Details of the Target
General Information of Target
| Target ID | LDTP13591 | |||||
|---|---|---|---|---|---|---|
| Target Name | A-kinase anchor protein 8-like (AKAP8L) | |||||
| Gene Name | AKAP8L | |||||
| Gene ID | 26993 | |||||
| Synonyms |
NAKAP; NAKAP95; A-kinase anchor protein 8-like; AKAP8-like protein; Helicase A-binding protein 95; HAP95; Homologous to AKAP95 protein; HA95; Neighbor of A-kinase-anchoring protein 95; Neighbor of AKAP95
|
|||||
| 3D Structure | ||||||
| Sequence |
MPRPALSVTSFCHRLGKRERKQSFMGNSGNSWSHTPFPKLELGLGPQPMAPRELPTCSIC
LERLRDPISLDCGHDFCIRCFSTHRLPGCEPPCCPECRKICKQKRGLRSLGEKMKLLPQR PLPPALQETCPVRAEPLLLVRINASGGLILRMGAINRCLKHPLARDTPVCLLAVLGEQHS GKSFLLNHLLQGLPGLESGEGGRPRGGEASLQGCRWGANGLARGIWMWSHPFLLGKEGKK VAVFLVDTGDAMSPELSRETRIKLCALTTMLSSYQILSTSQELKDTDLDYLEMFVHVAEV MGKHYGMVPIQHLDLLVRDSSHPNKAGQGHVGNIFQRLSGRYPKVQELLQGKRARCCLLP APGRRRMNQGHASPGDTDDDFRHLLGAYVSDVLSAAPQHAKSRCQGYWNEGRAVARGDRR LLTGQQLAQEIKNLSGWMGRTGPGFTSPDEMAAQLHDLRKVEAAKREFEEYVRQQDVATK RIFSALRVLPDTMRNLLSTQKDAILARHGVALLCKGRDQTLEALEAELQATAKAFMDSYT MRFCGHLAAVGGAVGAGLMGLAGGVVGAGMAAAALAAEAGMVAAGAAVGATGAAVVGGGV GAGLAATVGCMEKEEDERLLEGDREPLLQEE |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
AKAP95 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Could play a role in constitutive transport element (CTE)-mediated gene expression by association with DHX9. Increases CTE-dependent nuclear unspliced mRNA export. Proposed to target PRKACA to the nucleus but does not seem to be implicated in the binding of regulatory subunit II of PKA. May be involved in nuclear envelope breakdown and chromatin condensation. May be involved in anchoring nuclear membranes to chromatin in interphase and in releasing membranes from chromating at mitosis. May regulate the initiation phase of DNA replication when associated with TMPO isoform Beta. Required for cell cycle G2/M transition and histone deacetylation during mitosis. In mitotic cells recruits HDAC3 to the vicinity of chromatin leading to deacetylation and subsequent phosphorylation at 'Ser-10' of histone H3; in this function seems to act redundantly with AKAP8. May be involved in regulation of pre-mRNA splicing.; (Microbial infection) In case of EBV infection, may target PRKACA to EBNA-LP-containing nuclear sites to modulate transcription from specific promoters.; (Microbial infection) Can synergize with DHX9 to activate the CTE-mediated gene expression of type D retroviruses.; (Microbial infection) In case of HIV-1 infection, involved in the DHX9-promoted annealing of host tRNA(Lys3) to viral genomic RNA as a primer in reverse transcription; in vitro negatively regulates DHX9 annealing activity.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CAL33 | SNV: p.K441R | . | |||
| CASKI | SNV: p.E556Q | DBIA Probe Info | |||
| CHL1 | SNV: p.T65N | DBIA Probe Info | |||
| HCT15 | SNV: p.G350R | DBIA Probe Info | |||
| HT115 | SNV: p.T398P | DBIA Probe Info | |||
| MCC13 | Substitution: p.L489F | DBIA Probe Info | |||
| MDAMB231 | SNV: p.Q372H | IA-alkyne Probe Info | |||
| MDAMB453 | SNV: p.Q24E | DBIA Probe Info | |||
| MEC1 | SNV: p.P219L | DBIA Probe Info | |||
| P31FUJ | SNV: p.S68Y | DBIA Probe Info | |||
| REH | SNV: p.M472T | DBIA Probe Info | |||
| RL952 | SNV: p.M514V | DBIA Probe Info | |||
| TOV21G | SNV: p.E555Ter | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K264(6.67); K439(0.81); K503(5.36); K519(1.43) | LDD0277 | [2] | |
|
Probe 1 Probe Info |
 |
Y435(9.02) | LDD3495 | [3] | |
|
BTD Probe Info |
 |
C211(1.00) | LDD2090 | [4] | |
|
Sulforaphane-probe2 Probe Info |
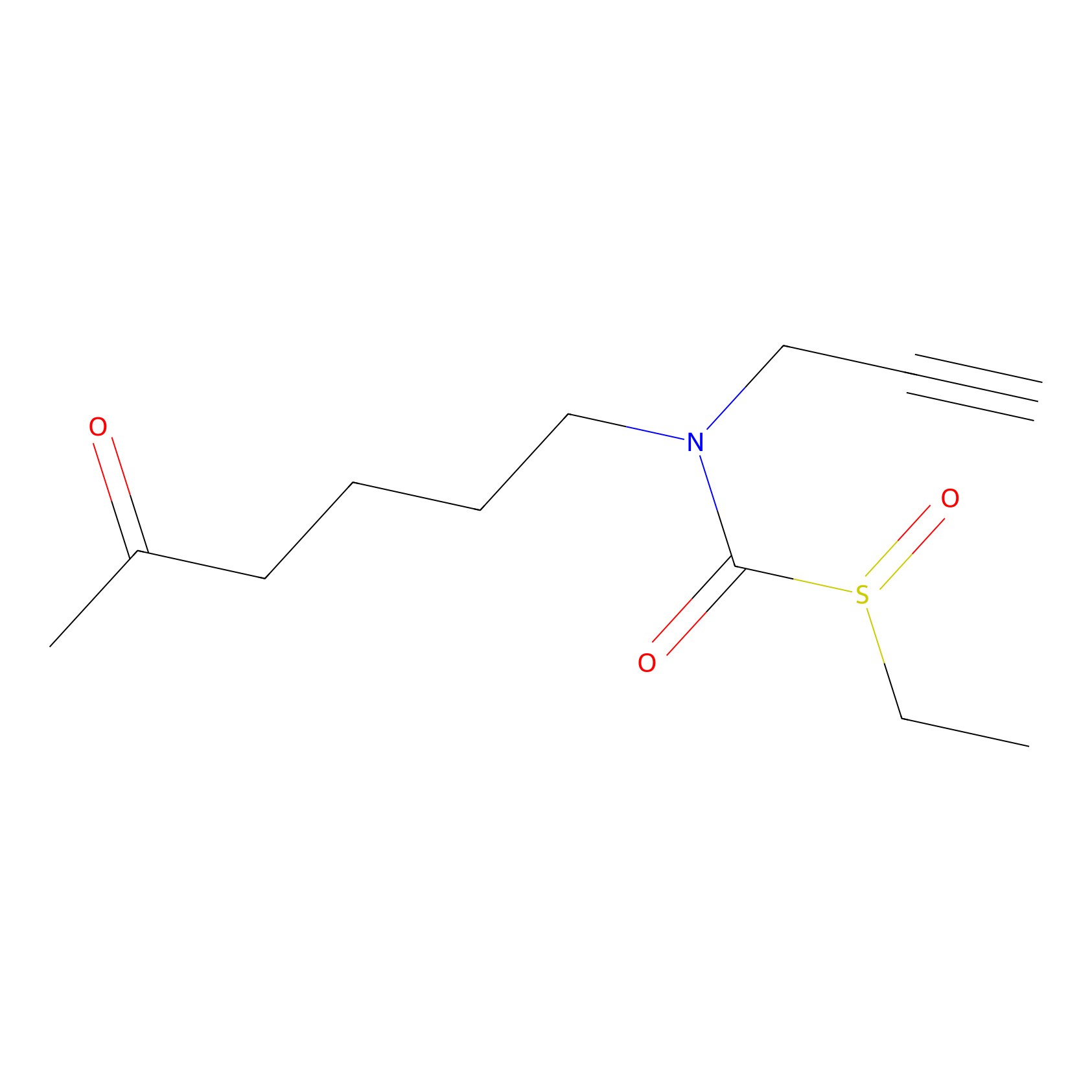 |
1.75 | LDD0160 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C128(2.97) | LDD0169 | [6] | |
|
HPAP Probe Info |
 |
3.06 | LDD0062 | [7] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [8] | |
|
DBIA Probe Info |
 |
C128(1.01); C211(0.85) | LDD0078 | [9] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C211(0.00); C128(0.00) | LDD0038 | [10] | |
|
IA-alkyne Probe Info |
 |
C211(0.00); C487(0.00); C484(0.00); C128(0.00) | LDD0036 | [10] | |
|
Lodoacetamide azide Probe Info |
 |
C128(0.00); C211(0.00) | LDD0037 | [10] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [11] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [11] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [12] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [12] | |
|
Compound 10 Probe Info |
 |
C128(0.00); C211(0.00) | LDD2216 | [13] | |
|
Compound 11 Probe Info |
 |
C128(0.00); C211(0.00) | LDD2213 | [13] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [12] | |
|
TFBX Probe Info |
 |
C128(0.00); C211(0.00) | LDD0148 | [14] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [12] | |
|
Phosphinate-6 Probe Info |
 |
C128(0.00); C211(0.00) | LDD0018 | [15] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [16] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [16] | |
|
Methacrolein Probe Info |
 |
C128(0.00); C211(0.00) | LDD0218 | [16] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [17] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [18] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C165 Probe Info |
 |
14.72 | LDD1845 | [19] | |
|
C206 Probe Info |
 |
15.24 | LDD1881 | [19] | |
|
C361 Probe Info |
 |
18.77 | LDD2022 | [19] | |
|
FFF probe11 Probe Info |
 |
13.14 | LDD0472 | [20] | |
|
FFF probe12 Probe Info |
 |
20.00 | LDD0473 | [20] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [20] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [20] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0465 | [20] | |
|
FFF probe6 Probe Info |
 |
5.55 | LDD0467 | [20] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C211(0.92) | LDD2117 | [4] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C211(1.03) | LDD2152 | [4] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C128(2.97) | LDD0169 | [6] |
| LDCM0214 | AC1 | HCT 116 | C128(0.94); C211(0.90); C391(1.10); C394(1.03) | LDD0531 | [9] |
| LDCM0215 | AC10 | HCT 116 | C128(1.02); C211(0.95); C394(1.00); C484(0.62) | LDD0532 | [9] |
| LDCM0216 | AC100 | HCT 116 | C128(0.95); C211(1.31); C391(0.99); C394(0.83) | LDD0533 | [9] |
| LDCM0217 | AC101 | HCT 116 | C128(1.08); C211(1.49); C391(0.96); C394(0.84) | LDD0534 | [9] |
| LDCM0218 | AC102 | HCT 116 | C128(1.12); C211(1.36); C391(0.86); C394(0.76) | LDD0535 | [9] |
| LDCM0219 | AC103 | HCT 116 | C128(1.41); C211(1.24); C391(0.93); C394(0.80) | LDD0536 | [9] |
| LDCM0220 | AC104 | HCT 116 | C128(1.09); C211(1.03); C391(1.11); C394(0.93) | LDD0537 | [9] |
| LDCM0221 | AC105 | HCT 116 | C128(1.17); C211(1.58); C391(0.97); C394(0.91) | LDD0538 | [9] |
| LDCM0222 | AC106 | HCT 116 | C128(1.40); C211(1.62); C391(0.93); C394(0.77) | LDD0539 | [9] |
| LDCM0223 | AC107 | HCT 116 | C128(1.21); C211(1.44); C391(0.97); C394(0.84) | LDD0540 | [9] |
| LDCM0224 | AC108 | HCT 116 | C128(1.29); C211(1.36); C391(1.19); C394(0.93) | LDD0541 | [9] |
| LDCM0225 | AC109 | HCT 116 | C128(1.33); C211(1.16); C391(1.23); C394(1.03) | LDD0542 | [9] |
| LDCM0226 | AC11 | HCT 116 | C128(1.36); C211(1.27); C394(0.96); C484(0.59) | LDD0543 | [9] |
| LDCM0227 | AC110 | HCT 116 | C128(1.25); C211(1.35); C391(0.85); C394(0.71) | LDD0544 | [9] |
| LDCM0228 | AC111 | HCT 116 | C128(1.22); C211(1.50); C391(0.99); C394(0.79) | LDD0545 | [9] |
| LDCM0229 | AC112 | HCT 116 | C128(1.20); C211(1.65); C391(0.98); C394(0.77) | LDD0546 | [9] |
| LDCM0230 | AC113 | HCT 116 | C128(1.07); C211(0.85); C394(1.01) | LDD0547 | [9] |
| LDCM0231 | AC114 | HCT 116 | C128(1.44); C211(0.80); C394(1.10) | LDD0548 | [9] |
| LDCM0232 | AC115 | HCT 116 | C128(1.50); C211(1.04); C394(1.00) | LDD0549 | [9] |
| LDCM0233 | AC116 | HCT 116 | C128(1.31); C211(0.85); C394(1.05) | LDD0550 | [9] |
| LDCM0234 | AC117 | HCT 116 | C128(1.31); C211(0.96); C394(0.96) | LDD0551 | [9] |
| LDCM0235 | AC118 | HCT 116 | C128(1.35); C211(0.87); C394(1.09) | LDD0552 | [9] |
| LDCM0236 | AC119 | HCT 116 | C128(1.28); C211(0.79); C394(1.00) | LDD0553 | [9] |
| LDCM0237 | AC12 | HCT 116 | C128(1.32); C211(0.99); C394(1.04); C484(0.65) | LDD0554 | [9] |
| LDCM0238 | AC120 | HCT 116 | C128(1.22); C211(0.96); C394(1.06) | LDD0555 | [9] |
| LDCM0239 | AC121 | HCT 116 | C128(1.35); C211(0.73); C394(0.94) | LDD0556 | [9] |
| LDCM0240 | AC122 | HCT 116 | C128(1.28); C211(0.78); C394(1.07) | LDD0557 | [9] |
| LDCM0241 | AC123 | HCT 116 | C128(1.32); C211(0.77); C394(0.92) | LDD0558 | [9] |
| LDCM0242 | AC124 | HCT 116 | C128(1.20); C211(0.81); C394(0.96) | LDD0559 | [9] |
| LDCM0243 | AC125 | HCT 116 | C128(1.38); C211(0.90); C394(1.22) | LDD0560 | [9] |
| LDCM0244 | AC126 | HCT 116 | C128(1.68); C211(0.67); C394(1.05) | LDD0561 | [9] |
| LDCM0245 | AC127 | HCT 116 | C128(1.59); C211(0.90); C394(1.02) | LDD0562 | [9] |
| LDCM0246 | AC128 | HCT 116 | C128(0.80); C211(0.64) | LDD0563 | [9] |
| LDCM0247 | AC129 | HCT 116 | C128(0.76); C211(0.75) | LDD0564 | [9] |
| LDCM0249 | AC130 | HCT 116 | C128(0.91); C211(0.80) | LDD0566 | [9] |
| LDCM0250 | AC131 | HCT 116 | C128(0.75); C211(0.71) | LDD0567 | [9] |
| LDCM0251 | AC132 | HCT 116 | C128(0.73); C211(0.87) | LDD0568 | [9] |
| LDCM0252 | AC133 | HCT 116 | C128(0.74); C211(0.76) | LDD0569 | [9] |
| LDCM0253 | AC134 | HCT 116 | C128(0.94); C211(0.87) | LDD0570 | [9] |
| LDCM0254 | AC135 | HCT 116 | C128(0.90); C211(0.70) | LDD0571 | [9] |
| LDCM0255 | AC136 | HCT 116 | C128(0.76); C211(0.89) | LDD0572 | [9] |
| LDCM0256 | AC137 | HCT 116 | C128(0.76); C211(0.86) | LDD0573 | [9] |
| LDCM0257 | AC138 | HCT 116 | C128(0.87); C211(0.73) | LDD0574 | [9] |
| LDCM0258 | AC139 | HCT 116 | C128(0.84); C211(0.90) | LDD0575 | [9] |
| LDCM0259 | AC14 | HCT 116 | C128(0.86); C211(0.91); C394(1.66); C484(0.63) | LDD0576 | [9] |
| LDCM0260 | AC140 | HCT 116 | C128(0.86); C211(1.05) | LDD0577 | [9] |
| LDCM0261 | AC141 | HCT 116 | C128(0.92); C211(0.95) | LDD0578 | [9] |
| LDCM0262 | AC142 | HCT 116 | C128(0.69); C211(0.81) | LDD0579 | [9] |
| LDCM0263 | AC143 | HCT 116 | C128(1.14); C211(1.28) | LDD0580 | [9] |
| LDCM0264 | AC144 | HCT 116 | C128(1.23); C211(1.37) | LDD0581 | [9] |
| LDCM0265 | AC145 | HCT 116 | C128(1.29); C211(1.30) | LDD0582 | [9] |
| LDCM0266 | AC146 | HCT 116 | C128(1.42); C211(1.43) | LDD0583 | [9] |
| LDCM0267 | AC147 | HCT 116 | C128(1.16); C211(1.43) | LDD0584 | [9] |
| LDCM0268 | AC148 | HCT 116 | C128(1.52); C211(1.60) | LDD0585 | [9] |
| LDCM0269 | AC149 | HCT 116 | C211(1.44); C128(1.63) | LDD0586 | [9] |
| LDCM0270 | AC15 | HCT 116 | C484(0.58); C487(0.58); C211(1.11); C128(1.11) | LDD0587 | [9] |
| LDCM0271 | AC150 | HCT 116 | C128(1.05); C211(1.31) | LDD0588 | [9] |
| LDCM0272 | AC151 | HCT 116 | C128(1.17); C211(1.50) | LDD0589 | [9] |
| LDCM0273 | AC152 | HCT 116 | C128(1.21); C211(1.62) | LDD0590 | [9] |
| LDCM0274 | AC153 | HCT 116 | C128(1.99); C211(2.34) | LDD0591 | [9] |
| LDCM0621 | AC154 | HCT 116 | C128(1.20); C211(1.66) | LDD2158 | [9] |
| LDCM0622 | AC155 | HCT 116 | C128(1.06); C211(1.84) | LDD2159 | [9] |
| LDCM0623 | AC156 | HCT 116 | C128(1.12); C211(1.70) | LDD2160 | [9] |
| LDCM0624 | AC157 | HCT 116 | C128(1.41); C211(1.42) | LDD2161 | [9] |
| LDCM0276 | AC17 | HCT 116 | C211(0.89); C484(0.96); C487(0.96); C394(1.05) | LDD0593 | [9] |
| LDCM0277 | AC18 | HCT 116 | C484(0.82); C487(0.82); C211(0.93); C394(1.01) | LDD0594 | [9] |
| LDCM0278 | AC19 | HCT 116 | C394(0.96); C211(1.14); C484(1.17); C487(1.17) | LDD0595 | [9] |
| LDCM0279 | AC2 | HCT 116 | C484(0.84); C487(0.84); C211(0.93); C128(1.12) | LDD0596 | [9] |
| LDCM0280 | AC20 | HCT 116 | C211(0.88); C394(0.88); C484(1.10); C487(1.10) | LDD0597 | [9] |
| LDCM0281 | AC21 | HCT 116 | C211(0.85); C394(0.89); C484(1.07); C487(1.07) | LDD0598 | [9] |
| LDCM0282 | AC22 | HCT 116 | C394(0.82); C211(0.99); C128(1.37); C484(1.40) | LDD0599 | [9] |
| LDCM0283 | AC23 | HCT 116 | C394(0.89); C211(0.98); C484(1.13); C487(1.13) | LDD0600 | [9] |
| LDCM0284 | AC24 | HCT 116 | C211(1.00); C128(1.08); C394(1.15); C484(1.16) | LDD0601 | [9] |
| LDCM0285 | AC25 | HCT 116 | C394(1.13); C211(1.18); C128(1.46) | LDD0602 | [9] |
| LDCM0286 | AC26 | HCT 116 | C394(0.80); C211(1.29); C128(1.51) | LDD0603 | [9] |
| LDCM0287 | AC27 | HCT 116 | C394(0.85); C211(1.06); C128(1.68) | LDD0604 | [9] |
| LDCM0288 | AC28 | HCT 116 | C394(0.86); C211(1.41); C128(2.06) | LDD0605 | [9] |
| LDCM0289 | AC29 | HCT 116 | C394(0.63); C211(1.60); C128(2.39) | LDD0606 | [9] |
| LDCM0290 | AC3 | HCT 116 | C484(0.75); C487(0.75); C128(0.87); C211(0.91) | LDD0607 | [9] |
| LDCM0291 | AC30 | HCT 116 | C394(0.98); C211(1.51); C128(1.68) | LDD0608 | [9] |
| LDCM0292 | AC31 | HCT 116 | C211(1.15); C394(1.41); C128(1.56) | LDD0609 | [9] |
| LDCM0293 | AC32 | HCT 116 | C394(1.14); C128(1.52); C211(1.57) | LDD0610 | [9] |
| LDCM0294 | AC33 | HCT 116 | C394(1.08); C211(1.16); C128(2.24) | LDD0611 | [9] |
| LDCM0295 | AC34 | HCT 116 | C394(0.69); C211(1.59); C128(2.52) | LDD0612 | [9] |
| LDCM0296 | AC35 | HCT 116 | C394(0.90); C128(1.07); C484(1.11); C487(1.11) | LDD0613 | [9] |
| LDCM0297 | AC36 | HCT 116 | C394(0.81); C128(0.97); C484(1.02); C487(1.02) | LDD0614 | [9] |
| LDCM0298 | AC37 | HCT 116 | C394(0.77); C128(0.95); C484(0.97); C487(0.97) | LDD0615 | [9] |
| LDCM0299 | AC38 | HCT 116 | C394(0.84); C484(1.03); C487(1.03); C128(1.12) | LDD0616 | [9] |
| LDCM0300 | AC39 | HCT 116 | C394(0.67); C484(0.78); C487(0.78); C128(1.02) | LDD0617 | [9] |
| LDCM0301 | AC4 | HCT 116 | C484(0.66); C487(0.66); C211(0.90); C128(0.90) | LDD0618 | [9] |
| LDCM0302 | AC40 | HCT 116 | C394(0.53); C484(1.05); C487(1.05); C128(1.11) | LDD0619 | [9] |
| LDCM0303 | AC41 | HCT 116 | C394(0.75); C484(0.83); C487(0.83); C128(1.06) | LDD0620 | [9] |
| LDCM0304 | AC42 | HCT 116 | C484(0.83); C487(0.83); C394(0.86); C128(1.05) | LDD0621 | [9] |
| LDCM0305 | AC43 | HCT 116 | C484(0.81); C487(0.81); C394(0.82); C128(1.05) | LDD0622 | [9] |
| LDCM0306 | AC44 | HCT 116 | C394(0.55); C484(0.92); C487(0.92); C128(1.01) | LDD0623 | [9] |
| LDCM0307 | AC45 | HCT 116 | C394(0.52); C484(0.81); C487(0.81); C128(1.24) | LDD0624 | [9] |
| LDCM0308 | AC46 | HCT 116 | C484(0.78); C487(0.78); C394(0.88); C391(1.02) | LDD0625 | [9] |
| LDCM0309 | AC47 | HCT 116 | C484(0.84); C487(0.84); C394(0.84); C391(0.94) | LDD0626 | [9] |
| LDCM0310 | AC48 | HCT 116 | C484(0.78); C487(0.78); C211(0.82); C394(0.88) | LDD0627 | [9] |
| LDCM0311 | AC49 | HCT 116 | C484(0.80); C487(0.80); C394(0.85); C391(0.90) | LDD0628 | [9] |
| LDCM0312 | AC5 | HCT 116 | C484(0.65); C487(0.65); C211(0.75); C128(0.88) | LDD0629 | [9] |
| LDCM0313 | AC50 | HCT 116 | C391(0.90); C394(0.91); C484(0.95); C487(0.95) | LDD0630 | [9] |
| LDCM0314 | AC51 | HCT 116 | C211(0.86); C484(0.98); C487(0.98); C128(1.17) | LDD0631 | [9] |
| LDCM0315 | AC52 | HCT 116 | C484(0.88); C487(0.88); C211(0.92); C394(1.01) | LDD0632 | [9] |
| LDCM0316 | AC53 | HCT 116 | C484(0.86); C487(0.86); C394(0.88); C391(1.03) | LDD0633 | [9] |
| LDCM0317 | AC54 | HCT 116 | C211(0.85); C394(0.86); C391(0.96); C484(1.03) | LDD0634 | [9] |
| LDCM0318 | AC55 | HCT 116 | C394(0.73); C391(0.90); C484(0.90); C487(0.90) | LDD0635 | [9] |
| LDCM0319 | AC56 | HCT 116 | C394(0.66); C391(0.78); C211(0.83); C484(0.88) | LDD0636 | [9] |
| LDCM0320 | AC57 | HCT 116 | C484(0.52); C487(0.52); C394(0.83); C391(0.86) | LDD0637 | [9] |
| LDCM0321 | AC58 | HCT 116 | C484(0.56); C487(0.56); C211(1.07); C391(1.12) | LDD0638 | [9] |
| LDCM0322 | AC59 | HCT 116 | C484(0.52); C487(0.52); C391(0.99); C394(1.02) | LDD0639 | [9] |
| LDCM0323 | AC6 | HCT 116 | C484(0.72); C487(0.72); C211(0.97); C394(1.14) | LDD0640 | [9] |
| LDCM0324 | AC60 | HCT 116 | C484(0.50); C487(0.50); C394(0.86); C391(0.88) | LDD0641 | [9] |
| LDCM0325 | AC61 | HCT 116 | C484(0.54); C487(0.54); C128(0.88); C211(1.00) | LDD0642 | [9] |
| LDCM0326 | AC62 | HCT 116 | C484(0.62); C487(0.62); C394(0.92); C391(0.92) | LDD0643 | [9] |
| LDCM0327 | AC63 | HCT 116 | C484(0.48); C487(0.48); C394(0.90); C391(0.92) | LDD0644 | [9] |
| LDCM0328 | AC64 | HCT 116 | C484(0.58); C487(0.58); C394(1.03); C391(1.07) | LDD0645 | [9] |
| LDCM0329 | AC65 | HCT 116 | C484(0.45); C487(0.45); C128(0.68); C394(1.05) | LDD0646 | [9] |
| LDCM0330 | AC66 | HCT 116 | C484(0.43); C487(0.43); C128(0.60); C391(1.11) | LDD0647 | [9] |
| LDCM0331 | AC67 | HCT 116 | C484(0.37); C487(0.37); C128(0.83); C394(0.95) | LDD0648 | [9] |
| LDCM0332 | AC68 | HCT 116 | C484(0.96); C487(0.96); C394(1.03); C391(1.04) | LDD0649 | [9] |
| LDCM0333 | AC69 | HCT 116 | C394(1.04); C484(1.05); C487(1.05); C391(1.08) | LDD0650 | [9] |
| LDCM0334 | AC7 | HCT 116 | C484(0.79); C487(0.79); C128(0.87); C211(0.91) | LDD0651 | [9] |
| LDCM0335 | AC70 | HCT 116 | C484(0.86); C487(0.86); C391(1.08); C394(1.12) | LDD0652 | [9] |
| LDCM0336 | AC71 | HCT 116 | C484(0.97); C487(0.97); C128(1.09); C391(1.10) | LDD0653 | [9] |
| LDCM0337 | AC72 | HCT 116 | C394(1.00); C391(1.08); C484(1.09); C487(1.09) | LDD0654 | [9] |
| LDCM0338 | AC73 | HCT 116 | C391(0.92); C394(0.98); C484(1.36); C487(1.36) | LDD0655 | [9] |
| LDCM0339 | AC74 | HCT 116 | C391(1.08); C394(1.14); C484(1.27); C487(1.27) | LDD0656 | [9] |
| LDCM0340 | AC75 | HCT 116 | C394(0.87); C391(0.94); C484(1.28); C487(1.28) | LDD0657 | [9] |
| LDCM0341 | AC76 | HCT 116 | C391(0.91); C394(1.08); C484(1.14); C487(1.14) | LDD0658 | [9] |
| LDCM0342 | AC77 | HCT 116 | C394(0.88); C391(0.90); C484(1.08); C487(1.08) | LDD0659 | [9] |
| LDCM0343 | AC78 | HCT 116 | C391(0.88); C394(1.03); C484(1.05); C487(1.05) | LDD0660 | [9] |
| LDCM0344 | AC79 | HCT 116 | C391(0.94); C394(0.99); C484(1.00); C487(1.00) | LDD0661 | [9] |
| LDCM0345 | AC8 | HCT 116 | C484(0.66); C487(0.66); C211(1.20); C394(1.20) | LDD0662 | [9] |
| LDCM0346 | AC80 | HCT 116 | C391(0.98); C484(1.03); C487(1.03); C394(1.08) | LDD0663 | [9] |
| LDCM0347 | AC81 | HCT 116 | C484(0.75); C487(0.75); C391(0.92); C394(0.95) | LDD0664 | [9] |
| LDCM0348 | AC82 | HCT 116 | C394(1.09); C391(1.11); C484(1.21); C487(1.21) | LDD0665 | [9] |
| LDCM0349 | AC83 | HCT 116 | C484(0.68); C487(0.68); C394(1.02); C211(1.70) | LDD0666 | [9] |
| LDCM0350 | AC84 | HCT 116 | C484(0.59); C487(0.59); C394(1.11); C128(1.57) | LDD0667 | [9] |
| LDCM0351 | AC85 | HCT 116 | C484(0.60); C487(0.60); C394(1.01); C211(1.28) | LDD0668 | [9] |
| LDCM0352 | AC86 | HCT 116 | C484(0.66); C487(0.66); C394(0.96); C128(1.23) | LDD0669 | [9] |
| LDCM0353 | AC87 | HCT 116 | C484(0.73); C487(0.73); C211(0.89); C394(1.02) | LDD0670 | [9] |
| LDCM0354 | AC88 | HCT 116 | C484(0.84); C487(0.84); C394(0.96); C211(1.28) | LDD0671 | [9] |
| LDCM0355 | AC89 | HCT 116 | C484(0.70); C487(0.70); C394(0.98); C211(1.43) | LDD0672 | [9] |
| LDCM0357 | AC90 | HCT 116 | C484(0.85); C487(0.85); C211(0.99); C394(1.01) | LDD0674 | [9] |
| LDCM0358 | AC91 | HCT 116 | C484(0.69); C487(0.69); C394(1.02); C211(1.92) | LDD0675 | [9] |
| LDCM0359 | AC92 | HCT 116 | C484(0.59); C487(0.59); C394(0.98); C128(1.72) | LDD0676 | [9] |
| LDCM0360 | AC93 | HCT 116 | C484(0.80); C487(0.80); C394(0.90); C211(1.22) | LDD0677 | [9] |
| LDCM0361 | AC94 | HCT 116 | C484(0.93); C487(0.93); C394(0.97); C211(1.30) | LDD0678 | [9] |
| LDCM0362 | AC95 | HCT 116 | C484(0.68); C487(0.68); C394(1.15); C211(1.28) | LDD0679 | [9] |
| LDCM0363 | AC96 | HCT 116 | C484(0.87); C487(0.87); C394(1.05); C128(1.75) | LDD0680 | [9] |
| LDCM0364 | AC97 | HCT 116 | C484(0.65); C487(0.65); C394(0.88); C211(1.67) | LDD0681 | [9] |
| LDCM0365 | AC98 | HCT 116 | C394(0.84); C391(1.00); C211(1.91); C128(1.98) | LDD0682 | [9] |
| LDCM0366 | AC99 | HCT 116 | C394(0.80); C391(0.91); C128(1.03); C211(1.15) | LDD0683 | [9] |
| LDCM0248 | AKOS034007472 | HCT 116 | C128(0.88); C211(0.80); C394(1.20); C484(0.61) | LDD0565 | [9] |
| LDCM0356 | AKOS034007680 | HCT 116 | C484(0.77); C487(0.77); C211(0.92); C394(0.95) | LDD0673 | [9] |
| LDCM0275 | AKOS034007705 | HCT 116 | C484(0.50); C487(0.50); C211(1.10); C394(1.41) | LDD0592 | [9] |
| LDCM0156 | Aniline | NCI-H1299 | 11.78 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C128(1.01); C211(0.85) | LDD0078 | [9] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [16] |
| LDCM0632 | CL-Sc | Hep-G2 | C128(20.00) | LDD2227 | [11] |
| LDCM0367 | CL1 | HCT 116 | C128(0.94); C394(1.10); C391(1.16); C484(1.21) | LDD0684 | [9] |
| LDCM0368 | CL10 | HCT 116 | C484(0.82); C487(0.82); C211(0.97); C394(1.24) | LDD0685 | [9] |
| LDCM0369 | CL100 | HCT 116 | C484(0.66); C487(0.66); C394(0.85); C391(0.94) | LDD0686 | [9] |
| LDCM0370 | CL101 | HCT 116 | C484(0.62); C487(0.62); C128(0.98); C211(1.09) | LDD0687 | [9] |
| LDCM0371 | CL102 | HCT 116 | C484(0.73); C487(0.73); C128(0.87); C211(1.13) | LDD0688 | [9] |
| LDCM0372 | CL103 | HCT 116 | C484(0.84); C487(0.84); C211(0.96); C128(0.97) | LDD0689 | [9] |
| LDCM0373 | CL104 | HCT 116 | C484(0.75); C487(0.75); C128(0.92); C211(1.06) | LDD0690 | [9] |
| LDCM0374 | CL105 | HCT 116 | C394(0.83); C211(0.88); C484(1.01); C487(1.01) | LDD0691 | [9] |
| LDCM0375 | CL106 | HCT 116 | C211(0.63); C484(0.81); C487(0.81); C394(0.89) | LDD0692 | [9] |
| LDCM0376 | CL107 | HCT 116 | C484(0.77); C487(0.77); C394(0.85); C211(1.02) | LDD0693 | [9] |
| LDCM0377 | CL108 | HCT 116 | C484(0.75); C487(0.75); C211(0.89); C394(1.10) | LDD0694 | [9] |
| LDCM0378 | CL109 | HCT 116 | C394(0.80); C484(0.96); C487(0.96); C211(1.20) | LDD0695 | [9] |
| LDCM0379 | CL11 | HCT 116 | C484(0.85); C487(0.85); C394(1.10); C391(1.26) | LDD0696 | [9] |
| LDCM0380 | CL110 | HCT 116 | C484(0.87); C487(0.87); C211(0.93); C394(1.01) | LDD0697 | [9] |
| LDCM0381 | CL111 | HCT 116 | C394(0.81); C211(0.90); C484(0.94); C487(0.94) | LDD0698 | [9] |
| LDCM0382 | CL112 | HCT 116 | C394(1.01); C128(1.22); C211(1.85) | LDD0699 | [9] |
| LDCM0383 | CL113 | HCT 116 | C394(0.98); C211(1.59); C128(1.92) | LDD0700 | [9] |
| LDCM0384 | CL114 | HCT 116 | C394(0.97); C128(1.81); C211(1.93) | LDD0701 | [9] |
| LDCM0385 | CL115 | HCT 116 | C394(0.81); C211(1.26); C128(1.79) | LDD0702 | [9] |
| LDCM0386 | CL116 | HCT 116 | C211(0.97); C394(1.17); C128(1.80) | LDD0703 | [9] |
| LDCM0387 | CL117 | HCT 116 | C394(0.66); C484(0.78); C487(0.78); C211(1.64) | LDD0704 | [9] |
| LDCM0388 | CL118 | HCT 116 | C484(0.76); C487(0.76); C394(0.86); C128(0.99) | LDD0705 | [9] |
| LDCM0389 | CL119 | HCT 116 | C394(0.71); C484(0.99); C487(0.99); C128(1.18) | LDD0706 | [9] |
| LDCM0390 | CL12 | HCT 116 | C484(0.79); C487(0.79); C394(0.98); C211(1.07) | LDD0707 | [9] |
| LDCM0391 | CL120 | HCT 116 | C484(0.79); C487(0.79); C394(0.81); C128(1.01) | LDD0708 | [9] |
| LDCM0392 | CL121 | HCT 116 | C211(0.91); C128(1.06); C391(1.15); C394(1.30) | LDD0709 | [9] |
| LDCM0393 | CL122 | HCT 116 | C484(0.82); C487(0.82); C394(0.93); C391(0.95) | LDD0710 | [9] |
| LDCM0394 | CL123 | HCT 116 | C394(0.73); C484(0.81); C487(0.81); C391(0.89) | LDD0711 | [9] |
| LDCM0395 | CL124 | HCT 116 | C394(0.73); C484(0.89); C487(0.89); C391(0.95) | LDD0712 | [9] |
| LDCM0396 | CL125 | HCT 116 | C484(0.67); C487(0.67); C128(0.86); C394(0.99) | LDD0713 | [9] |
| LDCM0397 | CL126 | HCT 116 | C128(0.85); C484(0.94); C487(0.94); C391(1.05) | LDD0714 | [9] |
| LDCM0398 | CL127 | HCT 116 | C484(0.75); C487(0.75); C128(0.83); C394(0.99) | LDD0715 | [9] |
| LDCM0399 | CL128 | HCT 116 | C484(0.61); C487(0.61); C394(1.02); C391(1.04) | LDD0716 | [9] |
| LDCM0400 | CL13 | HCT 116 | C484(0.85); C487(0.85); C211(1.20); C391(1.24) | LDD0717 | [9] |
| LDCM0401 | CL14 | HCT 116 | C484(0.99); C487(0.99); C211(1.06); C391(1.10) | LDD0718 | [9] |
| LDCM0402 | CL15 | HCT 116 | C484(0.85); C487(0.85); C128(1.34); C391(1.42) | LDD0719 | [9] |
| LDCM0403 | CL16 | HCT 116 | C211(0.73); C391(0.86); C394(0.86); C484(1.17) | LDD0720 | [9] |
| LDCM0404 | CL17 | HCT 116 | C211(0.29); C484(0.99); C487(0.99); C128(1.03) | LDD0721 | [9] |
| LDCM0405 | CL18 | HCT 116 | C211(0.42); C484(0.88); C487(0.88); C394(0.90) | LDD0722 | [9] |
| LDCM0406 | CL19 | HCT 116 | C211(0.37); C484(0.97); C487(0.97); C394(1.02) | LDD0723 | [9] |
| LDCM0407 | CL2 | HCT 116 | C128(0.94); C391(0.95); C394(1.06); C484(1.07) | LDD0724 | [9] |
| LDCM0408 | CL20 | HCT 116 | C211(0.43); C484(1.02); C487(1.02); C394(1.19) | LDD0725 | [9] |
| LDCM0409 | CL21 | HCT 116 | C211(0.59); C484(0.73); C487(0.73); C394(1.30) | LDD0726 | [9] |
| LDCM0410 | CL22 | HCT 116 | C211(0.30); C484(0.79); C487(0.79); C394(1.74) | LDD0727 | [9] |
| LDCM0411 | CL23 | HCT 116 | C211(0.35); C394(0.70); C484(0.85); C487(0.85) | LDD0728 | [9] |
| LDCM0412 | CL24 | HCT 116 | C211(0.50); C394(0.58); C484(0.96); C487(0.96) | LDD0729 | [9] |
| LDCM0413 | CL25 | HCT 116 | C128(1.49); C211(0.53); C394(0.68); C484(0.89) | LDD0730 | [9] |
| LDCM0414 | CL26 | HCT 116 | C128(1.27); C211(0.48); C394(0.84); C484(0.97) | LDD0731 | [9] |
| LDCM0415 | CL27 | HCT 116 | C128(1.32); C211(0.30); C394(1.00); C484(0.95) | LDD0732 | [9] |
| LDCM0416 | CL28 | HCT 116 | C128(1.87); C211(0.42); C394(0.98); C484(0.82) | LDD0733 | [9] |
| LDCM0417 | CL29 | HCT 116 | C128(1.66); C211(0.36); C394(0.93); C484(0.80) | LDD0734 | [9] |
| LDCM0418 | CL3 | HCT 116 | C128(1.10); C211(1.14); C391(1.27); C394(1.29) | LDD0735 | [9] |
| LDCM0419 | CL30 | HCT 116 | C128(1.30); C211(0.38); C394(1.11); C484(1.23) | LDD0736 | [9] |
| LDCM0420 | CL31 | HCT 116 | C128(1.19); C211(1.41); C484(0.91); C487(0.91) | LDD0737 | [9] |
| LDCM0421 | CL32 | HCT 116 | C128(1.29); C211(1.23); C391(0.87); C394(0.88) | LDD0738 | [9] |
| LDCM0422 | CL33 | HCT 116 | C128(1.37); C211(2.27); C391(0.83); C394(0.80) | LDD0739 | [9] |
| LDCM0423 | CL34 | HCT 116 | C128(1.44); C211(1.26); C391(0.96); C394(0.91) | LDD0740 | [9] |
| LDCM0424 | CL35 | HCT 116 | C128(1.96); C211(1.13); C391(0.97); C394(0.90) | LDD0741 | [9] |
| LDCM0425 | CL36 | HCT 116 | C128(1.43); C211(1.28); C391(1.18); C394(1.35) | LDD0742 | [9] |
| LDCM0426 | CL37 | HCT 116 | C128(1.77); C211(1.29); C391(0.98); C394(1.01) | LDD0743 | [9] |
| LDCM0428 | CL39 | HCT 116 | C128(1.66); C211(1.22); C391(0.85); C394(0.81) | LDD0745 | [9] |
| LDCM0429 | CL4 | HCT 116 | C128(1.23); C211(2.35); C391(1.08); C394(1.30) | LDD0746 | [9] |
| LDCM0430 | CL40 | HCT 116 | C128(1.58); C211(2.64); C391(0.93); C394(1.04) | LDD0747 | [9] |
| LDCM0431 | CL41 | HCT 116 | C128(1.53); C211(1.44); C391(1.10); C394(1.06) | LDD0748 | [9] |
| LDCM0432 | CL42 | HCT 116 | C128(2.64); C211(1.27); C391(1.30); C394(1.43) | LDD0749 | [9] |
| LDCM0433 | CL43 | HCT 116 | C128(1.97); C211(1.49); C391(1.09); C394(1.22) | LDD0750 | [9] |
| LDCM0434 | CL44 | HCT 116 | C128(1.45); C211(1.17); C391(1.21); C394(1.44) | LDD0751 | [9] |
| LDCM0435 | CL45 | HCT 116 | C128(1.92); C211(1.40); C391(1.11); C394(1.20) | LDD0752 | [9] |
| LDCM0436 | CL46 | HCT 116 | C128(0.88); C211(0.85); C391(0.96); C394(0.90) | LDD0753 | [9] |
| LDCM0437 | CL47 | HCT 116 | C128(0.81); C211(0.77); C391(1.27); C394(1.23) | LDD0754 | [9] |
| LDCM0438 | CL48 | HCT 116 | C128(0.84); C211(0.90); C391(1.17); C394(1.31) | LDD0755 | [9] |
| LDCM0439 | CL49 | HCT 116 | C128(0.78); C211(0.92); C391(1.34); C394(1.42) | LDD0756 | [9] |
| LDCM0440 | CL5 | HCT 116 | C128(1.12); C211(1.01); C391(1.61); C394(1.85) | LDD0757 | [9] |
| LDCM0441 | CL50 | HCT 116 | C128(0.91); C211(0.73); C391(1.34); C394(1.40) | LDD0758 | [9] |
| LDCM0442 | CL51 | HCT 116 | C128(1.03); C211(0.96); C391(1.35); C394(1.31) | LDD0759 | [9] |
| LDCM0443 | CL52 | HCT 116 | C128(1.05); C211(1.14); C391(1.61); C394(1.56) | LDD0760 | [9] |
| LDCM0444 | CL53 | HCT 116 | C128(0.96); C211(0.80); C391(0.95); C394(0.99) | LDD0761 | [9] |
| LDCM0445 | CL54 | HCT 116 | C128(0.95); C211(0.80); C391(1.50); C394(1.37) | LDD0762 | [9] |
| LDCM0446 | CL55 | HCT 116 | C128(0.91); C211(0.98); C391(0.92); C394(0.87) | LDD0763 | [9] |
| LDCM0447 | CL56 | HCT 116 | C128(1.10); C211(0.92); C391(1.47); C394(1.47) | LDD0764 | [9] |
| LDCM0448 | CL57 | HCT 116 | C128(0.89); C211(0.71); C391(1.62); C394(1.50) | LDD0765 | [9] |
| LDCM0449 | CL58 | HCT 116 | C128(1.12); C211(1.07); C391(1.31); C394(1.36) | LDD0766 | [9] |
| LDCM0450 | CL59 | HCT 116 | C128(0.99); C211(0.95); C391(1.38); C394(1.18) | LDD0767 | [9] |
| LDCM0451 | CL6 | HCT 116 | C128(1.31); C211(1.20); C391(1.23); C394(1.36) | LDD0768 | [9] |
| LDCM0452 | CL60 | HCT 116 | C128(0.97); C211(1.07); C391(1.62); C394(1.54) | LDD0769 | [9] |
| LDCM0453 | CL61 | HCT 116 | C128(1.08); C211(1.39); C394(0.98); C484(0.82) | LDD0770 | [9] |
| LDCM0454 | CL62 | HCT 116 | C128(1.25); C211(1.41); C394(1.05); C484(0.73) | LDD0771 | [9] |
| LDCM0455 | CL63 | HCT 116 | C128(1.26); C211(2.24); C394(1.25); C484(0.71) | LDD0772 | [9] |
| LDCM0456 | CL64 | HCT 116 | C128(1.19); C211(1.65); C394(1.03); C484(0.79) | LDD0773 | [9] |
| LDCM0457 | CL65 | HCT 116 | C128(1.12); C211(1.49); C394(1.12); C484(0.73) | LDD0774 | [9] |
| LDCM0458 | CL66 | HCT 116 | C128(1.48); C211(1.42); C394(1.24); C484(0.58) | LDD0775 | [9] |
| LDCM0459 | CL67 | HCT 116 | C128(1.24); C211(1.57); C394(1.22); C484(0.74) | LDD0776 | [9] |
| LDCM0460 | CL68 | HCT 116 | C128(1.63); C211(1.39); C394(1.10); C484(0.57) | LDD0777 | [9] |
| LDCM0461 | CL69 | HCT 116 | C128(1.21); C211(2.19); C394(1.06); C484(0.70) | LDD0778 | [9] |
| LDCM0462 | CL7 | HCT 116 | C128(1.41); C211(1.02); C391(1.45); C394(1.52) | LDD0779 | [9] |
| LDCM0463 | CL70 | HCT 116 | C128(1.25); C211(1.33); C394(0.76); C484(0.72) | LDD0780 | [9] |
| LDCM0464 | CL71 | HCT 116 | C128(1.19); C211(2.91); C394(0.90); C484(0.63) | LDD0781 | [9] |
| LDCM0465 | CL72 | HCT 116 | C128(1.06); C211(1.57); C394(0.82); C484(0.82) | LDD0782 | [9] |
| LDCM0466 | CL73 | HCT 116 | C128(1.30); C211(1.98); C394(1.07); C484(0.80) | LDD0783 | [9] |
| LDCM0467 | CL74 | HCT 116 | C128(1.60); C211(1.48); C394(1.00); C484(0.75) | LDD0784 | [9] |
| LDCM0469 | CL76 | HCT 116 | C128(1.43); C211(0.92); C391(0.79); C394(0.84) | LDD0786 | [9] |
| LDCM0470 | CL77 | HCT 116 | C128(1.20); C211(0.88); C391(1.27); C394(1.32) | LDD0787 | [9] |
| LDCM0471 | CL78 | HCT 116 | C128(1.37); C211(1.19); C391(0.86); C394(0.81) | LDD0788 | [9] |
| LDCM0472 | CL79 | HCT 116 | C128(1.99); C211(0.99); C391(0.84); C394(1.02) | LDD0789 | [9] |
| LDCM0473 | CL8 | HCT 116 | C128(1.88); C211(2.29); C391(1.07); C394(1.10) | LDD0790 | [9] |
| LDCM0474 | CL80 | HCT 116 | C128(1.44); C211(0.84); C391(1.00); C394(0.97) | LDD0791 | [9] |
| LDCM0475 | CL81 | HCT 116 | C128(2.15); C211(1.00); C391(1.26); C394(1.46) | LDD0792 | [9] |
| LDCM0476 | CL82 | HCT 116 | C128(2.22); C211(0.86); C391(1.10); C394(1.25) | LDD0793 | [9] |
| LDCM0477 | CL83 | HCT 116 | C128(1.59); C211(1.03); C391(0.83); C394(0.91) | LDD0794 | [9] |
| LDCM0478 | CL84 | HCT 116 | C128(2.03); C211(0.99); C391(1.03); C394(0.95) | LDD0795 | [9] |
| LDCM0479 | CL85 | HCT 116 | C128(1.32); C211(1.03); C391(0.94); C394(0.85) | LDD0796 | [9] |
| LDCM0480 | CL86 | HCT 116 | C128(1.18); C211(1.08); C391(0.88); C394(0.93) | LDD0797 | [9] |
| LDCM0481 | CL87 | HCT 116 | C128(1.30); C211(0.89); C391(1.09); C394(1.01) | LDD0798 | [9] |
| LDCM0482 | CL88 | HCT 116 | C128(1.62); C211(1.00); C391(1.15); C394(1.23) | LDD0799 | [9] |
| LDCM0483 | CL89 | HCT 116 | C128(4.60); C211(0.91); C391(1.07); C394(1.21) | LDD0800 | [9] |
| LDCM0484 | CL9 | HCT 116 | C128(1.27); C211(1.02); C391(1.19); C394(1.25) | LDD0801 | [9] |
| LDCM0485 | CL90 | HCT 116 | C128(1.04); C211(1.29); C391(0.99); C394(1.01) | LDD0802 | [9] |
| LDCM0486 | CL91 | HCT 116 | C128(1.28); C211(1.12); C391(0.92); C394(1.20) | LDD0803 | [9] |
| LDCM0487 | CL92 | HCT 116 | C128(1.12); C211(1.28); C391(1.03); C394(1.29) | LDD0804 | [9] |
| LDCM0488 | CL93 | HCT 116 | C128(1.05); C211(1.04); C391(1.14); C394(1.13) | LDD0805 | [9] |
| LDCM0489 | CL94 | HCT 116 | C128(1.40); C211(0.94); C391(1.22); C394(1.35) | LDD0806 | [9] |
| LDCM0490 | CL95 | HCT 116 | C128(1.32); C211(1.74); C391(1.12); C394(1.33) | LDD0807 | [9] |
| LDCM0491 | CL96 | HCT 116 | C128(1.19); C211(1.41); C391(1.29); C394(1.70) | LDD0808 | [9] |
| LDCM0492 | CL97 | HCT 116 | C128(0.97); C211(1.04); C391(1.31); C394(1.67) | LDD0809 | [9] |
| LDCM0493 | CL98 | HCT 116 | C128(0.94); C211(1.21); C391(0.81); C394(0.82) | LDD0810 | [9] |
| LDCM0494 | CL99 | HCT 116 | C128(1.06); C211(0.97); C391(1.02); C394(0.94) | LDD0811 | [9] |
| LDCM0495 | E2913 | HEK-293T | C211(1.07); C128(1.04); C391(0.92); C487(0.93) | LDD1698 | [21] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C211(3.83) | LDD1702 | [4] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [8] |
| LDCM0625 | F8 | Ramos | C211(1.63) | LDD2187 | [22] |
| LDCM0572 | Fragment10 | Ramos | C211(6.72) | LDD2189 | [22] |
| LDCM0573 | Fragment11 | Ramos | C211(2.96) | LDD2190 | [22] |
| LDCM0574 | Fragment12 | Ramos | C211(2.48) | LDD1394 | [23] |
| LDCM0575 | Fragment13 | MDA-MB-231 | C211(1.09) | LDD1395 | [23] |
| LDCM0576 | Fragment14 | Ramos | C211(1.45) | LDD2193 | [22] |
| LDCM0579 | Fragment20 | Ramos | C211(7.83) | LDD2194 | [22] |
| LDCM0580 | Fragment21 | Ramos | C211(1.63) | LDD1405 | [23] |
| LDCM0582 | Fragment23 | Ramos | C211(1.07) | LDD2196 | [22] |
| LDCM0584 | Fragment25 | MDA-MB-231 | C211(1.19) | LDD1411 | [23] |
| LDCM0578 | Fragment27 | Ramos | C211(0.97) | LDD2197 | [22] |
| LDCM0586 | Fragment28 | Ramos | C211(0.80) | LDD2198 | [22] |
| LDCM0588 | Fragment30 | Ramos | C211(1.98) | LDD2199 | [22] |
| LDCM0589 | Fragment31 | Ramos | C211(1.29) | LDD2200 | [22] |
| LDCM0590 | Fragment32 | Ramos | C211(5.56) | LDD2201 | [22] |
| LDCM0468 | Fragment33 | HCT 116 | C128(1.99); C211(1.54); C394(1.12); C484(0.74) | LDD0785 | [9] |
| LDCM0593 | Fragment35 | Ramos | C211(0.95) | LDD1430 | [23] |
| LDCM0596 | Fragment38 | MDA-MB-231 | C211(1.13) | LDD1433 | [23] |
| LDCM0566 | Fragment4 | Ramos | C211(2.44) | LDD2184 | [22] |
| LDCM0599 | Fragment41 | MDA-MB-231 | C211(1.36) | LDD1438 | [23] |
| LDCM0601 | Fragment43 | Ramos | C211(1.73) | LDD1442 | [23] |
| LDCM0427 | Fragment51 | HCT 116 | C128(3.02); C211(2.58); C391(0.90); C394(0.93) | LDD0744 | [9] |
| LDCM0610 | Fragment52 | Ramos | C211(1.91) | LDD2204 | [22] |
| LDCM0614 | Fragment56 | MDA-MB-231 | C211(2.07) | LDD1458 | [23] |
| LDCM0569 | Fragment7 | Ramos | C211(5.27) | LDD2186 | [22] |
| LDCM0570 | Fragment8 | Ramos | C211(1.79) | LDD1386 | [23] |
| LDCM0571 | Fragment9 | Ramos | C211(7.66) | LDD2188 | [22] |
| LDCM0015 | HNE | MDA-MB-231 | C211(1.03) | LDD0346 | [22] |
| LDCM0022 | KB02 | MDA-MB-231 | C211(20.00) | LDD1374 | [23] |
| LDCM0023 | KB03 | Jurkat | C128(10.56); C211(9.87) | LDD0209 | [24] |
| LDCM0024 | KB05 | COLO792 | C211(4.33) | LDD3310 | [25] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C211(0.47) | LDD2121 | [4] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [16] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C211(1.00) | LDD2090 | [4] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C211(0.81) | LDD2099 | [4] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C211(1.07) | LDD2105 | [4] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C211(0.72) | LDD2107 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C211(1.25) | LDD2109 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C211(0.27) | LDD2111 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C211(0.96) | LDD2123 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C211(0.74) | LDD2125 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C211(0.95) | LDD2127 | [4] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C211(1.15) | LDD2128 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C211(1.13) | LDD2129 | [4] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C211(1.15) | LDD2135 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C211(1.01) | LDD2136 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C211(0.89) | LDD2137 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C211(0.56) | LDD2140 | [4] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C211(2.20) | LDD2144 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C211(1.04) | LDD2146 | [4] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C211(0.59) | LDD2206 | [26] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C128(0.49) | LDD2207 | [26] |
| LDCM0014 | Panhematin | HEK-293T | 3.06 | LDD0062 | [7] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 1.75 | LDD0160 | [5] |
| LDCM0021 | THZ1 | HCT 116 | C211(1.02) | LDD2173 | [9] |
| LDCM0112 | W16 | Hep-G2 | C128(0.76) | LDD0239 | [17] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Metal transporter CNNM3 (CNNM3) | ACDP family | Q8NE01 | |||
| Cation channel sperm-associated protein 1 (CATSPER1) | Cation channel sperm-associated (TC 1.A.1.19) family | Q8NEC5 | |||
Transcription factor
Other
References
