Details of the Target
General Information of Target
| Target ID | LDTP13449 | |||||
|---|---|---|---|---|---|---|
| Target Name | Cleavage and polyadenylation specificity factor subunit 3 (CPSF3) | |||||
| Gene Name | CPSF3 | |||||
| Gene ID | 51692 | |||||
| Synonyms |
CPSF73; Cleavage and polyadenylation specificity factor subunit 3; EC 3.1.27.-; Cleavage and polyadenylation specificity factor 73 kDa subunit; CPSF 73 kDa subunit; mRNA 3'-end-processing endonuclease CPSF-73
|
|||||
| 3D Structure | ||||||
| Sequence |
MKMLLLLHCLGVFLSCSGHIQDEHPQYHSPPDVVIPVRITGTTRGMTPPGWLSYILPFGG
QKHIIHIKVKKLLFSKHLPVFTYTDQGAILEDQPFVQNNCYYHGYVEGDPESLVSLSTCF GGFQGILQINDFAYEIKPLAFSTTFEHLVYKMDSEEKQFSTMRSGFMQNEITCRMEFEEI DNSTQKQSSYVGWWIHFRIVEIVVVIDNYLYIRYERNDSKLLEDLYVIVNIVDSILDVIG VKVLLFGLEIWTNKNLIVVDDVRKSVHLYCKWKSENITPRMQHDTSHLFTTLGLRGLSGI GAFRGMCTPHRSCAIVTFMNKTLGTFSIAVAHHLGHNLGMNHDEDTCRCSQPRCIMHEGN PPITKFSNCSYGDFWEYTVERTKCLLETVHTKDIFNVKRCGNGVVEEGEECDCGPLKHCA KDPCCLSNCTLTDGSTCAFGLCCKDCKFLPSGKVCRKEVNECDLPEWCNGTSHKCPDDFY VEDGIPCKERGYCYEKSCHDRNEQCRRIFGAGANTASETCYKELNTLGDRVGHCGIKNAT YIKCNISDVQCGRIQCENVTEIPNMSDHTTVHWARFNDIMCWSTDYHLGMKGPDIGEVKD GTECGIDHICIHRHCVHITILNSNCSPAFCNKRGICNNKHHCHCNYLWDPPNCLIKGYGG SVDSGPPPKRKKKKKFCYLCILLLIVLFILLCCLYRLCKKSKPIKKQQDVQTPSAKEEEK IQRRPHELPPQSQPWVMPSQSQPPVTPSQSHPQVMPSQSQPPVTPSQSQPRVMPSQSQPP VMPSQSHPQLTPSQSQPPVTPSQRQPQLMPSQSQPPVTPS |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Metallo-beta-lactamase superfamily, RNA-metabolizing metallo-beta-lactamase-like family, CPSF3 subfamily
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Component of the cleavage and polyadenylation specificity factor (CPSF) complex that plays a key role in pre-mRNA 3'-end formation, recognizing the AAUAAA signal sequence and interacting with poly(A) polymerase and other factors to bring about cleavage and poly(A) addition. Has endonuclease activity, and functions as an mRNA 3'-end-processing endonuclease. Also involved in the histone 3'-end pre-mRNA processing. U7 snRNP-dependent protein that induces both the 3'-endoribonucleolytic cleavage of histone pre-mRNAs and acts as a 5' to 3' exonuclease for degrading the subsequent downstream cleavage product (DCP) of mature histone mRNAs. Cleavage occurs after the 5'-ACCCA-3' sequence in the histone pre-mRNA leaving a 3'hydroxyl group on the upstream fragment containing the stem loop (SL) and 5' phosphate on the downstream cleavage product (DCP) starting with CU nucleotides. The U7-dependent 5' to 3' exonuclease activity is processive and degrades the DCP RNA substrate even after complete removal of the U7-binding site. Binds to the downstream cleavage product (DCP) of histone pre-mRNAs and the cleaved DCP RNA substrate in a U7 snRNP dependent manner. Required for entering/progressing through S-phase of the cell cycle. Required for the selective processing of microRNAs (miRNAs) during embryonic stem cell differentiation via its interaction with ISY1. Required for the biogenesis of all miRNAs from the pri-miR-17-92 primary transcript except miR-92a. Only required for the biogenesis of miR-290 and miR-96 from the pri-miR-290-295 and pri-miR-96-183 primary transcripts, respectively.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K295(10.00); K612(9.09) | LDD0277 | [2] | |
|
P11 Probe Info |
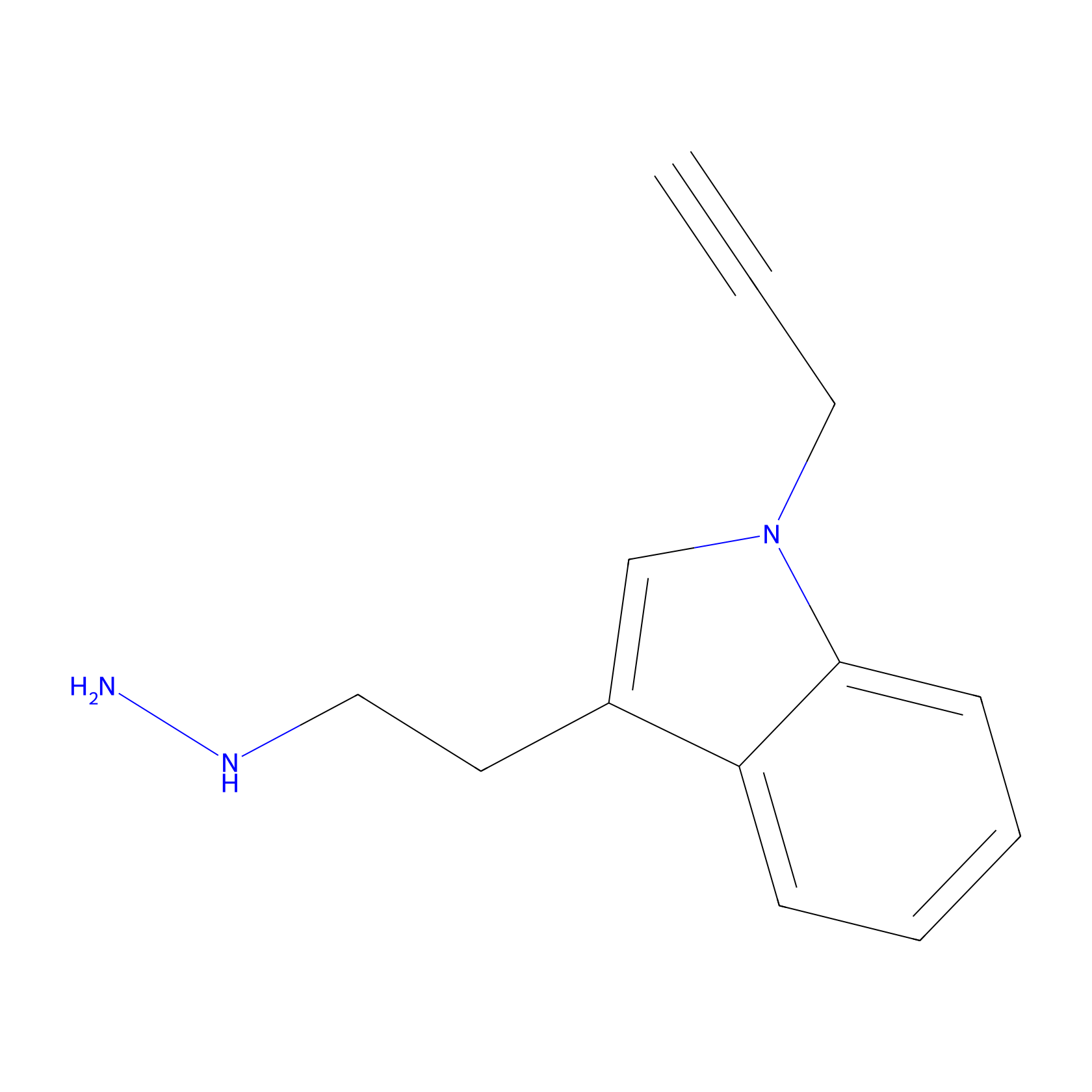 |
11.55 | LDD0201 | [3] | |
|
BTD Probe Info |
 |
C223(1.06) | LDD2089 | [4] | |
|
Sulforaphane-probe2 Probe Info |
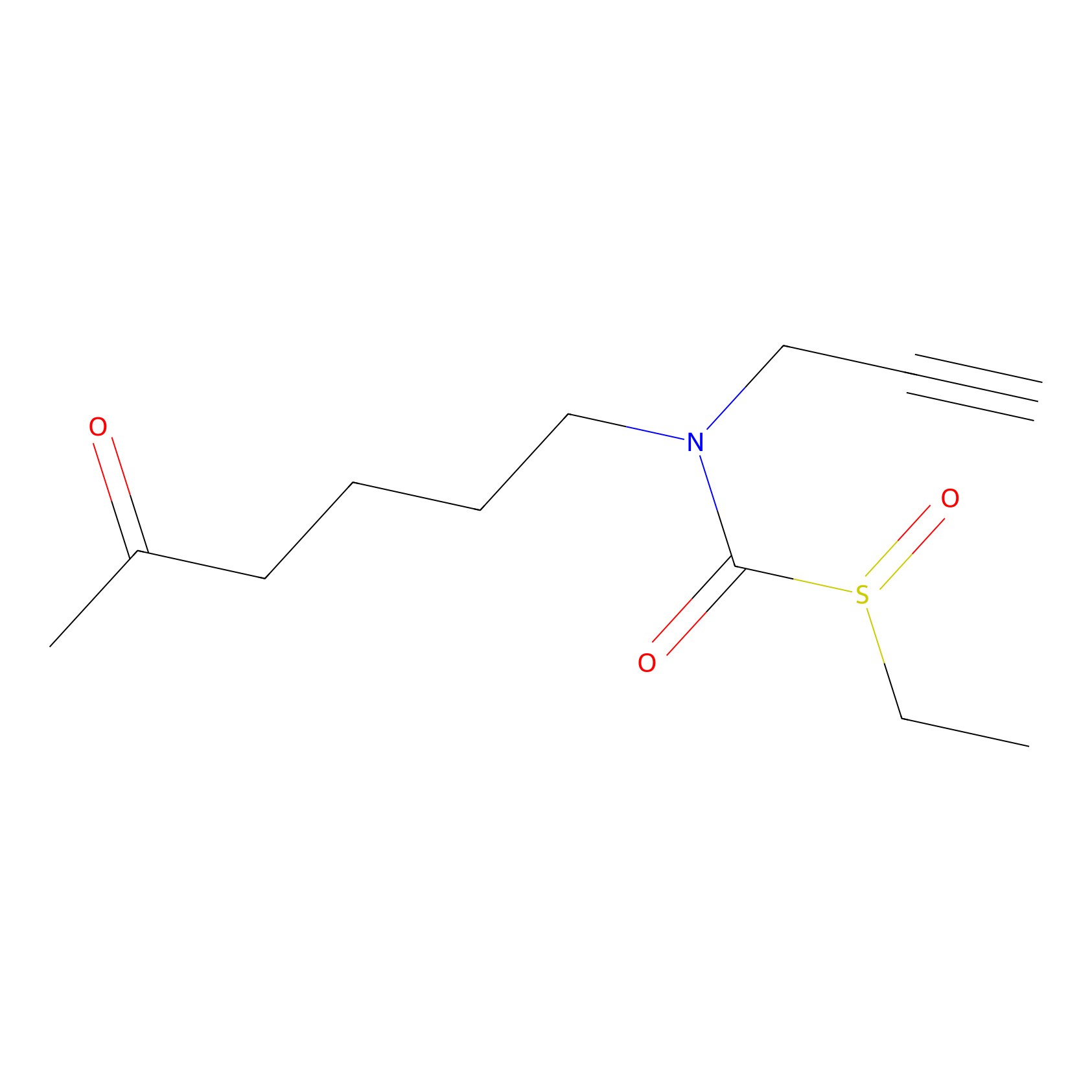 |
1.38 | LDD0160 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C655(4.20) | LDD0170 | [6] | |
|
HHS-475 Probe Info |
 |
Y528(0.76); Y519(1.10) | LDD0264 | [7] | |
|
HHS-465 Probe Info |
 |
Y528(5.95) | LDD2237 | [8] | |
|
DBIA Probe Info |
 |
C498(3.63); C527(1.13); C278(1.16) | LDD0080 | [9] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C345(0.00); C223(0.00); C498(0.00); C527(0.00) | LDD0038 | [10] | |
|
IA-alkyne Probe Info |
 |
C278(0.00); C345(0.00); C223(0.00); C498(0.00) | LDD0036 | [10] | |
|
IPIAA_L Probe Info |
 |
C655(0.00); C345(0.00); C27(0.00); C498(0.00) | LDD0031 | [11] | |
|
Lodoacetamide azide Probe Info |
 |
C345(0.00); C223(0.00); C278(0.00); C498(0.00) | LDD0037 | [10] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [12] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [13] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [13] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [14] | |
|
Phosphinate-6 Probe Info |
 |
C498(0.00); C655(0.00) | LDD0018 | [15] | |
|
Acrolein Probe Info |
 |
C655(0.00); C223(0.00); C345(0.00); H307(0.00) | LDD0217 | [16] | |
|
Crotonaldehyde Probe Info |
 |
C655(0.00); C223(0.00) | LDD0219 | [16] | |
|
Methacrolein Probe Info |
 |
C655(0.00); C223(0.00) | LDD0218 | [16] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [17] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [18] | |
|
NAIA_5 Probe Info |
 |
C345(0.00); C223(0.00); C358(0.00) | LDD2223 | [12] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
5.13 | LDD0471 | [19] | |
|
FFF probe2 Probe Info |
 |
13.63 | LDD0463 | [19] | |
|
FFF probe3 Probe Info |
 |
7.93 | LDD0465 | [19] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C223(1.09) | LDD2112 | [4] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C223(0.87) | LDD2130 | [4] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C223(1.21) | LDD2117 | [4] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C223(0.91) | LDD2132 | [4] |
| LDCM0025 | 4SU-RNA | DM93 | C655(4.20) | LDD0170 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C655(2.94) | LDD0171 | [6] |
| LDCM0214 | AC1 | HCT 116 | C223(1.33); C278(0.98); C345(0.77) | LDD0531 | [9] |
| LDCM0215 | AC10 | HCT 116 | C223(1.26); C278(0.85); C345(0.83); C527(0.99) | LDD0532 | [9] |
| LDCM0216 | AC100 | HCT 116 | C223(1.00); C278(0.84); C345(0.68); C527(0.75) | LDD0533 | [9] |
| LDCM0217 | AC101 | HCT 116 | C223(1.06); C278(0.90); C345(0.60); C527(0.79) | LDD0534 | [9] |
| LDCM0218 | AC102 | HCT 116 | C223(1.00); C278(0.90); C345(0.61); C527(0.75) | LDD0535 | [9] |
| LDCM0219 | AC103 | HCT 116 | C223(0.92); C278(1.01); C345(0.56); C527(0.92) | LDD0536 | [9] |
| LDCM0220 | AC104 | HCT 116 | C223(1.13); C278(0.86); C345(0.76); C527(0.96) | LDD0537 | [9] |
| LDCM0221 | AC105 | HCT 116 | C223(1.02); C278(0.86); C345(0.59); C527(0.71) | LDD0538 | [9] |
| LDCM0222 | AC106 | HCT 116 | C223(1.02); C278(0.92); C345(0.62); C527(0.60) | LDD0539 | [9] |
| LDCM0223 | AC107 | HCT 116 | C223(0.98); C278(0.82); C345(0.68); C527(0.92) | LDD0540 | [9] |
| LDCM0224 | AC108 | HCT 116 | C223(1.13); C278(0.86); C345(0.68); C527(0.70) | LDD0541 | [9] |
| LDCM0225 | AC109 | HCT 116 | C223(1.11); C278(0.81); C345(0.97); C527(0.73) | LDD0542 | [9] |
| LDCM0226 | AC11 | HCT 116 | C223(1.36); C278(0.97); C345(0.60); C527(1.07) | LDD0543 | [9] |
| LDCM0227 | AC110 | HCT 116 | C223(0.96); C278(0.92); C345(0.58); C527(0.78) | LDD0544 | [9] |
| LDCM0228 | AC111 | HCT 116 | C223(0.87); C278(1.14); C345(0.54); C527(0.81) | LDD0545 | [9] |
| LDCM0229 | AC112 | HCT 116 | C223(0.89); C278(0.92); C345(0.61); C527(0.79) | LDD0546 | [9] |
| LDCM0230 | AC113 | HCT 116 | C223(1.07); C278(0.93); C345(1.00) | LDD0547 | [9] |
| LDCM0231 | AC114 | HCT 116 | C223(1.06); C278(0.99); C345(0.98) | LDD0548 | [9] |
| LDCM0232 | AC115 | HCT 116 | C223(1.03); C278(0.95); C345(1.01) | LDD0549 | [9] |
| LDCM0233 | AC116 | HCT 116 | C223(0.89); C278(1.11); C345(2.12) | LDD0550 | [9] |
| LDCM0234 | AC117 | HCT 116 | C223(0.97); C278(0.98); C345(1.16) | LDD0551 | [9] |
| LDCM0235 | AC118 | HCT 116 | C223(1.21); C278(0.93); C345(0.92) | LDD0552 | [9] |
| LDCM0236 | AC119 | HCT 116 | C223(1.03); C278(0.84); C345(1.15) | LDD0553 | [9] |
| LDCM0237 | AC12 | HCT 116 | C223(1.53); C278(0.83); C345(1.03); C527(1.21) | LDD0554 | [9] |
| LDCM0238 | AC120 | HCT 116 | C223(1.04); C278(0.94); C345(0.89) | LDD0555 | [9] |
| LDCM0239 | AC121 | HCT 116 | C223(1.07); C278(0.93); C345(1.04) | LDD0556 | [9] |
| LDCM0240 | AC122 | HCT 116 | C223(1.10); C278(0.93); C345(0.97) | LDD0557 | [9] |
| LDCM0241 | AC123 | HCT 116 | C223(1.19); C278(1.01); C345(1.07) | LDD0558 | [9] |
| LDCM0242 | AC124 | HCT 116 | C223(0.93); C278(0.89); C345(1.03) | LDD0559 | [9] |
| LDCM0243 | AC125 | HCT 116 | C223(0.94); C278(1.13); C345(0.95) | LDD0560 | [9] |
| LDCM0244 | AC126 | HCT 116 | C223(1.09); C278(1.02); C345(0.93) | LDD0561 | [9] |
| LDCM0245 | AC127 | HCT 116 | C223(1.00); C278(1.02); C345(0.94) | LDD0562 | [9] |
| LDCM0246 | AC128 | HCT 116 | C223(1.05); C278(1.38); C527(0.92) | LDD0563 | [9] |
| LDCM0247 | AC129 | HCT 116 | C223(1.18); C278(0.92); C527(0.99) | LDD0564 | [9] |
| LDCM0249 | AC130 | HCT 116 | C223(1.18); C278(1.05); C527(0.70) | LDD0566 | [9] |
| LDCM0250 | AC131 | HCT 116 | C223(1.02); C278(1.08); C527(1.25) | LDD0567 | [9] |
| LDCM0251 | AC132 | HCT 116 | C223(1.25); C278(0.99); C527(0.72) | LDD0568 | [9] |
| LDCM0252 | AC133 | HCT 116 | C223(1.11); C278(1.08); C527(0.93) | LDD0569 | [9] |
| LDCM0253 | AC134 | HCT 116 | C223(0.95); C278(1.17); C527(0.79) | LDD0570 | [9] |
| LDCM0254 | AC135 | HCT 116 | C223(1.14); C278(1.34); C527(0.78) | LDD0571 | [9] |
| LDCM0255 | AC136 | HCT 116 | C223(1.13); C278(0.98); C527(0.85) | LDD0572 | [9] |
| LDCM0256 | AC137 | HCT 116 | C223(1.20); C278(0.96); C527(0.63) | LDD0573 | [9] |
| LDCM0257 | AC138 | HCT 116 | C223(1.01); C278(1.01); C527(0.78) | LDD0574 | [9] |
| LDCM0258 | AC139 | HCT 116 | C223(1.20); C278(1.01); C527(0.73) | LDD0575 | [9] |
| LDCM0259 | AC14 | HCT 116 | C223(1.03); C278(1.06); C345(0.71); C527(1.05) | LDD0576 | [9] |
| LDCM0260 | AC140 | HCT 116 | C223(1.12); C278(1.09); C527(0.61) | LDD0577 | [9] |
| LDCM0261 | AC141 | HCT 116 | C223(1.11); C278(1.03); C527(0.83) | LDD0578 | [9] |
| LDCM0262 | AC142 | HCT 116 | C223(1.10); C278(0.91); C527(0.80) | LDD0579 | [9] |
| LDCM0263 | AC143 | HCT 116 | C223(0.94); C278(0.97); C527(0.99) | LDD0580 | [9] |
| LDCM0264 | AC144 | HCT 116 | C527(0.88); C278(1.20); C223(1.43) | LDD0581 | [9] |
| LDCM0265 | AC145 | HCT 116 | C527(0.82); C278(1.08); C223(1.48) | LDD0582 | [9] |
| LDCM0266 | AC146 | HCT 116 | C527(0.69); C278(1.07); C223(1.11) | LDD0583 | [9] |
| LDCM0267 | AC147 | HCT 116 | C527(0.88); C278(1.21); C223(1.34) | LDD0584 | [9] |
| LDCM0268 | AC148 | HCT 116 | C527(0.84); C278(1.28); C223(1.54) | LDD0585 | [9] |
| LDCM0269 | AC149 | HCT 116 | C527(0.68); C223(1.07); C278(1.10) | LDD0586 | [9] |
| LDCM0270 | AC15 | HCT 116 | C223(0.89); C527(1.09); C278(1.11); C345(1.13) | LDD0587 | [9] |
| LDCM0271 | AC150 | HCT 116 | C527(0.81); C278(0.96); C223(1.12) | LDD0588 | [9] |
| LDCM0272 | AC151 | HCT 116 | C527(0.81); C223(0.90); C278(1.06) | LDD0589 | [9] |
| LDCM0273 | AC152 | HCT 116 | C527(0.90); C223(0.91); C278(1.24) | LDD0590 | [9] |
| LDCM0274 | AC153 | HCT 116 | C527(0.87); C278(1.47); C223(1.71) | LDD0591 | [9] |
| LDCM0621 | AC154 | HCT 116 | C223(1.09); C278(1.21); C527(0.98) | LDD2158 | [9] |
| LDCM0622 | AC155 | HCT 116 | C223(1.03); C278(1.18); C527(0.81) | LDD2159 | [9] |
| LDCM0623 | AC156 | HCT 116 | C223(0.82); C278(1.01); C527(0.91) | LDD2160 | [9] |
| LDCM0624 | AC157 | HCT 116 | C223(1.07); C278(1.02); C527(1.24) | LDD2161 | [9] |
| LDCM0276 | AC17 | HCT 116 | C655(0.77); C278(0.98); C223(1.12); C527(1.34) | LDD0593 | [9] |
| LDCM0277 | AC18 | HCT 116 | C223(1.01); C655(1.05); C278(1.19); C527(1.57) | LDD0594 | [9] |
| LDCM0278 | AC19 | HCT 116 | C655(0.76); C223(1.00); C527(1.13); C278(1.14) | LDD0595 | [9] |
| LDCM0279 | AC2 | HCT 116 | C345(0.60); C278(0.91); C223(1.04) | LDD0596 | [9] |
| LDCM0280 | AC20 | HCT 116 | C655(0.71); C223(0.92); C278(1.00); C527(1.09) | LDD0597 | [9] |
| LDCM0281 | AC21 | HCT 116 | C655(0.73); C278(1.00); C223(1.03); C527(1.29) | LDD0598 | [9] |
| LDCM0282 | AC22 | HCT 116 | C655(0.65); C223(0.96); C278(1.01); C527(1.40) | LDD0599 | [9] |
| LDCM0283 | AC23 | HCT 116 | C655(0.71); C223(1.00); C278(1.04); C527(1.16) | LDD0600 | [9] |
| LDCM0284 | AC24 | HCT 116 | C655(0.75); C223(0.98); C278(1.03); C527(1.24) | LDD0601 | [9] |
| LDCM0285 | AC25 | HCT 116 | C527(0.82); C278(0.94); C223(0.97); C345(1.18) | LDD0602 | [9] |
| LDCM0286 | AC26 | HCT 116 | C527(0.77); C345(0.80); C278(0.95); C223(1.01) | LDD0603 | [9] |
| LDCM0287 | AC27 | HCT 116 | C345(0.77); C223(0.93); C527(0.94); C278(1.13) | LDD0604 | [9] |
| LDCM0288 | AC28 | HCT 116 | C345(0.84); C527(0.87); C223(1.03); C278(1.07) | LDD0605 | [9] |
| LDCM0289 | AC29 | HCT 116 | C345(0.61); C223(0.87); C527(0.91); C278(1.17) | LDD0606 | [9] |
| LDCM0290 | AC3 | HCT 116 | C345(0.75); C278(0.83); C223(1.34) | LDD0607 | [9] |
| LDCM0291 | AC30 | HCT 116 | C345(0.64); C223(0.87); C527(0.95); C278(1.18) | LDD0608 | [9] |
| LDCM0292 | AC31 | HCT 116 | C345(0.66); C527(0.80); C278(1.10); C223(1.17) | LDD0609 | [9] |
| LDCM0293 | AC32 | HCT 116 | C345(0.60); C527(0.81); C223(1.06); C278(1.11) | LDD0610 | [9] |
| LDCM0294 | AC33 | HCT 116 | C345(0.70); C223(0.90); C527(0.90); C278(1.20) | LDD0611 | [9] |
| LDCM0295 | AC34 | HCT 116 | C345(0.65); C223(0.73); C527(0.96); C278(1.42) | LDD0612 | [9] |
| LDCM0296 | AC35 | HCT 116 | C278(0.88); C223(0.95); C655(1.03); C345(1.09) | LDD0613 | [9] |
| LDCM0297 | AC36 | HCT 116 | C345(0.78); C278(0.98); C223(0.99); C527(1.00) | LDD0614 | [9] |
| LDCM0298 | AC37 | HCT 116 | C278(0.92); C527(0.99); C223(1.03); C345(1.07) | LDD0615 | [9] |
| LDCM0299 | AC38 | HCT 116 | C655(0.87); C527(0.91); C278(1.00); C223(1.05) | LDD0616 | [9] |
| LDCM0300 | AC39 | HCT 116 | C527(0.66); C278(0.86); C655(0.89); C223(0.91) | LDD0617 | [9] |
| LDCM0301 | AC4 | HCT 116 | C345(0.85); C278(0.94); C223(1.63) | LDD0618 | [9] |
| LDCM0302 | AC40 | HCT 116 | C223(0.86); C527(0.92); C655(1.07); C278(1.07) | LDD0619 | [9] |
| LDCM0303 | AC41 | HCT 116 | C527(0.73); C223(0.78); C345(0.91); C655(0.96) | LDD0620 | [9] |
| LDCM0304 | AC42 | HCT 116 | C527(0.94); C345(0.94); C655(0.95); C223(0.96) | LDD0621 | [9] |
| LDCM0305 | AC43 | HCT 116 | C223(0.82); C527(0.89); C278(0.93); C345(1.03) | LDD0622 | [9] |
| LDCM0306 | AC44 | HCT 116 | C527(0.89); C278(1.02); C223(1.03); C655(1.04) | LDD0623 | [9] |
| LDCM0307 | AC45 | HCT 116 | C527(0.83); C223(0.88); C345(0.97); C655(1.11) | LDD0624 | [9] |
| LDCM0308 | AC46 | HCT 116 | C527(0.73); C223(0.97); C345(1.06); C278(1.08) | LDD0625 | [9] |
| LDCM0309 | AC47 | HCT 116 | C527(0.67); C345(0.93); C223(0.98); C278(1.01) | LDD0626 | [9] |
| LDCM0310 | AC48 | HCT 116 | C527(0.44); C278(0.87); C223(0.89); C345(0.96) | LDD0627 | [9] |
| LDCM0311 | AC49 | HCT 116 | C527(0.80); C223(0.95); C345(0.95); C278(1.09) | LDD0628 | [9] |
| LDCM0312 | AC5 | HCT 116 | C345(0.82); C278(0.94); C223(1.52) | LDD0629 | [9] |
| LDCM0313 | AC50 | HCT 116 | C527(0.66); C345(0.89); C223(0.92); C278(1.11) | LDD0630 | [9] |
| LDCM0314 | AC51 | HCT 116 | C527(0.64); C223(0.88); C345(0.91); C278(1.02) | LDD0631 | [9] |
| LDCM0315 | AC52 | HCT 116 | C527(0.84); C278(0.85); C345(0.92); C223(0.93) | LDD0632 | [9] |
| LDCM0316 | AC53 | HCT 116 | C345(0.71); C527(0.90); C223(0.90); C278(1.13) | LDD0633 | [9] |
| LDCM0317 | AC54 | HCT 116 | C527(0.85); C223(0.86); C345(0.91); C278(1.07) | LDD0634 | [9] |
| LDCM0318 | AC55 | HCT 116 | C345(0.82); C527(0.85); C278(1.06); C223(1.10) | LDD0635 | [9] |
| LDCM0319 | AC56 | HCT 116 | C345(0.71); C223(0.91); C278(0.91); C527(0.94) | LDD0636 | [9] |
| LDCM0320 | AC57 | HCT 116 | C527(0.50); C345(0.82); C223(1.06); C278(1.16) | LDD0637 | [9] |
| LDCM0321 | AC58 | HCT 116 | C345(0.70); C527(0.83); C223(0.88); C278(1.50) | LDD0638 | [9] |
| LDCM0322 | AC59 | HCT 116 | C527(0.50); C345(0.86); C223(1.03); C278(1.34) | LDD0639 | [9] |
| LDCM0323 | AC6 | HCT 116 | C345(0.76); C223(0.92); C278(0.98); C655(1.01) | LDD0640 | [9] |
| LDCM0324 | AC60 | HCT 116 | C345(0.77); C527(0.83); C223(0.93); C278(1.19) | LDD0641 | [9] |
| LDCM0325 | AC61 | HCT 116 | C527(0.40); C345(0.76); C223(0.89); C278(1.13) | LDD0642 | [9] |
| LDCM0326 | AC62 | HCT 116 | C527(0.77); C345(0.78); C223(0.98); C278(1.35) | LDD0643 | [9] |
| LDCM0327 | AC63 | HCT 116 | C345(0.77); C527(0.87); C223(1.05); C278(1.24) | LDD0644 | [9] |
| LDCM0328 | AC64 | HCT 116 | C527(0.53); C345(0.83); C223(1.00); C278(1.31) | LDD0645 | [9] |
| LDCM0329 | AC65 | HCT 116 | C527(0.51); C223(1.00); C345(1.07); C278(1.26) | LDD0646 | [9] |
| LDCM0330 | AC66 | HCT 116 | C527(0.49); C223(0.98); C345(1.07); C278(1.39) | LDD0647 | [9] |
| LDCM0331 | AC67 | HCT 116 | C527(0.42); C345(0.67); C223(0.77); C278(1.32) | LDD0648 | [9] |
| LDCM0332 | AC68 | HCT 116 | C527(0.87); C223(0.98); C278(1.07) | LDD0649 | [9] |
| LDCM0333 | AC69 | HCT 116 | C527(0.86); C223(1.06); C278(1.09) | LDD0650 | [9] |
| LDCM0334 | AC7 | HCT 116 | C655(0.85); C345(0.91); C278(0.93); C223(0.96) | LDD0651 | [9] |
| LDCM0335 | AC70 | HCT 116 | C527(0.73); C278(0.97); C223(1.05) | LDD0652 | [9] |
| LDCM0336 | AC71 | HCT 116 | C527(0.87); C278(0.98); C223(1.01) | LDD0653 | [9] |
| LDCM0337 | AC72 | HCT 116 | C527(0.98); C278(1.02); C223(1.07) | LDD0654 | [9] |
| LDCM0338 | AC73 | HCT 116 | C223(0.97); C527(0.98); C278(1.22) | LDD0655 | [9] |
| LDCM0339 | AC74 | HCT 116 | C527(0.95); C278(1.26); C223(1.34) | LDD0656 | [9] |
| LDCM0340 | AC75 | HCT 116 | C527(0.98); C223(1.02); C278(1.32) | LDD0657 | [9] |
| LDCM0341 | AC76 | HCT 116 | C527(0.89); C223(0.98); C278(1.11) | LDD0658 | [9] |
| LDCM0342 | AC77 | HCT 116 | C527(0.77); C223(0.91); C278(1.09) | LDD0659 | [9] |
| LDCM0343 | AC78 | HCT 116 | C527(0.82); C278(0.89); C223(1.08) | LDD0660 | [9] |
| LDCM0344 | AC79 | HCT 116 | C527(0.86); C278(0.97); C223(1.06) | LDD0661 | [9] |
| LDCM0345 | AC8 | HCT 116 | C345(0.80); C278(0.97); C223(1.00); C527(1.20) | LDD0662 | [9] |
| LDCM0346 | AC80 | HCT 116 | C278(1.03); C223(1.05); C527(1.05) | LDD0663 | [9] |
| LDCM0347 | AC81 | HCT 116 | C527(0.56); C278(0.90); C223(1.23) | LDD0664 | [9] |
| LDCM0348 | AC82 | HCT 116 | C527(0.71); C278(1.04); C223(1.10) | LDD0665 | [9] |
| LDCM0349 | AC83 | HCT 116 | C345(0.87); C223(1.09); C278(1.25) | LDD0666 | [9] |
| LDCM0350 | AC84 | HCT 116 | C345(0.95); C223(1.05); C278(1.22) | LDD0667 | [9] |
| LDCM0351 | AC85 | HCT 116 | C278(0.99); C345(1.11); C223(1.13) | LDD0668 | [9] |
| LDCM0352 | AC86 | HCT 116 | C345(0.92); C278(1.04); C223(1.18) | LDD0669 | [9] |
| LDCM0353 | AC87 | HCT 116 | C345(0.86); C278(0.88); C223(1.04) | LDD0670 | [9] |
| LDCM0354 | AC88 | HCT 116 | C345(0.84); C278(0.96); C223(1.22) | LDD0671 | [9] |
| LDCM0355 | AC89 | HCT 116 | C223(1.10); C345(1.15); C278(1.19) | LDD0672 | [9] |
| LDCM0357 | AC90 | HCT 116 | C278(0.90); C345(0.92); C223(1.23) | LDD0674 | [9] |
| LDCM0358 | AC91 | HCT 116 | C223(1.05); C278(1.33); C345(1.34) | LDD0675 | [9] |
| LDCM0359 | AC92 | HCT 116 | C223(1.00); C278(1.10); C345(1.25) | LDD0676 | [9] |
| LDCM0360 | AC93 | HCT 116 | C278(0.91); C345(0.97); C223(1.04) | LDD0677 | [9] |
| LDCM0361 | AC94 | HCT 116 | C278(1.07); C223(1.08); C345(1.52) | LDD0678 | [9] |
| LDCM0362 | AC95 | HCT 116 | C278(0.90); C345(0.97); C223(1.10) | LDD0679 | [9] |
| LDCM0363 | AC96 | HCT 116 | C278(1.02); C223(1.09); C345(1.47) | LDD0680 | [9] |
| LDCM0364 | AC97 | HCT 116 | C345(0.84); C223(1.00); C278(1.04) | LDD0681 | [9] |
| LDCM0365 | AC98 | HCT 116 | C345(0.59); C527(0.63); C278(0.89); C223(0.95) | LDD0682 | [9] |
| LDCM0366 | AC99 | HCT 116 | C345(0.60); C527(0.70); C278(0.84); C223(1.10) | LDD0683 | [9] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C223(0.82) | LDD2113 | [4] |
| LDCM0248 | AKOS034007472 | HCT 116 | C223(1.15); C278(0.86); C345(0.91); C527(1.18) | LDD0565 | [9] |
| LDCM0356 | AKOS034007680 | HCT 116 | C345(0.71); C527(0.95); C278(0.95); C655(1.02) | LDD0673 | [9] |
| LDCM0275 | AKOS034007705 | HCT 116 | C223(0.89); C345(0.99); C527(1.13); C278(1.18) | LDD0592 | [9] |
| LDCM0156 | Aniline | NCI-H1299 | 11.44 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C498(1.09) | LDD2171 | [9] |
| LDCM0106 | BDHI 18 | Jurkat | C27(18.80) | LDD0208 | [20] |
| LDCM0102 | BDHI 8 | Jurkat | C27(18.80) | LDD0204 | [20] |
| LDCM0088 | C45 | HEK-293T | 11.55 | LDD0201 | [3] |
| LDCM0108 | Chloroacetamide | HeLa | C655(0.00); C498(0.00); C223(0.00); C345(0.00) | LDD0222 | [16] |
| LDCM0632 | CL-Sc | Hep-G2 | C498(2.72); C527(1.09) | LDD2227 | [12] |
| LDCM0367 | CL1 | HCT 116 | C223(0.74); C278(0.85); C527(1.04) | LDD0684 | [9] |
| LDCM0368 | CL10 | HCT 116 | C527(0.74); C223(0.91); C278(0.93) | LDD0685 | [9] |
| LDCM0369 | CL100 | HCT 116 | C345(0.82); C223(1.05); C278(1.06) | LDD0686 | [9] |
| LDCM0370 | CL101 | HCT 116 | C345(0.82); C527(0.96); C223(1.01); C278(1.13) | LDD0687 | [9] |
| LDCM0371 | CL102 | HCT 116 | C345(0.74); C655(0.87); C278(0.94); C223(1.03) | LDD0688 | [9] |
| LDCM0372 | CL103 | HCT 116 | C345(0.84); C278(0.88); C527(0.97); C655(1.47) | LDD0689 | [9] |
| LDCM0373 | CL104 | HCT 116 | C655(0.83); C345(0.85); C278(0.88); C223(1.04) | LDD0690 | [9] |
| LDCM0374 | CL105 | HCT 116 | C655(0.83); C223(1.02); C527(1.02); C278(1.14) | LDD0691 | [9] |
| LDCM0375 | CL106 | HCT 116 | C655(0.95); C223(1.21); C527(1.23); C278(1.27) | LDD0692 | [9] |
| LDCM0376 | CL107 | HCT 116 | C655(0.93); C278(1.05); C223(1.09); C527(1.11) | LDD0693 | [9] |
| LDCM0377 | CL108 | HCT 116 | C655(1.05); C223(1.23); C527(1.23); C278(1.38) | LDD0694 | [9] |
| LDCM0378 | CL109 | HCT 116 | C223(0.88); C655(1.02); C278(1.13); C527(1.38) | LDD0695 | [9] |
| LDCM0379 | CL11 | HCT 116 | C527(0.86); C278(0.95); C223(0.96) | LDD0696 | [9] |
| LDCM0380 | CL110 | HCT 116 | C655(0.95); C527(1.06); C223(1.10); C278(1.14) | LDD0697 | [9] |
| LDCM0381 | CL111 | HCT 116 | C655(0.81); C223(0.89); C278(1.20); C527(1.31) | LDD0698 | [9] |
| LDCM0382 | CL112 | HCT 116 | C527(0.79); C345(0.81); C278(1.02); C223(1.16) | LDD0699 | [9] |
| LDCM0383 | CL113 | HCT 116 | C345(0.50); C527(0.90); C223(0.95); C278(1.31) | LDD0700 | [9] |
| LDCM0384 | CL114 | HCT 116 | C345(0.42); C223(0.81); C527(0.82); C278(1.22) | LDD0701 | [9] |
| LDCM0385 | CL115 | HCT 116 | C345(0.57); C527(0.74); C223(0.88); C278(1.08) | LDD0702 | [9] |
| LDCM0386 | CL116 | HCT 116 | C345(0.71); C527(0.77); C223(0.93); C278(1.04) | LDD0703 | [9] |
| LDCM0387 | CL117 | HCT 116 | C223(0.79); C527(0.93); C345(0.98); C655(1.20) | LDD0704 | [9] |
| LDCM0388 | CL118 | HCT 116 | C527(0.87); C223(0.94); C345(0.96); C278(0.97) | LDD0705 | [9] |
| LDCM0389 | CL119 | HCT 116 | C223(0.85); C278(0.97); C527(1.03); C345(1.06) | LDD0706 | [9] |
| LDCM0390 | CL12 | HCT 116 | C223(1.00); C527(1.00); C278(1.04) | LDD0707 | [9] |
| LDCM0391 | CL120 | HCT 116 | C527(0.74); C278(0.89); C223(0.93); C345(1.11) | LDD0708 | [9] |
| LDCM0392 | CL121 | HCT 116 | C345(0.67); C527(0.84); C223(0.86); C278(0.96) | LDD0709 | [9] |
| LDCM0393 | CL122 | HCT 116 | C527(0.69); C345(0.75); C223(0.80); C278(0.86) | LDD0710 | [9] |
| LDCM0394 | CL123 | HCT 116 | C527(0.75); C345(0.80); C223(0.86); C278(0.89) | LDD0711 | [9] |
| LDCM0395 | CL124 | HCT 116 | C527(0.81); C345(0.81); C223(0.85); C278(0.96) | LDD0712 | [9] |
| LDCM0396 | CL125 | HCT 116 | C527(0.78); C278(0.98); C223(1.03); C345(1.19) | LDD0713 | [9] |
| LDCM0397 | CL126 | HCT 116 | C223(0.89); C527(1.06); C345(1.06); C278(1.09) | LDD0714 | [9] |
| LDCM0398 | CL127 | HCT 116 | C527(0.51); C345(0.92); C223(0.99); C278(1.18) | LDD0715 | [9] |
| LDCM0399 | CL128 | HCT 116 | C527(0.49); C345(0.89); C223(0.98); C278(1.30) | LDD0716 | [9] |
| LDCM0400 | CL13 | HCT 116 | C527(0.91); C278(1.03); C223(1.06) | LDD0717 | [9] |
| LDCM0401 | CL14 | HCT 116 | C527(0.84); C278(0.94); C223(1.07) | LDD0718 | [9] |
| LDCM0402 | CL15 | HCT 116 | C527(0.75); C278(1.07); C223(1.16) | LDD0719 | [9] |
| LDCM0403 | CL16 | HCT 116 | C223(0.78); C527(1.05); C278(1.15) | LDD0720 | [9] |
| LDCM0404 | CL17 | HCT 116 | C223(0.76); C527(0.93); C278(1.15) | LDD0721 | [9] |
| LDCM0405 | CL18 | HCT 116 | C223(0.87); C527(1.24); C278(1.43) | LDD0722 | [9] |
| LDCM0406 | CL19 | HCT 116 | C223(0.84); C527(1.10); C278(1.28) | LDD0723 | [9] |
| LDCM0407 | CL2 | HCT 116 | C278(0.80); C223(0.83); C527(0.92) | LDD0724 | [9] |
| LDCM0408 | CL20 | HCT 116 | C223(0.90); C527(1.14); C278(1.29) | LDD0725 | [9] |
| LDCM0409 | CL21 | HCT 116 | C223(0.87); C527(1.26); C278(1.44) | LDD0726 | [9] |
| LDCM0410 | CL22 | HCT 116 | C223(1.03); C527(1.22); C278(1.81) | LDD0727 | [9] |
| LDCM0411 | CL23 | HCT 116 | C223(0.88); C527(1.17); C278(1.36) | LDD0728 | [9] |
| LDCM0412 | CL24 | HCT 116 | C223(0.82); C527(1.17); C278(1.37) | LDD0729 | [9] |
| LDCM0413 | CL25 | HCT 116 | C223(0.92); C278(1.41); C527(1.02) | LDD0730 | [9] |
| LDCM0414 | CL26 | HCT 116 | C223(0.80); C278(1.22); C527(1.17) | LDD0731 | [9] |
| LDCM0415 | CL27 | HCT 116 | C223(1.04); C278(1.24); C527(0.78) | LDD0732 | [9] |
| LDCM0416 | CL28 | HCT 116 | C223(0.87); C278(1.18); C527(0.96) | LDD0733 | [9] |
| LDCM0417 | CL29 | HCT 116 | C223(0.99); C278(1.32); C527(1.06) | LDD0734 | [9] |
| LDCM0418 | CL3 | HCT 116 | C223(1.14); C278(0.86); C527(0.92) | LDD0735 | [9] |
| LDCM0419 | CL30 | HCT 116 | C223(0.84); C278(1.28); C527(1.11) | LDD0736 | [9] |
| LDCM0420 | CL31 | HCT 116 | C223(0.91); C278(1.17) | LDD0737 | [9] |
| LDCM0421 | CL32 | HCT 116 | C223(0.78); C278(1.30) | LDD0738 | [9] |
| LDCM0422 | CL33 | HCT 116 | C223(0.80); C278(1.23) | LDD0739 | [9] |
| LDCM0423 | CL34 | HCT 116 | C223(0.83); C278(1.38) | LDD0740 | [9] |
| LDCM0424 | CL35 | HCT 116 | C223(0.89); C278(1.55) | LDD0741 | [9] |
| LDCM0425 | CL36 | HCT 116 | C223(0.91); C278(1.44) | LDD0742 | [9] |
| LDCM0426 | CL37 | HCT 116 | C223(0.94); C278(1.78) | LDD0743 | [9] |
| LDCM0428 | CL39 | HCT 116 | C223(0.91); C278(1.46) | LDD0745 | [9] |
| LDCM0429 | CL4 | HCT 116 | C223(0.91); C278(1.00); C527(0.91) | LDD0746 | [9] |
| LDCM0430 | CL40 | HCT 116 | C223(0.81); C278(1.47) | LDD0747 | [9] |
| LDCM0431 | CL41 | HCT 116 | C223(0.88); C278(1.43) | LDD0748 | [9] |
| LDCM0432 | CL42 | HCT 116 | C223(0.78); C278(1.78) | LDD0749 | [9] |
| LDCM0433 | CL43 | HCT 116 | C223(0.81); C278(1.63) | LDD0750 | [9] |
| LDCM0434 | CL44 | HCT 116 | C223(0.78); C278(1.42) | LDD0751 | [9] |
| LDCM0435 | CL45 | HCT 116 | C223(0.80); C278(1.54) | LDD0752 | [9] |
| LDCM0436 | CL46 | HCT 116 | C223(1.10); C278(0.88); C345(0.85) | LDD0753 | [9] |
| LDCM0437 | CL47 | HCT 116 | C223(0.96); C278(0.86); C345(0.73) | LDD0754 | [9] |
| LDCM0438 | CL48 | HCT 116 | C223(0.98); C278(0.95); C345(1.04) | LDD0755 | [9] |
| LDCM0439 | CL49 | HCT 116 | C223(1.11); C278(0.94); C345(1.11) | LDD0756 | [9] |
| LDCM0440 | CL5 | HCT 116 | C223(1.06); C278(0.81); C527(0.84) | LDD0757 | [9] |
| LDCM0441 | CL50 | HCT 116 | C223(1.11); C278(0.92); C345(1.04) | LDD0758 | [9] |
| LDCM0442 | CL51 | HCT 116 | C223(1.16); C278(0.90); C345(1.00) | LDD0759 | [9] |
| LDCM0443 | CL52 | HCT 116 | C223(1.10); C278(1.07); C345(0.77) | LDD0760 | [9] |
| LDCM0444 | CL53 | HCT 116 | C223(1.00); C278(1.02); C345(0.61) | LDD0761 | [9] |
| LDCM0445 | CL54 | HCT 116 | C223(0.91); C278(0.95); C345(0.77) | LDD0762 | [9] |
| LDCM0446 | CL55 | HCT 116 | C223(0.91); C278(0.91); C345(0.90) | LDD0763 | [9] |
| LDCM0447 | CL56 | HCT 116 | C223(0.94); C278(1.01); C345(0.71) | LDD0764 | [9] |
| LDCM0448 | CL57 | HCT 116 | C223(1.05); C278(0.96); C345(0.73) | LDD0765 | [9] |
| LDCM0449 | CL58 | HCT 116 | C223(1.03); C278(0.97); C345(0.91) | LDD0766 | [9] |
| LDCM0450 | CL59 | HCT 116 | C223(1.00); C278(0.93); C345(0.94) | LDD0767 | [9] |
| LDCM0451 | CL6 | HCT 116 | C223(0.95); C278(0.92); C527(0.69) | LDD0768 | [9] |
| LDCM0452 | CL60 | HCT 116 | C223(0.96); C278(0.96); C345(0.95) | LDD0769 | [9] |
| LDCM0453 | CL61 | HCT 116 | C223(1.06); C278(1.08); C527(0.99); C655(1.04) | LDD0770 | [9] |
| LDCM0454 | CL62 | HCT 116 | C223(1.04); C278(1.08); C527(0.81); C655(0.87) | LDD0771 | [9] |
| LDCM0455 | CL63 | HCT 116 | C223(1.06); C278(1.10); C527(0.74); C655(1.27) | LDD0772 | [9] |
| LDCM0456 | CL64 | HCT 116 | C223(1.06); C278(1.23); C527(0.80); C655(1.03) | LDD0773 | [9] |
| LDCM0457 | CL65 | HCT 116 | C223(1.05); C278(1.05); C527(0.76); C655(0.87) | LDD0774 | [9] |
| LDCM0458 | CL66 | HCT 116 | C223(0.92); C278(1.23); C527(0.75); C655(1.35) | LDD0775 | [9] |
| LDCM0459 | CL67 | HCT 116 | C223(1.22); C278(1.32); C527(1.00); C655(0.98) | LDD0776 | [9] |
| LDCM0460 | CL68 | HCT 116 | C223(0.81); C278(1.50); C527(0.76); C655(1.47) | LDD0777 | [9] |
| LDCM0461 | CL69 | HCT 116 | C223(0.82); C278(1.27); C527(0.86); C655(0.91) | LDD0778 | [9] |
| LDCM0462 | CL7 | HCT 116 | C223(0.92); C278(0.85); C527(0.92) | LDD0779 | [9] |
| LDCM0463 | CL70 | HCT 116 | C223(0.86); C278(1.19); C527(0.82); C655(1.26) | LDD0780 | [9] |
| LDCM0464 | CL71 | HCT 116 | C223(0.89); C278(1.13); C527(0.81); C655(0.90) | LDD0781 | [9] |
| LDCM0465 | CL72 | HCT 116 | C223(0.84); C278(1.10); C527(0.88); C655(0.92) | LDD0782 | [9] |
| LDCM0466 | CL73 | HCT 116 | C223(0.94); C278(1.96); C527(1.15); C655(1.10) | LDD0783 | [9] |
| LDCM0467 | CL74 | HCT 116 | C223(0.96); C278(1.53); C527(1.09); C655(1.22) | LDD0784 | [9] |
| LDCM0469 | CL76 | HCT 116 | C223(1.09); C278(1.02) | LDD0786 | [9] |
| LDCM0470 | CL77 | HCT 116 | C223(1.10); C278(1.64) | LDD0787 | [9] |
| LDCM0471 | CL78 | HCT 116 | C223(1.11); C278(1.04) | LDD0788 | [9] |
| LDCM0472 | CL79 | HCT 116 | C223(1.12); C278(1.30) | LDD0789 | [9] |
| LDCM0473 | CL8 | HCT 116 | C223(1.11); C278(1.11); C527(0.94) | LDD0790 | [9] |
| LDCM0474 | CL80 | HCT 116 | C223(1.26); C278(1.14) | LDD0791 | [9] |
| LDCM0475 | CL81 | HCT 116 | C223(1.52); C278(1.01) | LDD0792 | [9] |
| LDCM0476 | CL82 | HCT 116 | C223(1.50); C278(1.35) | LDD0793 | [9] |
| LDCM0477 | CL83 | HCT 116 | C223(1.26); C278(1.17) | LDD0794 | [9] |
| LDCM0478 | CL84 | HCT 116 | C223(0.93); C278(1.34) | LDD0795 | [9] |
| LDCM0479 | CL85 | HCT 116 | C223(1.10); C278(1.02) | LDD0796 | [9] |
| LDCM0480 | CL86 | HCT 116 | C223(1.15); C278(0.96) | LDD0797 | [9] |
| LDCM0481 | CL87 | HCT 116 | C223(1.46); C278(1.09) | LDD0798 | [9] |
| LDCM0482 | CL88 | HCT 116 | C223(1.58); C278(1.24) | LDD0799 | [9] |
| LDCM0483 | CL89 | HCT 116 | C223(1.36); C278(1.26) | LDD0800 | [9] |
| LDCM0484 | CL9 | HCT 116 | C223(0.76); C278(0.89); C527(1.04) | LDD0801 | [9] |
| LDCM0485 | CL90 | HCT 116 | C223(1.11); C278(0.94) | LDD0802 | [9] |
| LDCM0486 | CL91 | HCT 116 | C223(1.11); C278(0.88); C345(0.69) | LDD0803 | [9] |
| LDCM0487 | CL92 | HCT 116 | C223(1.24); C278(1.08); C345(1.04) | LDD0804 | [9] |
| LDCM0488 | CL93 | HCT 116 | C223(1.56); C278(1.24); C345(0.91) | LDD0805 | [9] |
| LDCM0489 | CL94 | HCT 116 | C223(1.25); C278(1.08); C345(0.88) | LDD0806 | [9] |
| LDCM0490 | CL95 | HCT 116 | C223(1.19); C278(1.07); C345(0.77) | LDD0807 | [9] |
| LDCM0491 | CL96 | HCT 116 | C223(1.13); C278(0.91); C345(0.67) | LDD0808 | [9] |
| LDCM0492 | CL97 | HCT 116 | C223(1.16); C278(1.01); C345(0.66) | LDD0809 | [9] |
| LDCM0493 | CL98 | HCT 116 | C223(1.23); C278(0.90); C345(0.68) | LDD0810 | [9] |
| LDCM0494 | CL99 | HCT 116 | C223(1.21); C278(0.83); C345(0.76) | LDD0811 | [9] |
| LDCM0495 | E2913 | HEK-293T | C655(0.97); C223(0.95); C278(0.98); C358(1.10) | LDD1698 | [21] |
| LDCM0625 | F8 | Ramos | C655(0.99); C278(1.63); C498(0.76) | LDD2187 | [22] |
| LDCM0572 | Fragment10 | MDA-MB-231 | C498(4.50) | LDD1389 | [23] |
| LDCM0573 | Fragment11 | MDA-MB-231 | C498(2.08) | LDD1391 | [23] |
| LDCM0574 | Fragment12 | MDA-MB-231 | C498(2.08) | LDD1393 | [23] |
| LDCM0575 | Fragment13 | MDA-MB-231 | C498(0.94) | LDD1395 | [23] |
| LDCM0576 | Fragment14 | MDA-MB-231 | C498(1.97) | LDD1397 | [23] |
| LDCM0577 | Fragment15 | MDA-MB-231 | C498(0.79) | LDD1399 | [23] |
| LDCM0579 | Fragment20 | Ramos | C498(2.51) | LDD1403 | [23] |
| LDCM0580 | Fragment21 | MDA-MB-231 | C498(1.40) | LDD1404 | [23] |
| LDCM0582 | Fragment23 | MDA-MB-231 | C498(1.18) | LDD1408 | [23] |
| LDCM0583 | Fragment24 | Ramos | C498(1.42) | LDD1410 | [23] |
| LDCM0584 | Fragment25 | MDA-MB-231 | C498(1.06) | LDD1411 | [23] |
| LDCM0585 | Fragment26 | Ramos | C498(1.03) | LDD1412 | [23] |
| LDCM0578 | Fragment27 | MDA-MB-231 | C498(1.33) | LDD1401 | [23] |
| LDCM0586 | Fragment28 | MDA-MB-231 | C498(1.06) | LDD1415 | [23] |
| LDCM0587 | Fragment29 | MDA-MB-231 | C498(1.82) | LDD1417 | [23] |
| LDCM0588 | Fragment30 | MDA-MB-231 | C498(0.90) | LDD1419 | [23] |
| LDCM0589 | Fragment31 | Ramos | C498(1.77) | LDD1422 | [23] |
| LDCM0590 | Fragment32 | MDA-MB-231 | C498(3.76) | LDD1423 | [23] |
| LDCM0468 | Fragment33 | HCT 116 | C223(0.90); C278(1.28); C527(0.76); C655(0.86) | LDD0785 | [9] |
| LDCM0593 | Fragment35 | MDA-MB-231 | C498(1.25) | LDD1429 | [23] |
| LDCM0594 | Fragment36 | MDA-MB-231 | C498(2.07) | LDD1431 | [23] |
| LDCM0595 | Fragment37 | Ramos | C498(0.92) | LDD1432 | [23] |
| LDCM0596 | Fragment38 | MDA-MB-231 | C498(1.36) | LDD1433 | [23] |
| LDCM0566 | Fragment4 | MDA-MB-231 | C498(2.00) | LDD1378 | [23] |
| LDCM0599 | Fragment41 | MDA-MB-231 | C498(1.09) | LDD1438 | [23] |
| LDCM0600 | Fragment42 | Ramos | C498(0.96) | LDD1440 | [23] |
| LDCM0601 | Fragment43 | MDA-MB-231 | C498(13.88) | LDD1441 | [23] |
| LDCM0603 | Fragment45 | MDA-MB-231 | C498(20.00) | LDD1482 | [23] |
| LDCM0604 | Fragment46 | MDA-MB-231 | C498(0.90) | LDD1445 | [23] |
| LDCM0607 | Fragment49 | MDA-MB-231 | C498(3.55) | LDD1448 | [23] |
| LDCM0608 | Fragment50 | MDA-MB-231 | C498(2.09) | LDD1449 | [23] |
| LDCM0427 | Fragment51 | HCT 116 | C223(0.79); C278(1.81) | LDD0744 | [9] |
| LDCM0610 | Fragment52 | MDA-MB-231 | C498(1.10) | LDD1452 | [23] |
| LDCM0611 | Fragment53 | MDA-MB-231 | C498(1.17) | LDD1454 | [23] |
| LDCM0614 | Fragment56 | MDA-MB-231 | C498(1.17) | LDD1458 | [23] |
| LDCM0568 | Fragment6 | MDA-MB-231 | C498(1.53) | LDD1382 | [23] |
| LDCM0569 | Fragment7 | MDA-MB-231 | C498(2.06) | LDD1383 | [23] |
| LDCM0570 | Fragment8 | MDA-MB-231 | C498(12.73) | LDD1385 | [23] |
| LDCM0571 | Fragment9 | MDA-MB-231 | C498(1.27) | LDD1387 | [23] |
| LDCM0116 | HHS-0101 | DM93 | Y528(0.76); Y519(1.10) | LDD0264 | [7] |
| LDCM0117 | HHS-0201 | DM93 | Y528(0.59); Y519(1.56) | LDD0265 | [7] |
| LDCM0118 | HHS-0301 | DM93 | Y528(0.63); Y519(1.99) | LDD0266 | [7] |
| LDCM0119 | HHS-0401 | DM93 | Y528(0.67); Y519(1.31) | LDD0267 | [7] |
| LDCM0120 | HHS-0701 | DM93 | Y528(0.63); Y519(1.85) | LDD0268 | [7] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [16] |
| LDCM0022 | KB02 | HCT 116 | C498(3.63); C527(1.13); C278(1.16) | LDD0080 | [9] |
| LDCM0023 | KB03 | HCT 116 | C498(1.82); C527(1.08); C278(0.99) | LDD0081 | [9] |
| LDCM0024 | KB05 | HCT 116 | C498(2.64); C527(1.66); C278(1.46) | LDD0082 | [9] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [16] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C223(1.06) | LDD2089 | [4] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C223(1.40) | LDD2099 | [4] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C223(0.65) | LDD2100 | [4] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C223(1.14) | LDD2108 | [4] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C223(0.70) | LDD2114 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C223(1.38) | LDD2123 | [4] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C223(0.89) | LDD2134 | [4] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C223(1.71) | LDD2135 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C223(1.05) | LDD2137 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C223(1.27) | LDD2140 | [4] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C223(0.66) | LDD2141 | [4] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C223(2.39) | LDD2144 | [4] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C223(1.20) | LDD2145 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C223(1.31) | LDD2146 | [4] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C223(0.99) | LDD2148 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C223(0.88) | LDD2150 | [4] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C498(10.26) | LDD2206 | [24] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C498(8.52); C498(0.90) | LDD2207 | [24] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 1.38 | LDD0160 | [5] |
| LDCM0021 | THZ1 | HCT 116 | C498(1.09) | LDD2173 | [9] |
| LDCM0112 | W16 | Hep-G2 | C498(0.63) | LDD0239 | [17] |
The Interaction Atlas With This Target
References
