Details of the Target
General Information of Target
| Target ID | LDTP01100 | |||||
|---|---|---|---|---|---|---|
| Target Name | Iron-sulfur clusters transporter ABCB7, mitochondrial (ABCB7) | |||||
| Gene Name | ABCB7 | |||||
| Gene ID | 22 | |||||
| Synonyms |
ABC7; Iron-sulfur clusters transporter ABCB7, mitochondrial; ATP-binding cassette sub-family B member 7, mitochondrial; ATP-binding cassette transporter 7; ABC transporter 7 protein |
|||||
| 3D Structure | ||||||
| Sequence |
MALLAMHSWRWAAAAAAFEKRRHSAILIRPLVSVSGSGPQWRPHQLGALGTARAYQIPES
LKSITWQRLGKGNSGQFLDAAKALQVWPLIEKRTCWHGHAGGGLHTDPKEGLKDVDTRKI IKAMLSYVWPKDRPDLRARVAISLGFLGGAKAMNIVVPFMFKYAVDSLNQMSGNMLNLSD APNTVATMATAVLIGYGVSRAGAAFFNEVRNAVFGKVAQNSIRRIAKNVFLHLHNLDLGF HLSRQTGALSKAIDRGTRGISFVLSALVFNLLPIMFEVMLVSGVLYYKCGAQFALVTLGT LGTYTAFTVAVTRWRTRFRIEMNKADNDAGNAAIDSLLNYETVKYFNNERYEAQRYDGFL KTYETASLKSTSTLAMLNFGQSAIFSVGLTAIMVLASQGIVAGTLTVGDLVMVNGLLFQL SLPLNFLGTVYRETRQALIDMNTLFTLLKVDTQIKDKVMASPLQITPQTATVAFDNVHFE YIEGQKVLSGISFEVPAGKKVAIVGGSGSGKSTIVRLLFRFYEPQKGSIYLAGQNIQDVS LESLRRAVGVVPQDAVLFHNTIYYNLLYGNISASPEEVYAVAKLAGLHDAILRMPHGYDT QVGERGLKLSGGEKQRVAIARAILKDPPVILYDEATSSLDSITEETILGAMKDVVKHRTS IFIAHRLSTVVDADEIIVLDQGKVAERGTHHGLLANPHSIYSEMWHTQSSRVQNHDNPKW EAKKENISKEEERKKLQEEIVNSVKGCGNCSC |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
ABC transporter superfamily, ABCB family, Heavy Metal importer (TC 3.A.1.210) subfamily
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Exports glutathione-coordinated iron-sulfur clusters such as [2Fe-2S]-(GS)4 cluster from the mitochondria to the cytosol in an ATP-dependent manner allowing the assembly of the cytosolic iron-sulfur (Fe/S) cluster-containing proteins and participates in iron homeostasis. Moreover, through a functional complex formed of ABCB7, FECH and ABCB10, also plays a role in the cellular iron homeostasis, mitochondrial function and heme biosynthesis. In cardiomyocytes, regulates cellular iron homeostasis and cellular reactive oxygen species (ROS) levels through its interaction with COX4I1. May also play a role in hematopoiesis.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
N1 Probe Info |
 |
100.00 | LDD0242 | [1] | |
|
m-APA Probe Info |
 |
11.61 | LDD0403 | [2] | |
|
Acrolein Probe Info |
 |
H665(0.00); H596(0.00) | LDD0222 | [3] | |
|
DBIA Probe Info |
 |
C752(1.02) | LDD1492 | [4] | |
|
ATP probe Probe Info |
 |
K719(0.00); K511(0.00); K735(0.00) | LDD0199 | [5] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [6] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [6] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [3] | |
|
AOyne Probe Info |
 |
8.90 | LDD0443 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C056 Probe Info |
 |
26.35 | LDD1753 | [8] | |
|
C070 Probe Info |
 |
9.92 | LDD1766 | [8] | |
|
C091 Probe Info |
 |
13.45 | LDD1782 | [8] | |
|
C112 Probe Info |
 |
18.51 | LDD1799 | [8] | |
|
C134 Probe Info |
 |
25.63 | LDD1816 | [8] | |
|
C143 Probe Info |
 |
12.55 | LDD1825 | [8] | |
|
C158 Probe Info |
 |
19.16 | LDD1838 | [8] | |
|
C159 Probe Info |
 |
7.41 | LDD1839 | [8] | |
|
C161 Probe Info |
 |
10.85 | LDD1841 | [8] | |
|
C173 Probe Info |
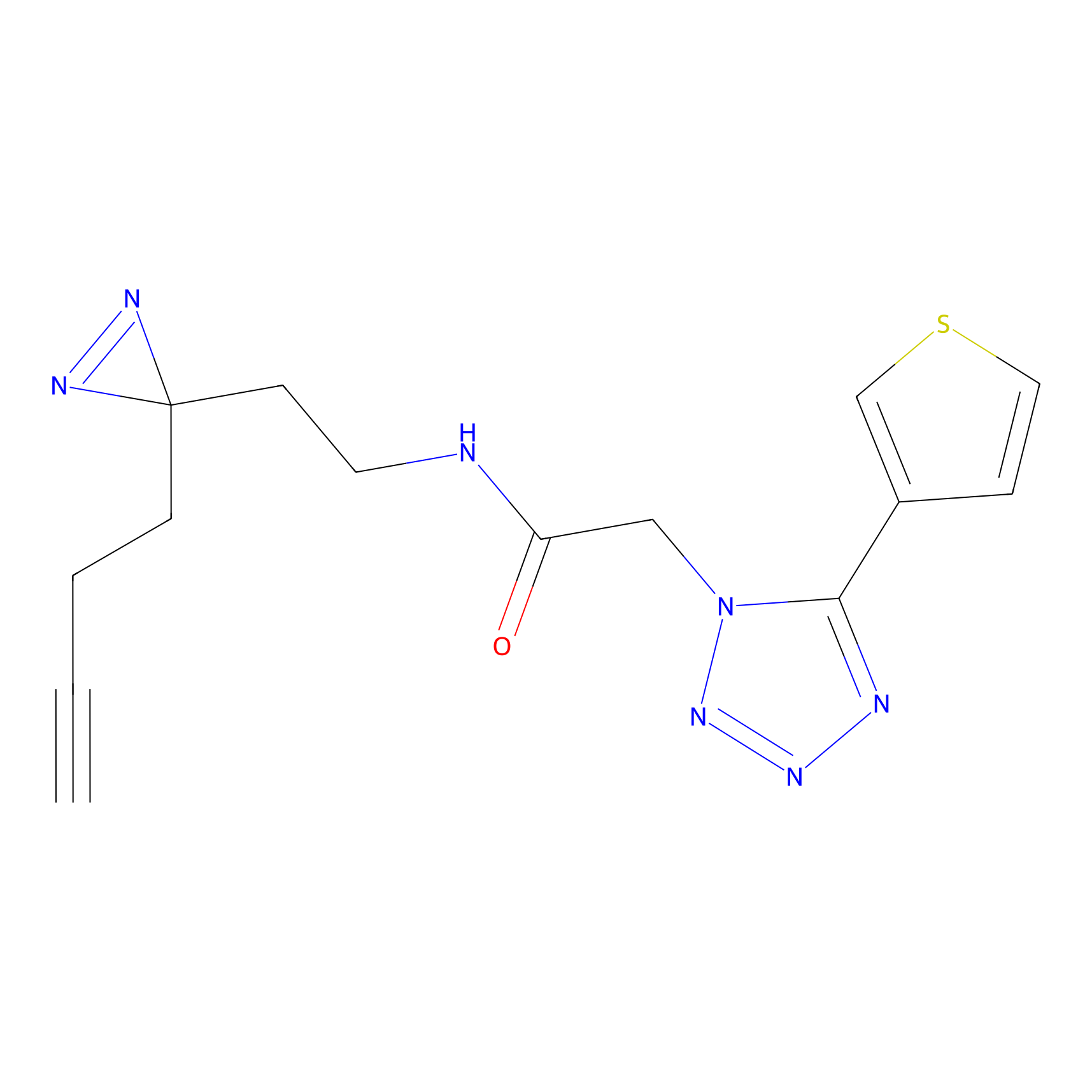 |
16.00 | LDD1853 | [8] | |
|
C198 Probe Info |
 |
23.92 | LDD1874 | [8] | |
|
C220 Probe Info |
 |
13.83 | LDD1894 | [8] | |
|
C235 Probe Info |
 |
16.91 | LDD1908 | [8] | |
|
C249 Probe Info |
 |
21.26 | LDD1922 | [8] | |
|
C284 Probe Info |
 |
51.63 | LDD1954 | [8] | |
|
C288 Probe Info |
 |
13.18 | LDD1958 | [8] | |
|
C289 Probe Info |
 |
26.54 | LDD1959 | [8] | |
|
C305 Probe Info |
 |
16.00 | LDD1974 | [8] | |
|
C338 Probe Info |
 |
14.93 | LDD2001 | [8] | |
|
C348 Probe Info |
 |
14.83 | LDD2009 | [8] | |
|
C349 Probe Info |
 |
9.25 | LDD2010 | [8] | |
|
C350 Probe Info |
 |
23.43 | LDD2011 | [8] | |
|
C356 Probe Info |
 |
11.47 | LDD2017 | [8] | |
|
C367 Probe Info |
 |
6.92 | LDD2028 | [8] | |
|
C382 Probe Info |
 |
12.21 | LDD2041 | [8] | |
|
C388 Probe Info |
 |
54.95 | LDD2047 | [8] | |
|
C390 Probe Info |
 |
45.57 | LDD2049 | [8] | |
|
C391 Probe Info |
 |
14.93 | LDD2050 | [8] | |
|
C394 Probe Info |
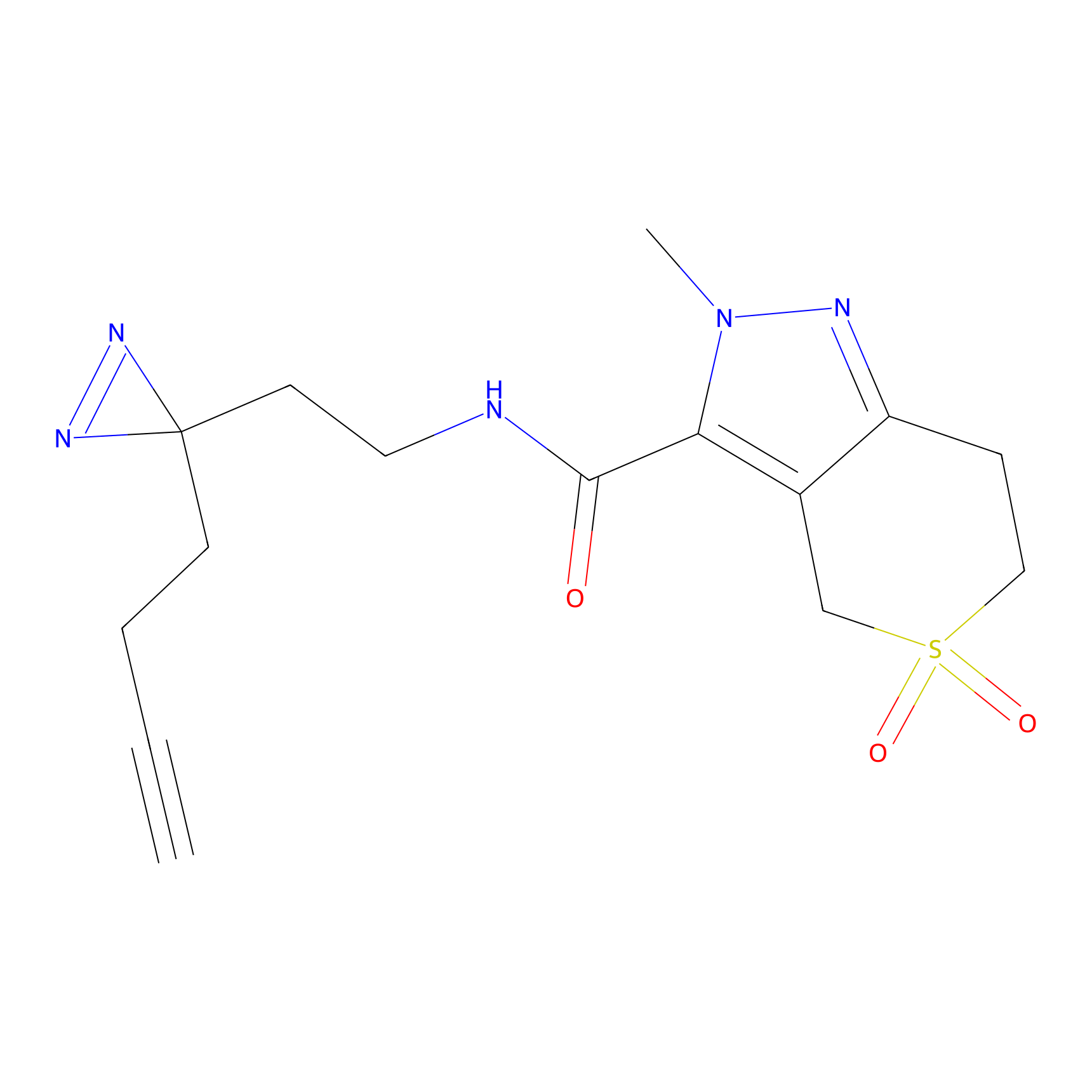 |
5.50 | LDD2053 | [8] | |
|
C413 Probe Info |
 |
16.68 | LDD2069 | [8] | |
|
FFF probe11 Probe Info |
 |
10.48 | LDD0471 | [9] | |
|
FFF probe13 Probe Info |
 |
16.91 | LDD0475 | [9] | |
|
FFF probe14 Probe Info |
 |
12.46 | LDD0477 | [9] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [9] | |
|
JN0003 Probe Info |
 |
16.95 | LDD0469 | [9] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [10] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [10] | |
|
OEA-DA Probe Info |
 |
4.65 | LDD0046 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0237 | AC12 | HEK-293T | C95(1.14) | LDD1510 | [4] |
| LDCM0280 | AC20 | HEK-293T | C95(0.95) | LDD1519 | [4] |
| LDCM0288 | AC28 | HEK-293T | C95(0.98) | LDD1527 | [4] |
| LDCM0297 | AC36 | HEK-293T | C95(1.10) | LDD1536 | [4] |
| LDCM0301 | AC4 | HEK-293T | C95(0.97) | LDD1540 | [4] |
| LDCM0306 | AC44 | HEK-293T | C95(1.02) | LDD1545 | [4] |
| LDCM0315 | AC52 | HEK-293T | C95(1.09) | LDD1554 | [4] |
| LDCM0324 | AC60 | HEK-293T | C95(1.06) | LDD1563 | [4] |
| LDCM0156 | Aniline | NCI-H1299 | 11.61 | LDD0403 | [2] |
| LDCM0108 | Chloroacetamide | HeLa | H665(0.00); H596(0.00) | LDD0222 | [3] |
| LDCM0408 | CL20 | HEK-293T | C95(1.13) | LDD1612 | [4] |
| LDCM0421 | CL32 | HEK-293T | C95(1.06) | LDD1625 | [4] |
| LDCM0434 | CL44 | HEK-293T | C95(1.01) | LDD1638 | [4] |
| LDCM0447 | CL56 | HEK-293T | C95(1.26) | LDD1650 | [4] |
| LDCM0460 | CL68 | HEK-293T | C95(1.33) | LDD1663 | [4] |
| LDCM0473 | CL8 | HEK-293T | C95(2.11) | LDD1676 | [4] |
| LDCM0474 | CL80 | HEK-293T | C95(1.40) | LDD1677 | [4] |
| LDCM0487 | CL92 | HEK-293T | C95(1.10) | LDD1690 | [4] |
| LDCM0022 | KB02 | HEK-293T | C752(1.02) | LDD1492 | [4] |
| LDCM0023 | KB03 | HEK-293T | C752(1.00) | LDD1497 | [4] |
| LDCM0024 | KB05 | HEK-293T | C752(1.07) | LDD1502 | [4] |
| LDCM0109 | NEM | HeLa | H588(0.00); H596(0.00) | LDD0223 | [3] |
References
