Details of the Target
General Information of Target
| Target ID | LDTP00890 | |||||
|---|---|---|---|---|---|---|
| Target Name | S-adenosylhomocysteine hydrolase-like protein 1 (AHCYL1) | |||||
| Gene Name | AHCYL1 | |||||
| Gene ID | 10768 | |||||
| Synonyms |
DCAL; IRBIT; XPVKONA; S-adenosylhomocysteine hydrolase-like protein 1; DC-expressed AHCY-like molecule; IP(3)Rs binding protein released with IP(3); IRBIT; Putative adenosylhomocysteinase 2; S-adenosyl-L-homocysteine hydrolase 2; AdoHcyase 2
|
|||||
| 3D Structure | ||||||
| Sequence |
MSMPDAMPLPGVGEELKQAKEIEDAEKYSFMATVTKAPKKQIQFADDMQEFTKFPTKTGR
RSLSRSISQSSTDSYSSAASYTDSSDDEVSPREKQQTNSKGSSNFCVKNIKQAEFGRREI EIAEQDMSALISLRKRAQGEKPLAGAKIVGCTHITAQTAVLIETLCALGAQCRWSACNIY STQNEVAAALAEAGVAVFAWKGESEDDFWWCIDRCVNMDGWQANMILDDGGDLTHWVYKK YPNVFKKIRGIVEESVTGVHRLYQLSKAGKLCVPAMNVNDSVTKQKFDNLYCCRESILDG LKRTTDVMFGGKQVVVCGYGEVGKGCCAALKALGAIVYITEIDPICALQACMDGFRVVKL NEVIRQVDVVITCTGNKNVVTREHLDRMKNSCIVCNMGHSNTEIDVTSLRTPELTWERVR SQVDHVIWPDGKRVVLLAEGRLLNLSCSTVPTFVLSITATTQALALIELYNAPEGRYKQD VYLLPKKMDEYVASLHLPSFDAHLTELTDDQAKYLGLNKNGPFKPNYYRY |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Adenosylhomocysteinase family
|
|||||
| Subcellular location |
Endoplasmic reticulum
|
|||||
| Function |
Multifaceted cellular regulator which coordinates several essential cellular functions including regulation of epithelial HCO3(-) and fluid secretion, mRNA processing and DNA replication. Regulates ITPR1 sensitivity to inositol 1,4,5-trisphosphate, competing for the common binding site and acting as endogenous 'pseudoligand' whose inhibitory activity can be modulated by its phosphorylation status. Promotes the formation of contact points between the endoplasmic reticulum (ER) and mitochondria, facilitating transfer of Ca(2+) from the ER to mitochondria. Under normal cellular conditions, functions cooperatively with BCL2L10 to limit ITPR1-mediated Ca(2+) release but, under apoptotic stress conditions, dephosphorylated which promotes dissociation of both AHCYL1 and BCL2L10 from mitochondria-associated endoplasmic reticulum membranes, inhibits BCL2L10 interaction with ITPR1 and leads to increased Ca(2+) transfer to mitochondria which promotes apoptosis. In the pancreatic and salivary ducts, at resting state, attenuates inositol 1,4,5-trisphosphate-induced calcium release by interacting with ITPR1. When extracellular stimuli induce ITPR1 phosphorylation or inositol 1,4,5-trisphosphate production, dissociates from ITPR1 to interact with CFTR and SLC26A6, mediating their synergistic activation by calcium and cAMP that stimulates the epithelial secretion of electrolytes and fluid. Also activates basolateral SLC4A4 isoform 1 to coordinate fluid and HCO3(-) secretion. Inhibits the effect of STK39 on SLC4A4 and CFTR by recruiting PP1 phosphatase which activates SLC4A4, SLC26A6 and CFTR through dephosphorylation. Mediates the induction of SLC9A3 surface expression produced by Angiotensin-2. Depending on the cell type, activates SLC9A3 in response to calcium or reverses SLC9A3R2-dependent calcium inhibition. May modulate the polyadenylation state of specific mRNAs, both by controlling the subcellular location of FIP1L1 and by inhibiting PAPOLA activity, in response to a stimulus that alters its phosphorylation state. Acts as a (dATP)-dependent inhibitor of ribonucleotide reductase large subunit RRM1, controlling the endogenous dNTP pool and ensuring normal cell cycle progression. In vitro does not exhibit any S-adenosyl-L-homocysteine hydrolase activity.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| EFO27 | SNV: p.S400A | DBIA Probe Info | |||
| HCT15 | SNV: p.R410C | DBIA Probe Info | |||
| HEC1 | SNV: p.R410H | DBIA Probe Info | |||
| HEC1B | SNV: p.S74R | . | |||
| HT29 | SNV: p.I393L | DBIA Probe Info | |||
| IM95 | Deletion: p.E140RfsTer9 | DBIA Probe Info | |||
| JURKAT | SNV: p.H235Q | . | |||
| KYSE410 | SNV: p.A329S | DBIA Probe Info | |||
| MV411 | SNV: p.C392Y; p.I393L | DBIA Probe Info | |||
| NCIH1048 | SNV: p.C392Y; p.I393L | DBIA Probe Info | |||
| NCIH1975 | SNV: p.C392Y | . | |||
| SW756 | SNV: p.S181Ter | DBIA Probe Info | |||
| TOV21G | SNV: p.R410S | DBIA Probe Info | |||
| U2OS | SNV: p.Q49Ter | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
TH211 Probe Info |
 |
Y514(20.00); Y491(7.58) | LDD0257 | [2] | |
|
STPyne Probe Info |
 |
K359(9.18) | LDD0277 | [3] | |
|
ONAyne Probe Info |
 |
K57(10.00) | LDD0275 | [3] | |
|
OPA-S-S-alkyne Probe Info |
 |
K302(1.25) | LDD3494 | [4] | |
|
Probe 1 Probe Info |
 |
Y28(16.69); Y241(41.97); Y482(8.85); Y514(25.66) | LDD3495 | [5] | |
|
BTD Probe Info |
 |
C106(1.52) | LDD1700 | [6] | |
|
MCL-14 Probe Info |
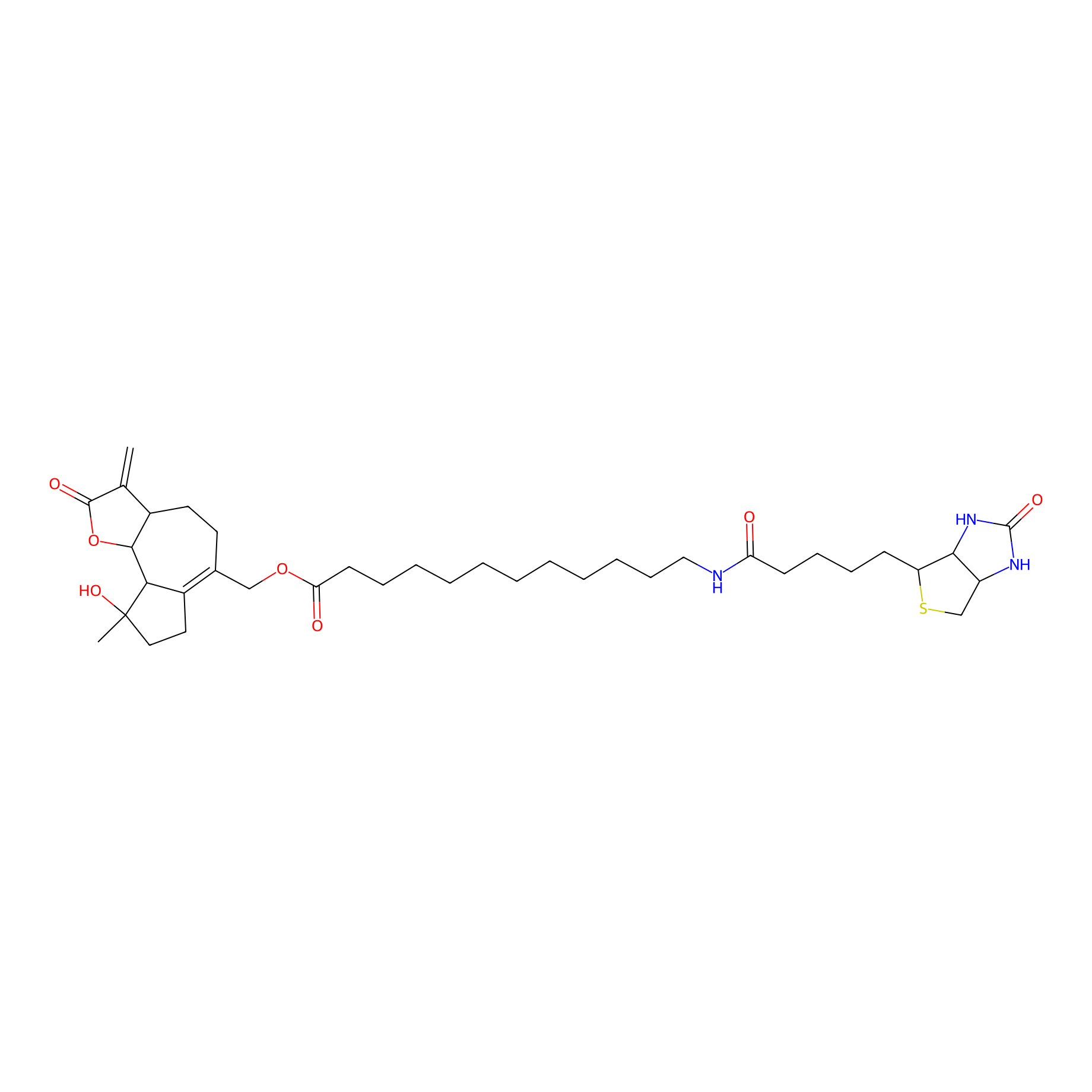 |
2.00 | LDD0050 | [7] | |
|
AHL-Pu-1 Probe Info |
 |
C272(2.12) | LDD0169 | [8] | |
|
HHS-475 Probe Info |
 |
Y491(0.75); Y514(0.90); Y527(0.95) | LDD0264 | [9] | |
|
HHS-465 Probe Info |
 |
Y514(10.00) | LDD2237 | [10] | |
|
DBIA Probe Info |
 |
C292(0.96) | LDD0078 | [11] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [12] | |
|
ATP probe Probe Info |
 |
K284(0.00); K286(0.00); K267(0.00); K270(0.00) | LDD0199 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C373(0.00); C106(0.00) | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [15] | |
|
Lodoacetamide azide Probe Info |
 |
C215(0.00); C211(0.00); C106(0.00); C272(0.00) | LDD0037 | [14] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [16] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [16] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [17] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [17] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [17] | |
|
IPM Probe Info |
 |
C293(0.00); C292(0.00) | LDD0005 | [17] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [17] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [18] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [17] | |
|
Phosphinate-6 Probe Info |
 |
C293(0.00); C272(0.00) | LDD0018 | [19] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [20] | |
|
W1 Probe Info |
 |
C292(0.00); C293(0.00) | LDD0236 | [21] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [22] | |
|
NAIA_5 Probe Info |
 |
C272(0.00); C215(0.00); C106(0.00); C211(0.00) | LDD2223 | [23] | |
|
HHS-482 Probe Info |
 |
Y491(1.04) | LDD2239 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
6.54 | LDD1740 | [24] | |
|
C087 Probe Info |
 |
7.84 | LDD1779 | [24] | |
|
C166 Probe Info |
 |
7.16 | LDD1846 | [24] | |
|
C174 Probe Info |
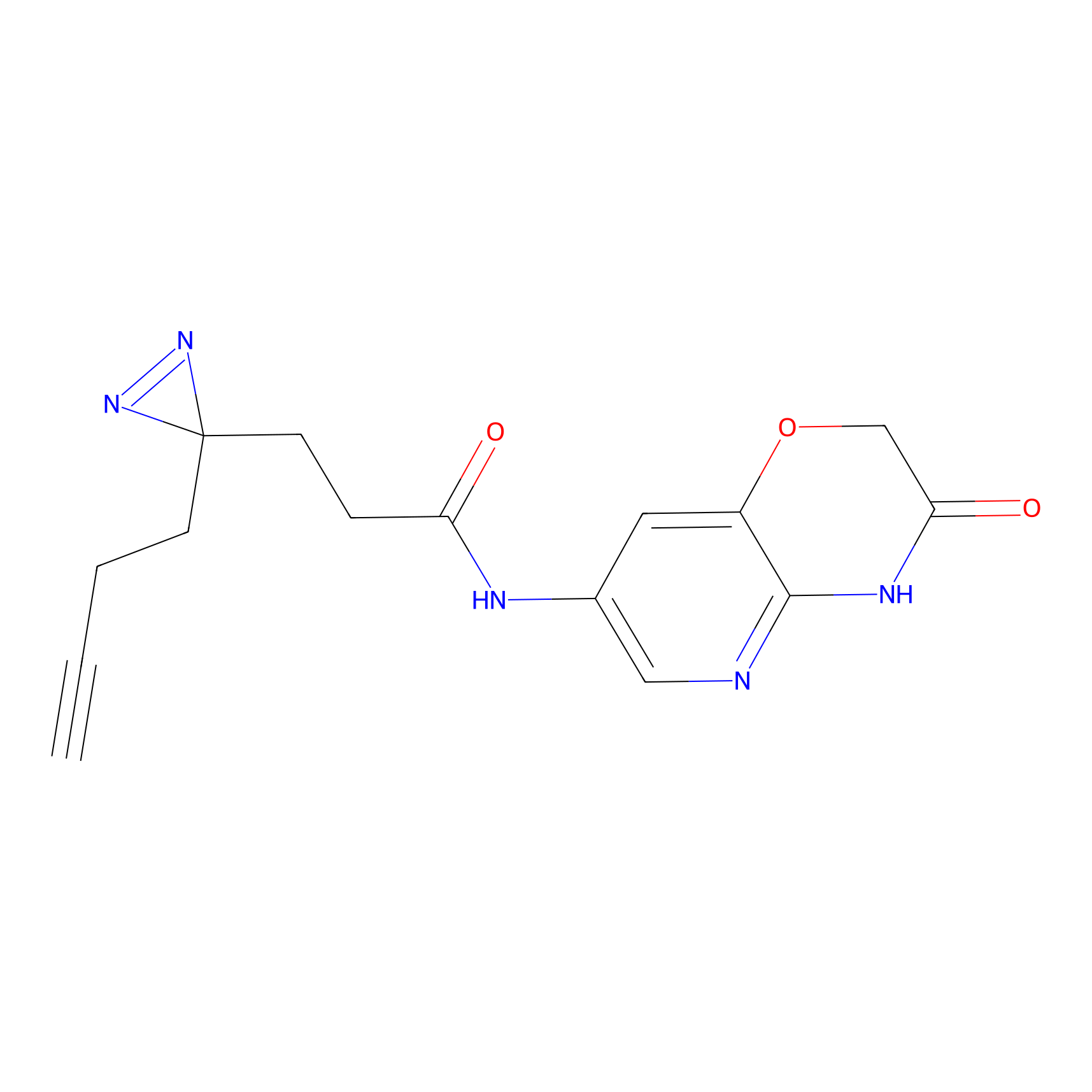 |
5.31 | LDD1854 | [24] | |
|
C187 Probe Info |
 |
15.24 | LDD1865 | [24] | |
|
C210 Probe Info |
 |
49.18 | LDD1884 | [24] | |
|
C216 Probe Info |
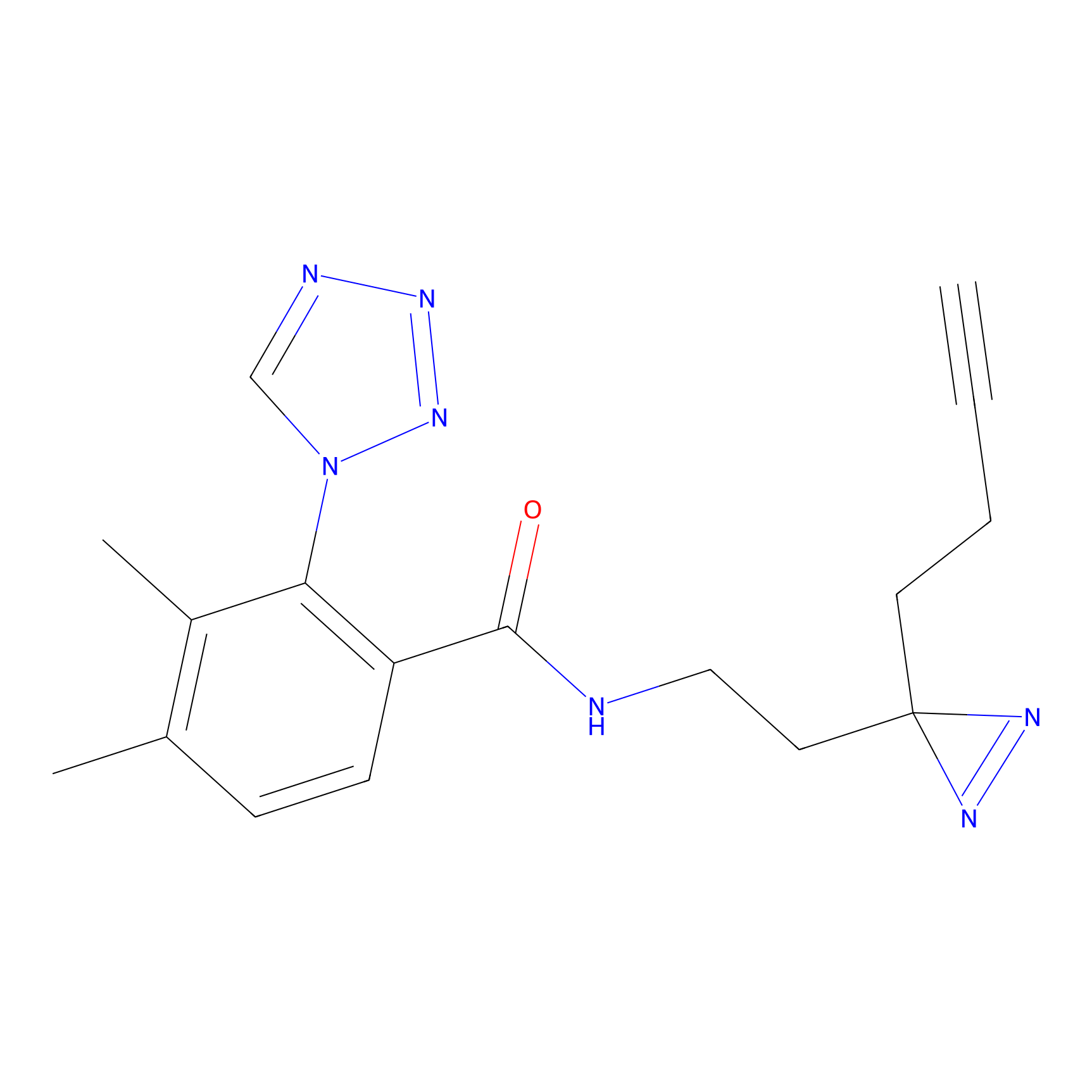 |
5.62 | LDD1890 | [24] | |
|
C218 Probe Info |
 |
11.00 | LDD1892 | [24] | |
|
C258 Probe Info |
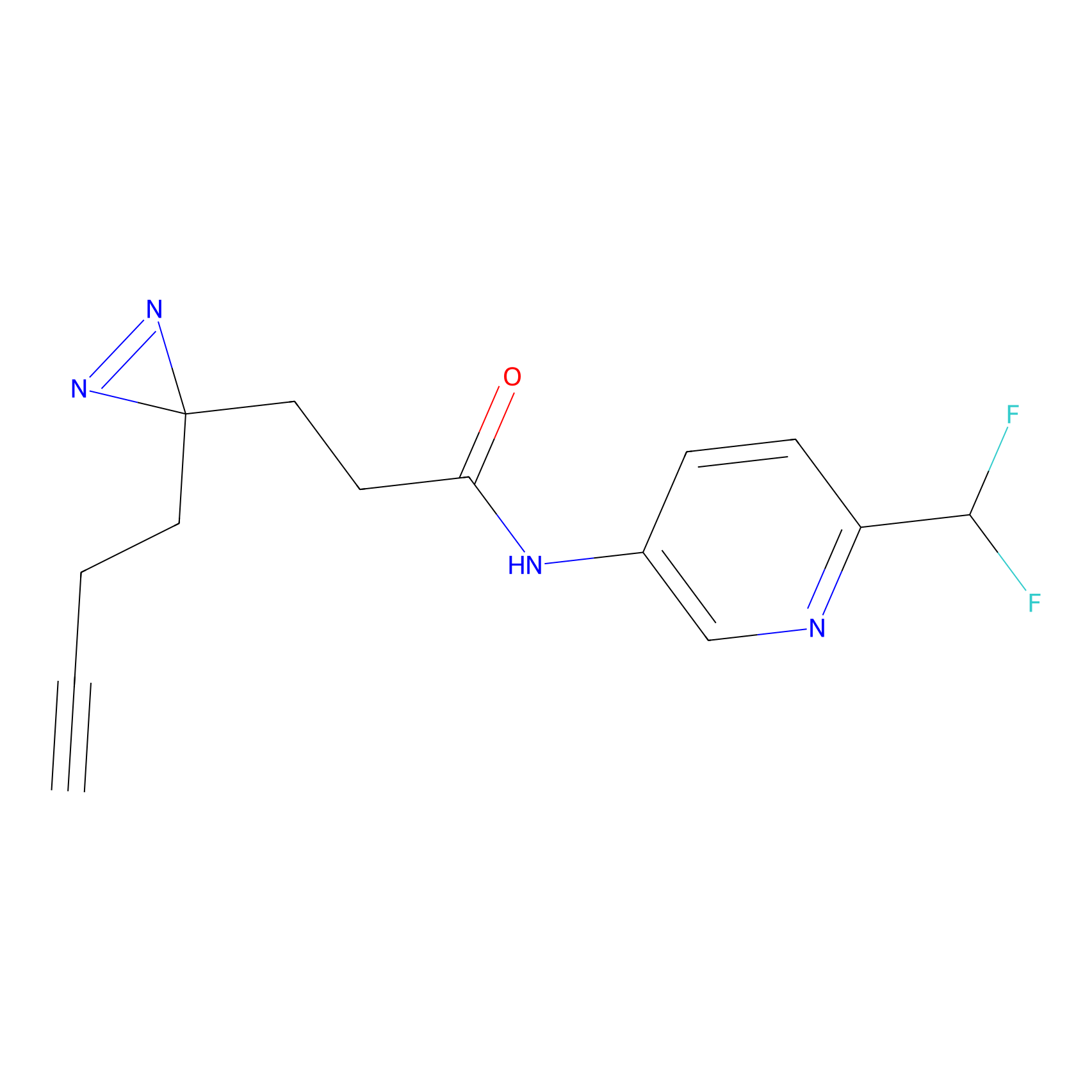 |
5.21 | LDD1931 | [24] | |
|
C269 Probe Info |
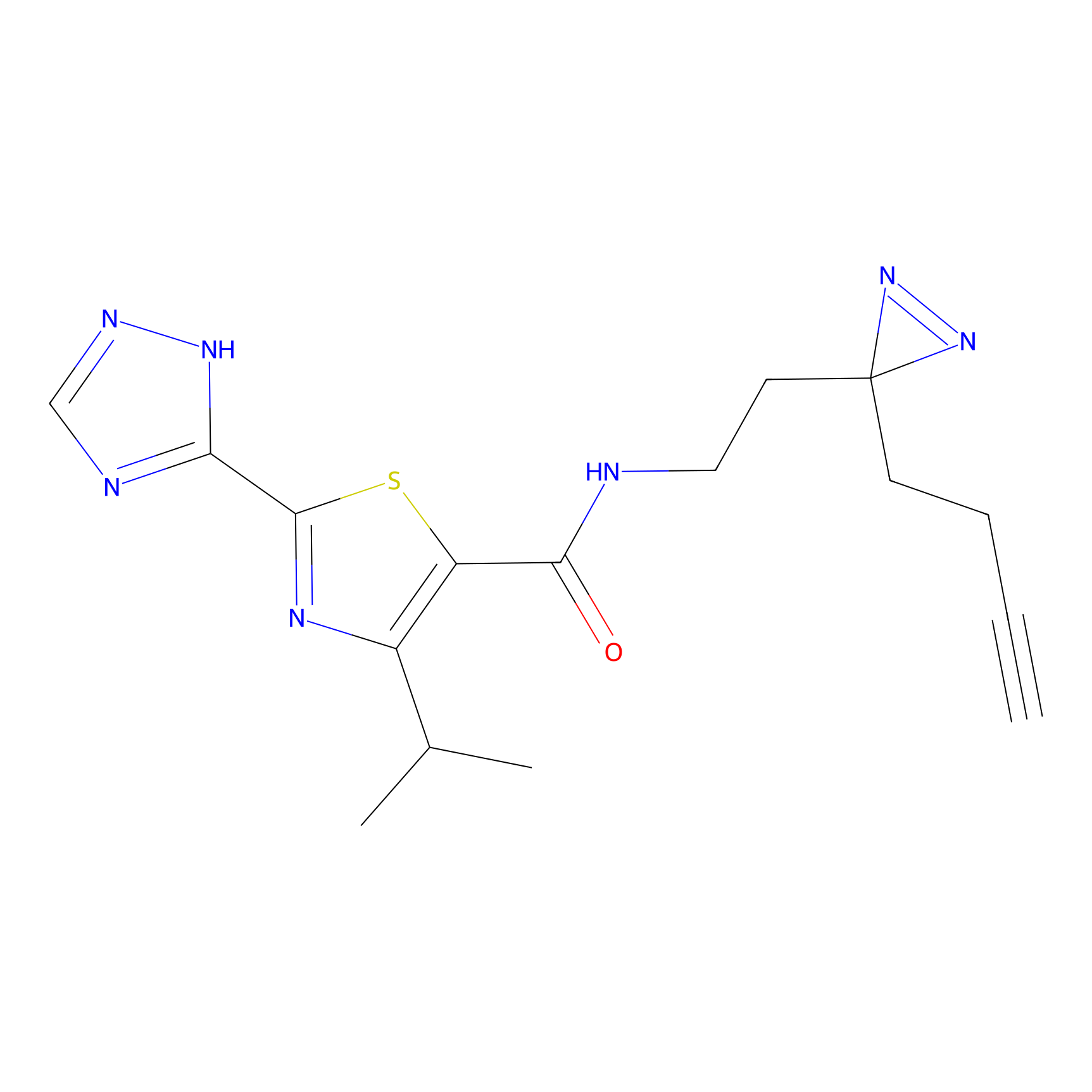 |
5.70 | LDD1939 | [24] | |
|
C403 Probe Info |
 |
14.22 | LDD2061 | [24] | |
|
C420 Probe Info |
 |
11.16 | LDD2075 | [24] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [25] | |
|
FFF probe13 Probe Info |
 |
8.01 | LDD0476 | [25] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [25] | |
|
FFF probe6 Probe Info |
 |
17.03 | LDD0467 | [25] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [26] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C106(0.92) | LDD2130 | [6] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C106(1.01); C327(0.96) | LDD2117 | [6] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C106(1.71) | LDD2152 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C272(2.12) | LDD0169 | [8] |
| LDCM0214 | AC1 | HCT 116 | C106(2.60); C211(1.03); C272(1.11); C292(0.80) | LDD0531 | [11] |
| LDCM0215 | AC10 | HCT 116 | C211(0.94); C272(0.87); C292(0.62); C293(0.64) | LDD0532 | [11] |
| LDCM0216 | AC100 | HCT 116 | C106(0.38); C211(0.59); C292(1.09); C293(1.23) | LDD0533 | [11] |
| LDCM0217 | AC101 | HCT 116 | C106(0.43); C211(0.66); C292(1.03); C293(1.11) | LDD0534 | [11] |
| LDCM0218 | AC102 | HCT 116 | C106(0.58); C211(0.67); C292(0.95); C293(1.14) | LDD0535 | [11] |
| LDCM0219 | AC103 | HCT 116 | C106(0.38); C211(0.53); C292(1.06); C293(1.18) | LDD0536 | [11] |
| LDCM0220 | AC104 | HCT 116 | C106(0.61); C211(0.77); C292(0.89); C293(1.00) | LDD0537 | [11] |
| LDCM0221 | AC105 | HCT 116 | C106(0.44); C211(0.63); C292(1.04); C293(1.13) | LDD0538 | [11] |
| LDCM0222 | AC106 | HCT 116 | C106(0.45); C211(0.65); C292(1.13); C293(1.27) | LDD0539 | [11] |
| LDCM0223 | AC107 | HCT 116 | C106(0.53); C211(0.63); C292(1.08); C293(1.13) | LDD0540 | [11] |
| LDCM0224 | AC108 | HCT 116 | C106(0.40); C211(0.92); C292(1.17); C293(1.34) | LDD0541 | [11] |
| LDCM0225 | AC109 | HCT 116 | C106(0.70); C211(0.89); C292(1.00); C293(1.06) | LDD0542 | [11] |
| LDCM0226 | AC11 | HCT 116 | C211(0.96); C272(0.91); C292(0.59); C293(0.67) | LDD0543 | [11] |
| LDCM0227 | AC110 | HCT 116 | C106(0.43); C211(0.66); C292(1.07); C293(1.18) | LDD0544 | [11] |
| LDCM0228 | AC111 | HCT 116 | C106(0.42); C211(0.66); C292(1.14); C293(1.24) | LDD0545 | [11] |
| LDCM0229 | AC112 | HCT 116 | C106(0.44); C211(0.58); C292(1.10); C293(1.23) | LDD0546 | [11] |
| LDCM0230 | AC113 | HCT 116 | C106(1.19); C211(1.11); C272(0.99); C292(1.11) | LDD0547 | [11] |
| LDCM0231 | AC114 | HCT 116 | C106(1.03); C211(1.05); C272(1.01); C292(1.00) | LDD0548 | [11] |
| LDCM0232 | AC115 | HCT 116 | C106(0.84); C211(0.95); C272(0.94); C292(0.93) | LDD0549 | [11] |
| LDCM0233 | AC116 | HCT 116 | C106(1.16); C211(1.18); C272(1.40); C292(0.96) | LDD0550 | [11] |
| LDCM0234 | AC117 | HCT 116 | C106(1.02); C211(0.97); C272(1.01); C292(0.93) | LDD0551 | [11] |
| LDCM0235 | AC118 | HCT 116 | C106(1.12); C211(0.94); C272(1.30); C292(0.97) | LDD0552 | [11] |
| LDCM0236 | AC119 | HCT 116 | C106(1.44); C211(1.01); C272(0.97); C292(0.96) | LDD0553 | [11] |
| LDCM0237 | AC12 | HCT 116 | C211(0.97); C272(0.79); C292(0.75); C293(0.85) | LDD0554 | [11] |
| LDCM0238 | AC120 | HCT 116 | C106(1.29); C211(1.06); C272(1.31); C292(1.15) | LDD0555 | [11] |
| LDCM0239 | AC121 | HCT 116 | C106(1.18); C211(0.93); C272(1.03); C292(0.99) | LDD0556 | [11] |
| LDCM0240 | AC122 | HCT 116 | C106(1.34); C211(0.95); C272(1.25); C292(0.94) | LDD0557 | [11] |
| LDCM0241 | AC123 | HCT 116 | C106(1.07); C211(0.96); C272(0.90); C292(1.05) | LDD0558 | [11] |
| LDCM0242 | AC124 | HCT 116 | C106(1.34); C211(1.04); C272(1.15); C292(0.95) | LDD0559 | [11] |
| LDCM0243 | AC125 | HCT 116 | C106(1.40); C211(1.07); C272(1.27); C292(1.07) | LDD0560 | [11] |
| LDCM0244 | AC126 | HCT 116 | C106(1.35); C211(0.98); C272(1.36); C292(1.05) | LDD0561 | [11] |
| LDCM0245 | AC127 | HCT 116 | C106(1.19); C211(1.00); C272(1.37); C292(0.90) | LDD0562 | [11] |
| LDCM0246 | AC128 | HCT 116 | C106(0.95); C211(2.46); C272(1.11); C292(2.14) | LDD0563 | [11] |
| LDCM0247 | AC129 | HCT 116 | C106(0.94); C211(2.52); C272(1.16); C292(1.83) | LDD0564 | [11] |
| LDCM0249 | AC130 | HCT 116 | C106(0.66); C211(1.61); C272(1.22); C292(2.02) | LDD0566 | [11] |
| LDCM0250 | AC131 | HCT 116 | C106(0.66); C211(1.33); C272(1.01); C292(1.43) | LDD0567 | [11] |
| LDCM0251 | AC132 | HCT 116 | C106(0.54); C211(1.20); C272(1.18); C292(2.12) | LDD0568 | [11] |
| LDCM0252 | AC133 | HCT 116 | C106(0.63); C211(1.39); C272(1.36); C292(1.70) | LDD0569 | [11] |
| LDCM0253 | AC134 | HCT 116 | C106(0.44); C211(0.79); C272(1.11); C292(1.70) | LDD0570 | [11] |
| LDCM0254 | AC135 | HCT 116 | C106(0.75); C211(1.47); C272(0.99); C292(1.78) | LDD0571 | [11] |
| LDCM0255 | AC136 | HCT 116 | C106(0.71); C211(1.25); C272(1.34); C292(2.12) | LDD0572 | [11] |
| LDCM0256 | AC137 | HCT 116 | C106(0.60); C211(1.39); C272(1.09); C292(1.76) | LDD0573 | [11] |
| LDCM0257 | AC138 | HCT 116 | C106(0.37); C211(0.71); C272(1.26); C292(2.70) | LDD0574 | [11] |
| LDCM0258 | AC139 | HCT 116 | C106(0.46); C211(0.85); C272(1.25); C292(2.23) | LDD0575 | [11] |
| LDCM0259 | AC14 | HCT 116 | C211(0.88); C272(0.89); C292(0.74); C293(0.79) | LDD0576 | [11] |
| LDCM0260 | AC140 | HCT 116 | C106(0.36); C211(0.65); C272(1.18); C292(1.87) | LDD0577 | [11] |
| LDCM0261 | AC141 | HCT 116 | C106(0.38); C211(0.64); C272(1.25); C292(1.82) | LDD0578 | [11] |
| LDCM0262 | AC142 | HCT 116 | C106(0.47); C211(0.90); C272(1.42); C292(1.79) | LDD0579 | [11] |
| LDCM0263 | AC143 | HCT 116 | C106(0.68); C272(1.13); C292(1.03); C293(1.15) | LDD0580 | [11] |
| LDCM0264 | AC144 | HCT 116 | C293(0.93); C272(0.97); C106(0.97); C292(1.07) | LDD0581 | [11] |
| LDCM0265 | AC145 | HCT 116 | C272(1.00); C292(1.00); C293(1.02); C106(1.10) | LDD0582 | [11] |
| LDCM0266 | AC146 | HCT 116 | C106(0.67); C293(1.00); C292(1.01); C272(1.10) | LDD0583 | [11] |
| LDCM0267 | AC147 | HCT 116 | C292(0.90); C272(0.91); C293(1.03); C106(1.07) | LDD0584 | [11] |
| LDCM0268 | AC148 | HCT 116 | C106(0.64); C292(0.91); C293(1.09); C272(2.70) | LDD0585 | [11] |
| LDCM0269 | AC149 | HCT 116 | C106(0.70); C292(0.99); C293(1.22); C272(1.58) | LDD0586 | [11] |
| LDCM0270 | AC15 | HCT 116 | C211(0.80); C272(0.87); C292(0.94); C293(1.03) | LDD0587 | [11] |
| LDCM0271 | AC150 | HCT 116 | C106(0.65); C292(0.98); C272(1.02); C293(1.23) | LDD0588 | [11] |
| LDCM0272 | AC151 | HCT 116 | C106(0.76); C272(0.91); C293(1.02); C292(1.22) | LDD0589 | [11] |
| LDCM0273 | AC152 | HCT 116 | C106(0.62); C292(0.89); C272(1.06); C293(1.36) | LDD0590 | [11] |
| LDCM0274 | AC153 | HCT 116 | C106(0.54); C292(1.04); C293(1.12); C272(1.40) | LDD0591 | [11] |
| LDCM0621 | AC154 | HCT 116 | C106(0.64); C272(0.91); C292(0.95); C293(0.98) | LDD2158 | [11] |
| LDCM0622 | AC155 | HCT 116 | C106(0.57); C272(1.06); C292(1.11); C293(0.95) | LDD2159 | [11] |
| LDCM0623 | AC156 | HCT 116 | C106(0.69); C272(0.95); C292(1.03); C293(1.04) | LDD2160 | [11] |
| LDCM0624 | AC157 | HCT 116 | C106(0.63); C272(0.97); C292(0.96); C293(1.04) | LDD2161 | [11] |
| LDCM0276 | AC17 | HCT 116 | C373(0.79); C292(0.90); C293(0.97); C272(1.00) | LDD0593 | [11] |
| LDCM0277 | AC18 | HCT 116 | C392(0.34); C395(0.34); C373(0.47); C211(0.82) | LDD0594 | [11] |
| LDCM0278 | AC19 | HCT 116 | C373(0.74); C292(0.83); C293(1.01); C272(1.02) | LDD0595 | [11] |
| LDCM0279 | AC2 | HCT 116 | C392(0.71); C395(0.71); C292(0.74); C211(0.83) | LDD0596 | [11] |
| LDCM0280 | AC20 | HCT 116 | C373(0.71); C317(0.95); C272(1.02); C211(1.16) | LDD0597 | [11] |
| LDCM0281 | AC21 | HCT 116 | C373(0.66); C392(0.76); C395(0.76); C292(0.94) | LDD0598 | [11] |
| LDCM0282 | AC22 | HCT 116 | C373(0.79); C317(0.95); C272(1.07); C211(1.10) | LDD0599 | [11] |
| LDCM0283 | AC23 | HCT 116 | C373(0.69); C392(0.98); C395(0.98); C211(0.98) | LDD0600 | [11] |
| LDCM0284 | AC24 | HCT 116 | C373(0.91); C211(1.00); C272(1.01); C317(1.08) | LDD0601 | [11] |
| LDCM0285 | AC25 | HCT 116 | C272(0.80); C211(0.91); C392(0.93); C395(0.93) | LDD0602 | [11] |
| LDCM0286 | AC26 | HCT 116 | C392(0.57); C395(0.57); C326(0.66); C327(0.66) | LDD0603 | [11] |
| LDCM0287 | AC27 | HCT 116 | C373(0.77); C326(0.78); C327(0.78); C211(0.84) | LDD0604 | [11] |
| LDCM0288 | AC28 | HCT 116 | C392(0.58); C395(0.58); C326(0.70); C327(0.70) | LDD0605 | [11] |
| LDCM0289 | AC29 | HCT 116 | C326(0.50); C327(0.50); C392(0.60); C395(0.60) | LDD0606 | [11] |
| LDCM0290 | AC3 | HCT 116 | C292(0.72); C293(0.73); C211(1.01); C392(1.12) | LDD0607 | [11] |
| LDCM0291 | AC30 | HCT 116 | C326(0.52); C327(0.52); C392(0.57); C395(0.57) | LDD0608 | [11] |
| LDCM0292 | AC31 | HCT 116 | C326(0.62); C327(0.62); C392(0.64); C395(0.64) | LDD0609 | [11] |
| LDCM0293 | AC32 | HCT 116 | C392(0.43); C395(0.43); C326(0.46); C327(0.46) | LDD0610 | [11] |
| LDCM0294 | AC33 | HCT 116 | C211(0.59); C392(0.76); C395(0.76); C292(0.87) | LDD0611 | [11] |
| LDCM0295 | AC34 | HCT 116 | C326(0.55); C327(0.55); C373(0.79); C392(0.81) | LDD0612 | [11] |
| LDCM0296 | AC35 | HCT 116 | C293(0.73); C272(0.91); C292(0.92); C373(1.18) | LDD0613 | [11] |
| LDCM0297 | AC36 | HCT 116 | C293(0.83); C272(0.99); C292(1.04); C326(1.08) | LDD0614 | [11] |
| LDCM0298 | AC37 | HCT 116 | C293(0.80); C292(0.99); C272(0.99); C373(1.04) | LDD0615 | [11] |
| LDCM0299 | AC38 | HCT 116 | C326(0.82); C327(0.82); C373(0.94); C272(1.03) | LDD0616 | [11] |
| LDCM0300 | AC39 | HCT 116 | C326(0.91); C327(0.91); C293(0.93); C373(0.98) | LDD0617 | [11] |
| LDCM0301 | AC4 | HCT 116 | C292(0.73); C293(0.77); C211(1.02); C272(1.11) | LDD0618 | [11] |
| LDCM0302 | AC40 | HCT 116 | C373(0.70); C272(0.97); C292(0.99); C293(1.00) | LDD0619 | [11] |
| LDCM0303 | AC41 | HCT 116 | C272(0.91); C392(0.96); C395(0.96); C292(1.01) | LDD0620 | [11] |
| LDCM0304 | AC42 | HCT 116 | C373(0.76); C292(1.03); C272(1.07); C326(1.09) | LDD0621 | [11] |
| LDCM0305 | AC43 | HCT 116 | C373(0.92); C272(1.05); C292(1.09); C326(1.14) | LDD0622 | [11] |
| LDCM0306 | AC44 | HCT 116 | C272(0.84); C373(0.89); C292(1.00); C392(1.44) | LDD0623 | [11] |
| LDCM0307 | AC45 | HCT 116 | C373(0.80); C272(0.89); C292(1.01); C293(1.37) | LDD0624 | [11] |
| LDCM0308 | AC46 | HCT 116 | C326(0.33); C327(0.33); C211(0.42); C392(0.71) | LDD0625 | [11] |
| LDCM0309 | AC47 | HCT 116 | C326(0.64); C327(0.64); C392(0.77); C395(0.77) | LDD0626 | [11] |
| LDCM0310 | AC48 | HCT 116 | C326(0.40); C327(0.40); C211(0.51); C392(0.75) | LDD0627 | [11] |
| LDCM0311 | AC49 | HCT 116 | C326(0.33); C327(0.33); C211(0.42); C392(0.52) | LDD0628 | [11] |
| LDCM0312 | AC5 | HCT 116 | C293(0.79); C292(0.81); C272(1.12); C211(1.13) | LDD0629 | [11] |
| LDCM0313 | AC50 | HCT 116 | C326(0.34); C327(0.34); C211(0.46); C373(0.74) | LDD0630 | [11] |
| LDCM0314 | AC51 | HCT 116 | C326(0.33); C327(0.33); C211(0.42); C392(0.91) | LDD0631 | [11] |
| LDCM0315 | AC52 | HCT 116 | C326(0.36); C327(0.36); C211(0.53); C292(0.90) | LDD0632 | [11] |
| LDCM0316 | AC53 | HCT 116 | C373(0.77); C211(0.79); C392(0.88); C395(0.88) | LDD0633 | [11] |
| LDCM0317 | AC54 | HCT 116 | C326(0.39); C327(0.39); C211(0.53); C392(0.66) | LDD0634 | [11] |
| LDCM0318 | AC55 | HCT 116 | C373(0.73); C293(0.99); C211(1.13); C292(1.13) | LDD0635 | [11] |
| LDCM0319 | AC56 | HCT 116 | C326(0.67); C327(0.67); C373(0.74); C211(0.74) | LDD0636 | [11] |
| LDCM0320 | AC57 | HCT 116 | C373(0.79); C392(0.85); C395(0.85); C272(0.89) | LDD0637 | [11] |
| LDCM0321 | AC58 | HCT 116 | C373(0.69); C326(0.70); C327(0.70); C272(0.80) | LDD0638 | [11] |
| LDCM0322 | AC59 | HCT 116 | C373(0.62); C392(0.93); C395(0.93); C272(0.98) | LDD0639 | [11] |
| LDCM0323 | AC6 | HCT 116 | C326(0.71); C327(0.71); C392(0.75); C395(0.75) | LDD0640 | [11] |
| LDCM0324 | AC60 | HCT 116 | C373(0.85); C272(0.99); C211(1.09); C392(1.24) | LDD0641 | [11] |
| LDCM0325 | AC61 | HCT 116 | C373(0.85); C272(0.93); C392(1.08); C395(1.08) | LDD0642 | [11] |
| LDCM0326 | AC62 | HCT 116 | C373(0.60); C293(1.01); C272(1.03); C292(1.11) | LDD0643 | [11] |
| LDCM0327 | AC63 | HCT 116 | C373(0.90); C272(0.98); C392(1.15); C395(1.15) | LDD0644 | [11] |
| LDCM0328 | AC64 | HCT 116 | C373(0.68); C293(0.99); C272(1.02); C292(1.04) | LDD0645 | [11] |
| LDCM0329 | AC65 | HCT 116 | C392(0.71); C395(0.71); C326(0.91); C327(0.91) | LDD0646 | [11] |
| LDCM0330 | AC66 | HCT 116 | C392(0.72); C395(0.72); C272(0.82); C373(0.94) | LDD0647 | [11] |
| LDCM0331 | AC67 | HCT 116 | C373(0.61); C392(0.62); C395(0.62); C272(1.03) | LDD0648 | [11] |
| LDCM0332 | AC68 | HCT 116 | C211(0.54); C317(0.66); C392(0.77); C395(0.77) | LDD0649 | [11] |
| LDCM0333 | AC69 | HCT 116 | C211(0.68); C317(0.77); C272(0.87); C292(0.97) | LDD0650 | [11] |
| LDCM0334 | AC7 | HCT 116 | C392(0.83); C395(0.83); C211(0.83); C292(0.85) | LDD0651 | [11] |
| LDCM0335 | AC70 | HCT 116 | C317(0.48); C211(0.50); C326(0.78); C327(0.78) | LDD0652 | [11] |
| LDCM0336 | AC71 | HCT 116 | C211(0.75); C272(0.89); C292(0.98); C317(1.00) | LDD0653 | [11] |
| LDCM0337 | AC72 | HCT 116 | C211(0.55); C317(0.56); C326(0.82); C327(0.82) | LDD0654 | [11] |
| LDCM0338 | AC73 | HCT 116 | C317(0.55); C211(0.56); C326(0.71); C327(0.71) | LDD0655 | [11] |
| LDCM0339 | AC74 | HCT 116 | C272(0.87); C292(0.90); C211(0.95); C317(0.95) | LDD0656 | [11] |
| LDCM0340 | AC75 | HCT 116 | C317(0.53); C211(0.55); C293(0.82); C326(0.87) | LDD0657 | [11] |
| LDCM0341 | AC76 | HCT 116 | C272(0.86); C293(0.89); C211(0.89); C317(0.89) | LDD0658 | [11] |
| LDCM0342 | AC77 | HCT 116 | C317(0.58); C211(0.62); C326(0.81); C327(0.81) | LDD0659 | [11] |
| LDCM0343 | AC78 | HCT 116 | C211(0.64); C317(0.71); C272(0.90); C326(0.93) | LDD0660 | [11] |
| LDCM0344 | AC79 | HCT 116 | C211(0.49); C317(0.53); C326(0.85); C327(0.85) | LDD0661 | [11] |
| LDCM0345 | AC8 | HCT 116 | C326(0.85); C327(0.85); C292(0.87); C211(0.88) | LDD0662 | [11] |
| LDCM0346 | AC80 | HCT 116 | C293(0.80); C211(0.84); C272(0.92); C292(0.95) | LDD0663 | [11] |
| LDCM0347 | AC81 | HCT 116 | C211(0.50); C317(0.60); C272(0.88); C292(0.94) | LDD0664 | [11] |
| LDCM0348 | AC82 | HCT 116 | C211(0.63); C272(0.88); C293(0.99); C292(1.00) | LDD0665 | [11] |
| LDCM0349 | AC83 | HCT 116 | C392(0.27); C395(0.27); C211(0.74); C292(0.81) | LDD0666 | [11] |
| LDCM0350 | AC84 | HCT 116 | C392(0.24); C395(0.24); C211(0.67); C106(0.75) | LDD0667 | [11] |
| LDCM0351 | AC85 | HCT 116 | C392(0.31); C395(0.31); C106(0.70); C211(0.75) | LDD0668 | [11] |
| LDCM0352 | AC86 | HCT 116 | C392(0.43); C395(0.43); C211(0.64); C106(0.71) | LDD0669 | [11] |
| LDCM0353 | AC87 | HCT 116 | C106(0.55); C211(0.67); C392(0.73); C395(0.73) | LDD0670 | [11] |
| LDCM0354 | AC88 | HCT 116 | C106(0.54); C211(0.57); C392(0.75); C395(0.75) | LDD0671 | [11] |
| LDCM0355 | AC89 | HCT 116 | C106(0.59); C392(0.62); C395(0.62); C211(0.72) | LDD0672 | [11] |
| LDCM0357 | AC90 | HCT 116 | C211(0.54); C106(0.65); C293(0.86); C292(0.89) | LDD0674 | [11] |
| LDCM0358 | AC91 | HCT 116 | C392(0.28); C395(0.28); C293(0.85); C292(0.94) | LDD0675 | [11] |
| LDCM0359 | AC92 | HCT 116 | C392(0.34); C395(0.34); C106(0.69); C292(0.83) | LDD0676 | [11] |
| LDCM0360 | AC93 | HCT 116 | C392(0.37); C395(0.37); C106(0.75); C211(0.81) | LDD0677 | [11] |
| LDCM0361 | AC94 | HCT 116 | C392(0.33); C395(0.33); C293(0.82); C292(0.96) | LDD0678 | [11] |
| LDCM0362 | AC95 | HCT 116 | C106(0.43); C211(0.46); C292(0.93); C293(0.96) | LDD0679 | [11] |
| LDCM0363 | AC96 | HCT 116 | C392(0.52); C395(0.52); C293(0.78); C292(0.93) | LDD0680 | [11] |
| LDCM0364 | AC97 | HCT 116 | C392(0.28); C395(0.28); C211(0.49); C106(0.61) | LDD0681 | [11] |
| LDCM0365 | AC98 | HCT 116 | C326(0.29); C327(0.29); C106(0.36); C392(0.41) | LDD0682 | [11] |
| LDCM0366 | AC99 | HCT 116 | C392(0.32); C395(0.32); C326(0.33); C327(0.33) | LDD0683 | [11] |
| LDCM0545 | Acetamide | MDA-MB-231 | C106(0.45) | LDD2138 | [6] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C106(0.90) | LDD2113 | [6] |
| LDCM0248 | AKOS034007472 | HCT 116 | C211(0.81); C272(0.99); C292(0.73); C293(0.82) | LDD0565 | [11] |
| LDCM0356 | AKOS034007680 | HCT 116 | C292(0.75); C293(0.79); C272(0.92); C211(0.94) | LDD0673 | [11] |
| LDCM0275 | AKOS034007705 | HCT 116 | C392(0.57); C395(0.57); C326(0.70); C327(0.70) | LDD0592 | [11] |
| LDCM0156 | Aniline | NCI-H1299 | 11.51 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C292(0.96) | LDD0078 | [11] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C272(0.59) | LDD2091 | [6] |
| LDCM0630 | CCW28-3 | 231MFP | C272(1.18) | LDD2214 | [27] |
| LDCM0108 | Chloroacetamide | HeLa | C106(0.00); C395(0.00) | LDD0222 | [20] |
| LDCM0367 | CL1 | HCT 116 | C392(0.72); C395(0.72); C292(0.85); C211(0.91) | LDD0684 | [11] |
| LDCM0368 | CL10 | HCT 116 | C292(0.66); C392(0.68); C395(0.68); C293(0.92) | LDD0685 | [11] |
| LDCM0369 | CL100 | HCT 116 | C292(0.75); C293(0.82); C211(0.86); C272(1.11) | LDD0686 | [11] |
| LDCM0370 | CL101 | HCT 116 | C292(0.75); C293(0.77); C392(0.92); C395(0.92) | LDD0687 | [11] |
| LDCM0371 | CL102 | HCT 116 | C292(0.62); C293(0.66); C392(0.68); C395(0.68) | LDD0688 | [11] |
| LDCM0372 | CL103 | HCT 116 | C293(0.64); C292(0.65); C272(0.97); C211(1.04) | LDD0689 | [11] |
| LDCM0373 | CL104 | HCT 116 | C392(0.73); C395(0.73); C211(0.79); C292(0.88) | LDD0690 | [11] |
| LDCM0374 | CL105 | HCT 116 | C373(0.62); C392(0.77); C395(0.77); C211(0.80) | LDD0691 | [11] |
| LDCM0375 | CL106 | HCT 116 | C373(0.45); C392(0.48); C395(0.48); C211(0.77) | LDD0692 | [11] |
| LDCM0376 | CL107 | HCT 116 | C392(0.43); C395(0.43); C373(0.48); C211(0.88) | LDD0693 | [11] |
| LDCM0377 | CL108 | HCT 116 | C392(0.31); C395(0.31); C373(0.44); C272(0.96) | LDD0694 | [11] |
| LDCM0378 | CL109 | HCT 116 | C373(0.62); C392(0.80); C395(0.80); C211(0.88) | LDD0695 | [11] |
| LDCM0379 | CL11 | HCT 116 | C292(0.69); C211(0.82); C293(0.89); C272(0.93) | LDD0696 | [11] |
| LDCM0380 | CL110 | HCT 116 | C373(0.45); C392(0.57); C395(0.57); C211(0.88) | LDD0697 | [11] |
| LDCM0381 | CL111 | HCT 116 | C211(0.55); C373(0.62); C392(0.78); C395(0.78) | LDD0698 | [11] |
| LDCM0382 | CL112 | HCT 116 | C392(0.66); C395(0.66); C211(0.84); C292(0.85) | LDD0699 | [11] |
| LDCM0383 | CL113 | HCT 116 | C326(0.46); C327(0.46); C392(0.50); C395(0.50) | LDD0700 | [11] |
| LDCM0384 | CL114 | HCT 116 | C326(0.37); C327(0.37); C373(0.43); C392(0.65) | LDD0701 | [11] |
| LDCM0385 | CL115 | HCT 116 | C326(0.54); C327(0.54); C392(0.54); C395(0.54) | LDD0702 | [11] |
| LDCM0386 | CL116 | HCT 116 | C326(0.63); C327(0.63); C392(0.73); C395(0.73) | LDD0703 | [11] |
| LDCM0387 | CL117 | HCT 116 | C373(0.73); C326(0.80); C327(0.80); C392(0.84) | LDD0704 | [11] |
| LDCM0388 | CL118 | HCT 116 | C373(0.94); C272(1.01); C293(1.04); C292(1.15) | LDD0705 | [11] |
| LDCM0389 | CL119 | HCT 116 | C373(0.85); C272(0.97); C293(1.10); C292(1.17) | LDD0706 | [11] |
| LDCM0390 | CL12 | HCT 116 | C292(0.72); C211(0.91); C293(0.96); C272(1.04) | LDD0707 | [11] |
| LDCM0391 | CL120 | HCT 116 | C272(1.00); C392(1.04); C395(1.04); C326(1.05) | LDD0708 | [11] |
| LDCM0392 | CL121 | HCT 116 | C211(0.65); C326(0.69); C327(0.69); C373(0.81) | LDD0709 | [11] |
| LDCM0393 | CL122 | HCT 116 | C326(0.53); C327(0.53); C211(0.58); C392(0.82) | LDD0710 | [11] |
| LDCM0394 | CL123 | HCT 116 | C326(0.59); C327(0.59); C392(0.65); C395(0.65) | LDD0711 | [11] |
| LDCM0395 | CL124 | HCT 116 | C326(0.46); C327(0.46); C211(0.58); C392(0.65) | LDD0712 | [11] |
| LDCM0396 | CL125 | HCT 116 | C392(0.76); C395(0.76); C326(0.80); C327(0.80) | LDD0713 | [11] |
| LDCM0397 | CL126 | HCT 116 | C326(0.77); C327(0.77); C373(0.83); C272(0.88) | LDD0714 | [11] |
| LDCM0398 | CL127 | HCT 116 | C272(1.01); C373(1.05); C326(1.05); C327(1.05) | LDD0715 | [11] |
| LDCM0399 | CL128 | HCT 116 | C373(0.67); C392(0.71); C395(0.71); C326(0.96) | LDD0716 | [11] |
| LDCM0400 | CL13 | HCT 116 | C392(0.92); C395(0.92); C211(0.93); C292(0.94) | LDD0717 | [11] |
| LDCM0401 | CL14 | HCT 116 | C292(0.87); C272(0.96); C293(1.02); C211(1.05) | LDD0718 | [11] |
| LDCM0402 | CL15 | HCT 116 | C272(0.83); C211(0.90); C292(0.99); C293(1.08) | LDD0719 | [11] |
| LDCM0403 | CL16 | HCT 116 | C373(0.75); C272(1.02); C293(1.14); C292(1.15) | LDD0720 | [11] |
| LDCM0404 | CL17 | HCT 116 | C373(0.61); C293(0.72); C292(0.89) | LDD0721 | [11] |
| LDCM0405 | CL18 | HCT 116 | C373(0.70); C293(0.75); C292(0.88) | LDD0722 | [11] |
| LDCM0406 | CL19 | HCT 116 | C373(0.89); C293(0.91); C292(1.01) | LDD0723 | [11] |
| LDCM0407 | CL2 | HCT 116 | C392(0.83); C395(0.83); C292(0.92); C211(0.94) | LDD0724 | [11] |
| LDCM0408 | CL20 | HCT 116 | C373(0.54); C293(0.92); C292(1.02) | LDD0725 | [11] |
| LDCM0409 | CL21 | HCT 116 | C373(0.28); C293(0.71); C292(0.89) | LDD0726 | [11] |
| LDCM0410 | CL22 | HCT 116 | C373(0.50); C292(1.04); C293(1.16) | LDD0727 | [11] |
| LDCM0411 | CL23 | HCT 116 | C373(0.61); C292(1.13); C293(1.68) | LDD0728 | [11] |
| LDCM0412 | CL24 | HCT 116 | C373(0.33); C293(0.98); C292(0.99) | LDD0729 | [11] |
| LDCM0413 | CL25 | HCT 116 | C292(0.98); C293(0.96); C373(0.39) | LDD0730 | [11] |
| LDCM0414 | CL26 | HCT 116 | C292(1.02); C293(0.99); C373(0.49) | LDD0731 | [11] |
| LDCM0415 | CL27 | HCT 116 | C292(1.06); C293(1.00); C373(0.69) | LDD0732 | [11] |
| LDCM0416 | CL28 | HCT 116 | C292(0.98); C293(0.84); C373(0.56) | LDD0733 | [11] |
| LDCM0417 | CL29 | HCT 116 | C292(1.10); C293(1.12); C373(0.43) | LDD0734 | [11] |
| LDCM0418 | CL3 | HCT 116 | C211(0.88); C272(0.91); C292(0.86); C293(0.98) | LDD0735 | [11] |
| LDCM0419 | CL30 | HCT 116 | C292(0.95); C293(0.76); C373(0.49) | LDD0736 | [11] |
| LDCM0420 | CL31 | HCT 116 | C106(0.74); C272(0.92); C292(0.98); C293(0.90) | LDD0737 | [11] |
| LDCM0421 | CL32 | HCT 116 | C106(0.70); C272(1.06); C292(1.13); C293(1.15) | LDD0738 | [11] |
| LDCM0422 | CL33 | HCT 116 | C106(0.69); C272(1.04); C292(1.08); C293(1.06) | LDD0739 | [11] |
| LDCM0423 | CL34 | HCT 116 | C106(0.54); C272(1.05); C292(1.11); C293(1.09) | LDD0740 | [11] |
| LDCM0424 | CL35 | HCT 116 | C106(0.42); C272(1.04); C292(1.23); C293(1.19) | LDD0741 | [11] |
| LDCM0425 | CL36 | HCT 116 | C106(0.45); C272(0.92); C292(1.16); C293(1.25) | LDD0742 | [11] |
| LDCM0426 | CL37 | HCT 116 | C106(0.32); C272(0.94); C292(1.24); C293(1.22) | LDD0743 | [11] |
| LDCM0428 | CL39 | HCT 116 | C106(0.41); C272(1.02); C292(1.17); C293(1.12) | LDD0745 | [11] |
| LDCM0429 | CL4 | HCT 116 | C211(0.82); C272(0.85); C292(0.98); C293(1.16) | LDD0746 | [11] |
| LDCM0430 | CL40 | HCT 116 | C106(0.54); C272(0.93); C292(1.04); C293(1.05) | LDD0747 | [11] |
| LDCM0431 | CL41 | HCT 116 | C106(0.52); C272(0.92); C292(1.19); C293(1.02) | LDD0748 | [11] |
| LDCM0432 | CL42 | HCT 116 | C106(0.31); C272(0.97); C292(1.10); C293(1.07) | LDD0749 | [11] |
| LDCM0433 | CL43 | HCT 116 | C106(0.41); C272(0.99); C292(1.12); C293(1.09) | LDD0750 | [11] |
| LDCM0434 | CL44 | HCT 116 | C106(0.54); C272(0.98); C292(1.19); C293(1.17) | LDD0751 | [11] |
| LDCM0435 | CL45 | HCT 116 | C106(0.43); C272(0.90); C292(1.22); C293(1.24) | LDD0752 | [11] |
| LDCM0436 | CL46 | HCT 116 | C272(1.12); C292(1.08); C293(1.23); C392(1.20) | LDD0753 | [11] |
| LDCM0437 | CL47 | HCT 116 | C272(1.06); C292(1.10); C293(1.14); C392(0.90) | LDD0754 | [11] |
| LDCM0438 | CL48 | HCT 116 | C272(1.00); C292(1.12); C293(1.17); C392(0.94) | LDD0755 | [11] |
| LDCM0439 | CL49 | HCT 116 | C272(1.21); C292(1.13); C293(1.25); C392(0.93) | LDD0756 | [11] |
| LDCM0440 | CL5 | HCT 116 | C211(1.06); C272(0.91); C292(0.82); C293(0.94) | LDD0757 | [11] |
| LDCM0441 | CL50 | HCT 116 | C272(0.96); C292(1.13); C293(1.25); C392(0.86) | LDD0758 | [11] |
| LDCM0442 | CL51 | HCT 116 | C272(0.97); C292(1.15); C293(1.26); C392(1.12) | LDD0759 | [11] |
| LDCM0443 | CL52 | HCT 116 | C272(1.19); C292(1.03); C293(1.27); C392(0.92) | LDD0760 | [11] |
| LDCM0444 | CL53 | HCT 116 | C272(0.83); C292(1.16); C293(1.16); C392(0.75) | LDD0761 | [11] |
| LDCM0445 | CL54 | HCT 116 | C272(1.00); C292(1.06); C293(1.17); C392(0.87) | LDD0762 | [11] |
| LDCM0446 | CL55 | HCT 116 | C272(0.95); C292(0.99); C293(1.04); C392(0.92) | LDD0763 | [11] |
| LDCM0447 | CL56 | HCT 116 | C272(1.00); C292(1.11); C293(1.07); C392(0.96) | LDD0764 | [11] |
| LDCM0448 | CL57 | HCT 116 | C272(1.03); C292(1.15); C293(1.13); C392(0.95) | LDD0765 | [11] |
| LDCM0449 | CL58 | HCT 116 | C272(1.17); C292(1.15); C293(1.19); C392(0.87) | LDD0766 | [11] |
| LDCM0450 | CL59 | HCT 116 | C272(1.07); C292(1.06); C293(1.17); C392(0.87) | LDD0767 | [11] |
| LDCM0451 | CL6 | HCT 116 | C211(0.88); C272(0.97); C292(0.79); C293(0.97) | LDD0768 | [11] |
| LDCM0452 | CL60 | HCT 116 | C272(1.05); C292(1.12); C293(1.13); C392(0.90) | LDD0769 | [11] |
| LDCM0453 | CL61 | HCT 116 | C292(0.85); C293(1.19); C317(0.73); C392(1.17) | LDD0770 | [11] |
| LDCM0454 | CL62 | HCT 116 | C292(0.81); C293(0.94); C317(0.79); C392(0.67) | LDD0771 | [11] |
| LDCM0455 | CL63 | HCT 116 | C292(0.96); C293(1.32); C317(0.75); C392(0.68) | LDD0772 | [11] |
| LDCM0456 | CL64 | HCT 116 | C292(0.76); C293(0.93); C317(0.84); C392(0.57) | LDD0773 | [11] |
| LDCM0457 | CL65 | HCT 116 | C292(0.85); C293(0.98); C317(1.20); C392(0.78) | LDD0774 | [11] |
| LDCM0458 | CL66 | HCT 116 | C292(0.86); C293(1.05); C317(1.04); C392(0.47) | LDD0775 | [11] |
| LDCM0459 | CL67 | HCT 116 | C292(0.85); C293(0.91); C317(1.15); C392(0.61) | LDD0776 | [11] |
| LDCM0460 | CL68 | HCT 116 | C292(1.20); C293(1.41); C317(0.57); C392(0.63) | LDD0777 | [11] |
| LDCM0461 | CL69 | HCT 116 | C292(0.92); C293(1.08); C317(0.49); C392(1.16) | LDD0778 | [11] |
| LDCM0462 | CL7 | HCT 116 | C211(0.85); C272(1.00); C292(0.66); C293(0.87) | LDD0779 | [11] |
| LDCM0463 | CL70 | HCT 116 | C292(1.09); C293(1.25); C317(0.71); C392(1.39) | LDD0780 | [11] |
| LDCM0464 | CL71 | HCT 116 | C292(0.97); C293(1.18); C317(0.61); C392(0.57) | LDD0781 | [11] |
| LDCM0465 | CL72 | HCT 116 | C292(1.03); C293(1.26); C317(0.51); C392(0.61) | LDD0782 | [11] |
| LDCM0466 | CL73 | HCT 116 | C292(0.96); C293(1.04); C317(0.65); C392(0.63) | LDD0783 | [11] |
| LDCM0467 | CL74 | HCT 116 | C292(1.02); C293(1.15); C317(0.71); C392(0.92) | LDD0784 | [11] |
| LDCM0469 | CL76 | HCT 116 | C272(0.79); C292(1.30); C293(1.26); C392(1.04) | LDD0786 | [11] |
| LDCM0470 | CL77 | HCT 116 | C272(0.77); C292(1.54); C293(1.46); C392(1.12) | LDD0787 | [11] |
| LDCM0471 | CL78 | HCT 116 | C272(0.84); C292(1.10); C293(1.09); C392(1.12) | LDD0788 | [11] |
| LDCM0472 | CL79 | HCT 116 | C272(0.87); C292(1.15); C293(1.11); C392(0.79) | LDD0789 | [11] |
| LDCM0473 | CL8 | HCT 116 | C211(0.86); C272(0.84); C292(0.90); C293(1.02) | LDD0790 | [11] |
| LDCM0474 | CL80 | HCT 116 | C272(0.95); C292(1.19); C293(1.29); C392(1.21) | LDD0791 | [11] |
| LDCM0475 | CL81 | HCT 116 | C272(0.92); C292(1.26); C293(1.24); C392(1.30) | LDD0792 | [11] |
| LDCM0476 | CL82 | HCT 116 | C272(0.82); C292(1.10); C293(0.98); C392(1.02) | LDD0793 | [11] |
| LDCM0477 | CL83 | HCT 116 | C272(0.85); C292(0.97); C293(1.06); C392(0.65) | LDD0794 | [11] |
| LDCM0478 | CL84 | HCT 116 | C272(0.88); C292(0.99); C293(1.05); C392(0.72) | LDD0795 | [11] |
| LDCM0479 | CL85 | HCT 116 | C272(0.88); C292(1.19); C293(1.20); C392(0.94) | LDD0796 | [11] |
| LDCM0480 | CL86 | HCT 116 | C272(0.81); C292(1.23); C293(1.20); C392(0.93) | LDD0797 | [11] |
| LDCM0481 | CL87 | HCT 116 | C272(0.94); C292(1.08); C293(1.14); C392(0.96) | LDD0798 | [11] |
| LDCM0482 | CL88 | HCT 116 | C272(0.92); C292(1.24); C293(1.18); C392(0.98) | LDD0799 | [11] |
| LDCM0483 | CL89 | HCT 116 | C272(0.80); C292(1.14); C293(1.17); C392(0.80) | LDD0800 | [11] |
| LDCM0484 | CL9 | HCT 116 | C211(0.94); C272(0.96); C292(1.02); C293(1.15) | LDD0801 | [11] |
| LDCM0485 | CL90 | HCT 116 | C272(0.90); C292(1.07); C293(1.15); C392(1.10) | LDD0802 | [11] |
| LDCM0486 | CL91 | HCT 116 | C106(0.63); C211(0.72); C272(1.25); C292(0.94) | LDD0803 | [11] |
| LDCM0487 | CL92 | HCT 116 | C106(1.01); C211(0.78); C272(1.14); C292(1.11) | LDD0804 | [11] |
| LDCM0488 | CL93 | HCT 116 | C106(1.36); C211(1.06); C272(0.95); C292(0.88) | LDD0805 | [11] |
| LDCM0489 | CL94 | HCT 116 | C106(1.75); C211(1.10); C272(0.89); C292(0.81) | LDD0806 | [11] |
| LDCM0490 | CL95 | HCT 116 | C106(0.87); C211(0.74); C272(1.09); C292(0.90) | LDD0807 | [11] |
| LDCM0491 | CL96 | HCT 116 | C106(0.88); C211(0.85); C272(1.28); C292(0.87) | LDD0808 | [11] |
| LDCM0492 | CL97 | HCT 116 | C106(0.84); C211(0.86); C272(1.14); C292(0.89) | LDD0809 | [11] |
| LDCM0493 | CL98 | HCT 116 | C106(1.39); C211(0.72); C272(1.13); C292(0.81) | LDD0810 | [11] |
| LDCM0494 | CL99 | HCT 116 | C106(1.99); C211(0.85); C272(1.10); C292(0.76) | LDD0811 | [11] |
| LDCM0191 | Compound 21 | HEK-293T | 12.12 | LDD0508 | [25] |
| LDCM0192 | Compound 35 | HEK-293T | 5.01 | LDD0509 | [25] |
| LDCM0193 | Compound 36 | HEK-293T | 8.40 | LDD0511 | [25] |
| LDCM0634 | CY-0357 | Hep-G2 | C272(1.15) | LDD2228 | [23] |
| LDCM0495 | E2913 | HEK-293T | C106(0.96); C373(0.95) | LDD1698 | [28] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C292(14.50); C293(6.29); C272(2.71); C373(1.47) | LDD1702 | [6] |
| LDCM0625 | F8 | Ramos | C272(1.28) | LDD2187 | [29] |
| LDCM0572 | Fragment10 | Ramos | C272(1.23) | LDD2189 | [29] |
| LDCM0573 | Fragment11 | Ramos | C272(0.00) | LDD2190 | [29] |
| LDCM0574 | Fragment12 | Ramos | C272(0.86) | LDD2191 | [29] |
| LDCM0575 | Fragment13 | Ramos | C272(0.96) | LDD2192 | [29] |
| LDCM0576 | Fragment14 | Ramos | C272(0.65) | LDD2193 | [29] |
| LDCM0579 | Fragment20 | Ramos | C272(0.69) | LDD2194 | [29] |
| LDCM0580 | Fragment21 | Ramos | C272(0.97) | LDD2195 | [29] |
| LDCM0582 | Fragment23 | Ramos | C272(1.14) | LDD2196 | [29] |
| LDCM0578 | Fragment27 | Ramos | C272(1.03) | LDD2197 | [29] |
| LDCM0586 | Fragment28 | Ramos | C272(0.78) | LDD2198 | [29] |
| LDCM0588 | Fragment30 | Ramos | C272(1.19) | LDD2199 | [29] |
| LDCM0589 | Fragment31 | Ramos | C272(0.91) | LDD2200 | [29] |
| LDCM0590 | Fragment32 | Ramos | C272(0.74) | LDD2201 | [29] |
| LDCM0468 | Fragment33 | HCT 116 | C292(1.01); C293(1.09); C317(0.71); C392(0.69) | LDD0785 | [11] |
| LDCM0596 | Fragment38 | Ramos | C272(0.81) | LDD2203 | [29] |
| LDCM0566 | Fragment4 | Ramos | C272(0.85) | LDD2184 | [29] |
| LDCM0427 | Fragment51 | HCT 116 | C106(0.52); C272(0.85); C292(1.00); C293(0.96) | LDD0744 | [11] |
| LDCM0610 | Fragment52 | Ramos | C272(1.13) | LDD2204 | [29] |
| LDCM0614 | Fragment56 | Ramos | C272(1.22) | LDD2205 | [29] |
| LDCM0569 | Fragment7 | Ramos | C272(1.31) | LDD2186 | [29] |
| LDCM0571 | Fragment9 | Ramos | C272(0.98) | LDD2188 | [29] |
| LDCM0116 | HHS-0101 | DM93 | Y491(0.75); Y514(0.90); Y527(0.95) | LDD0264 | [9] |
| LDCM0117 | HHS-0201 | DM93 | Y514(0.84); Y491(0.94); Y527(0.97) | LDD0265 | [9] |
| LDCM0118 | HHS-0301 | DM93 | Y491(0.77); Y527(0.77); Y514(0.78) | LDD0266 | [9] |
| LDCM0119 | HHS-0401 | DM93 | Y491(0.53); Y527(0.85); Y514(0.86) | LDD0267 | [9] |
| LDCM0120 | HHS-0701 | DM93 | Y491(0.76); Y514(0.93); Y527(1.75) | LDD0268 | [9] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [20] |
| LDCM0022 | KB02 | HCT 116 | C392(2.79); C395(2.79); C211(1.79); C292(3.18) | LDD0080 | [11] |
| LDCM0023 | KB03 | HCT 116 | C392(2.79); C395(2.79); C211(3.19); C292(2.33) | LDD0081 | [11] |
| LDCM0024 | KB05 | HCT 116 | C392(2.50); C395(2.50); C211(2.50); C292(2.43) | LDD0082 | [11] |
| LDCM0006 | Micheliolide | M9-ENL1 | 2.00 | LDD0050 | [7] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C106(0.47) | LDD2089 | [6] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C106(0.54) | LDD2096 | [6] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C106(1.05) | LDD2097 | [6] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C106(0.81) | LDD2099 | [6] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C106(1.08) | LDD2101 | [6] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C106(0.63) | LDD2109 | [6] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C106(1.00) | LDD2115 | [6] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C106(0.89) | LDD2118 | [6] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C106(0.89) | LDD2120 | [6] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C106(0.57); C272(0.32) | LDD2122 | [6] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C106(0.94); C327(0.66) | LDD2123 | [6] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C106(0.68); C272(0.31) | LDD2124 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C106(0.99); C327(1.17) | LDD2125 | [6] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C106(0.73); C272(0.23) | LDD2126 | [6] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C106(1.07) | LDD2127 | [6] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C106(0.91) | LDD2129 | [6] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C106(1.01) | LDD2133 | [6] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C106(0.56) | LDD2134 | [6] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C106(1.14) | LDD2135 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C106(1.25); C327(1.05) | LDD2136 | [6] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C106(1.11); C327(1.00) | LDD2137 | [6] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C106(1.52) | LDD1700 | [6] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C106(1.02) | LDD2143 | [6] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C106(3.49) | LDD2144 | [6] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C106(1.04) | LDD2146 | [6] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C272(0.62) | LDD2150 | [6] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C272(0.80) | LDD2206 | [30] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C272(0.70); C272(0.07) | LDD2207 | [30] |
| LDCM0021 | THZ1 | HCT 116 | C292(1.02); C293(1.02) | LDD2173 | [11] |
| LDCM0112 | W16 | Hep-G2 | C292(0.73); C293(0.73) | LDD0239 | [21] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
Other
References
