82 | HCT 116 | C453(0.90) | LDD0793 | [10] |
| LDCM0477 | CL83 | HCT 116 | C453(1.12) | LDD0794 | [10] |
| LDCM0478 | CL84 | HCT 116 | C453(0.97) | LDD0795 | [10] |
| LDCM0479 | CL85 | HCT 116 | C453(1.16) | LDD0796 | [10] |
| LDCM0480 | CL86 | HCT 116 | C453(1.19) | LDD0797 | [10] |
| LDCM0481 | CL87 | HCT 116 | C453(1.19) | LDD0798 | [10] |
| LDCM0482 | CL88 | HCT 116 | C453(1.14) | LDD0799 | [10] |
| LDCM0483 | CL89 | HCT 116 | C453(0.76) | LDD0800 | [10] |
| LDCM0484 | CL9 | HCT 116 | C453(1.35); C380(1.14) | LDD0801 | [10] |
| LDCM0485 | CL90 | HCT 116 | C453(1.38) | LDD0802 | [10] |
| LDCM0486 | CL91 | HCT 116 | C453(1.03) | LDD0803 | [10] |
| LDCM0487 | CL92 | HCT 116 | C453(1.77) | LDD0804 | [10] |
| LDCM0488 | CL93 | HCT 116 | C453(1.10) | LDD0805 | [10] |
| LDCM0489 | CL94 | HCT 116 | C453(0.92) | LDD0806 | [10] |
| LDCM0490 | CL95 | HCT 116 | C453(1.20) | LDD0807 | [10] |
| LDCM0491 | CL96 | HCT 116 | C453(1.04) | LDD0808 | [10] |
| LDCM0492 | CL97 | HCT 116 | C453(1.15) | LDD0809 | [10] |
| LDCM0493 | CL98 | HCT 116 | C453(0.74) | LDD0810 | [10] |
| LDCM0494 | CL99 | HCT 116 | C453(0.74) | LDD0811 | [10] |
| LDCM0495 | E2913 | HEK-293T | C380(0.92); C410(0.99); C69(1.04); C453(1.00) | LDD1698 | [34] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C410(11.90); C268(3.66); C154(1.94) | LDD1702 | [32] |
| LDCM0175 | Ethacrynic acid | HeLa | C453(0.78) | LDD2210 | [9] |
| LDCM0625 | F8 | Ramos | C410(0.37); C154(0.52); C380(0.39); C268(0.53) | LDD2187 | [35] |
| LDCM0572 | Fragment10 | Ramos | C410(0.64); C154(0.61); C380(0.37); C268(0.41) | LDD2189 | [35] |
| LDCM0573 | Fragment11 | Ramos | C410(0.11); C154(0.32); C380(0.05) | LDD2190 | [35] |
| LDCM0574 | Fragment12 | Ramos | C410(0.66); C154(0.52); C380(0.25); C268(0.29) | LDD2191 | [35] |
| LDCM0575 | Fragment13 | Ramos | C410(0.85); C154(0.90); C380(0.69); C268(0.55) | LDD2192 | [35] |
| LDCM0576 | Fragment14 | Ramos | C410(0.23); C154(0.36); C380(0.16); C268(0.28) | LDD2193 | [35] |
| LDCM0579 | Fragment20 | Ramos | C410(0.66); C154(0.68); C380(0.33); C268(0.27) | LDD2194 | [35] |
| LDCM0580 | Fragment21 | Ramos | C410(0.57); C154(1.14); C380(0.74); C268(0.32) | LDD2195 | [35] |
| LDCM0582 | Fragment23 | Ramos | C410(1.02); C154(1.30); C380(0.55); C268(0.62) | LDD2196 | [35] |
| LDCM0578 | Fragment27 | Ramos | C410(0.81); C154(1.30); C380(0.93); C268(0.65) | LDD2197 | [35] |
| LDCM0586 | Fragment28 | Ramos | C410(0.75); C154(0.67); C380(0.47); C268(1.07) | LDD2198 | [35] |
| LDCM0588 | Fragment30 | Ramos | C410(0.93); C154(0.83); C380(0.81); C268(0.45) | LDD2199 | [35] |
| LDCM0589 | Fragment31 | Ramos | C410(0.44); C154(0.63); C380(0.40); C268(0.32) | LDD2200 | [35] |
| LDCM0590 | Fragment32 | Ramos | C410(0.59); C154(0.58); C380(0.28); C268(0.43) | LDD2201 | [35] |
| LDCM0468 | Fragment33 | HCT 116 | C453(0.97) | LDD0785 | [10] |
| LDCM0596 | Fragment38 | Ramos | C410(0.49); C154(0.53); C380(0.50); C268(0.40) | LDD2203 | [35] |
| LDCM0566 | Fragment4 | Ramos | C410(0.77); C154(0.90); C380(0.59); C268(0.66) | LDD2184 | [35] |
| LDCM0427 | Fragment51 | HCT 116 | C453(1.77); C268(0.28); C380(2.06) | LDD0744 | [10] |
| LDCM0610 | Fragment52 | Ramos | C410(1.00); C154(0.53); C380(0.76); C268(0.63) | LDD2204 | [35] |
| LDCM0614 | Fragment56 | Ramos | C410(1.13); C154(0.78); C380(0.77); C268(0.62) | LDD2205 | [35] |
| LDCM0569 | Fragment7 | Ramos | C410(0.47); C154(0.66); C380(0.26); C268(0.61) | LDD2186 | [35] |
| LDCM0571 | Fragment9 | Ramos | C410(0.61); C154(0.58); C380(0.28); C268(0.35) | LDD2188 | [35] |
| LDCM0107 | IAA | HeLa | C154(0.00); C410(0.00); H402(0.00); C380(0.00) | LDD0221 | [24] |
| LDCM0179 | JZ128 | PC-3 | C453(0.00); C410(0.00) | LDD0462 | [8] |
| LDCM0022 | KB02 | HCT 116 | C453(1.58) | LDD0080 | [10] |
| LDCM0023 | KB03 | HCT 116 | C453(1.02) | LDD0081 | [10] |
| LDCM0024 | KB05 | HCT 116 | C453(1.24) | LDD0082 | [10] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C410(0.92) | LDD2102 | [32] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C380(0.87) | LDD2121 | [32] |
| LDCM0109 | NEM | HeLa | C410(0.00); C453(0.00); H402(0.00); H223(0.00) | LDD0223 | [24] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C380(0.86); C154(1.02) | LDD2089 | [32] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C410(0.99) | LDD2092 | [32] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C410(1.21) | LDD2093 | [32] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C410(0.90) | LDD2096 | [32] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C410(1.08) | LDD2097 | [32] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C410(1.13) | LDD2098 | [32] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C380(1.07); C410(1.08) | LDD2099 | [32] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C380(0.73); C410(1.19) | LDD2100 | [32] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C380(0.91); C410(1.00) | LDD2101 | [32] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C410(0.79) | LDD2104 | [32] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C410(1.19) | LDD2105 | [32] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C380(1.17) | LDD2107 | [32] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C380(0.90) | LDD2108 | [32] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C410(1.20) | LDD2109 | [32] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C380(1.43); C410(1.24) | LDD2111 | [32] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C380(0.78); C410(0.64) | LDD2114 | [32] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C410(0.49) | LDD2118 | [32] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C380(1.73); C410(2.25) | LDD2119 | [32] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C410(0.67) | LDD2120 | [32] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C380(0.35) | LDD2122 | [32] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C410(1.31) | LDD2123 | [32] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C380(0.51); C410(0.48) | LDD2124 | [32] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C380(0.96); C410(1.04) | LDD2125 | [32] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C410(0.51) | LDD2126 | [32] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C410(0.99) | LDD2127 | [32] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C410(0.74) | LDD2128 | [32] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C380(1.31); C410(1.47) | LDD2129 | [32] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C380(0.39); C410(0.49) | LDD2133 | [32] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C410(0.50) | LDD2134 | [32] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C380(1.00); C410(1.34) | LDD2135 | [32] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C380(1.19); C410(1.40) | LDD2136 | [32] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C380(0.96); C410(1.05) | LDD2137 | [32] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C410(1.01) | LDD1700 | [32] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C380(0.95); C410(0.98) | LDD2140 | [32] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C380(0.81); C410(0.90) | LDD2141 | [32] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C410(0.80) | LDD2143 | [32] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C410(1.30) | LDD2144 | [32] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C410(1.13) | LDD2147 | [32] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C410(0.51) | LDD2148 | [32] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C380(0.42) | LDD2150 | [32] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C453(0.88) | LDD2206 | [36] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C410(1.26); C380(0.57); C453(0.49); C453(0.32) | LDD2207 | [36] |
| LDCM0131 | RA190 | MM1.R | C268(1.44); C453(1.40); C154(1.34); C380(1.17) | LDD0304 | [37] |
| LDCM0021 | THZ1 | HCT 116 | C380(1.05); C453(0.96); C268(1.00) | LDD2173 | [10] |
| LDCM0111 | W14 | Hep-G2 | W401(0.69) | LDD0238 | [3] |
| LDCM0112 | W16 | Hep-G2 | C453(0.56) | LDD0239 | [3] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Beclin-1 (BECN1) | Beclin family | Q14457 | |||
| Phosphatidylinositol transfer protein alpha isoform (PITPNA) | PtdIns transfer protein family | Q00169 | |||
Transcription factor
Other
The Drug(s) Related To This Target
Investigative
References
