Details of the Target
General Information of Target
| Target ID | LDTP00590 | |||||
|---|---|---|---|---|---|---|
| Target Name | Protein RER1 (RER1) | |||||
| Gene Name | RER1 | |||||
| Gene ID | 11079 | |||||
| Synonyms |
Protein RER1 |
|||||
| 3D Structure | ||||||
| Sequence |
MSEGDSVGESVHGKPSVVYRFFTRLGQIYQSWLDKSTPYTAVRWVVTLGLSFVYMIRVYL
LQGWYIVTYALGIYHLNLFIAFLSPKVDPSLMEDSDDGPSLPTKQNEEFRPFIRRLPEFK FWHAATKGILVAMVCTFFDAFNVPVFWPILVMYFIMLFCITMKRQIKHMIKYRYIPFTHG KRRYRGKEDAGKAFAS |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
RER1 family
|
|||||
| Subcellular location |
Golgi apparatus membrane
|
|||||
| Function | Involved in the retrieval of endoplasmic reticulum membrane proteins from the early Golgi compartment. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
HDSF-alk Probe Info |
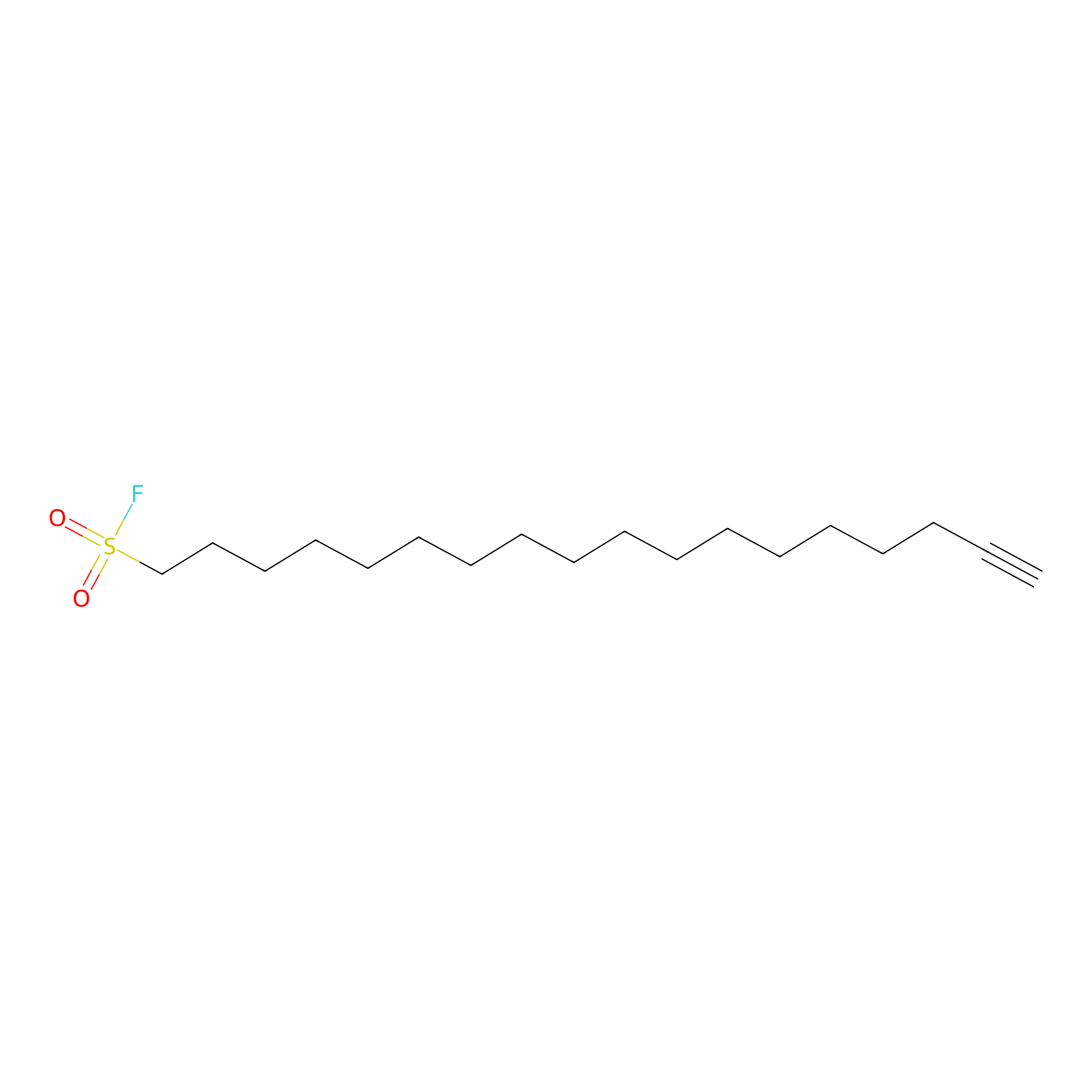 |
1.71 | LDD0197 | [1] | |
|
CHEMBL5175495 Probe Info |
 |
5.69 | LDD0196 | [2] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [3] | |
|
TH211 Probe Info |
 |
Y174(10.08); Y29(9.41) | LDD0260 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
OPA-S-S-alkyne Probe Info |
 |
K181(7.46) | LDD3494 | [6] | |
|
HPAP Probe Info |
 |
3.60 | LDD0064 | [7] | |
|
Alkyne-RA190 Probe Info |
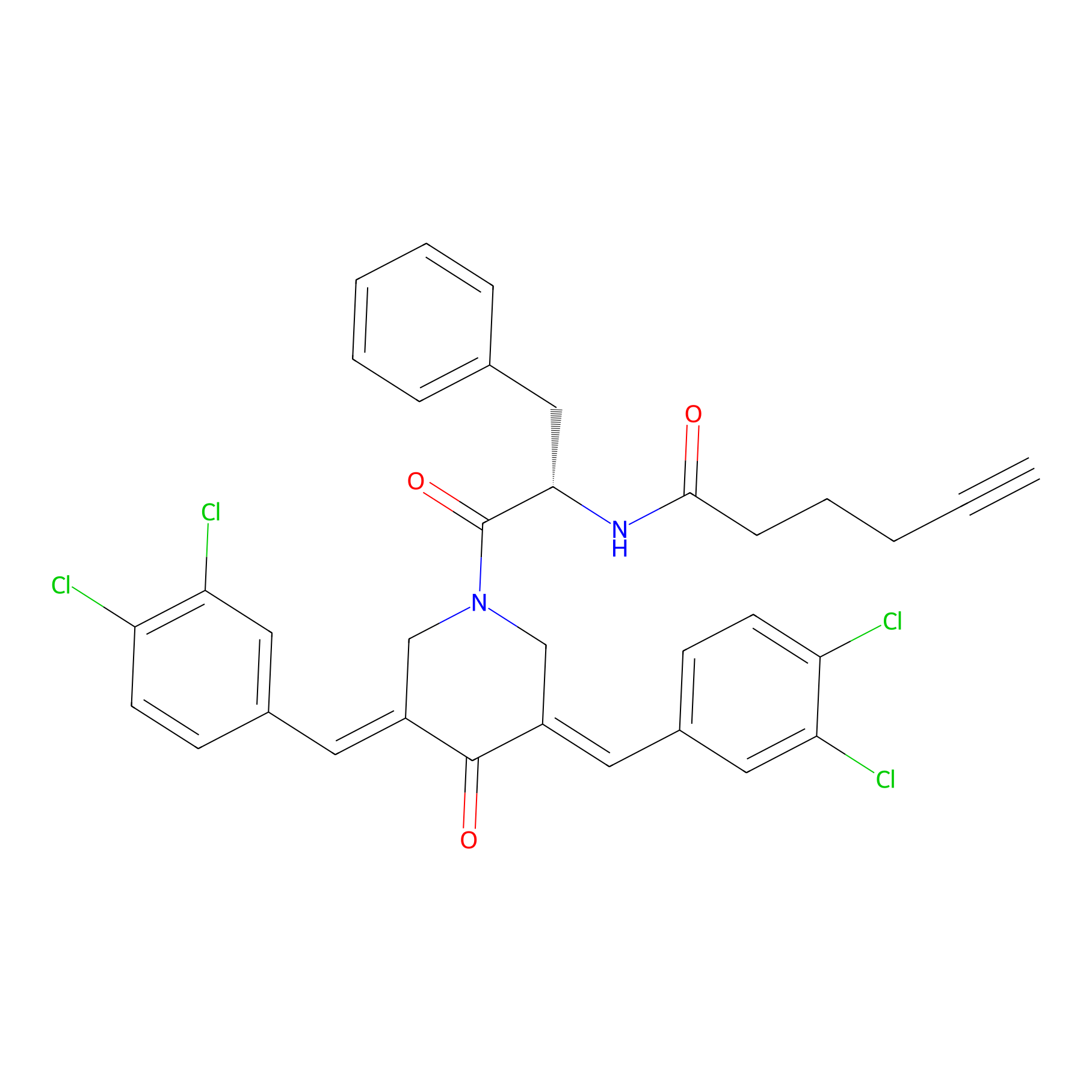 |
3.95 | LDD0300 | [8] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0227 | [9] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [10] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [11] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [11] | |
|
AOyne Probe Info |
 |
14.40 | LDD0443 | [12] | |
PAL-AfBPP Probe
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References



























