Details of the Target
General Information of Target
| Target ID | LDTP05033 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ubiquitin-ribosomal protein eS31 fusion protein (RPS27A) | |||||
| Gene Name | RPS27A | |||||
| Gene ID | 6233 | |||||
| Synonyms |
UBA80; UBCEP1; Ubiquitin-ribosomal protein eS31 fusion protein; Ubiquitin carboxyl extension protein 80) [Cleaved into: Ubiquitin; Small ribosomal subunit protein eS31; 40S ribosomal protein S27a)] |
|||||
| 3D Structure | ||||||
| Sequence |
MQIFVKTLTGKTITLEVEPSDTIENVKAKIQDKEGIPPDQQRLIFAGKQLEDGRTLSDYN
IQKESTLHLVLRLRGGAKKRKKKSYTTPKKNKHKRKKVKLAVLKYYKVDENGKISRLRRE CPSDECGAGVFMASHFDRHYCGKCCLTYCFNKPEDK |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Ubiquitin family; Eukaryotic ribosomal protein eS31 family
|
|||||
| Subcellular location |
Cytoplasm; Nucleus, nucleolus
|
|||||
| Function |
[Ubiquitin]: Exists either covalently attached to another protein, or free (unanchored). When covalently bound, it is conjugated to target proteins via an isopeptide bond either as a monomer (monoubiquitin), a polymer linked via different Lys residues of the ubiquitin (polyubiquitin chains) or a linear polymer linked via the initiator Met of the ubiquitin (linear polyubiquitin chains). Polyubiquitin chains, when attached to a target protein, have different functions depending on the Lys residue of the ubiquitin that is linked: Lys-6-linked may be involved in DNA repair; Lys-11-linked is involved in ERAD (endoplasmic reticulum-associated degradation) and in cell-cycle regulation; Lys-29-linked is involved in proteotoxic stress response and cell cycle; Lys-33-linked is involved in kinase modification; Lys-48-linked is involved in protein degradation via the proteasome; Lys-63-linked is involved in endocytosis, DNA-damage responses as well as in signaling processes leading to activation of the transcription factor NF-kappa-B. Linear polymer chains formed via attachment by the initiator Met lead to cell signaling. Ubiquitin is usually conjugated to Lys residues of target proteins, however, in rare cases, conjugation to Cys or Ser residues has been observed. When polyubiquitin is free (unanchored-polyubiquitin), it also has distinct roles, such as in activation of protein kinases, and in signaling.; [Small ribosomal subunit protein eS31]: Component of the 40S subunit of the ribosome. Part of the small subunit (SSU) processome, first precursor of the small eukaryotic ribosomal subunit. During the assembly of the SSU processome in the nucleolus, many ribosome biogenesis factors, an RNA chaperone and ribosomal proteins associate with the nascent pre-rRNA and work in concert to generate RNA folding, modifications, rearrangements and cleavage as well as targeted degradation of pre-ribosomal RNA by the RNA exosome.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
11.66 | LDD0402 | [1] | |
|
P1 Probe Info |
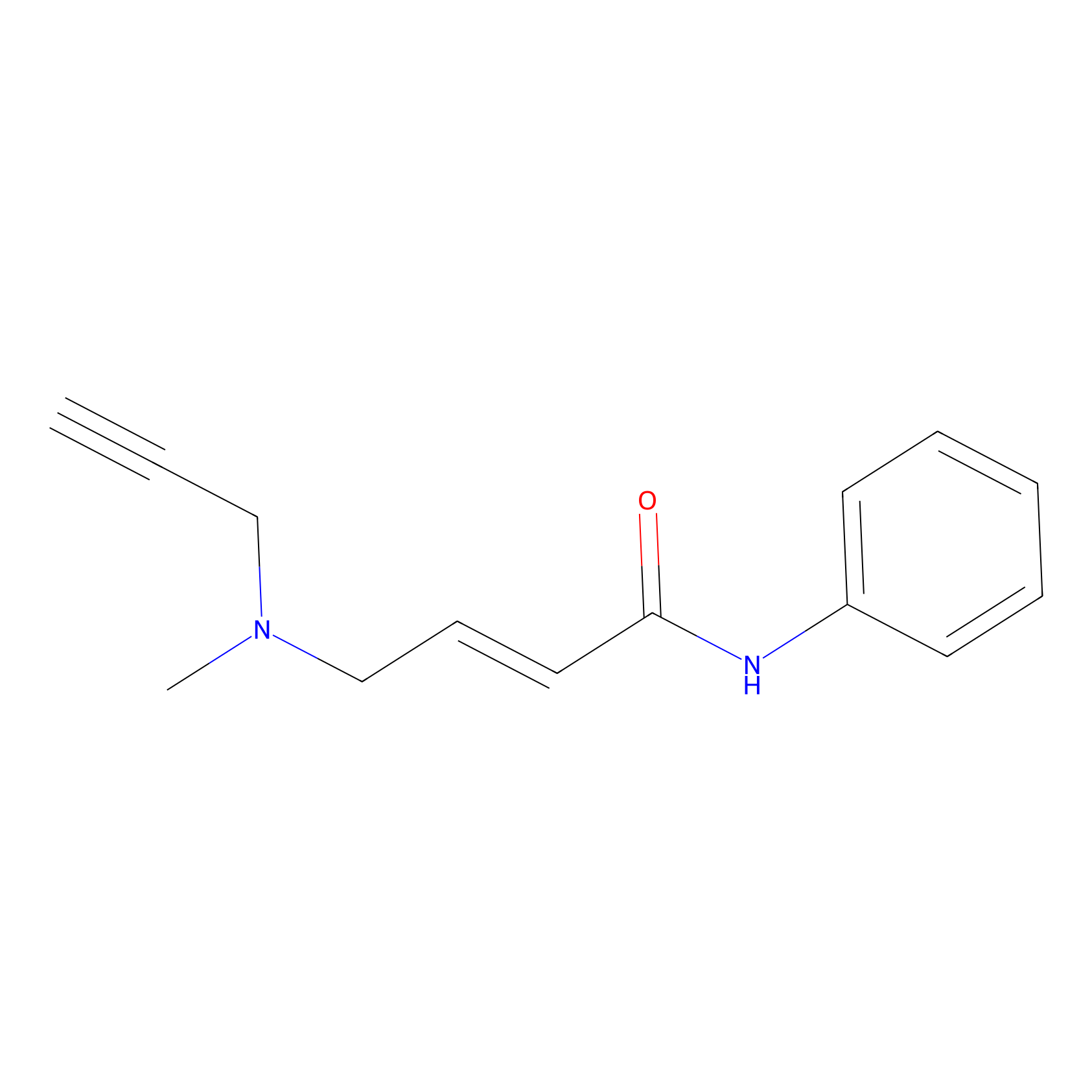 |
2.98 | LDD0448 | [2] | |
|
P2 Probe Info |
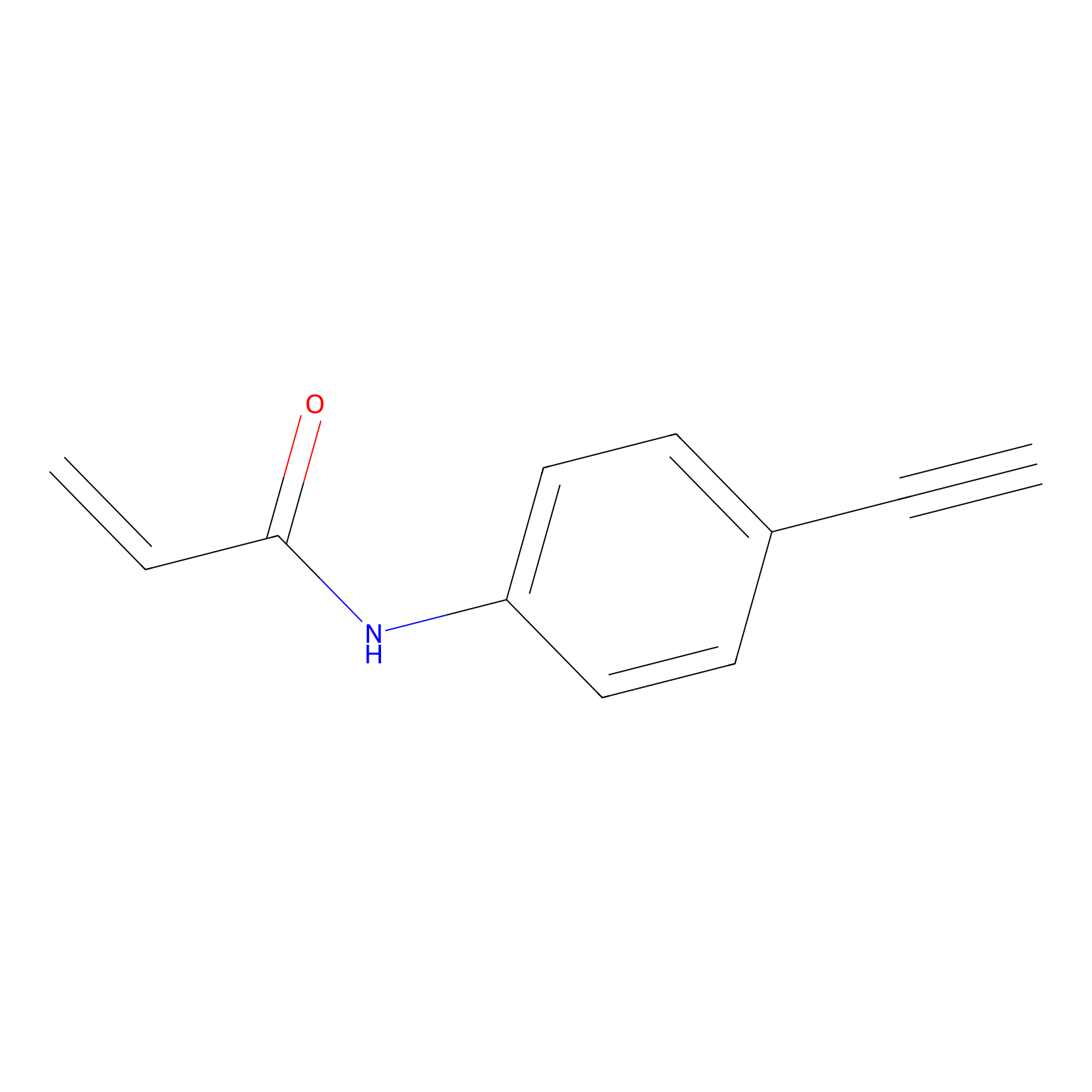 |
2.21 | LDD0449 | [2] | |
|
P3 Probe Info |
 |
1.95 | LDD0450 | [2] | |
|
A-EBA Probe Info |
 |
6.42 | LDD0215 | [3] | |
|
CY-1 Probe Info |
 |
4.26 | LDD0243 | [4] | |
|
N1 Probe Info |
 |
3.94 | LDD0242 | [4] | |
|
TH211 Probe Info |
 |
Y105(20.00); Y106(20.00); Y148(20.00); Y140(15.49) | LDD0257 | [5] | |
|
TH214 Probe Info |
 |
Y105(20.00) | LDD0258 | [5] | |
|
TH216 Probe Info |
 |
Y105(20.00); Y148(20.00); Y85(14.55); Y106(13.83) | LDD0259 | [5] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [6] | |
|
STPyne Probe Info |
 |
K104(0.39); K107(8.19); K11(10.00); K113(5.25) | LDD0277 | [7] | |
|
ONAyne Probe Info |
 |
K107(7.88) | LDD0275 | [7] | |
|
OPA-S-S-alkyne Probe Info |
 |
K113(1.38) | LDD3494 | [8] | |
|
Probe 1 Probe Info |
 |
Y85(184.83) | LDD3495 | [9] | |
|
AF-1 Probe Info |
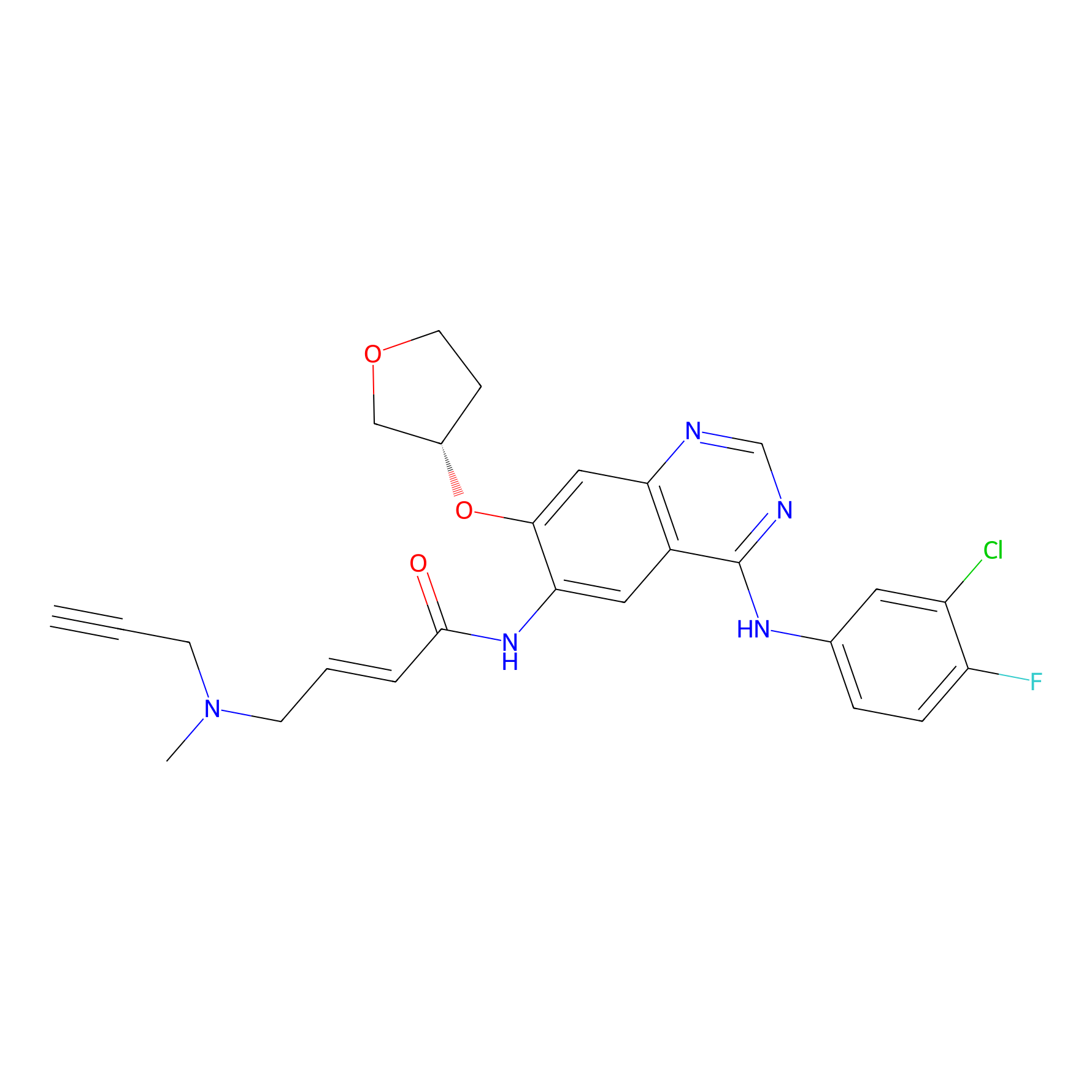 |
1.54 | LDD0421 | [10] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [11] | |
|
AHL-Pu-1 Probe Info |
 |
C121(3.22) | LDD0169 | [12] | |
|
HHS-475 Probe Info |
 |
Y148(0.86) | LDD0264 | [13] | |
|
HHS-465 Probe Info |
 |
Y59(4.27) | LDD2237 | [14] | |
|
DBIA Probe Info |
 |
C144(1.05); C149(1.05); C121(1.03); C126(1.03) | LDD0078 | [15] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [16] | |
|
AMP probe Probe Info |
 |
K48(0.00); K6(0.00) | LDD0200 | [17] | |
|
ATP probe Probe Info |
 |
K48(0.00); K6(0.00); K63(0.00); K11(0.00) | LDD0199 | [17] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C126(0.00); C121(0.00); C144(0.00); C145(0.00) | LDD0038 | [18] | |
|
IA-alkyne Probe Info |
 |
C145(0.00); C144(0.00); C121(0.00) | LDD0032 | [19] | |
|
IPIAA_L Probe Info |
 |
C121(0.00); C144(0.00) | LDD0031 | [20] | |
|
Lodoacetamide azide Probe Info |
 |
C121(0.00); C126(0.00); C144(0.00); C145(0.00) | LDD0037 | [18] | |
|
ATP probe Probe Info |
 |
K6(0.00); K107(0.00); K48(0.00); K29(0.00) | LDD0035 | [21] | |
|
AZ-9 Probe Info |
 |
E24(0.00); E18(0.00) | LDD0395 | [22] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [23] | |
|
JW-RF-010 Probe Info |
 |
C126(0.00); C141(0.00); C145(0.00); C144(0.00) | LDD0026 | [24] | |
|
NAIA_4 Probe Info |
 |
C126(0.00); C121(0.00); 0.00 | LDD2226 | [25] | |
|
TFBX Probe Info |
 |
C149(0.00); C144(0.00); C126(0.00); C121(0.00) | LDD0027 | [24] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [23] | |
|
1d-yne Probe Info |
 |
K6(0.00); K104(0.00); K11(0.00) | LDD0356 | [26] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [23] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [23] | |
|
SF Probe Info |
 |
Y105(0.00); K107(0.00); K48(0.00); K6(0.00) | LDD0028 | [27] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [23] | |
|
Phosphinate-6 Probe Info |
 |
C141(0.00); C121(0.00); C149(0.00); C126(0.00) | LDD0018 | [28] | |
|
1c-yne Probe Info |
 |
K33(0.00); K104(0.00); K11(0.00); K63(0.00) | LDD0228 | [26] | |
|
Acrolein Probe Info |
 |
C149(0.00); C126(0.00) | LDD0217 | [29] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [29] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [29] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [30] | |
|
AOyne Probe Info |
 |
5.70 | LDD0443 | [31] | |
|
NAIA_5 Probe Info |
 |
C144(0.00); C126(0.00); C121(0.00); C149(0.00) | LDD2223 | [25] | |
|
HHS-482 Probe Info |
 |
Y148(1.03); Y85(1.00) | LDD2239 | [14] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
6.00 | LDD0471 | [32] | |
|
FFF probe3 Probe Info |
 |
6.15 | LDD0464 | [32] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [33] | |
|
Alk-rapa Probe Info |
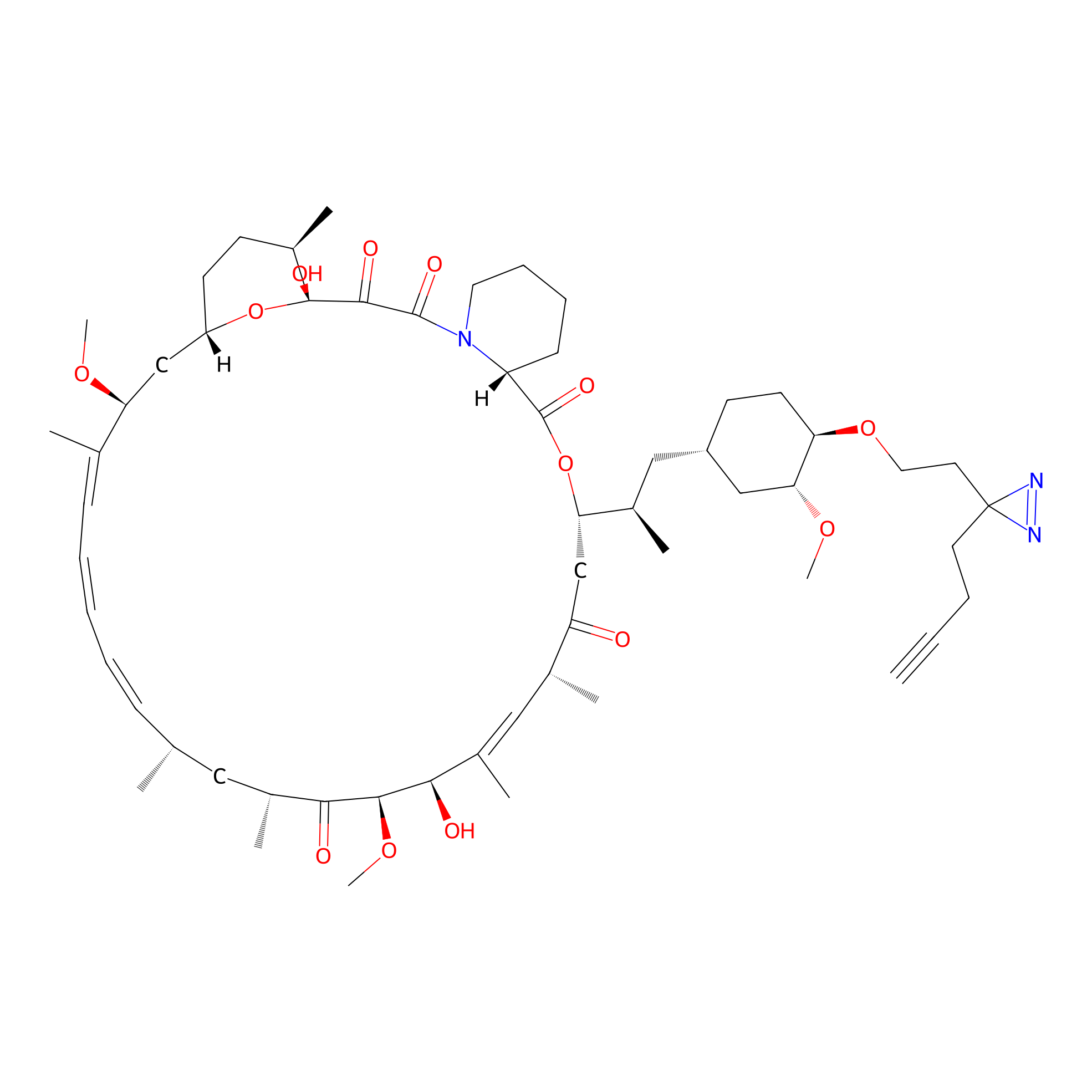 |
4.49 | LDD0213 | [34] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [35] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [36] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C149(1.13) | LDD2112 | [37] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C149(1.14) | LDD2130 | [37] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C149(0.89); C145(1.28); C126(1.19) | LDD2117 | [37] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C149(0.99) | LDD2152 | [37] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C149(1.20) | LDD2103 | [37] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C149(1.44) | LDD2132 | [37] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C121(3.22) | LDD0169 | [12] |
| LDCM0214 | AC1 | HCT 116 | C121(1.62); C126(0.98); C144(0.76); C145(0.73) | LDD0531 | [15] |
| LDCM0215 | AC10 | HCT 116 | C121(0.46); C126(0.43); C144(0.73); C145(0.68) | LDD0532 | [15] |
| LDCM0216 | AC100 | HCT 116 | C121(0.73); C126(0.55); C144(0.41); C145(0.44) | LDD0533 | [15] |
| LDCM0217 | AC101 | HCT 116 | C121(0.64); C126(0.58); C144(0.52); C145(0.60) | LDD0534 | [15] |
| LDCM0218 | AC102 | HCT 116 | C121(0.55); C126(0.47); C144(0.51); C145(0.54) | LDD0535 | [15] |
| LDCM0219 | AC103 | HCT 116 | C121(0.59); C126(0.50); C144(0.41); C145(0.46) | LDD0536 | [15] |
| LDCM0220 | AC104 | HCT 116 | C121(0.62); C126(0.56); C144(0.48); C145(0.50) | LDD0537 | [15] |
| LDCM0221 | AC105 | HCT 116 | C121(0.49); C126(0.44); C144(0.42); C145(0.47) | LDD0538 | [15] |
| LDCM0222 | AC106 | HCT 116 | C121(0.51); C126(0.41); C144(0.45); C145(0.49) | LDD0539 | [15] |
| LDCM0223 | AC107 | HCT 116 | C121(0.62); C126(0.50); C144(0.42); C145(0.54) | LDD0540 | [15] |
| LDCM0224 | AC108 | HCT 116 | C121(0.55); C126(0.46); C144(0.51); C145(0.53) | LDD0541 | [15] |
| LDCM0225 | AC109 | HCT 116 | C121(0.91); C126(1.06); C144(0.69); C145(0.65) | LDD0542 | [15] |
| LDCM0226 | AC11 | HCT 116 | C121(0.46); C126(0.43); C144(0.73); C145(0.68) | LDD0543 | [15] |
| LDCM0227 | AC110 | HCT 116 | C121(0.84); C126(0.60); C144(0.60); C145(0.62) | LDD0544 | [15] |
| LDCM0228 | AC111 | HCT 116 | C121(0.76); C126(0.75); C144(0.56); C145(0.62) | LDD0545 | [15] |
| LDCM0229 | AC112 | HCT 116 | C121(0.65); C126(0.52); C144(0.49); C145(0.55) | LDD0546 | [15] |
| LDCM0230 | AC113 | HCT 116 | C121(0.87); C126(0.88); C144(0.94); C145(0.95) | LDD0547 | [15] |
| LDCM0231 | AC114 | HCT 116 | C121(0.72); C126(0.72); C144(0.67); C145(0.66) | LDD0548 | [15] |
| LDCM0232 | AC115 | HCT 116 | C121(0.51); C126(0.47); C144(0.55); C145(0.49) | LDD0549 | [15] |
| LDCM0233 | AC116 | HCT 116 | C121(0.40); C126(0.39); C144(0.57); C145(0.50) | LDD0550 | [15] |
| LDCM0234 | AC117 | HCT 116 | C121(0.61); C126(0.62); C144(0.70); C145(0.63) | LDD0551 | [15] |
| LDCM0235 | AC118 | HCT 116 | C121(0.55); C126(0.57); C144(0.73); C145(0.66) | LDD0552 | [15] |
| LDCM0236 | AC119 | HCT 116 | C121(0.42); C126(0.45); C144(0.62); C145(0.56) | LDD0553 | [15] |
| LDCM0237 | AC12 | HCT 116 | C121(0.64); C126(0.61); C144(0.88); C145(0.84) | LDD0554 | [15] |
| LDCM0238 | AC120 | HCT 116 | C121(0.98); C126(0.95); C144(0.60); C145(0.53) | LDD0555 | [15] |
| LDCM0239 | AC121 | HCT 116 | C121(0.71); C126(0.74); C144(0.69); C145(0.68) | LDD0556 | [15] |
| LDCM0240 | AC122 | HCT 116 | C121(0.60); C126(0.60); C144(0.88); C145(0.80) | LDD0557 | [15] |
| LDCM0241 | AC123 | HCT 116 | C121(1.24); C126(1.27); C144(1.00); C145(0.96) | LDD0558 | [15] |
| LDCM0242 | AC124 | HCT 116 | C121(0.81); C126(0.80); C144(0.80); C145(0.79) | LDD0559 | [15] |
| LDCM0243 | AC125 | HCT 116 | C121(0.89); C126(0.87); C144(0.84); C145(0.82) | LDD0560 | [15] |
| LDCM0244 | AC126 | HCT 116 | C121(0.58); C126(0.59); C144(0.57); C145(0.53) | LDD0561 | [15] |
| LDCM0245 | AC127 | HCT 116 | C121(0.55); C126(0.54); C144(0.59); C145(0.54) | LDD0562 | [15] |
| LDCM0246 | AC128 | HCT 116 | C121(0.62); C126(0.69); C144(0.56); C145(0.55) | LDD0563 | [15] |
| LDCM0247 | AC129 | HCT 116 | C121(1.35); C126(1.23); C144(1.28); C145(1.29) | LDD0564 | [15] |
| LDCM0249 | AC130 | HCT 116 | C121(0.69); C126(0.68); C144(0.85); C145(0.85) | LDD0566 | [15] |
| LDCM0250 | AC131 | HCT 116 | C121(2.56); C126(1.78); C144(1.42); C145(1.39) | LDD0567 | [15] |
| LDCM0251 | AC132 | HCT 116 | C121(0.50); C126(0.50); C144(0.92); C145(0.90) | LDD0568 | [15] |
| LDCM0252 | AC133 | HCT 116 | C121(0.40); C126(0.39); C144(0.71); C145(0.70) | LDD0569 | [15] |
| LDCM0253 | AC134 | HCT 116 | C121(0.46); C126(0.51); C144(0.69); C145(0.67) | LDD0570 | [15] |
| LDCM0254 | AC135 | HCT 116 | C121(0.37); C126(0.37); C144(0.76); C145(0.74) | LDD0571 | [15] |
| LDCM0255 | AC136 | HCT 116 | C121(0.40); C126(0.39); C144(0.60); C145(0.68) | LDD0572 | [15] |
| LDCM0256 | AC137 | HCT 116 | C121(0.47); C126(0.43); C144(0.65); C145(0.67) | LDD0573 | [15] |
| LDCM0257 | AC138 | HCT 116 | C121(0.47); C126(0.51); C144(0.71); C145(0.74) | LDD0574 | [15] |
| LDCM0258 | AC139 | HCT 116 | C121(0.39); C126(0.39); C144(0.76); C145(0.64) | LDD0575 | [15] |
| LDCM0259 | AC14 | HCT 116 | C121(1.45); C126(1.22); C144(0.93); C145(0.87) | LDD0576 | [15] |
| LDCM0260 | AC140 | HCT 116 | C121(0.30); C126(0.32); C144(0.65); C145(0.60) | LDD0577 | [15] |
| LDCM0261 | AC141 | HCT 116 | C121(0.41); C126(0.42); C144(0.68); C145(0.67) | LDD0578 | [15] |
| LDCM0262 | AC142 | HCT 116 | C121(0.63); C126(0.58); C144(1.19); C145(1.23) | LDD0579 | [15] |
| LDCM0263 | AC143 | HCT 116 | C126(0.64); C121(0.70); C144(0.81); C145(0.83) | LDD0580 | [15] |
| LDCM0264 | AC144 | HCT 116 | C126(0.59); C144(0.64); C121(0.64); C145(0.67) | LDD0581 | [15] |
| LDCM0265 | AC145 | HCT 116 | C126(0.69); C121(0.76); C145(0.83); C144(0.83) | LDD0582 | [15] |
| LDCM0266 | AC146 | HCT 116 | C121(0.56); C126(0.59); C144(0.70); C145(0.70) | LDD0583 | [15] |
| LDCM0267 | AC147 | HCT 116 | C144(0.54); C145(0.55); C126(0.59); C121(0.64) | LDD0584 | [15] |
| LDCM0268 | AC148 | HCT 116 | C126(0.52); C121(0.60); C144(0.62); C145(0.75) | LDD0585 | [15] |
| LDCM0269 | AC149 | HCT 116 | C121(0.53); C126(0.55); C144(0.65); C145(0.69) | LDD0586 | [15] |
| LDCM0270 | AC15 | HCT 116 | C126(0.81); C145(0.84); C121(0.87); C149(0.87) | LDD0587 | [15] |
| LDCM0271 | AC150 | HCT 116 | C126(0.64); C121(0.64); C144(0.80); C145(0.82) | LDD0588 | [15] |
| LDCM0272 | AC151 | HCT 116 | C126(0.77); C121(0.86); C149(0.92); C144(0.96) | LDD0589 | [15] |
| LDCM0273 | AC152 | HCT 116 | C144(0.63); C126(0.67); C145(0.68); C121(0.84) | LDD0590 | [15] |
| LDCM0274 | AC153 | HCT 116 | C126(0.49); C121(0.54); C144(0.72); C145(0.77) | LDD0591 | [15] |
| LDCM0621 | AC154 | HCT 116 | C121(0.69); C126(0.59); C144(0.69); C145(0.65) | LDD2158 | [15] |
| LDCM0622 | AC155 | HCT 116 | C121(0.63); C126(0.59); C144(0.80); C145(0.81) | LDD2159 | [15] |
| LDCM0623 | AC156 | HCT 116 | C121(1.18); C126(1.06); C144(1.06); C145(1.07) | LDD2160 | [15] |
| LDCM0624 | AC157 | HCT 116 | C121(2.56); C126(2.07); C144(1.64); C145(1.62) | LDD2161 | [15] |
| LDCM0276 | AC17 | HCT 116 | C149(0.85); C144(0.91); C145(0.92); C126(1.56) | LDD0593 | [15] |
| LDCM0277 | AC18 | HCT 116 | C126(0.56); C121(0.61); C145(0.64); C144(0.66) | LDD0594 | [15] |
| LDCM0278 | AC19 | HCT 116 | C149(0.76); C145(0.76); C144(0.78); C121(1.16) | LDD0595 | [15] |
| LDCM0279 | AC2 | HCT 116 | C145(0.51); C144(0.60); C126(0.62); C149(0.66) | LDD0596 | [15] |
| LDCM0280 | AC20 | HCT 116 | C149(0.99); C144(1.01); C145(1.06); C126(1.30) | LDD0597 | [15] |
| LDCM0281 | AC21 | HCT 116 | C144(0.94); C145(0.96); C149(0.99); C126(1.05) | LDD0598 | [15] |
| LDCM0282 | AC22 | HCT 116 | C144(1.18); C149(1.19); C145(1.22); C126(1.29) | LDD0599 | [15] |
| LDCM0283 | AC23 | HCT 116 | C144(0.91); C145(0.97); C149(1.03); C126(1.29) | LDD0600 | [15] |
| LDCM0284 | AC24 | HCT 116 | C149(1.06); C144(1.09); C145(1.09); C121(1.60) | LDD0601 | [15] |
| LDCM0285 | AC25 | HCT 116 | C126(0.83); C121(0.86); C149(1.04); C144(1.08) | LDD0602 | [15] |
| LDCM0286 | AC26 | HCT 116 | C121(0.58); C126(0.60); C145(0.88); C144(0.94) | LDD0603 | [15] |
| LDCM0287 | AC27 | HCT 116 | C126(0.73); C121(0.77); C145(0.91); C149(0.96) | LDD0604 | [15] |
| LDCM0288 | AC28 | HCT 116 | C121(0.60); C126(0.62); C145(0.87); C149(0.88) | LDD0605 | [15] |
| LDCM0289 | AC29 | HCT 116 | C126(0.87); C121(0.90); C145(0.90); C149(1.00) | LDD0606 | [15] |
| LDCM0290 | AC3 | HCT 116 | C145(0.57); C126(0.60); C149(0.64); C144(0.65) | LDD0607 | [15] |
| LDCM0291 | AC30 | HCT 116 | C126(0.69); C121(0.74); C145(0.86); C149(0.93) | LDD0608 | [15] |
| LDCM0292 | AC31 | HCT 116 | C121(0.69); C126(0.76); C145(0.81); C149(0.90) | LDD0609 | [15] |
| LDCM0293 | AC32 | HCT 116 | C126(0.53); C121(0.59); C145(0.88); C144(0.98) | LDD0610 | [15] |
| LDCM0294 | AC33 | HCT 116 | C126(0.82); C121(0.86); C145(0.93); C149(1.07) | LDD0611 | [15] |
| LDCM0295 | AC34 | HCT 116 | C121(0.81); C126(0.82); C145(0.95); C149(1.17) | LDD0612 | [15] |
| LDCM0296 | AC35 | HCT 116 | C126(0.92); C144(0.94); C121(0.98); C149(0.98) | LDD0613 | [15] |
| LDCM0297 | AC36 | HCT 116 | C121(0.74); C126(0.83); C144(0.99); C145(1.05) | LDD0614 | [15] |
| LDCM0298 | AC37 | HCT 116 | C121(0.73); C126(0.83); C144(0.96); C145(1.05) | LDD0615 | [15] |
| LDCM0299 | AC38 | HCT 116 | C121(0.88); C126(0.94); C144(1.02); C145(1.06) | LDD0616 | [15] |
| LDCM0300 | AC39 | HCT 116 | C126(0.46); C121(0.47); C149(0.73); C145(0.75) | LDD0617 | [15] |
| LDCM0301 | AC4 | HCT 116 | C145(0.53); C126(0.57); C144(0.58); C149(0.62) | LDD0618 | [15] |
| LDCM0302 | AC40 | HCT 116 | C126(0.65); C121(0.66); C145(0.80); C149(0.82) | LDD0619 | [15] |
| LDCM0303 | AC41 | HCT 116 | C121(0.67); C126(0.73); C149(0.95); C145(1.01) | LDD0620 | [15] |
| LDCM0304 | AC42 | HCT 116 | C121(0.81); C126(0.82); C149(0.95); C145(0.99) | LDD0621 | [15] |
| LDCM0305 | AC43 | HCT 116 | C121(0.68); C126(0.70); C145(0.97); C144(1.05) | LDD0622 | [15] |
| LDCM0306 | AC44 | HCT 116 | C121(0.68); C126(0.69); C149(0.92); C145(0.93) | LDD0623 | [15] |
| LDCM0307 | AC45 | HCT 116 | C126(0.64); C121(0.64); C149(0.84); C144(0.85) | LDD0624 | [15] |
| LDCM0308 | AC46 | HCT 116 | C144(0.90); C149(0.99); C145(1.10); C126(1.14) | LDD0625 | [15] |
| LDCM0309 | AC47 | HCT 116 | C126(0.71); C121(0.73); C145(0.90); C144(0.90) | LDD0626 | [15] |
| LDCM0310 | AC48 | HCT 116 | C149(0.83); C126(0.97); C144(0.97); C121(1.00) | LDD0627 | [15] |
| LDCM0311 | AC49 | HCT 116 | C145(1.03); C144(1.24); C126(1.50); C121(1.51) | LDD0628 | [15] |
| LDCM0312 | AC5 | HCT 116 | C126(0.39); C145(0.53); C144(0.59); C121(0.63) | LDD0629 | [15] |
| LDCM0313 | AC50 | HCT 116 | C145(1.07); C144(1.15); C126(1.23); C121(1.49) | LDD0630 | [15] |
| LDCM0314 | AC51 | HCT 116 | C149(1.17); C144(1.39); C145(1.57); C126(2.01) | LDD0631 | [15] |
| LDCM0315 | AC52 | HCT 116 | C126(1.07); C121(1.07); C144(1.20); C145(1.27) | LDD0632 | [15] |
| LDCM0316 | AC53 | HCT 116 | C126(1.02); C145(1.04); C144(1.09); C121(1.10) | LDD0633 | [15] |
| LDCM0317 | AC54 | HCT 116 | C126(0.89); C145(0.99); C121(1.02); C144(1.17) | LDD0634 | [15] |
| LDCM0318 | AC55 | HCT 116 | C126(0.82); C121(0.88); C145(1.00); C144(1.02) | LDD0635 | [15] |
| LDCM0319 | AC56 | HCT 116 | C126(1.15); C121(1.25); C144(1.31); C145(1.34) | LDD0636 | [15] |
| LDCM0320 | AC57 | HCT 116 | C121(0.40); C126(0.46); C145(0.50); C144(0.55) | LDD0637 | [15] |
| LDCM0321 | AC58 | HCT 116 | C145(0.66); C121(0.72); C144(0.78); C149(0.80) | LDD0638 | [15] |
| LDCM0322 | AC59 | HCT 116 | C121(0.43); C145(0.46); C126(0.51); C144(0.54) | LDD0639 | [15] |
| LDCM0323 | AC6 | HCT 116 | C126(0.54); C121(0.57); C145(0.78); C144(0.84) | LDD0640 | [15] |
| LDCM0324 | AC60 | HCT 116 | C121(0.53); C126(0.63); C145(0.68); C144(0.80) | LDD0641 | [15] |
| LDCM0325 | AC61 | HCT 116 | C121(0.62); C126(0.63); C145(0.99); C149(1.01) | LDD0642 | [15] |
| LDCM0326 | AC62 | HCT 116 | C121(0.39); C126(0.46); C145(0.55); C144(0.57) | LDD0643 | [15] |
| LDCM0327 | AC63 | HCT 116 | C121(0.50); C126(0.62); C145(0.66); C144(0.69) | LDD0644 | [15] |
| LDCM0328 | AC64 | HCT 116 | C121(0.42); C145(0.49); C126(0.49); C144(0.52) | LDD0645 | [15] |
| LDCM0329 | AC65 | HCT 116 | C121(0.38); C126(0.45); C145(0.46); C144(0.48) | LDD0646 | [15] |
| LDCM0330 | AC66 | HCT 116 | C145(0.49); C144(0.51); C149(0.56); C121(0.57) | LDD0647 | [15] |
| LDCM0331 | AC67 | HCT 116 | C121(0.43); C145(0.43); C126(0.44); C144(0.46) | LDD0648 | [15] |
| LDCM0332 | AC68 | HCT 116 | C121(0.91); C144(1.01); C126(1.02); C145(1.14) | LDD0649 | [15] |
| LDCM0333 | AC69 | HCT 116 | C126(1.07); C144(1.12); C121(1.12); C145(1.27) | LDD0650 | [15] |
| LDCM0334 | AC7 | HCT 116 | C126(0.54); C121(0.55); C145(0.80); C149(0.86) | LDD0651 | [15] |
| LDCM0335 | AC70 | HCT 116 | C121(0.62); C126(0.73); C144(0.96); C145(0.99) | LDD0652 | [15] |
| LDCM0336 | AC71 | HCT 116 | C121(0.86); C126(0.97); C144(1.14); C149(1.22) | LDD0653 | [15] |
| LDCM0337 | AC72 | HCT 116 | C126(0.89); C121(0.95); C144(0.97); C149(1.07) | LDD0654 | [15] |
| LDCM0338 | AC73 | HCT 116 | C121(0.69); C126(0.87); C144(1.03); C149(1.10) | LDD0655 | [15] |
| LDCM0339 | AC74 | HCT 116 | C121(0.70); C126(0.84); C144(0.98); C145(1.03) | LDD0656 | [15] |
| LDCM0340 | AC75 | HCT 116 | C121(0.77); C126(0.87); C144(1.03); C149(1.17) | LDD0657 | [15] |
| LDCM0341 | AC76 | HCT 116 | C121(0.80); C126(0.87); C144(0.98); C145(1.12) | LDD0658 | [15] |
| LDCM0342 | AC77 | HCT 116 | C121(1.10); C144(1.11); C126(1.14); C149(1.30) | LDD0659 | [15] |
| LDCM0343 | AC78 | HCT 116 | C126(1.21); C121(1.28); C149(1.50); C144(1.51) | LDD0660 | [15] |
| LDCM0344 | AC79 | HCT 116 | C121(0.87); C126(0.94); C144(1.11); C145(1.28) | LDD0661 | [15] |
| LDCM0345 | AC8 | HCT 116 | C126(0.56); C121(0.62); C145(0.84); C144(0.85) | LDD0662 | [15] |
| LDCM0346 | AC80 | HCT 116 | C121(0.78); C126(0.84); C144(1.02); C145(1.14) | LDD0663 | [15] |
| LDCM0347 | AC81 | HCT 116 | C121(0.95); C126(1.00); C144(1.28); C149(1.48) | LDD0664 | [15] |
| LDCM0348 | AC82 | HCT 116 | C121(0.62); C126(0.77); C144(0.99); C145(1.12) | LDD0665 | [15] |
| LDCM0349 | AC83 | HCT 116 | C144(0.27); C145(0.49); C126(0.50); C149(0.53) | LDD0666 | [15] |
| LDCM0350 | AC84 | HCT 116 | C144(0.28); C145(0.51); C149(0.54); C121(0.57) | LDD0667 | [15] |
| LDCM0351 | AC85 | HCT 116 | C144(0.46); C126(0.53); C121(0.59); C145(0.63) | LDD0668 | [15] |
| LDCM0352 | AC86 | HCT 116 | C144(0.36); C145(0.51); C149(0.55); C126(0.65) | LDD0669 | [15] |
| LDCM0353 | AC87 | HCT 116 | C144(0.79); C149(0.92); C145(0.93); C126(1.13) | LDD0670 | [15] |
| LDCM0354 | AC88 | HCT 116 | C144(0.49); C145(0.59); C149(0.61); C126(0.83) | LDD0671 | [15] |
| LDCM0355 | AC89 | HCT 116 | C144(0.40); C145(0.53); C149(0.58); C126(0.76) | LDD0672 | [15] |
| LDCM0357 | AC90 | HCT 116 | C149(0.73); C145(0.79); C144(0.88); C126(1.15) | LDD0674 | [15] |
| LDCM0358 | AC91 | HCT 116 | C144(0.28); C145(0.45); C149(0.48); C121(0.60) | LDD0675 | [15] |
| LDCM0359 | AC92 | HCT 116 | C144(0.28); C145(0.43); C149(0.46); C126(0.49) | LDD0676 | [15] |
| LDCM0360 | AC93 | HCT 116 | C144(0.43); C145(0.57); C149(0.58); C126(0.69) | LDD0677 | [15] |
| LDCM0361 | AC94 | HCT 116 | C144(0.65); C149(0.67); C145(0.70); C126(1.05) | LDD0678 | [15] |
| LDCM0362 | AC95 | HCT 116 | C144(0.81); C145(1.02); C149(1.04); C121(1.48) | LDD0679 | [15] |
| LDCM0363 | AC96 | HCT 116 | C144(0.49); C145(0.57); C149(0.62); C126(0.80) | LDD0680 | [15] |
| LDCM0364 | AC97 | HCT 116 | C144(0.30); C145(0.44); C149(0.47); C126(0.58) | LDD0681 | [15] |
| LDCM0365 | AC98 | HCT 116 | C126(0.29); C121(0.34); C144(0.38); C145(0.43) | LDD0682 | [15] |
| LDCM0366 | AC99 | HCT 116 | C144(0.51); C145(0.54); C149(0.56); C126(0.69) | LDD0683 | [15] |
| LDCM0166 | Afatinib | A431 | 1.54 | LDD0421 | [10] |
| LDCM0248 | AKOS034007472 | HCT 116 | C121(0.54); C126(0.52); C144(0.90); C145(0.89) | LDD0565 | [15] |
| LDCM0356 | AKOS034007680 | HCT 116 | C126(0.45); C121(0.46); C145(0.77); C149(0.83) | LDD0673 | [15] |
| LDCM0275 | AKOS034007705 | HCT 116 | C145(0.57); C126(0.60); C121(0.64); C144(0.68) | LDD0592 | [15] |
| LDCM0156 | Aniline | NCI-H1299 | N.A. | LDD0404 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C144(1.05); C149(1.05); C121(1.03); C126(1.03) | LDD0078 | [15] |
| LDCM0102 | BDHI 8 | Jurkat | C144(4.63); C145(4.63); C149(4.63) | LDD0204 | [38] |
| LDCM0632 | CL-Sc | Hep-G2 | C121(1.78); C144(0.76); C121(0.20) | LDD2227 | [25] |
| LDCM0367 | CL1 | HCT 116 | C121(1.02); C126(1.06); C149(1.11); C144(1.13) | LDD0684 | [15] |
| LDCM0368 | CL10 | HCT 116 | C121(0.73); C126(0.80); C144(0.86); C145(1.01) | LDD0685 | [15] |
| LDCM0369 | CL100 | HCT 116 | C145(0.54); C144(0.58); C149(0.60); C126(0.69) | LDD0686 | [15] |
| LDCM0370 | CL101 | HCT 116 | C145(0.84); C149(0.87); C144(0.96); C126(1.75) | LDD0687 | [15] |
| LDCM0371 | CL102 | HCT 116 | C145(0.95); C144(0.99); C149(1.03); C126(2.64) | LDD0688 | [15] |
| LDCM0372 | CL103 | HCT 116 | C144(1.02); C145(1.05); C149(1.11); C126(2.05) | LDD0689 | [15] |
| LDCM0373 | CL104 | HCT 116 | C145(0.96); C144(1.06); C149(1.06); C126(1.69) | LDD0690 | [15] |
| LDCM0374 | CL105 | HCT 116 | C126(0.53); C121(0.60); C145(0.66); C144(0.72) | LDD0691 | [15] |
| LDCM0375 | CL106 | HCT 116 | C126(0.55); C121(0.60); C144(0.72); C145(0.74) | LDD0692 | [15] |
| LDCM0376 | CL107 | HCT 116 | C126(0.42); C121(0.46); C145(0.59); C144(0.60) | LDD0693 | [15] |
| LDCM0377 | CL108 | HCT 116 | C126(0.40); C121(0.46); C145(0.60); C144(0.63) | LDD0694 | [15] |
| LDCM0378 | CL109 | HCT 116 | C144(0.70); C145(0.72); C149(0.80); C126(0.86) | LDD0695 | [15] |
| LDCM0379 | CL11 | HCT 116 | C121(0.88); C144(0.90); C126(0.92); C145(1.04) | LDD0696 | [15] |
| LDCM0380 | CL110 | HCT 116 | C126(0.46); C121(0.50); C144(0.52); C145(0.55) | LDD0697 | [15] |
| LDCM0381 | CL111 | HCT 116 | C145(0.67); C144(0.67); C126(0.83); C149(0.87) | LDD0698 | [15] |
| LDCM0382 | CL112 | HCT 116 | C126(0.91); C121(0.96); C144(0.99); C145(1.01) | LDD0699 | [15] |
| LDCM0383 | CL113 | HCT 116 | C126(0.74); C121(0.78); C145(0.96); C149(1.13) | LDD0700 | [15] |
| LDCM0384 | CL114 | HCT 116 | C121(0.85); C126(0.91); C145(0.96); C149(1.06) | LDD0701 | [15] |
| LDCM0385 | CL115 | HCT 116 | C121(0.84); C126(0.85); C145(0.94); C149(1.01) | LDD0702 | [15] |
| LDCM0386 | CL116 | HCT 116 | C126(0.57); C121(0.58); C145(0.87); C149(0.95) | LDD0703 | [15] |
| LDCM0387 | CL117 | HCT 116 | C145(0.86); C149(0.93); C144(0.97); C126(1.00) | LDD0704 | [15] |
| LDCM0388 | CL118 | HCT 116 | C126(0.50); C121(0.50); C149(0.77); C144(0.77) | LDD0705 | [15] |
| LDCM0389 | CL119 | HCT 116 | C121(0.66); C126(0.69); C144(0.82); C145(0.83) | LDD0706 | [15] |
| LDCM0390 | CL12 | HCT 116 | C144(0.89); C145(0.94); C126(0.97); C121(1.00) | LDD0707 | [15] |
| LDCM0391 | CL120 | HCT 116 | C126(0.51); C121(0.52); C144(0.84); C145(0.85) | LDD0708 | [15] |
| LDCM0392 | CL121 | HCT 116 | C149(1.04); C144(1.19); C145(1.27); C126(1.84) | LDD0709 | [15] |
| LDCM0393 | CL122 | HCT 116 | C126(0.93); C121(0.94); C144(1.20); C145(1.23) | LDD0710 | [15] |
| LDCM0394 | CL123 | HCT 116 | C126(0.73); C121(0.80); C145(0.89); C144(0.90) | LDD0711 | [15] |
| LDCM0395 | CL124 | HCT 116 | C126(0.69); C121(0.74); C144(0.90); C145(0.96) | LDD0712 | [15] |
| LDCM0396 | CL125 | HCT 116 | C121(0.61); C126(0.71); C145(0.73); C149(0.76) | LDD0713 | [15] |
| LDCM0397 | CL126 | HCT 116 | C121(0.75); C126(0.77); C149(1.54); C145(1.55) | LDD0714 | [15] |
| LDCM0398 | CL127 | HCT 116 | C121(0.68); C126(0.78); C145(0.84); C144(0.91) | LDD0715 | [15] |
| LDCM0399 | CL128 | HCT 116 | C121(0.47); C145(0.49); C126(0.51); C144(0.55) | LDD0716 | [15] |
| LDCM0400 | CL13 | HCT 116 | C144(1.05); C145(1.07); C126(1.18); C149(1.18) | LDD0717 | [15] |
| LDCM0401 | CL14 | HCT 116 | C144(1.06); C145(1.10); C149(1.10); C121(1.20) | LDD0718 | [15] |
| LDCM0402 | CL15 | HCT 116 | C121(1.11); C144(1.13); C126(1.14); C145(1.34) | LDD0719 | [15] |
| LDCM0403 | CL16 | HCT 116 | C121(0.92); C126(0.93); C145(1.04); C144(1.05) | LDD0720 | [15] |
| LDCM0404 | CL17 | HCT 116 | C126(0.99); C121(1.05); C144(1.14); C145(1.23) | LDD0721 | [15] |
| LDCM0405 | CL18 | HCT 116 | C149(2.05); C144(2.41); C145(2.47); C126(2.59) | LDD0722 | [15] |
| LDCM0406 | CL19 | HCT 116 | C126(1.13); C121(1.17); C145(1.24); C144(1.29) | LDD0723 | [15] |
| LDCM0407 | CL2 | HCT 116 | C126(1.07); C149(1.12); C144(1.13); C121(1.15) | LDD0724 | [15] |
| LDCM0408 | CL20 | HCT 116 | C121(0.87); C126(0.89); C144(1.02); C145(1.03) | LDD0725 | [15] |
| LDCM0409 | CL21 | HCT 116 | C126(0.75); C121(0.77); C145(0.93); C144(0.96) | LDD0726 | [15] |
| LDCM0410 | CL22 | HCT 116 | C121(1.01); C145(1.06); C126(1.07); C144(1.10) | LDD0727 | [15] |
| LDCM0411 | CL23 | HCT 116 | C145(1.04); C144(1.08); C149(1.17); C121(1.64) | LDD0728 | [15] |
| LDCM0412 | CL24 | HCT 116 | C121(0.76); C126(0.76); C144(0.89); C145(0.93) | LDD0729 | [15] |
| LDCM0413 | CL25 | HCT 116 | C121(0.60); C126(0.58); C144(0.79); C145(0.79) | LDD0730 | [15] |
| LDCM0414 | CL26 | HCT 116 | C121(1.02); C126(1.01); C144(0.99); C145(1.05) | LDD0731 | [15] |
| LDCM0415 | CL27 | HCT 116 | C121(0.64); C126(0.62); C144(0.93); C145(0.93) | LDD0732 | [15] |
| LDCM0416 | CL28 | HCT 116 | C121(0.95); C126(0.95); C144(1.05); C145(1.04) | LDD0733 | [15] |
| LDCM0417 | CL29 | HCT 116 | C121(0.83); C126(0.82); C144(1.02); C145(1.00) | LDD0734 | [15] |
| LDCM0418 | CL3 | HCT 116 | C121(1.32); C126(1.20); C144(1.02); C145(1.20) | LDD0735 | [15] |
| LDCM0419 | CL30 | HCT 116 | C121(1.26); C126(1.19); C144(1.30); C145(1.28) | LDD0736 | [15] |
| LDCM0420 | CL31 | HCT 116 | C121(0.83); C126(0.86); C144(0.91); C145(0.90) | LDD0737 | [15] |
| LDCM0421 | CL32 | HCT 116 | C121(1.39); C126(1.40); C144(1.59); C145(1.51) | LDD0738 | [15] |
| LDCM0422 | CL33 | HCT 116 | C121(1.29); C126(1.46); C144(1.19); C145(1.20) | LDD0739 | [15] |
| LDCM0423 | CL34 | HCT 116 | C121(0.81); C126(0.87); C144(0.92); C145(0.85) | LDD0740 | [15] |
| LDCM0424 | CL35 | HCT 116 | C121(0.95); C126(1.02); C144(0.79); C145(0.85) | LDD0741 | [15] |
| LDCM0425 | CL36 | HCT 116 | C121(1.35); C126(1.49); C144(1.25); C145(1.04) | LDD0742 | [15] |
| LDCM0426 | CL37 | HCT 116 | C121(0.81); C126(0.94); C144(0.70); C145(0.78) | LDD0743 | [15] |
| LDCM0428 | CL39 | HCT 116 | C121(0.68); C126(0.78); C144(0.78); C145(0.80) | LDD0745 | [15] |
| LDCM0429 | CL4 | HCT 116 | C121(1.05); C126(1.03); C144(1.01); C145(1.07) | LDD0746 | [15] |
| LDCM0430 | CL40 | HCT 116 | C121(0.70); C126(0.77); C144(0.83); C145(0.84) | LDD0747 | [15] |
| LDCM0431 | CL41 | HCT 116 | C121(1.38); C126(1.30); C144(1.17); C145(1.01) | LDD0748 | [15] |
| LDCM0432 | CL42 | HCT 116 | C121(0.90); C126(0.93); C144(0.91); C145(1.01) | LDD0749 | [15] |
| LDCM0433 | CL43 | HCT 116 | C121(1.11); C126(1.25); C144(0.98); C145(0.92) | LDD0750 | [15] |
| LDCM0434 | CL44 | HCT 116 | C121(1.06); C126(1.18); C144(1.04); C145(1.01) | LDD0751 | [15] |
| LDCM0435 | CL45 | HCT 116 | C121(0.72); C126(0.89); C144(0.80); C145(0.79) | LDD0752 | [15] |
| LDCM0436 | CL46 | HCT 116 | C121(2.81); C126(2.80); C144(1.10); C145(1.12) | LDD0753 | [15] |
| LDCM0437 | CL47 | HCT 116 | C121(2.14); C126(2.12); C144(1.02); C145(1.10) | LDD0754 | [15] |
| LDCM0438 | CL48 | HCT 116 | C121(2.06); C126(2.05); C144(1.51); C145(1.36) | LDD0755 | [15] |
| LDCM0439 | CL49 | HCT 116 | C121(2.71); C126(2.70); C144(0.97); C145(0.92) | LDD0756 | [15] |
| LDCM0440 | CL5 | HCT 116 | C121(1.08); C126(1.06); C144(1.09); C145(1.09) | LDD0757 | [15] |
| LDCM0441 | CL50 | HCT 116 | C121(1.91); C126(1.91); C144(1.04); C145(1.04) | LDD0758 | [15] |
| LDCM0442 | CL51 | HCT 116 | C121(1.64); C126(1.63); C144(0.89); C145(0.96) | LDD0759 | [15] |
| LDCM0443 | CL52 | HCT 116 | C121(1.74); C126(1.73); C144(1.18); C145(1.10) | LDD0760 | [15] |
| LDCM0444 | CL53 | HCT 116 | C121(3.61); C126(3.61); C144(1.47); C145(1.59) | LDD0761 | [15] |
| LDCM0445 | CL54 | HCT 116 | C121(2.75); C126(2.73); C144(1.08); C145(1.06) | LDD0762 | [15] |
| LDCM0446 | CL55 | HCT 116 | C121(3.07); C126(3.08); C144(1.26); C145(1.31) | LDD0763 | [15] |
| LDCM0447 | CL56 | HCT 116 | C121(1.33); C126(1.35); C144(1.04); C145(1.11) | LDD0764 | [15] |
| LDCM0448 | CL57 | HCT 116 | C121(1.95); C126(1.94); C144(1.42); C145(1.31) | LDD0765 | [15] |
| LDCM0449 | CL58 | HCT 116 | C121(2.71); C126(2.71); C144(1.26); C145(1.23) | LDD0766 | [15] |
| LDCM0450 | CL59 | HCT 116 | C121(1.44); C126(1.43); C144(1.02); C145(1.03) | LDD0767 | [15] |
| LDCM0451 | CL6 | HCT 116 | C121(1.16); C126(1.20); C144(1.30); C145(1.16) | LDD0768 | [15] |
| LDCM0452 | CL60 | HCT 116 | C121(1.78); C126(1.77); C144(0.96); C145(0.97) | LDD0769 | [15] |
| LDCM0453 | CL61 | HCT 116 | C121(0.73); C126(0.76); C144(1.09); C145(0.92) | LDD0770 | [15] |
| LDCM0454 | CL62 | HCT 116 | C121(0.79); C126(0.79); C144(0.97); C145(0.83) | LDD0771 | [15] |
| LDCM0455 | CL63 | HCT 116 | C121(0.56); C126(0.60); C144(0.88); C145(0.73) | LDD0772 | [15] |
| LDCM0456 | CL64 | HCT 116 | C121(0.57); C126(0.61); C144(0.83); C145(0.63) | LDD0773 | [15] |
| LDCM0457 | CL65 | HCT 116 | C121(0.73); C126(0.77); C144(0.87); C145(0.71) | LDD0774 | [15] |
| LDCM0458 | CL66 | HCT 116 | C121(0.48); C126(0.55); C144(0.77); C145(0.65) | LDD0775 | [15] |
| LDCM0459 | CL67 | HCT 116 | C121(0.80); C126(0.68); C144(1.25); C145(1.01) | LDD0776 | [15] |
| LDCM0460 | CL68 | HCT 116 | C121(1.01); C126(0.87); C144(1.38); C145(1.02) | LDD0777 | [15] |
| LDCM0461 | CL69 | HCT 116 | C121(1.13); C126(1.10); C144(1.98); C145(1.36) | LDD0778 | [15] |
| LDCM0462 | CL7 | HCT 116 | C121(0.54); C126(0.63); C144(0.91); C145(0.91) | LDD0779 | [15] |
| LDCM0463 | CL70 | HCT 116 | C121(0.90); C126(0.88); C144(1.19); C145(0.95) | LDD0780 | [15] |
| LDCM0464 | CL71 | HCT 116 | C121(0.62); C126(0.65); C144(0.79); C145(0.74) | LDD0781 | [15] |
| LDCM0465 | CL72 | HCT 116 | C121(0.73); C126(0.80); C144(1.05); C145(0.85) | LDD0782 | [15] |
| LDCM0466 | CL73 | HCT 116 | C121(0.86); C126(0.83); C144(1.13); C145(0.82) | LDD0783 | [15] |
| LDCM0467 | CL74 | HCT 116 | C121(0.86); C126(0.80); C144(1.16); C145(0.87) | LDD0784 | [15] |
| LDCM0469 | CL76 | HCT 116 | C121(1.08); C126(1.09); C144(0.96); C145(1.03) | LDD0786 | [15] |
| LDCM0470 | CL77 | HCT 116 | C121(1.09); C126(1.15); C144(1.50); C145(1.74) | LDD0787 | [15] |
| LDCM0471 | CL78 | HCT 116 | C121(0.65); C126(0.64); C144(0.71); C145(0.73) | LDD0788 | [15] |
| LDCM0472 | CL79 | HCT 116 | C121(0.69); C126(0.72); C144(0.66); C145(0.70) | LDD0789 | [15] |
| LDCM0473 | CL8 | HCT 116 | C121(0.88); C126(1.00); C144(0.99); C145(0.99) | LDD0790 | [15] |
| LDCM0474 | CL80 | HCT 116 | C121(1.11); C126(1.13); C144(1.16); C145(1.41) | LDD0791 | [15] |
| LDCM0475 | CL81 | HCT 116 | C121(0.65); C126(0.69); C144(0.66); C145(0.72) | LDD0792 | [15] |
| LDCM0476 | CL82 | HCT 116 | C121(0.57); C126(0.56); C144(0.64); C145(0.69) | LDD0793 | [15] |
| LDCM0477 | CL83 | HCT 116 | C121(0.61); C126(0.60); C144(0.66); C145(0.70) | LDD0794 | [15] |
| LDCM0478 | CL84 | HCT 116 | C121(0.60); C126(0.58); C144(0.69); C145(0.73) | LDD0795 | [15] |
| LDCM0479 | CL85 | HCT 116 | C121(0.80); C126(0.78); C144(0.76); C145(0.79) | LDD0796 | [15] |
| LDCM0480 | CL86 | HCT 116 | C121(0.90); C126(0.88); C144(0.86); C145(0.87) | LDD0797 | [15] |
| LDCM0481 | CL87 | HCT 116 | C121(0.65); C126(0.65); C144(0.81); C145(0.84) | LDD0798 | [15] |
| LDCM0482 | CL88 | HCT 116 | C121(0.81); C126(0.92); C144(1.05); C145(1.00) | LDD0799 | [15] |
| LDCM0483 | CL89 | HCT 116 | C121(0.63); C126(0.64); C144(0.77); C145(0.77) | LDD0800 | [15] |
| LDCM0484 | CL9 | HCT 116 | C121(0.94); C126(1.16); C144(0.94); C145(0.96) | LDD0801 | [15] |
| LDCM0485 | CL90 | HCT 116 | C121(0.86); C126(0.93); C144(0.76); C145(0.79) | LDD0802 | [15] |
| LDCM0486 | CL91 | HCT 116 | C121(0.80); C126(0.96); C144(0.67); C145(0.61) | LDD0803 | [15] |
| LDCM0487 | CL92 | HCT 116 | C121(0.75); C126(0.99); C144(0.76); C145(0.70) | LDD0804 | [15] |
| LDCM0488 | CL93 | HCT 116 | C121(0.83); C126(0.85); C144(0.90); C145(0.81) | LDD0805 | [15] |
| LDCM0489 | CL94 | HCT 116 | C121(0.65); C126(0.70); C144(0.78); C145(0.68) | LDD0806 | [15] |
| LDCM0490 | CL95 | HCT 116 | C121(0.59); C126(0.67); C144(0.74); C145(0.63) | LDD0807 | [15] |
| LDCM0491 | CL96 | HCT 116 | C121(0.72); C126(0.69); C144(0.64); C145(0.58) | LDD0808 | [15] |
| LDCM0492 | CL97 | HCT 116 | C121(0.77); C126(0.67); C144(0.62); C145(0.53) | LDD0809 | [15] |
| LDCM0493 | CL98 | HCT 116 | C121(0.85); C126(0.47); C144(0.50); C145(0.47) | LDD0810 | [15] |
| LDCM0494 | CL99 | HCT 116 | C121(0.76); C126(0.54); C144(0.53); C145(0.47) | LDD0811 | [15] |
| LDCM0495 | E2913 | HEK-293T | C121(1.02); C149(0.80) | LDD1698 | [39] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C126(1.68); C121(1.22); C144(1.06); C149(0.86) | LDD1702 | [37] |
| LDCM0573 | Fragment11 | Ramos | 0.19 | LDD2190 | [40] |
| LDCM0576 | Fragment14 | Ramos | 0.44 | LDD2193 | [40] |
| LDCM0468 | Fragment33 | HCT 116 | C121(0.55); C126(0.59); C144(0.95); C145(0.74) | LDD0785 | [15] |
| LDCM0427 | Fragment51 | HCT 116 | C121(0.81); C126(0.88); C144(0.85); C145(0.86) | LDD0744 | [15] |
| LDCM0116 | HHS-0101 | DM93 | Y148(0.86) | LDD0264 | [13] |
| LDCM0117 | HHS-0201 | DM93 | Y148(0.86) | LDD0265 | [13] |
| LDCM0118 | HHS-0301 | DM93 | Y148(0.73) | LDD0266 | [13] |
| LDCM0119 | HHS-0401 | DM93 | Y148(6.19) | LDD0267 | [13] |
| LDCM0120 | HHS-0701 | DM93 | Y148(2.28) | LDD0268 | [13] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [29] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [11] |
| LDCM0022 | KB02 | HCT 116 | C145(3.44); C121(2.23); C126(2.23); C149(1.22) | LDD0080 | [15] |
| LDCM0023 | KB03 | HCT 116 | C145(3.65); C121(2.85); C126(2.85); C149(0.95) | LDD0081 | [15] |
| LDCM0024 | KB05 | HCT 116 | C145(2.71); C121(2.55); C126(2.55); C149(1.29) | LDD0082 | [15] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C149(1.16) | LDD2102 | [37] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C149(1.40) | LDD2121 | [37] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0227 | [29] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C149(1.14) | LDD2090 | [37] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C149(0.89); C121(1.23) | LDD2092 | [37] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C149(1.18) | LDD2094 | [37] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C149(0.22) | LDD2096 | [37] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C149(1.68) | LDD2098 | [37] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C149(1.15) | LDD2099 | [37] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C149(1.85); C121(1.05) | LDD2105 | [37] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C149(1.04) | LDD2107 | [37] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C149(1.10) | LDD2109 | [37] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C149(1.22) | LDD2111 | [37] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C149(0.34) | LDD2116 | [37] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C149(0.31) | LDD2118 | [37] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C149(2.52) | LDD2119 | [37] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C149(0.22) | LDD2122 | [37] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C149(0.59) | LDD2123 | [37] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C149(0.19) | LDD2124 | [37] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C149(0.20) | LDD2126 | [37] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C149(0.52) | LDD2127 | [37] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C149(0.55) | LDD2129 | [37] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C149(1.28); C145(2.16) | LDD2136 | [37] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C149(0.59); C145(0.84); C126(1.08) | LDD2137 | [37] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C149(0.50) | LDD2141 | [37] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C149(3.81) | LDD2144 | [37] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C149(0.24) | LDD2149 | [37] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C149(1.35) | LDD2150 | [37] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C149(0.29) | LDD2151 | [37] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C149(2.05) | LDD2153 | [37] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C144(0.84) | LDD2207 | [41] |
| LDCM0131 | RA190 | MM1.R | C121(1.45); C126(1.45); C144(1.10); C145(1.10) | LDD0304 | [42] |
| LDCM0090 | Rapamycin | JHH-7 | 4.49 | LDD0213 | [34] |
| LDCM0021 | THZ1 | HCT 116 | C145(1.13); C149(1.13); C121(1.17); C126(1.17) | LDD2173 | [15] |
| LDCM0110 | W12 | Hep-G2 | K6(0.85) | LDD0237 | [30] |
| LDCM0111 | W14 | Hep-G2 | K6(1.33) | LDD0238 | [30] |
| LDCM0113 | W17 | Hep-G2 | K6(1.28) | LDD0240 | [30] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Desumoylating isopeptidase 1 (DESI1) | DeSI family | Q6ICB0 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
| Phospholipid scramblase 4 (PLSCR4) | Phospholipid scramblase family | Q9NRQ2 | |||
| Wolframin (WFS1) | . | O76024 | |||
Other
References
