Details of the Target
General Information of Target
| Target ID | LDTP11140 | |||||
|---|---|---|---|---|---|---|
| Target Name | Peroxiredoxin-like 2A (PRXL2A) | |||||
| Gene Name | PRXL2A | |||||
| Gene ID | 84293 | |||||
| Synonyms |
C10orf58; FAM213A; PAMM; Peroxiredoxin-like 2A; Peroxiredoxin-like 2 activated in M-CSF stimulated monocytes; Protein PAMM; Redox-regulatory protein FAM213A |
|||||
| 3D Structure | ||||||
| Sequence |
MSEAYFRVESGALGPEENFLSLDDILMSHEKLPVRTETAMPRLGAFFLERSAGAETDNAV
PQGSKLELPLWLAKGLFDNKRRILSVELPKIYQEGWRTVFSADPNVVDLHKMGPHFYGFG SQLLHFDSPENADISQSLLQTFIGRFRRIMDSSQNAYNEDTSALVARLDEMERGLFQTGQ KGLNDFQCWEKGQASQITASNLVQNYKKRKFTDMED |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Peroxiredoxin-like PRXL2 family, PRXL2A subfamily
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Involved in redox regulation of the cell. Acts as an antioxidant. Inhibits TNFSF11-induced NFKB1 and JUN activation and osteoclast differentiation. May affect bone resorption and help to maintain bone mass. Acts as a negative regulator of macrophage-mediated inflammation by inhibiting macrophage production of inflammatory cytokines, probably through suppression of the MAPK signaling pathway.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
EN219-alkyne Probe Info |
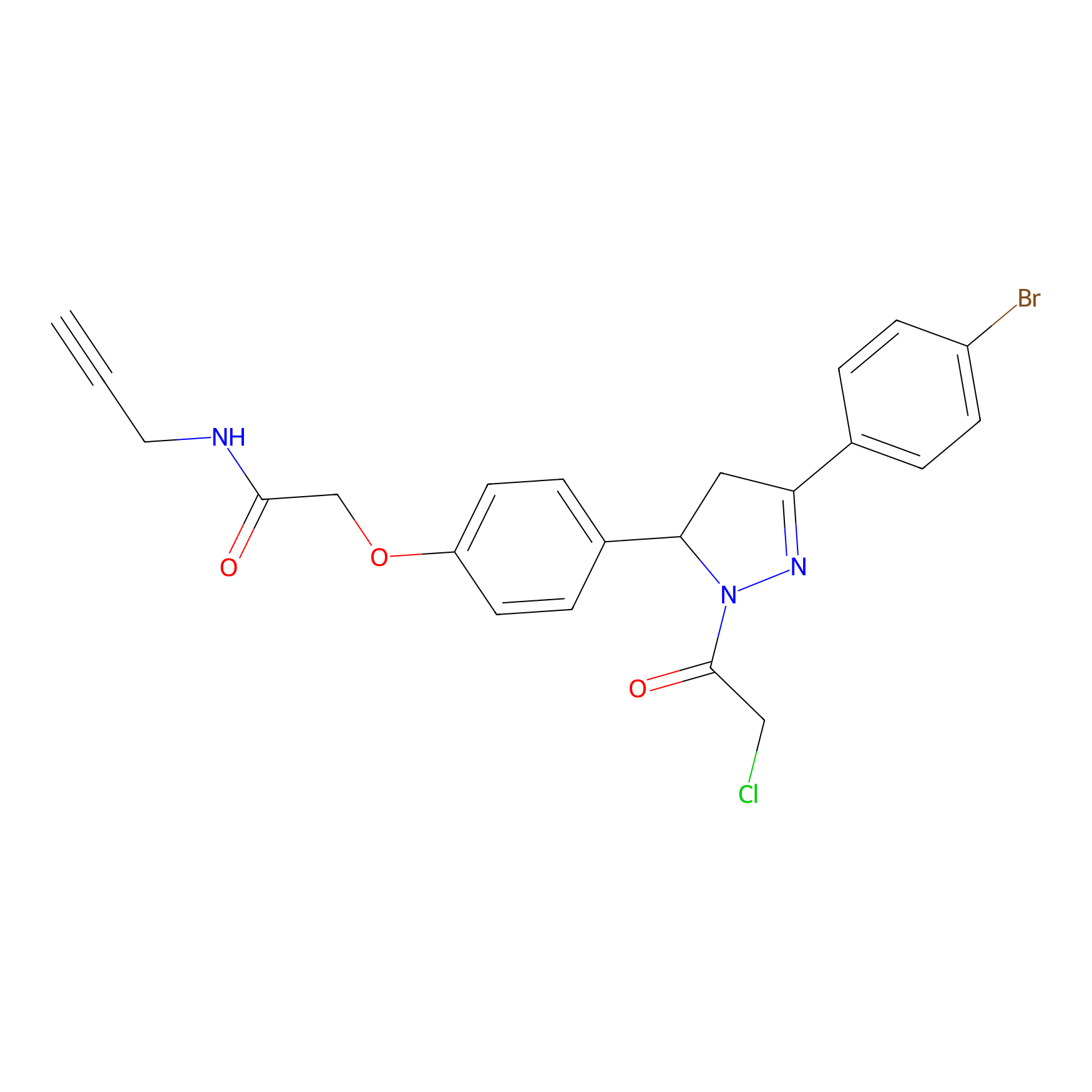 |
4.85 | LDD0297 | [1] | |
|
CHEMBL5175495 Probe Info |
 |
16.09 | LDD0196 | [2] | |
|
C-Sul Probe Info |
 |
2.30 | LDD0066 | [3] | |
|
FBP2 Probe Info |
 |
3.84 | LDD0317 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [5] | |
|
STPyne Probe Info |
 |
K122(10.00) | LDD0277 | [6] | |
|
CCW36 Probe Info |
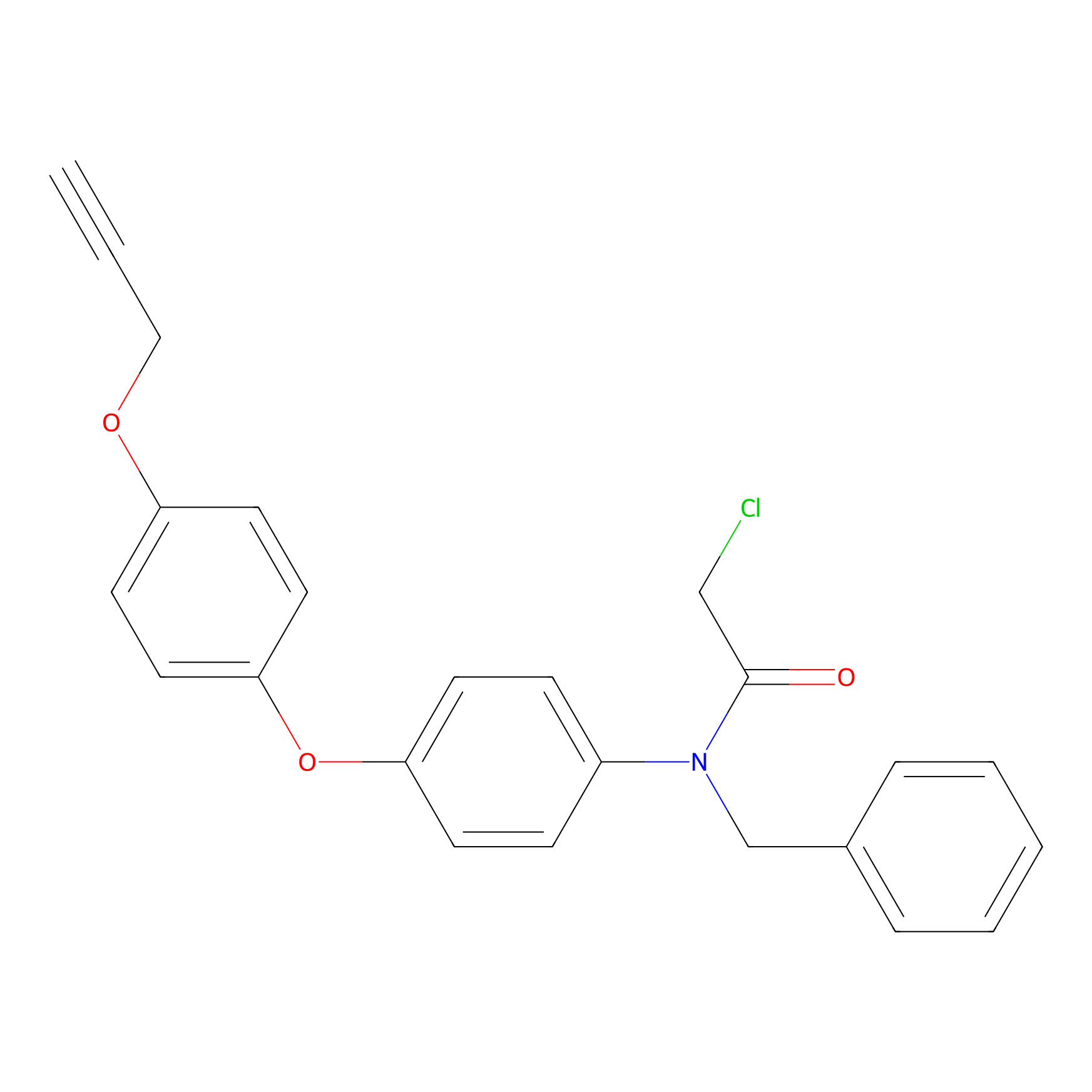 |
10.24 | LDD2215 | [7] | |
|
P21 Probe Info |
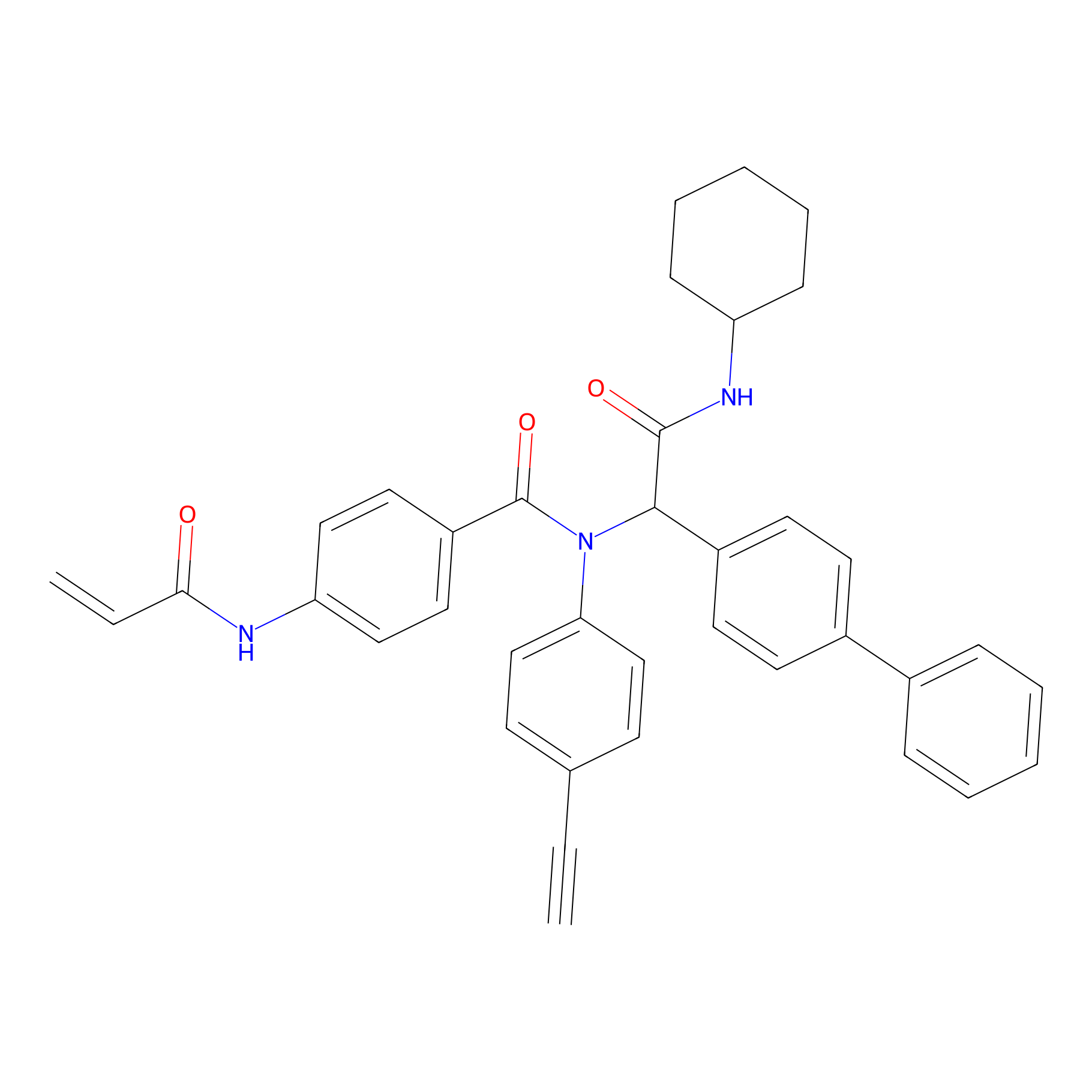 |
2.53 | LDD0408 | [8] | |
|
BTD Probe Info |
 |
C85(1.09) | LDD2092 | [9] | |
|
Johansson_61 Probe Info |
 |
_(20.00) | LDD1485 | [10] | |
|
YY4-yne Probe Info |
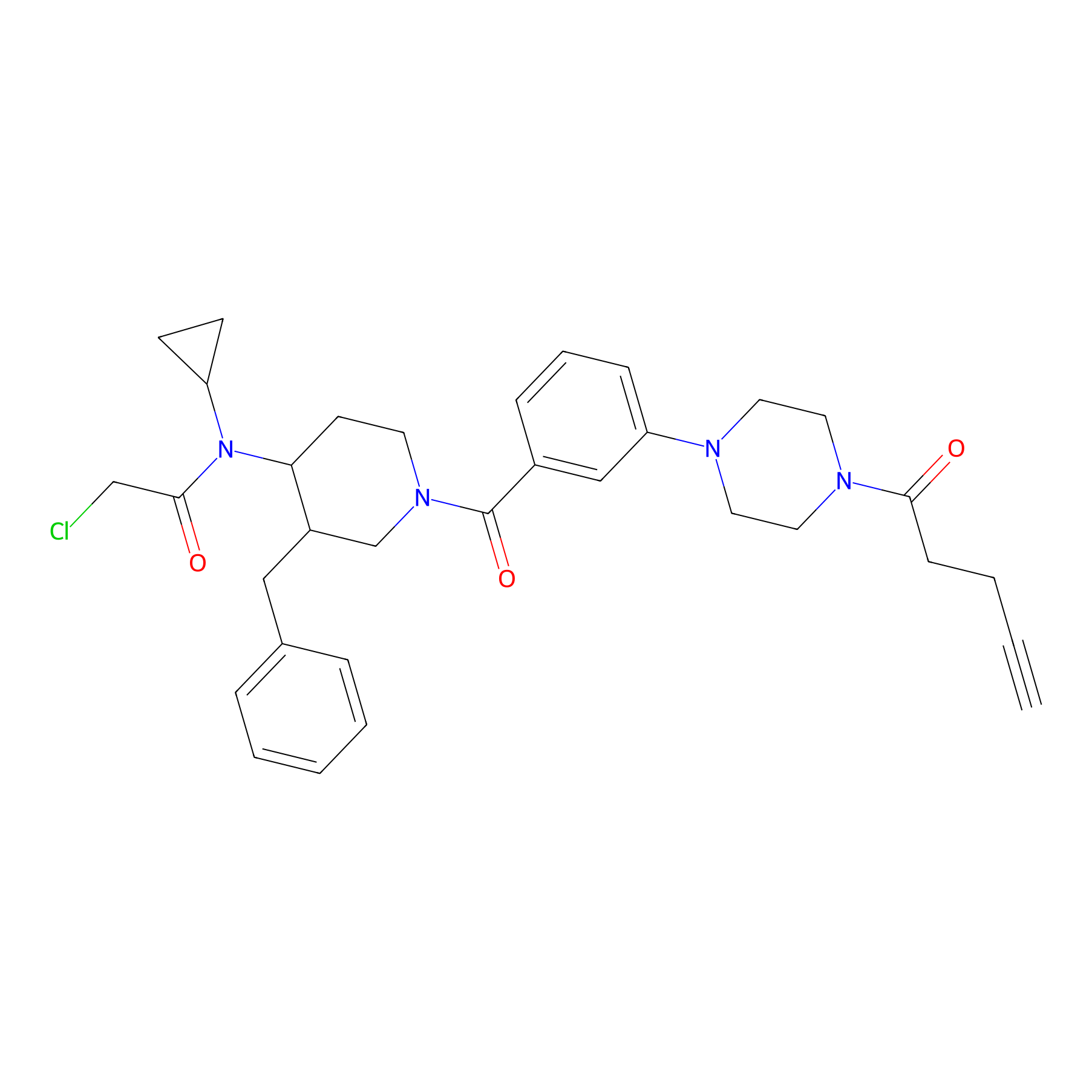 |
2.51 | LDD0400 | [11] | |
|
IA-alkyne Probe Info |
 |
C85(20.00) | LDD1704 | [12] | |
|
DBIA Probe Info |
 |
C85(3.72) | LDD0078 | [13] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [14] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [14] | |
|
Phosphinate-6 Probe Info |
 |
C85(0.00); C88(0.00) | LDD0018 | [15] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C027 Probe Info |
 |
6.45 | LDD1733 | [16] | |
|
C040 Probe Info |
 |
8.69 | LDD1740 | [16] | |
|
C094 Probe Info |
 |
23.10 | LDD1785 | [16] | |
|
C163 Probe Info |
 |
13.36 | LDD1843 | [16] | |
|
C235 Probe Info |
 |
20.25 | LDD1908 | [16] | |
|
C299 Probe Info |
 |
5.70 | LDD1968 | [16] | |
|
C362 Probe Info |
 |
37.27 | LDD2023 | [16] | |
|
C364 Probe Info |
 |
22.01 | LDD2025 | [16] | |
|
FFF probe11 Probe Info |
 |
14.01 | LDD0471 | [17] | |
|
JN0003 Probe Info |
 |
13.29 | LDD0469 | [17] | |
|
A-DA Probe Info |
 |
2.26 | LDD0145 | [18] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C85(0.89) | LDD2142 | [9] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C85(1.52) | LDD2112 | [9] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C85(1.06) | LDD2103 | [9] |
| LDCM0214 | AC1 | HEK-293T | C85(2.61) | LDD0812 | [13] |
| LDCM0216 | AC100 | HEK-293T | C85(0.78) | LDD0814 | [13] |
| LDCM0217 | AC101 | HEK-293T | C85(0.92) | LDD0815 | [13] |
| LDCM0218 | AC102 | HEK-293T | C85(0.92) | LDD0816 | [13] |
| LDCM0219 | AC103 | HEK-293T | C85(1.19) | LDD0817 | [13] |
| LDCM0220 | AC104 | HEK-293T | C85(0.92) | LDD0818 | [13] |
| LDCM0221 | AC105 | HEK-293T | C85(1.01) | LDD0819 | [13] |
| LDCM0222 | AC106 | HEK-293T | C85(0.96) | LDD0820 | [13] |
| LDCM0223 | AC107 | HEK-293T | C85(0.79) | LDD0821 | [13] |
| LDCM0224 | AC108 | HEK-293T | C85(0.95) | LDD0822 | [13] |
| LDCM0225 | AC109 | HEK-293T | C85(1.49) | LDD0823 | [13] |
| LDCM0227 | AC110 | HEK-293T | C85(1.03) | LDD0825 | [13] |
| LDCM0228 | AC111 | HEK-293T | C85(1.58) | LDD0826 | [13] |
| LDCM0229 | AC112 | HEK-293T | C85(0.85) | LDD0827 | [13] |
| LDCM0237 | AC12 | HEK-293T | C85(1.01) | LDD1510 | [19] |
| LDCM0263 | AC143 | HEK-293T | C85(1.06) | LDD0861 | [13] |
| LDCM0264 | AC144 | HEK-293T | C85(1.06) | LDD0862 | [13] |
| LDCM0265 | AC145 | HEK-293T | C85(1.32) | LDD0863 | [13] |
| LDCM0266 | AC146 | HEK-293T | C85(1.61) | LDD0864 | [13] |
| LDCM0267 | AC147 | HEK-293T | C85(1.27) | LDD0865 | [13] |
| LDCM0268 | AC148 | HEK-293T | C85(2.04) | LDD0866 | [13] |
| LDCM0269 | AC149 | HEK-293T | C85(0.94) | LDD0867 | [13] |
| LDCM0271 | AC150 | HEK-293T | C85(1.06) | LDD0869 | [13] |
| LDCM0272 | AC151 | HEK-293T | C85(1.18) | LDD0870 | [13] |
| LDCM0273 | AC152 | HEK-293T | C85(0.86) | LDD0871 | [13] |
| LDCM0274 | AC153 | HEK-293T | C85(1.34) | LDD0872 | [13] |
| LDCM0621 | AC154 | HEK-293T | C85(1.10) | LDD2162 | [13] |
| LDCM0622 | AC155 | HEK-293T | C85(1.16) | LDD2163 | [13] |
| LDCM0623 | AC156 | HEK-293T | C85(1.13) | LDD2164 | [13] |
| LDCM0624 | AC157 | HEK-293T | C85(0.88) | LDD2165 | [13] |
| LDCM0276 | AC17 | HEK-293T | C85(1.60) | LDD0874 | [13] |
| LDCM0277 | AC18 | HEK-293T | C85(1.67) | LDD0875 | [13] |
| LDCM0278 | AC19 | HEK-293T | C85(1.36) | LDD0876 | [13] |
| LDCM0279 | AC2 | HEK-293T | C85(1.22) | LDD0877 | [13] |
| LDCM0280 | AC20 | HEK-293T | C85(1.23) | LDD0878 | [13] |
| LDCM0281 | AC21 | HEK-293T | C85(5.52) | LDD0879 | [13] |
| LDCM0282 | AC22 | HEK-293T | C85(1.03) | LDD0880 | [13] |
| LDCM0283 | AC23 | HEK-293T | C85(3.11) | LDD0881 | [13] |
| LDCM0284 | AC24 | HEK-293T | C85(1.12) | LDD0882 | [13] |
| LDCM0288 | AC28 | HEK-293T | C85(1.00) | LDD1527 | [19] |
| LDCM0290 | AC3 | HEK-293T | C85(1.82) | LDD0888 | [13] |
| LDCM0296 | AC35 | HEK-293T | C85(1.16) | LDD0894 | [13] |
| LDCM0297 | AC36 | HEK-293T | C85(1.08) | LDD0895 | [13] |
| LDCM0298 | AC37 | HEK-293T | C85(1.10) | LDD0896 | [13] |
| LDCM0299 | AC38 | HEK-293T | C85(1.53) | LDD0897 | [13] |
| LDCM0300 | AC39 | HEK-293T | C85(1.12) | LDD0898 | [13] |
| LDCM0301 | AC4 | HEK-293T | C85(0.99) | LDD0899 | [13] |
| LDCM0302 | AC40 | HEK-293T | C85(2.72) | LDD0900 | [13] |
| LDCM0303 | AC41 | HEK-293T | C85(1.21) | LDD0901 | [13] |
| LDCM0304 | AC42 | HEK-293T | C85(2.03) | LDD0902 | [13] |
| LDCM0305 | AC43 | HEK-293T | C85(1.15) | LDD0903 | [13] |
| LDCM0306 | AC44 | HEK-293T | C85(1.25) | LDD0904 | [13] |
| LDCM0307 | AC45 | HEK-293T | C85(1.14) | LDD0905 | [13] |
| LDCM0312 | AC5 | HEK-293T | C85(1.38) | LDD0910 | [13] |
| LDCM0315 | AC52 | HEK-293T | C85(0.95) | LDD1554 | [19] |
| LDCM0320 | AC57 | HEK-293T | C85(9.07) | LDD0918 | [13] |
| LDCM0321 | AC58 | HEK-293T | C85(1.17) | LDD0919 | [13] |
| LDCM0322 | AC59 | HEK-293T | C85(7.04) | LDD0920 | [13] |
| LDCM0324 | AC60 | HEK-293T | C85(1.51) | LDD0922 | [13] |
| LDCM0325 | AC61 | HEK-293T | C85(0.97) | LDD0923 | [13] |
| LDCM0326 | AC62 | HEK-293T | C85(1.54) | LDD0924 | [13] |
| LDCM0327 | AC63 | HEK-293T | C85(1.53) | LDD0925 | [13] |
| LDCM0328 | AC64 | HEK-293T | C85(1.12) | LDD0926 | [13] |
| LDCM0329 | AC65 | HEK-293T | C85(1.06) | LDD0927 | [13] |
| LDCM0330 | AC66 | HEK-293T | C85(0.97) | LDD0928 | [13] |
| LDCM0331 | AC67 | HEK-293T | C85(0.95) | LDD0929 | [13] |
| LDCM0365 | AC98 | HEK-293T | C85(4.84) | LDD0963 | [13] |
| LDCM0366 | AC99 | HEK-293T | C85(0.91) | LDD0964 | [13] |
| LDCM0020 | ARS-1620 | HCC44 | C85(3.72) | LDD0078 | [13] |
| LDCM0631 | CCW16 | 231MFP | 10.24 | LDD2215 | [7] |
| LDCM0367 | CL1 | HEK-293T | C85(10.00) | LDD0965 | [13] |
| LDCM0368 | CL10 | HEK-293T | C85(5.43) | LDD0966 | [13] |
| LDCM0369 | CL100 | HEK-293T | C85(3.07) | LDD0967 | [13] |
| LDCM0374 | CL105 | HEK-293T | C85(6.58) | LDD0972 | [13] |
| LDCM0375 | CL106 | HEK-293T | C85(10.00) | LDD0973 | [13] |
| LDCM0376 | CL107 | HEK-293T | C85(1.32) | LDD0974 | [13] |
| LDCM0377 | CL108 | HEK-293T | C85(1.33) | LDD0975 | [13] |
| LDCM0378 | CL109 | HEK-293T | C85(7.07) | LDD0976 | [13] |
| LDCM0379 | CL11 | HEK-293T | C85(2.60) | LDD0977 | [13] |
| LDCM0380 | CL110 | HEK-293T | C85(8.24) | LDD0978 | [13] |
| LDCM0381 | CL111 | HEK-293T | C85(8.18) | LDD0979 | [13] |
| LDCM0387 | CL117 | HEK-293T | C85(1.30) | LDD0985 | [13] |
| LDCM0388 | CL118 | HEK-293T | C85(2.58) | LDD0986 | [13] |
| LDCM0389 | CL119 | HEK-293T | C85(1.17) | LDD0987 | [13] |
| LDCM0390 | CL12 | HEK-293T | C85(15.70) | LDD0988 | [13] |
| LDCM0391 | CL120 | HEK-293T | C85(1.26) | LDD0989 | [13] |
| LDCM0396 | CL125 | HEK-293T | C85(3.94) | LDD0994 | [13] |
| LDCM0397 | CL126 | HEK-293T | C85(2.20) | LDD0995 | [13] |
| LDCM0398 | CL127 | HEK-293T | C85(0.98) | LDD0996 | [13] |
| LDCM0399 | CL128 | HEK-293T | C85(1.73) | LDD0997 | [13] |
| LDCM0400 | CL13 | HEK-293T | C85(3.70) | LDD0998 | [13] |
| LDCM0401 | CL14 | HEK-293T | C85(1.09) | LDD0999 | [13] |
| LDCM0402 | CL15 | HEK-293T | C85(12.60) | LDD1000 | [13] |
| LDCM0403 | CL16 | HEK-293T | C85(0.87) | LDD1001 | [13] |
| LDCM0404 | CL17 | HEK-293T | C85(3.89) | LDD1002 | [13] |
| LDCM0405 | CL18 | HEK-293T | C85(3.06) | LDD1003 | [13] |
| LDCM0406 | CL19 | HEK-293T | C85(2.74) | LDD1004 | [13] |
| LDCM0407 | CL2 | HEK-293T | C85(1.29) | LDD1005 | [13] |
| LDCM0408 | CL20 | HEK-293T | C85(1.19) | LDD1006 | [13] |
| LDCM0409 | CL21 | HEK-293T | C85(1.96) | LDD1007 | [13] |
| LDCM0410 | CL22 | HEK-293T | C85(1.46) | LDD1008 | [13] |
| LDCM0411 | CL23 | HEK-293T | C85(0.98) | LDD1009 | [13] |
| LDCM0412 | CL24 | HEK-293T | C85(2.15) | LDD1010 | [13] |
| LDCM0413 | CL25 | HEK-293T | C85(1.77) | LDD1011 | [13] |
| LDCM0414 | CL26 | HEK-293T | C85(3.04) | LDD1012 | [13] |
| LDCM0415 | CL27 | HEK-293T | C85(2.85) | LDD1013 | [13] |
| LDCM0416 | CL28 | HEK-293T | C85(0.96) | LDD1014 | [13] |
| LDCM0417 | CL29 | HEK-293T | C85(1.01) | LDD1015 | [13] |
| LDCM0418 | CL3 | HEK-293T | C85(11.80) | LDD1016 | [13] |
| LDCM0419 | CL30 | HEK-293T | C85(0.97) | LDD1017 | [13] |
| LDCM0420 | CL31 | HEK-293T | C85(1.10) | LDD1018 | [13] |
| LDCM0421 | CL32 | HEK-293T | C85(4.04) | LDD1019 | [13] |
| LDCM0422 | CL33 | HEK-293T | C85(2.32) | LDD1020 | [13] |
| LDCM0423 | CL34 | HEK-293T | C85(1.86) | LDD1021 | [13] |
| LDCM0424 | CL35 | HEK-293T | C85(3.78) | LDD1022 | [13] |
| LDCM0425 | CL36 | HEK-293T | C85(3.92) | LDD1023 | [13] |
| LDCM0426 | CL37 | HEK-293T | C85(4.38) | LDD1024 | [13] |
| LDCM0428 | CL39 | HEK-293T | C85(2.26) | LDD1026 | [13] |
| LDCM0429 | CL4 | HEK-293T | C85(8.48) | LDD1027 | [13] |
| LDCM0430 | CL40 | HEK-293T | C85(1.39) | LDD1028 | [13] |
| LDCM0431 | CL41 | HEK-293T | C85(2.55) | LDD1029 | [13] |
| LDCM0432 | CL42 | HEK-293T | C85(1.33) | LDD1030 | [13] |
| LDCM0433 | CL43 | HEK-293T | C85(1.15) | LDD1031 | [13] |
| LDCM0434 | CL44 | HEK-293T | C85(3.60) | LDD1032 | [13] |
| LDCM0435 | CL45 | HEK-293T | C85(3.61) | LDD1033 | [13] |
| LDCM0440 | CL5 | HEK-293T | C85(1.49) | LDD1038 | [13] |
| LDCM0447 | CL56 | HEK-293T | C85(1.15) | LDD1650 | [19] |
| LDCM0451 | CL6 | HEK-293T | C85(20.00) | LDD1049 | [13] |
| LDCM0460 | CL68 | HEK-293T | C85(1.03) | LDD1663 | [19] |
| LDCM0462 | CL7 | HEK-293T | C85(1.89) | LDD1060 | [13] |
| LDCM0473 | CL8 | HEK-293T | C85(1.75) | LDD1676 | [19] |
| LDCM0474 | CL80 | HEK-293T | C85(0.98) | LDD1677 | [19] |
| LDCM0486 | CL91 | HEK-293T | C85(1.18) | LDD1084 | [13] |
| LDCM0487 | CL92 | HEK-293T | C85(1.72) | LDD1085 | [13] |
| LDCM0488 | CL93 | HEK-293T | C85(2.93) | LDD1086 | [13] |
| LDCM0489 | CL94 | HEK-293T | C85(1.91) | LDD1087 | [13] |
| LDCM0490 | CL95 | HEK-293T | C85(2.82) | LDD1088 | [13] |
| LDCM0491 | CL96 | HEK-293T | C85(2.75) | LDD1089 | [13] |
| LDCM0492 | CL97 | HEK-293T | C85(2.58) | LDD1090 | [13] |
| LDCM0493 | CL98 | HEK-293T | C85(2.44) | LDD1091 | [13] |
| LDCM0494 | CL99 | HEK-293T | C85(2.54) | LDD1092 | [13] |
| LDCM0100 | EN219 | 231MFP | 4.90 | LDD0298 | [1] |
| LDCM0427 | Fragment51 | HEK-293T | C85(1.12) | LDD1025 | [13] |
| LDCM0616 | Fragment61 | Jurkat | _(20.00) | LDD1489 | [10] |
| LDCM0615 | Fragment63-R | Jurkat | _(15.97) | LDD1487 | [10] |
| LDCM0569 | Fragment7 | Jurkat | _(20.00) | LDD1485 | [10] |
| LDCM0022 | KB02 | HEK-293T | C88(0.51); C85(0.25); C88(0.25) | LDD1492 | [19] |
| LDCM0023 | KB03 | HEK-293T | C88(1.14); C85(0.94); C88(0.94) | LDD1497 | [19] |
| LDCM0024 | KB05 | HEK-293T | C88(1.01); C85(0.91); C88(0.91) | LDD1502 | [19] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C85(1.63) | LDD2102 | [9] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C85(1.09) | LDD2092 | [9] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C85(0.98) | LDD2093 | [9] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C85(1.09) | LDD2094 | [9] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C85(1.25) | LDD2098 | [9] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C85(0.75) | LDD2104 | [9] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C85(2.17) | LDD2106 | [9] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C88(0.17) | LDD2109 | [9] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C85(3.58) | LDD2122 | [9] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C85(0.45) | LDD2123 | [9] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C85(1.45) | LDD2135 | [9] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C88(0.88) | LDD2137 | [9] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C85(0.63) | LDD2141 | [9] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C85(4.61) | LDD2145 | [9] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C85(1.10) | LDD2146 | [9] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C85(1.25) | LDD2151 | [9] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C85(2.57) | LDD2153 | [9] |
| LDCM0158 | P27 | BxPC-3 | 2.53 | LDD0408 | [8] |
| LDCM0084 | Ro 48-8071 | A-549 | 2.26 | LDD0145 | [18] |
| LDCM0154 | YY4 | T cell | 2.51 | LDD0400 | [11] |
References
