Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y106(20.00); Y50(20.00); Y59(20.00); Y51(11.40) | LDD0257 | [1] | |
|
TH214 Probe Info |
 |
Y59(20.00); Y106(16.94) | LDD0258 | [1] | |
|
TH216 Probe Info |
 |
Y106(20.00); Y50(20.00); Y59(20.00); Y51(7.25) | LDD0259 | [1] | |
|
AZ-9 Probe Info |
 |
E383(0.90) | LDD2208 | [2] | |
|
Probe 1 Probe Info |
 |
Y36(10.21) | LDD3495 | [3] | |
|
THZ1-DTB Probe Info |
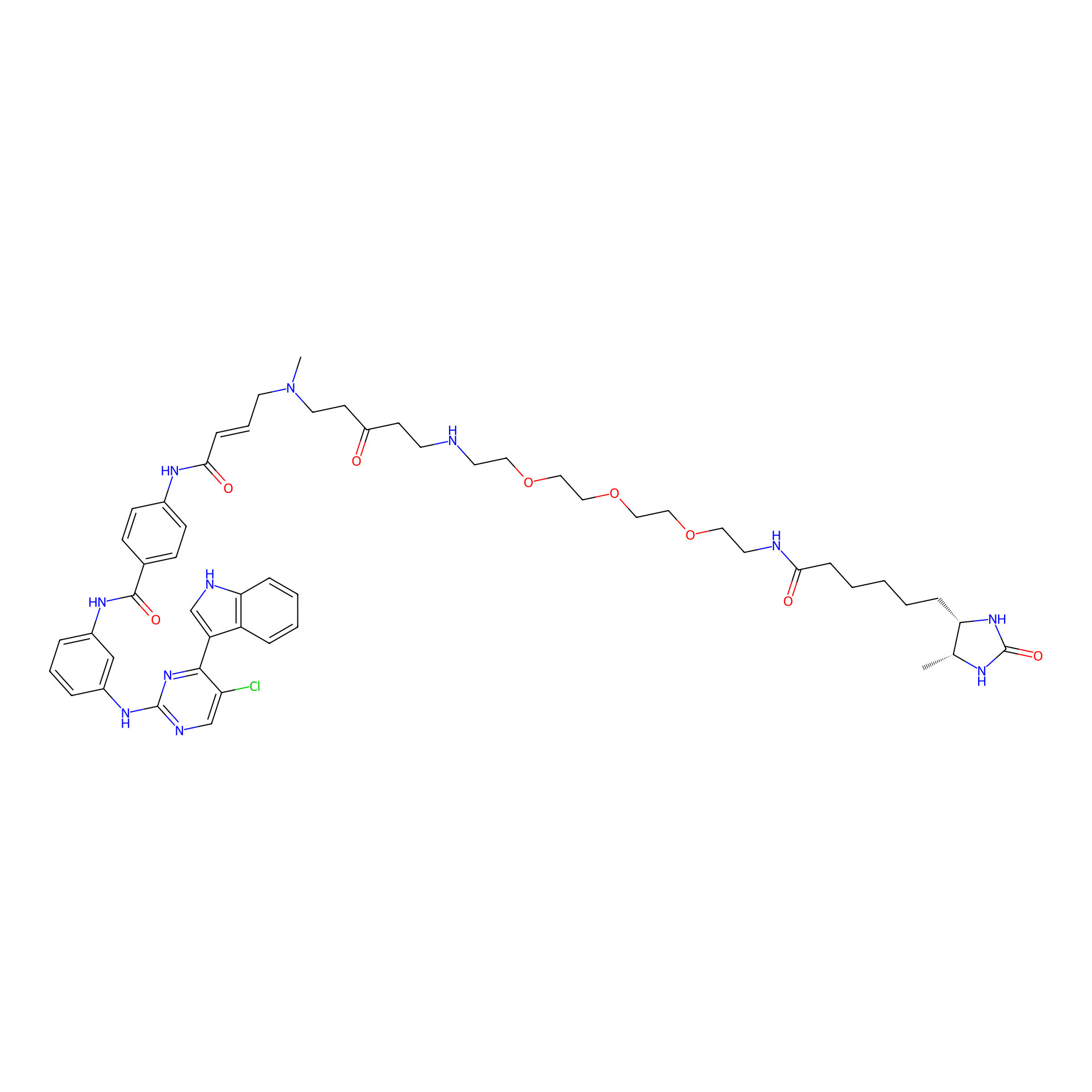 |
C239(1.11); C211(1.04); C239(0.99); C201(1.03) | LDD0460 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C129(2.47); C303(2.15) | LDD0168 | [5] | |
|
HPAP Probe Info |
 |
5.81 | LDD0062 | [6] | |
|
HHS-482 Probe Info |
 |
Y106(0.89); Y50(0.86); Y51(0.87); Y59(0.79) | LDD0285 | [7] | |
|
HHS-475 Probe Info |
 |
Y222(0.69); Y310(0.76); Y281(0.79); Y36(0.83) | LDD0264 | [8] | |
|
HHS-465 Probe Info |
 |
Y106(9.01); Y183(10.00); Y200(3.29); Y208(10.00) | LDD2237 | [9] | |
|
5E-2FA Probe Info |
 |
H6(0.00); H227(0.00) | LDD2235 | [10] | |
|
m-APA Probe Info |
 |
H6(0.00); H28(0.00); H227(0.00) | LDD2231 | [10] | |
|
N1 Probe Info |
 |
C239(0.00); C303(0.00); C354(0.00); D224(0.00) | LDD0245 | [11] | |
|
IA-alkyne Probe Info |
 |
C127(0.00); C239(0.00); C201(0.00); C354(0.00) | LDD0032 | [12] | |
|
IPIAA_H Probe Info |
 |
C239(0.00); C127(0.00); C303(0.00) | LDD0030 | [13] | |
|
IPIAA_L Probe Info |
 |
C127(0.00); C354(0.00); C239(0.00); C303(0.00) | LDD0031 | [13] | |
|
JW-RF-010 Probe Info |
 |
C129(0.00); C127(0.00); C354(0.00); C201(0.00) | LDD0026 | [14] | |
|
SF Probe Info |
 |
Y310(0.00); Y340(0.00); K252(0.00); Y222(0.00) | LDD0028 | [15] | |
|
VSF Probe Info |
 |
C239(0.00); C127(0.00); C129(0.00) | LDD0007 | [16] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [17] | |
|
Acrolein Probe Info |
 |
H28(0.00); H37(0.00) | LDD0217 | [18] | |
|
NAIA_5 Probe Info |
 |
C303(0.00); C12(0.00); C239(0.00) | LDD2223 | [19] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C049 Probe Info |
 |
8.06 | LDD1747 | [20] | |
|
C153 Probe Info |
 |
15.24 | LDD1834 | [20] | |
|
C159 Probe Info |
 |
23.75 | LDD1839 | [20] | |
|
C160 Probe Info |
 |
7.52 | LDD1840 | [20] | |
|
C198 Probe Info |
 |
41.64 | LDD1874 | [20] | |
|
C213 Probe Info |
 |
19.56 | LDD1887 | [20] | |
|
C255 Probe Info |
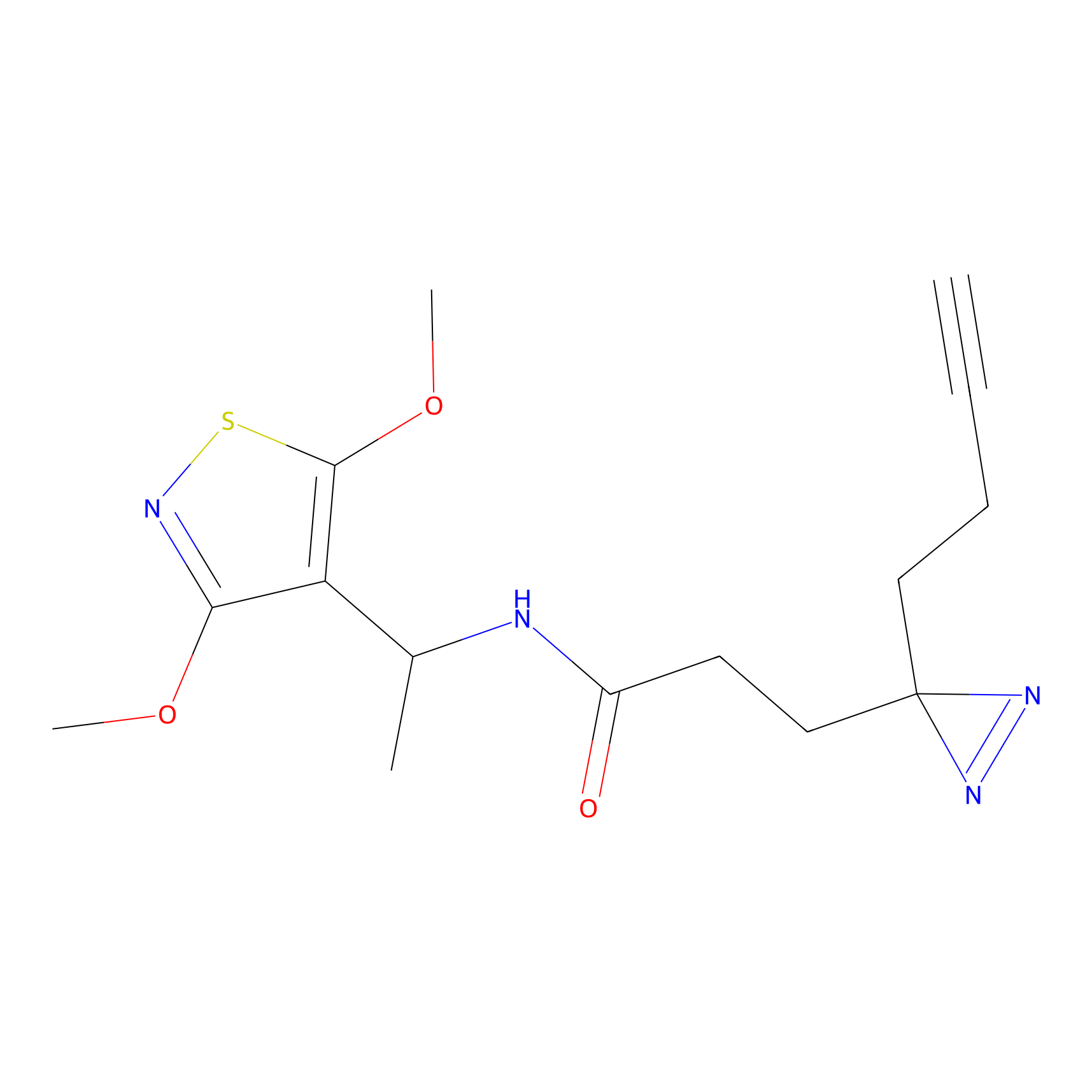 |
5.24 | LDD1928 | [20] | |
|
C264 Probe Info |
 |
53.82 | LDD1935 | [20] | |
|
C270 Probe Info |
 |
6.96 | LDD1940 | [20] | |
|
C287 Probe Info |
 |
17.51 | LDD1957 | [20] | |
|
C349 Probe Info |
 |
9.71 | LDD2010 | [20] | |
|
C363 Probe Info |
 |
17.15 | LDD2024 | [20] | |
|
C382 Probe Info |
 |
21.86 | LDD2041 | [20] | |
|
C413 Probe Info |
 |
11.71 | LDD2069 | [20] | |
|
FFF probe11 Probe Info |
 |
17.78 | LDD0472 | [21] | |
|
FFF probe13 Probe Info |
 |
16.59 | LDD0476 | [21] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0465 | [21] | |
|
FFF probe6 Probe Info |
 |
8.45 | LDD0468 | [21] | |
|
Alk-rapa Probe Info |
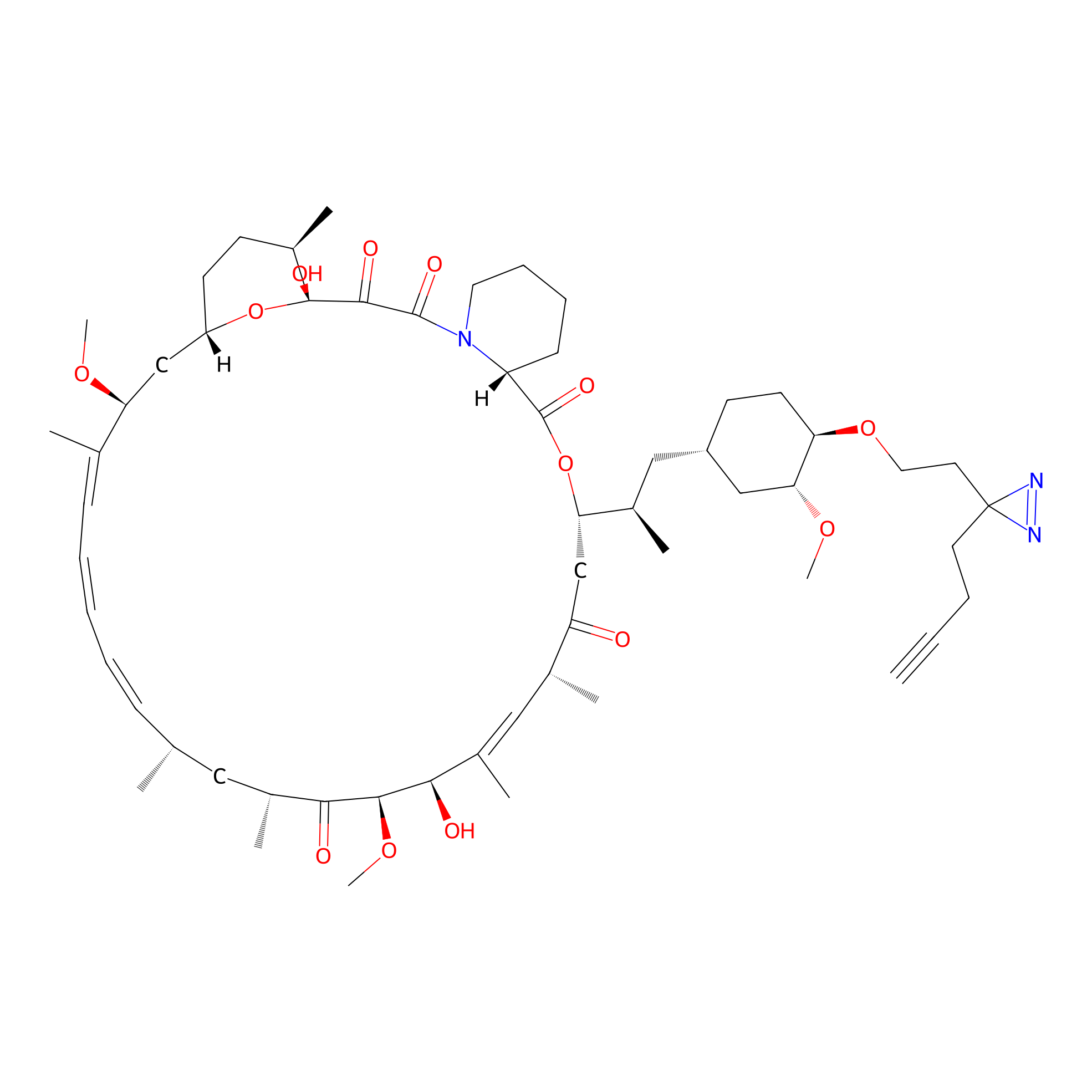 |
3.07 | LDD0213 | [22] | |
|
Photocelecoxib Probe Info |
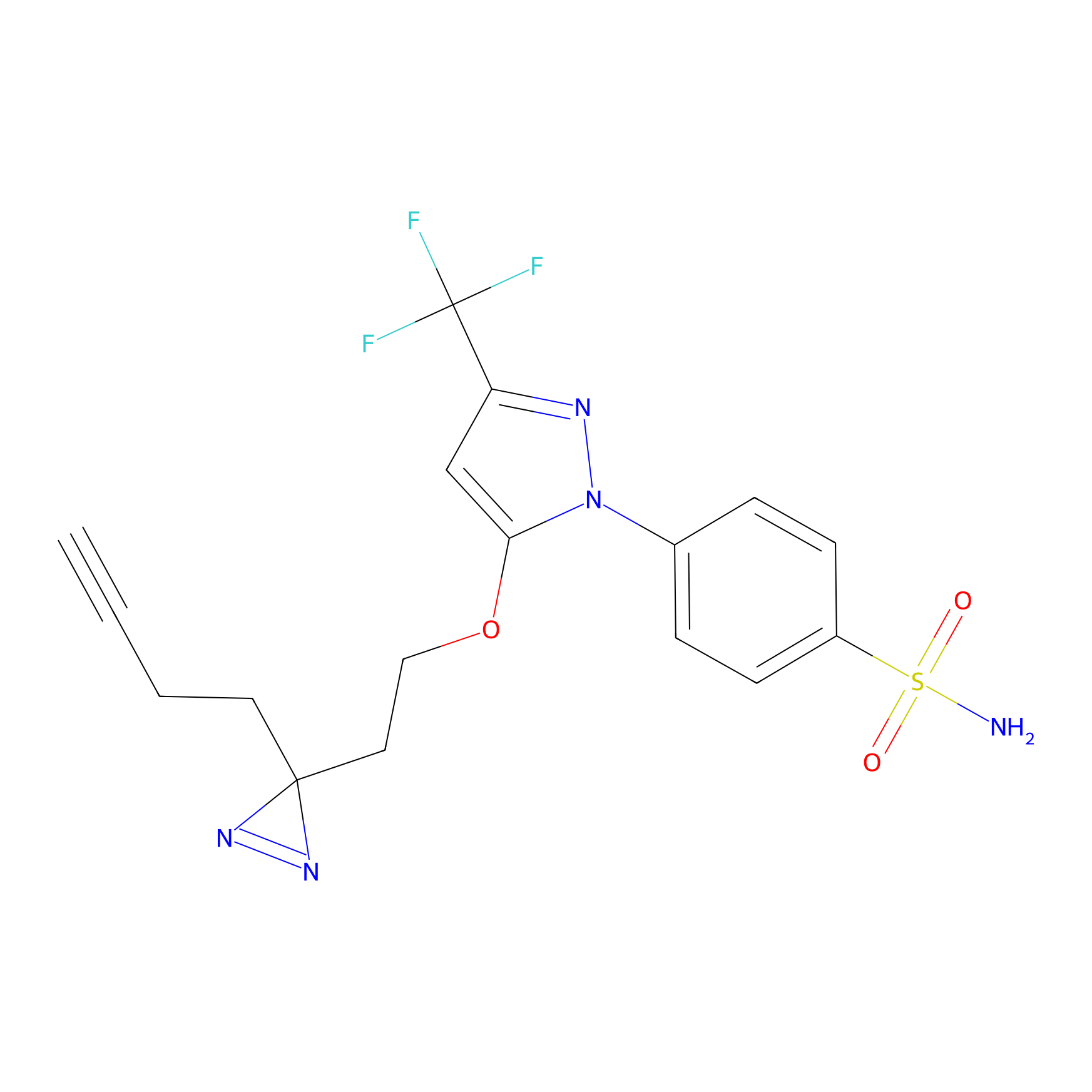 |
E123(0.00); S138(0.00) | LDD0019 | [23] | |
|
Photonaproxen Probe Info |
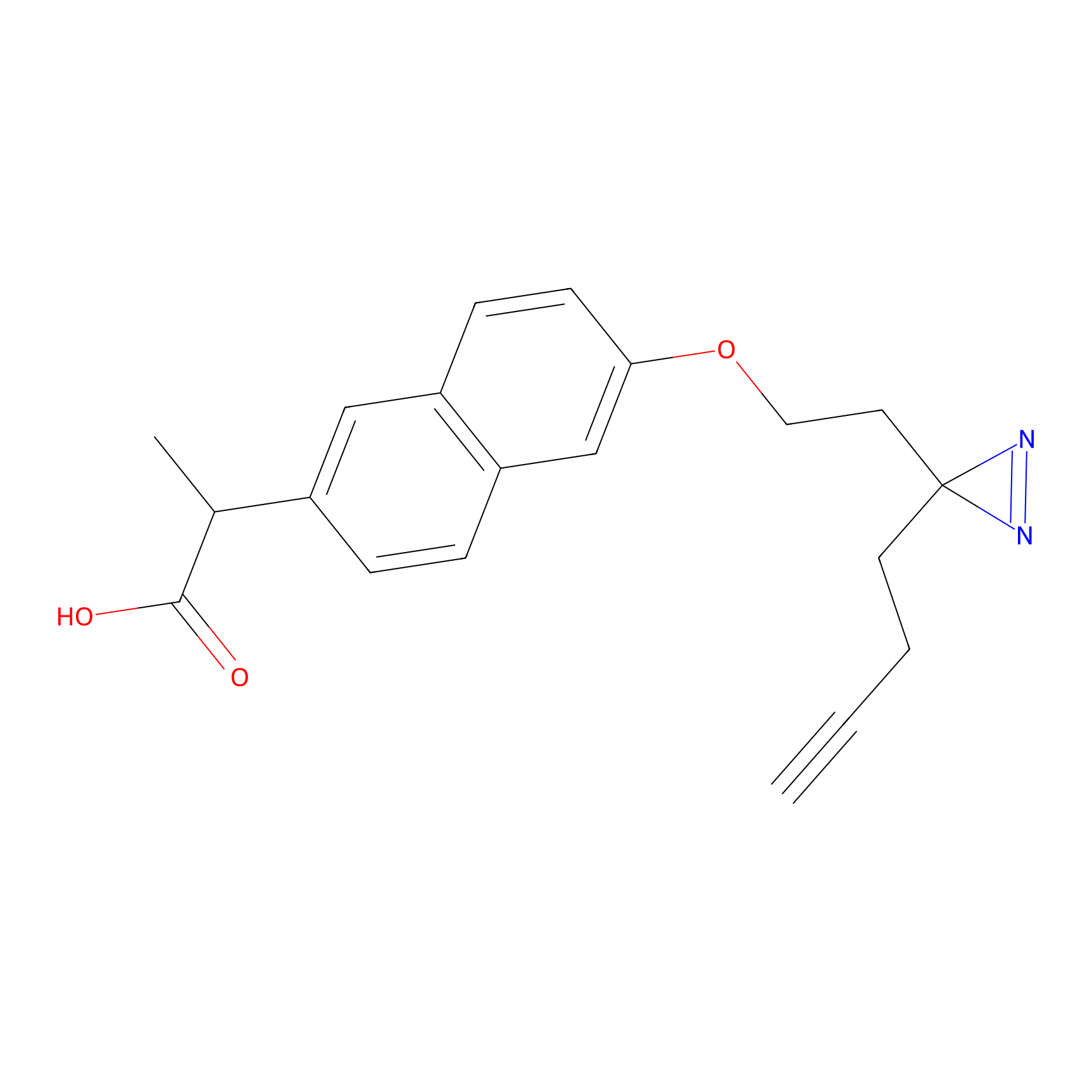 |
T218(0.00); S230(0.00); T219(0.00) | LDD0157 | [23] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [24] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [25] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C129(2.47); C303(2.15) | LDD0168 | [5] |
| LDCM0108 | Chloroacetamide | HeLa | H28(0.00); H37(0.00) | LDD0222 | [18] |
| LDCM0116 | HHS-0101 | DM93 | Y222(0.69); Y310(0.76); Y281(0.79); Y36(0.83) | LDD0264 | [8] |
| LDCM0117 | HHS-0201 | DM93 | Y222(0.59); Y310(0.74); Y281(0.80); Y340(0.92) | LDD0265 | [8] |
| LDCM0118 | HHS-0301 | DM93 | Y222(0.62); Y310(0.75); Y281(0.85); Y340(0.89) | LDD0266 | [8] |
| LDCM0119 | HHS-0401 | DM93 | Y222(0.63); Y310(0.74); Y281(0.91); Y36(1.03) | LDD0267 | [8] |
| LDCM0120 | HHS-0701 | DM93 | Y222(0.63); Y36(0.74); Y281(0.76); Y310(0.78) | LDD0268 | [8] |
| LDCM0107 | IAA | HeLa | H28(0.00); H37(0.00) | LDD0221 | [18] |
| LDCM0123 | JWB131 | DM93 | Y106(0.89); Y50(0.86); Y51(0.87); Y59(0.79) | LDD0285 | [7] |
| LDCM0124 | JWB142 | DM93 | Y106(0.57); Y50(0.61); Y51(1.32); Y59(0.53) | LDD0286 | [7] |
| LDCM0125 | JWB146 | DM93 | Y106(1.05); Y50(1.07); Y51(1.87); Y59(1.25) | LDD0287 | [7] |
| LDCM0126 | JWB150 | DM93 | Y106(2.48); Y50(3.41); Y51(2.57); Y59(2.36) | LDD0288 | [7] |
| LDCM0127 | JWB152 | DM93 | Y106(1.69); Y50(2.02); Y51(1.80); Y59(1.63) | LDD0289 | [7] |
| LDCM0128 | JWB198 | DM93 | Y106(0.93); Y50(0.95); Y51(1.01); Y59(1.15) | LDD0290 | [7] |
| LDCM0129 | JWB202 | DM93 | Y106(0.34); Y50(0.36); Y51(0.46); Y59(0.44) | LDD0291 | [7] |
| LDCM0130 | JWB211 | DM93 | Y106(0.87); Y50(1.07); Y51(0.92); Y59(0.92) | LDD0292 | [7] |
| LDCM0109 | NEM | HeLa | H37(0.00); H28(0.00) | LDD0223 | [18] |
| LDCM0014 | Panhematin | HEK-293T | 5.81 | LDD0062 | [6] |
| LDCM0090 | Rapamycin | JHH-7 | 3.07 | LDD0213 | [22] |
| LDCM0021 | THZ1 | HeLa S3 | C239(1.11); C211(1.04); C239(0.99); C201(1.03) | LDD0460 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Leucine-rich repeat serine/threonine-protein kinase 2 (LRRK2) | TKL Ser/Thr protein kinase family | Q5S007 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nuclear receptor subfamily 4immunitygroup A member 1 (NR4A1) | Nuclear hormone receptor family | P22736 | |||
References
