Details of the Target
General Information of Target
| Target ID | LDTP13802 | |||||
|---|---|---|---|---|---|---|
| Target Name | DNA replication complex GINS protein PSF2 (GINS2) | |||||
| Gene Name | GINS2 | |||||
| Gene ID | 51659 | |||||
| Synonyms |
PSF2; DNA replication complex GINS protein PSF2; GINS complex subunit 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MAQYKGTMREAGRAMHLLKKRERQREQMEVLKQRIAEETILKSQVDKRFSAHYDAVEAEL
KSSTVGLVTLNDMKARQEALVRERERQLAKRQHLEEQRLQQERQREQEQRRERKRKISCL SFALDDLDDQADAAEARRAGNLGKNPDVDTSFLPDRDREEEENRLREELRQEWEAQREKV KDEEMEVTFSYWDGSGHRRTVRVRKGNTVQQFLKKALQGLRKDFLELRSAGVEQLMFIKE DLILPHYHTFYDFIIARARGKSGPLFSFDVHDDVRLLSDATMEKDESHAGKVVLRSWYEK NKHIFPASRWEAYDPEKKWDKYTIR |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
GINS2/PSF2 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Required for correct functioning of the GINS complex, a complex that plays an essential role in the initiation of DNA replication, and progression of DNA replication forks. GINS complex is a core component of CDC45-MCM-GINS (CMG) helicase, the molecular machine that unwinds template DNA during replication, and around which the replisome is built.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
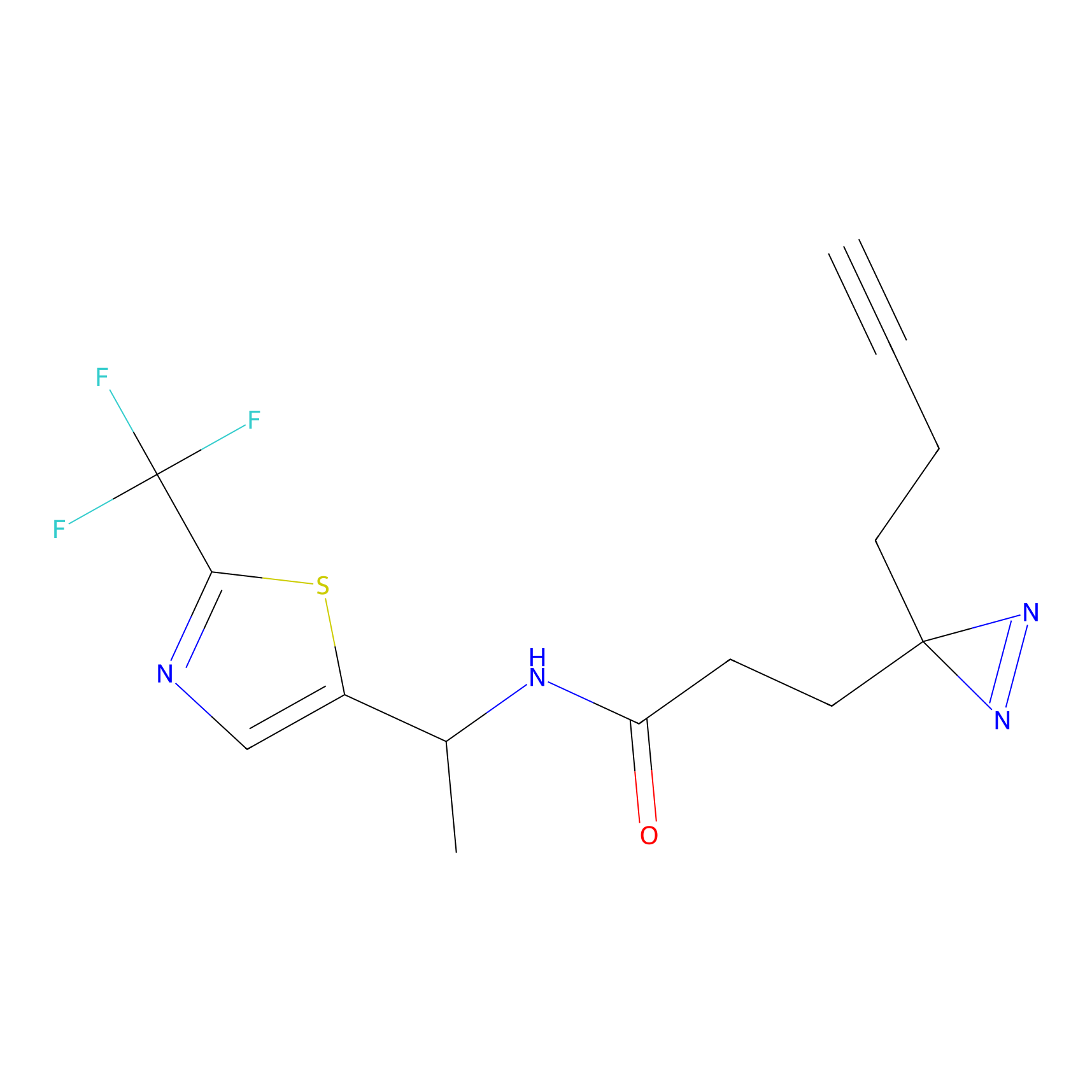
|
FBP2 Probe Info |
 |
3.36 | LDD0323 | [1] | |
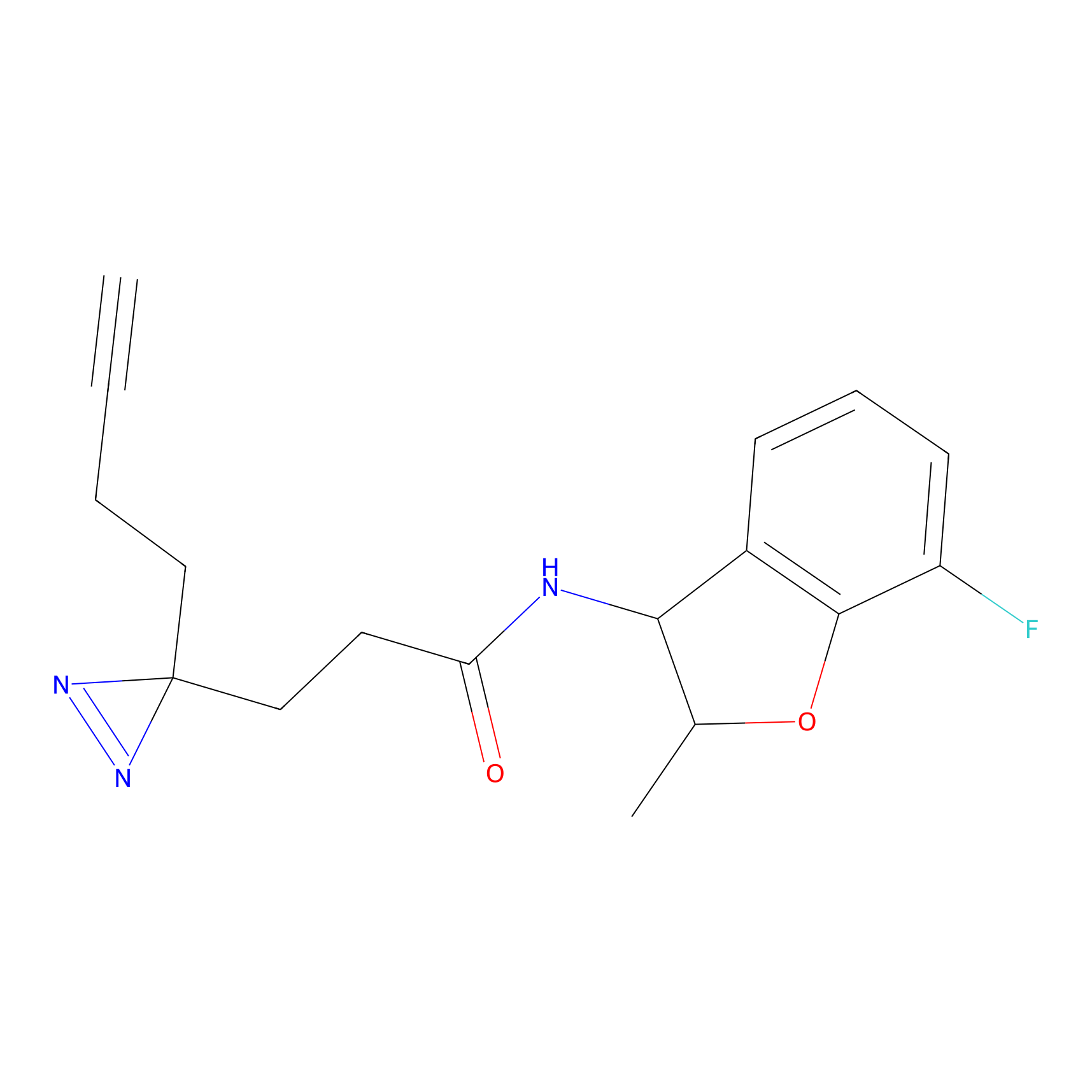
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [2] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [2] | |
PAL-AfBPP Probe
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Serine/threonine-protein kinase Chk2 (CHEK2) | CAMK Ser/Thr protein kinase family | O96017 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| DNA replication complex GINS protein SLD5 (GINS4) | GINS4/SLD5 family | Q9BRT9 | |||
| Mini-chromosome maintenance complex-binding protein (MCMBP) | MCMBP family | Q9BTE3 | |||
References