Details of the Target
General Information of Target
| Target ID | LDTP09899 | |||||
|---|---|---|---|---|---|---|
| Target Name | Probable ATP-dependent RNA helicase DDX17 (DDX17) | |||||
| Gene Name | DDX17 | |||||
| Gene ID | 10521 | |||||
| Synonyms |
Probable ATP-dependent RNA helicase DDX17; EC 3.6.4.13; DEAD box protein 17; DEAD box protein p72; DEAD box protein p82; RNA-dependent helicase p72 |
|||||
| 3D Structure | ||||||
| Sequence |
MRQQDAPKPTPAACRCSGLARRVLTIAFALLILGLMTWAYAAGVPLASDRYGLLAFGLYG
AFLSAHLVAQSLFAYLEHRRVAAAARGPLDAATARSVALTISAYQEDPAYLRQCLASARA LLYPRARLRVLMVVDGNRAEDLYMVDMFREVFADEDPATYVWDGNYHQPWEPAAAGAVGA GAYREVEAEDPGRLAVEALVRTRRCVCVAQRWGGKREVMYTAFKALGDSVDYVQVCDSDT RLDPMALLELVRVLDEDPRVGAVGGDVRILNPLDSWVSFLSSLRYWVAFNVERACQSYFH CVSCISGPLGLYRNNLLQQFLEAWYNQKFLGTHCTFGDDRHLTNRMLSMGYATKYTSRSR CYSETPSSFLRWLSQQTRWSKSYFREWLYNALWWHRHHAWMTYEAVVSGLFPFFVAATVL RLFYAGRPWALLWVLLCVQGVALAKAAFAAWLRGCLRMVLLSLYAPLYMCGLLPAKFLAL VTMNQSGWGTSGRRKLAANYVPLLPLALWALLLLGGLVRSVAHEARADWSGPSRAAEAYH LAAGAGAYVGYWVAMLTLYWVGVRRLCRRRTGGYRVQV |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
DEAD box helicase family, DDX5/DBP2 subfamily
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
As an RNA helicase, unwinds RNA and alters RNA structures through ATP binding and hydrolysis. Involved in multiple cellular processes, including pre-mRNA splicing, alternative splicing, ribosomal RNA processing and miRNA processing, as well as transcription regulation. Regulates the alternative splicing of exons exhibiting specific features. For instance, promotes the inclusion of AC-rich alternative exons in CD44 transcripts. This function requires the RNA helicase activity. Affects NFAT5 and histone macro-H2A.1/MACROH2A1 alternative splicing in a CDK9-dependent manner. In NFAT5, promotes the introduction of alternative exon 4, which contains 2 stop codons and may target NFAT5 exon 4-containing transcripts to nonsense-mediated mRNA decay, leading to the down-regulation of NFAT5 protein. Affects splicing of mediators of steroid hormone signaling pathway, including kinases that phosphorylates ESR1, such as CDK2, MAPK1 and GSK3B, and transcriptional regulators, such as CREBBP, MED1, NCOR1 and NCOR2. By affecting GSK3B splicing, participates in ESR1 and AR stabilization. In myoblasts and epithelial cells, cooperates with HNRNPH1 to control the splicing of specific subsets of exons. In addition to binding mature mRNAs, also interacts with certain pri-microRNAs, including MIR663/miR-663a, MIR99B/miR-99b, and MIR6087/miR-6087. Binds pri-microRNAs on the 3' segment flanking the stem loop via the 5'-[ACG]CAUC[ACU]-3' consensus sequence. Required for the production of subsets of microRNAs, including MIR21 and MIR125B1. May be involved not only in microRNA primary transcript processing, but also stabilization. Participates in MYC down-regulation at high cell density through the production of MYC-targeting microRNAs. Along with DDX5, may be involved in the processing of the 32S intermediate into the mature 28S ribosomal RNA. Promoter-specific transcription regulator, functioning as a coactivator or corepressor depending on the context of the promoter and the transcriptional complex in which it exists. Enhances NFAT5 transcriptional activity. Synergizes with TP53 in the activation of the MDM2 promoter; this activity requires acetylation on lysine residues. May also coactivate MDM2 transcription through a TP53-independent pathway. Coactivates MMP7 transcription. Along with CTNNB1, coactivates MYC, JUN, FOSL1 and cyclin D1/CCND1 transcription. Alone or in combination with DDX5 and/or SRA1 non-coding RNA, plays a critical role in promoting the assembly of proteins required for the formation of the transcription initiation complex and chromatin remodeling leading to coactivation of MYOD1-dependent transcription. This helicase-independent activity is required for skeletal muscle cells to properly differentiate into myotubes. During epithelial-to-mesenchymal transition, coregulates SMAD-dependent transcriptional activity, directly controlling key effectors of differentiation, including miRNAs which in turn directly repress its expression. Plays a role in estrogen and testosterone signaling pathway at several levels. Mediates the use of alternative promoters in estrogen-responsive genes and regulates transcription and splicing of a large number of steroid hormone target genes. Contrary to splicing regulation activity, transcriptional coregulation of the estrogen receptor ESR1 is helicase-independent. Plays a role in innate immunity. Specifically restricts bunyavirus infection, including Rift Valley fever virus (RVFV) or La Crosse virus (LACV), but not vesicular stomatitis virus (VSV), in an interferon- and DROSHA-independent manner. Binds to RVFV RNA, likely via structured viral RNA elements. Promotes mRNA degradation mediated by the antiviral zinc-finger protein ZC3HAV1, in an ATPase-dependent manner.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
10.37 | LDD0402 | [1] | |
|
P2 Probe Info |
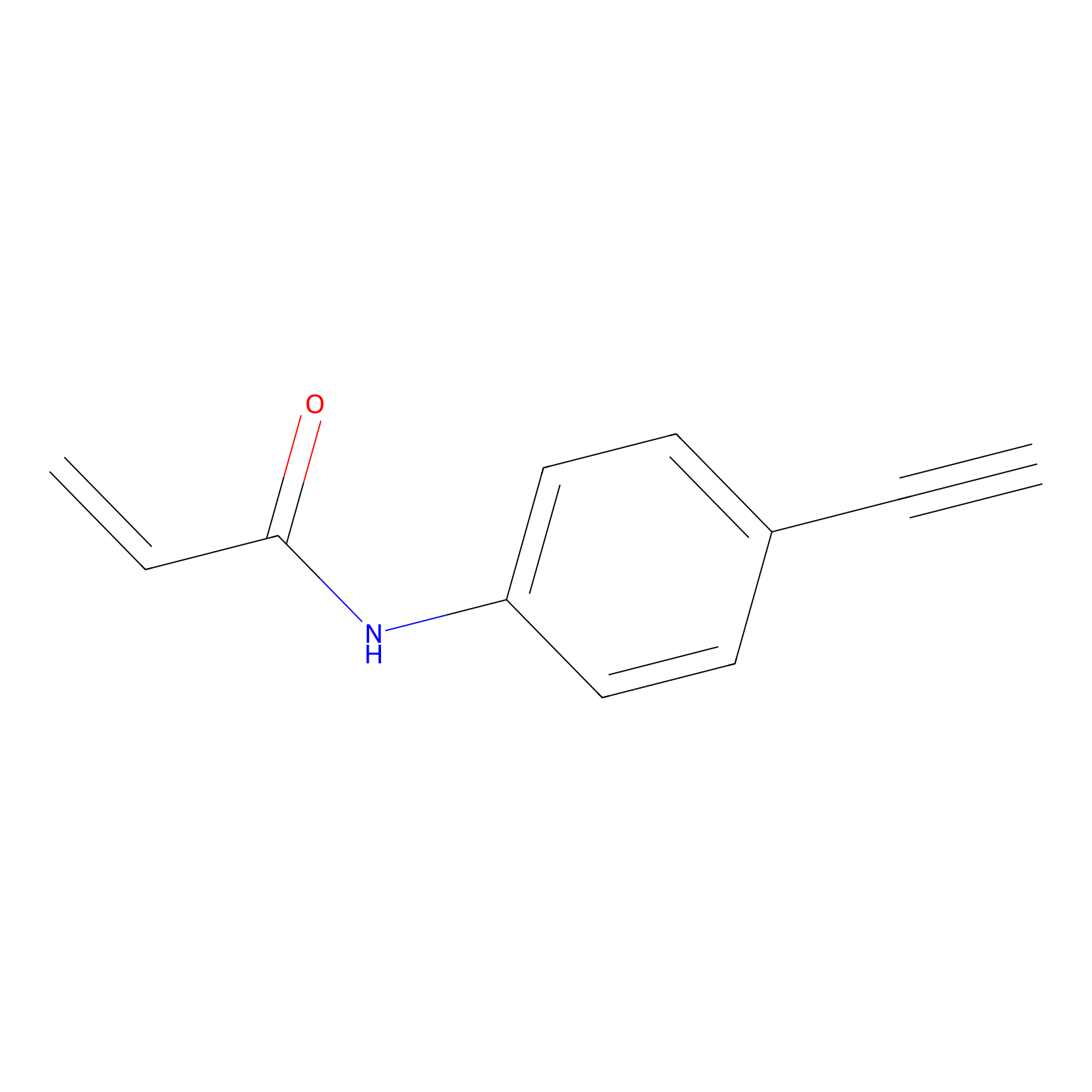 |
10.00 | LDD0449 | [2] | |
|
P8 Probe Info |
 |
10.00 | LDD0451 | [2] | |
|
A-EBA Probe Info |
 |
17.36 | LDD0215 | [3] | |
|
11RK73 Probe Info |
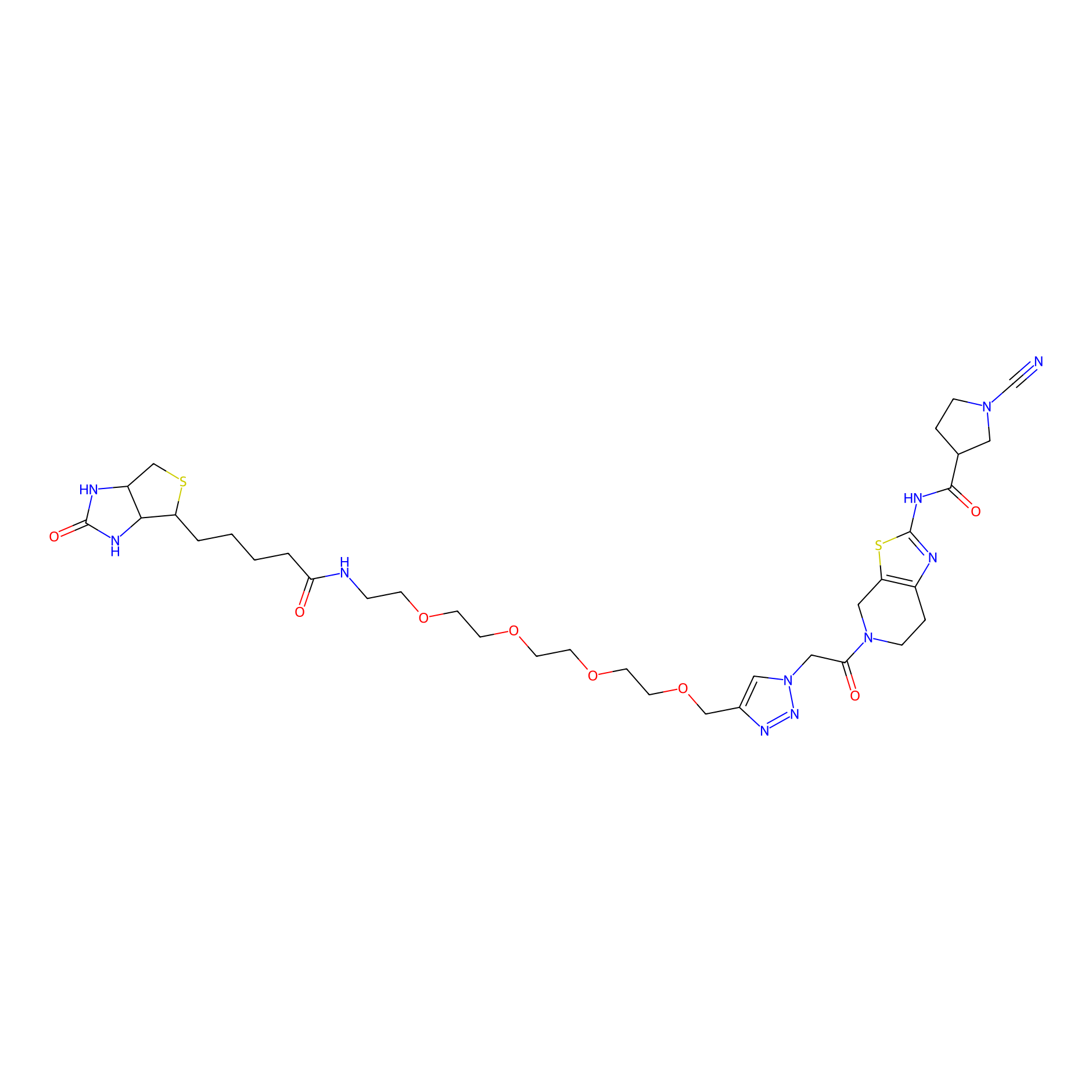 |
2.34 | LDD0328 | [4] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [5] | |
|
W1 Probe Info |
 |
14.23 | LDD0235 | [6] | |
|
TH211 Probe Info |
 |
Y580(20.00); Y519(13.27) | LDD0257 | [7] | |
|
TH216 Probe Info |
 |
Y279(10.26) | LDD0259 | [7] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [8] | |
|
BTD Probe Info |
 |
C584(1.45) | LDD1699 | [9] | |
|
ONAyne Probe Info |
 |
K592(0.00); K547(0.00); K109(0.00); K468(0.00) | LDD0273 | [10] | |
|
Probe 1 Probe Info |
 |
Y135(9.21); Y147(25.88); Y267(46.76); Y519(16.38) | LDD3495 | [11] | |
|
AZ-9 Probe Info |
 |
10.00 | LDD2154 | [12] | |
|
AHL-Pu-1 Probe Info |
 |
C298(2.11); C277(2.11) | LDD0169 | [13] | |
|
HHS-475 Probe Info |
 |
Y75(0.90) | LDD0264 | [14] | |
|
HHS-465 Probe Info |
 |
Y75(10.00) | LDD2237 | [15] | |
|
DBIA Probe Info |
 |
C277(3.18); C166(1.70); C584(1.93) | LDD0080 | [16] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [17] | |
|
ATP probe Probe Info |
 |
K528(0.00); K515(0.00); K452(0.00); K535(0.00) | LDD0199 | [18] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C431(0.00); C319(0.00); C447(0.00); C397(0.00) | LDD0038 | [19] | |
|
IA-alkyne Probe Info |
 |
C584(0.00); C298(0.00); C247(0.00); C319(0.00) | LDD0032 | [20] | |
|
IPIAA_H Probe Info |
 |
C319(0.00); C247(0.00) | LDD0030 | [21] | |
|
IPIAA_L Probe Info |
 |
C397(0.00); C247(0.00); C584(0.00); C298(0.00) | LDD0031 | [21] | |
|
Lodoacetamide azide Probe Info |
 |
C319(0.00); C447(0.00); C397(0.00); C247(0.00) | LDD0037 | [19] | |
|
JW-RF-010 Probe Info |
 |
C584(0.00); C270(0.00); C319(0.00); C277(0.00) | LDD0026 | [22] | |
|
NAIA_4 Probe Info |
 |
C270(0.00); C584(0.00) | LDD2226 | [23] | |
|
TFBX Probe Info |
 |
C277(0.00); C247(0.00); C584(0.00); C319(0.00) | LDD0027 | [22] | |
|
WYneN Probe Info |
 |
C277(0.00); C584(0.00); C247(0.00) | LDD0021 | [24] | |
|
WYneO Probe Info |
 |
C247(0.00); C584(0.00); C277(0.00) | LDD0022 | [24] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [25] | |
|
Compound 10 Probe Info |
 |
C247(0.00); C270(0.00); C277(0.00); C298(0.00) | LDD2216 | [26] | |
|
Compound 11 Probe Info |
 |
C247(0.00); C277(0.00); C298(0.00); C584(0.00) | LDD2213 | [26] | |
|
ENE Probe Info |
 |
C247(0.00); C584(0.00) | LDD0006 | [24] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [24] | |
|
NHS Probe Info |
 |
K515(0.00); K313(0.00); K547(0.00); K468(0.00) | LDD0010 | [24] | |
|
PPMS Probe Info |
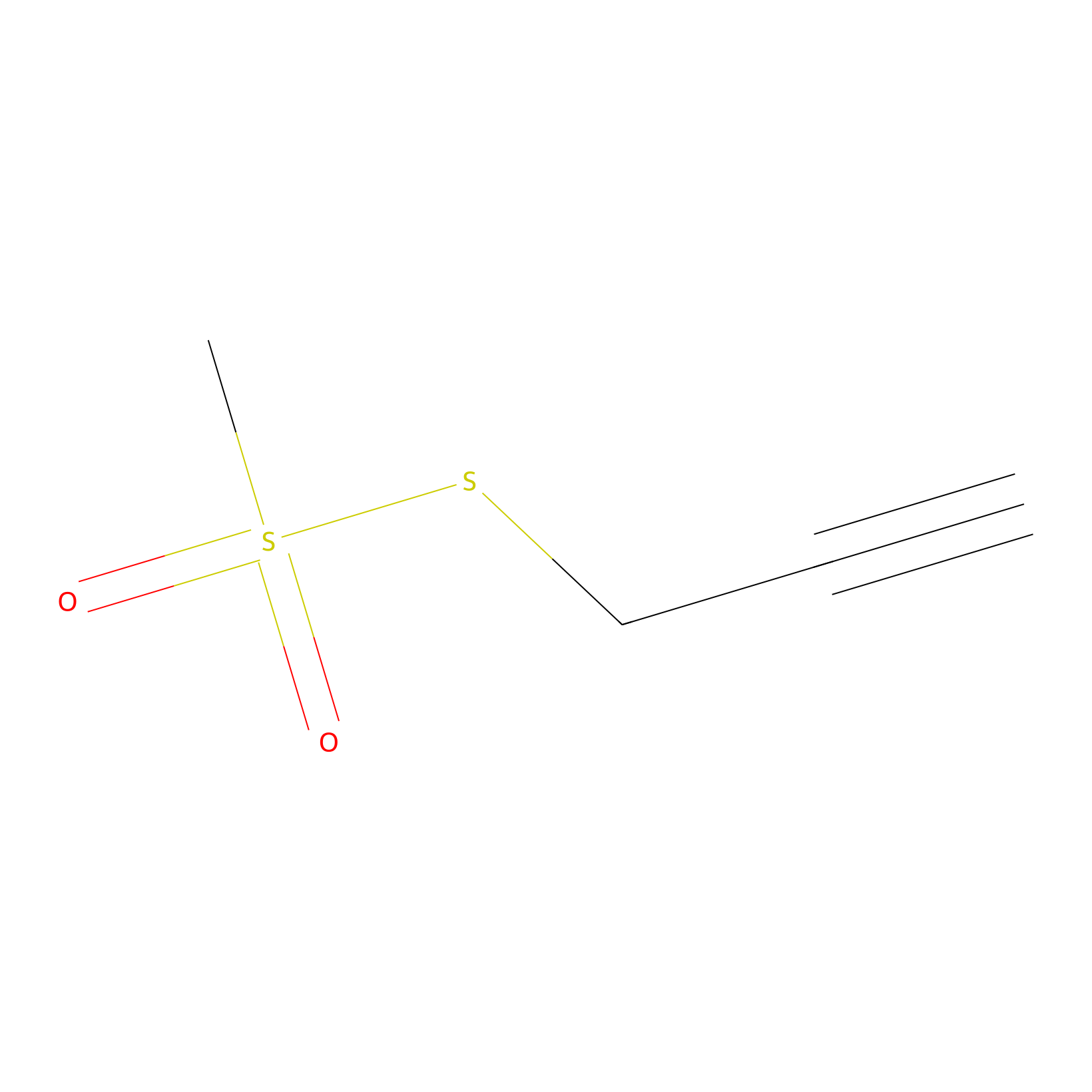 |
C298(0.00); C247(0.00) | LDD0008 | [24] | |
|
SF Probe Info |
 |
Y147(0.00); K109(0.00); Y279(0.00); Y238(0.00) | LDD0028 | [27] | |
|
STPyne Probe Info |
 |
K313(0.00); K284(0.00) | LDD0009 | [24] | |
|
VSF Probe Info |
 |
C298(0.00); C584(0.00) | LDD0007 | [24] | |
|
Phosphinate-6 Probe Info |
 |
C584(0.00); C277(0.00); C270(0.00) | LDD0018 | [28] | |
|
Ox-W18 Probe Info |
 |
W122(0.00); W443(0.00); W459(0.00) | LDD2175 | [29] | |
|
1c-yne Probe Info |
 |
K547(0.00); K535(0.00); K488(0.00); K221(0.00) | LDD0228 | [25] | |
|
Acrolein Probe Info |
 |
C584(0.00); C247(0.00); H504(0.00); C277(0.00) | LDD0217 | [30] | |
|
Cinnamaldehyde Probe Info |
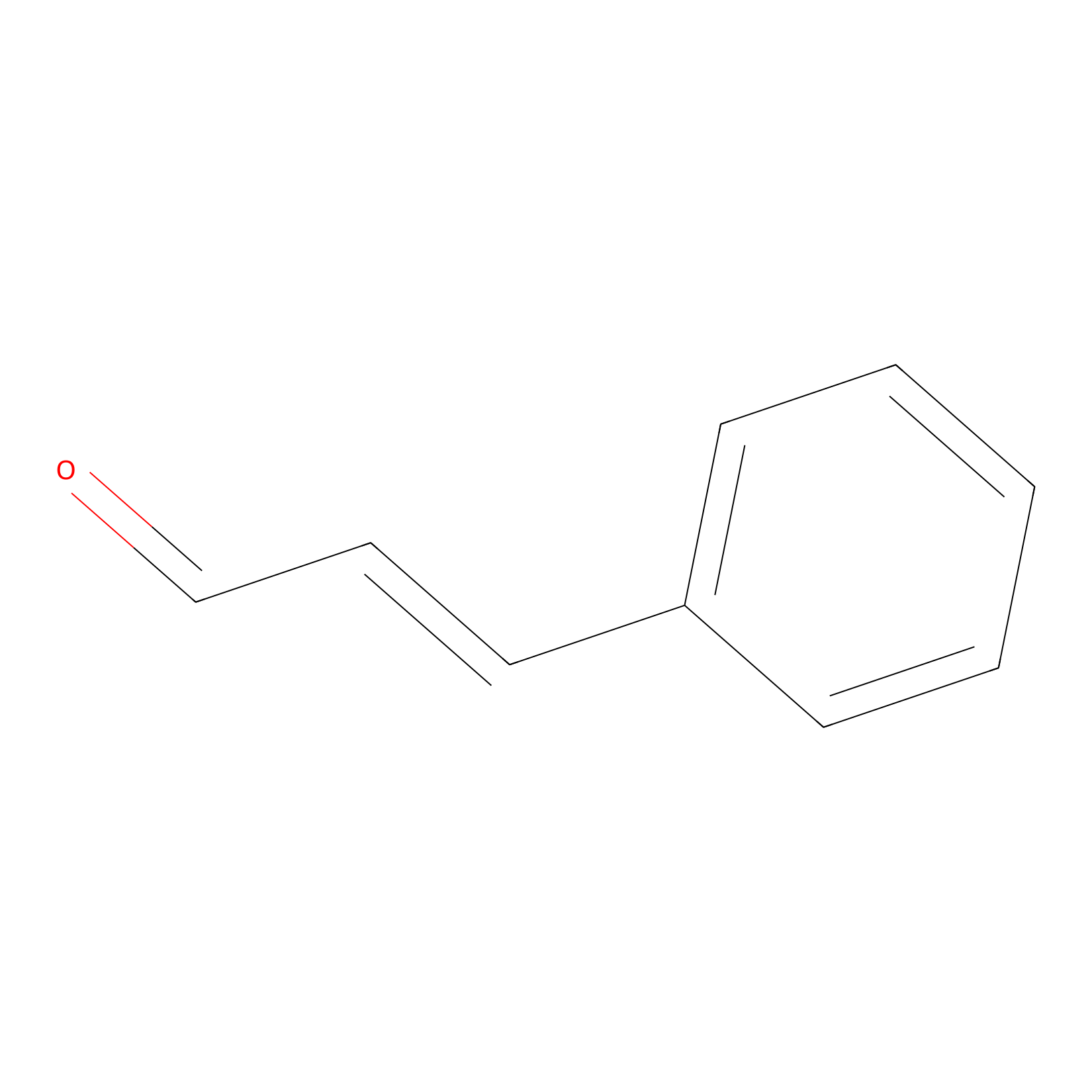 |
C298(0.00); C247(0.00); C584(0.00) | LDD0220 | [30] | |
|
Crotonaldehyde Probe Info |
 |
C298(0.00); C247(0.00); C584(0.00); C277(0.00) | LDD0219 | [30] | |
|
Methacrolein Probe Info |
 |
C584(0.00); C447(0.00); C298(0.00); C277(0.00) | LDD0218 | [30] | |
|
NAIA_5 Probe Info |
 |
C431(0.00); C319(0.00); C447(0.00); C270(0.00) | LDD2223 | [23] | |
|
HHS-482 Probe Info |
 |
Y279(0.41); Y580(0.91); Y75(1.19) | LDD2239 | [15] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C383 Probe Info |
 |
11.00 | LDD2042 | [31] | |
|
FFF probe11 Probe Info |
 |
5.53 | LDD0471 | [32] | |
|
FFF probe13 Probe Info |
 |
12.88 | LDD0475 | [32] | |
|
FFF probe14 Probe Info |
 |
9.50 | LDD0477 | [32] | |
|
FFF probe2 Probe Info |
 |
7.05 | LDD0463 | [32] | |
|
FFF probe3 Probe Info |
 |
7.09 | LDD0464 | [32] | |
|
JN0003 Probe Info |
 |
7.71 | LDD0469 | [32] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [33] | |
|
Diazir Probe Info |
 |
N.A. | LDD0011 | [24] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [34] | |
|
OEA-DA Probe Info |
 |
3.37 | LDD0046 | [35] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C166(0.82); C584(0.83); C431(0.82); C270(0.59) | LDD2142 | [9] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C166(0.92); C584(1.17); C431(1.05); C270(0.73) | LDD2112 | [9] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C166(0.75) | LDD2095 | [9] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C166(0.85) | LDD2130 | [9] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C166(0.92); C431(0.85); C270(0.87) | LDD2117 | [9] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C166(1.10); C584(1.22); C431(1.58) | LDD2152 | [9] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C166(0.79); C584(0.82) | LDD2103 | [9] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C431(0.69) | LDD2132 | [9] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C166(0.63) | LDD2131 | [9] |
| LDCM0025 | 4SU-RNA | HEK-293T | C247(2.31) | LDD0172 | [13] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C298(2.11); C277(2.11) | LDD0169 | [13] |
| LDCM0214 | AC1 | HCT 116 | C584(0.69); C166(1.38); C247(1.29); C277(0.95) | LDD0531 | [16] |
| LDCM0215 | AC10 | HCT 116 | C584(0.89); C166(1.56); C247(0.85); C277(0.90) | LDD0532 | [16] |
| LDCM0216 | AC100 | HCT 116 | C584(0.83); C166(1.68); C247(0.67); C277(0.79) | LDD0533 | [16] |
| LDCM0217 | AC101 | HCT 116 | C584(0.76); C166(0.97); C247(0.65); C277(0.62) | LDD0534 | [16] |
| LDCM0218 | AC102 | HCT 116 | C584(0.90); C166(1.19); C247(0.63); C277(0.65) | LDD0535 | [16] |
| LDCM0219 | AC103 | HCT 116 | C584(1.01); C166(1.22); C247(0.53); C277(0.57) | LDD0536 | [16] |
| LDCM0220 | AC104 | HCT 116 | C584(0.81); C166(1.31); C247(0.71); C277(0.62) | LDD0537 | [16] |
| LDCM0221 | AC105 | HCT 116 | C584(1.00); C166(1.38); C247(0.61); C277(0.66) | LDD0538 | [16] |
| LDCM0222 | AC106 | HCT 116 | C584(1.02); C166(1.56); C247(0.56); C277(0.52) | LDD0539 | [16] |
| LDCM0223 | AC107 | HCT 116 | C584(0.91); C166(1.75); C247(0.54); C277(0.57) | LDD0540 | [16] |
| LDCM0224 | AC108 | HCT 116 | C584(0.88); C166(1.29); C247(0.75); C277(0.70) | LDD0541 | [16] |
| LDCM0225 | AC109 | HCT 116 | C584(0.81); C166(1.33); C247(0.84); C277(0.62) | LDD0542 | [16] |
| LDCM0226 | AC11 | HCT 116 | C584(0.94); C166(2.45); C247(0.84); C277(0.83) | LDD0543 | [16] |
| LDCM0227 | AC110 | HCT 116 | C584(0.77); C166(1.57); C247(0.81); C277(0.67) | LDD0544 | [16] |
| LDCM0228 | AC111 | HCT 116 | C584(0.85); C166(0.93); C247(0.67); C277(0.58) | LDD0545 | [16] |
| LDCM0229 | AC112 | HCT 116 | C584(0.87); C166(0.94); C247(0.56); C277(0.55) | LDD0546 | [16] |
| LDCM0230 | AC113 | HCT 116 | C584(1.11); C166(1.08); C247(0.89); C277(1.08) | LDD0547 | [16] |
| LDCM0231 | AC114 | HCT 116 | C584(1.29); C166(0.95); C247(0.79); C277(0.96) | LDD0548 | [16] |
| LDCM0232 | AC115 | HCT 116 | C584(1.28); C166(0.93); C247(0.83); C277(0.83) | LDD0549 | [16] |
| LDCM0233 | AC116 | HCT 116 | C584(1.25); C166(1.16); C247(0.77); C277(0.79) | LDD0550 | [16] |
| LDCM0234 | AC117 | HCT 116 | C584(1.21); C166(1.24); C247(0.84); C277(0.92) | LDD0551 | [16] |
| LDCM0235 | AC118 | HCT 116 | C584(1.20); C166(1.19); C247(0.92); C277(0.99) | LDD0552 | [16] |
| LDCM0236 | AC119 | HCT 116 | C584(1.24); C166(1.04); C247(0.87); C277(1.16) | LDD0553 | [16] |
| LDCM0237 | AC12 | HCT 116 | C584(0.99); C166(3.02); C247(1.09); C277(1.10) | LDD0554 | [16] |
| LDCM0238 | AC120 | HCT 116 | C584(1.11); C166(1.15); C247(0.85); C277(1.05) | LDD0555 | [16] |
| LDCM0239 | AC121 | HCT 116 | C584(1.25); C166(1.18); C247(0.88); C277(1.15) | LDD0556 | [16] |
| LDCM0240 | AC122 | HCT 116 | C584(1.16); C166(1.20); C247(0.72); C277(0.94) | LDD0557 | [16] |
| LDCM0241 | AC123 | HCT 116 | C584(1.15); C166(1.29); C247(0.92); C277(1.38) | LDD0558 | [16] |
| LDCM0242 | AC124 | HCT 116 | C584(1.15); C166(1.15); C247(0.97); C277(0.96) | LDD0559 | [16] |
| LDCM0243 | AC125 | HCT 116 | C584(1.24); C166(0.98); C247(0.92); C277(0.92) | LDD0560 | [16] |
| LDCM0244 | AC126 | HCT 116 | C584(1.46); C166(1.04); C247(0.83); C277(1.08) | LDD0561 | [16] |
| LDCM0245 | AC127 | HCT 116 | C584(1.43); C166(1.26); C247(0.85); C277(0.97) | LDD0562 | [16] |
| LDCM0246 | AC128 | HCT 116 | C584(0.82); C166(1.62); C247(1.16); C277(0.81) | LDD0563 | [16] |
| LDCM0247 | AC129 | HCT 116 | C584(0.82); C166(1.31); C247(1.50); C277(1.23) | LDD0564 | [16] |
| LDCM0249 | AC130 | HCT 116 | C584(0.84); C166(1.15); C247(1.06); C277(0.93) | LDD0566 | [16] |
| LDCM0250 | AC131 | HCT 116 | C584(0.80); C166(0.99); C247(1.19); C277(0.99) | LDD0567 | [16] |
| LDCM0251 | AC132 | HCT 116 | C584(0.64); C166(0.89); C247(1.11); C277(1.23) | LDD0568 | [16] |
| LDCM0252 | AC133 | HCT 116 | C584(0.70); C166(1.07); C247(1.17); C277(0.95) | LDD0569 | [16] |
| LDCM0253 | AC134 | HCT 116 | C584(0.77); C166(0.75); C247(1.03); C277(0.89) | LDD0570 | [16] |
| LDCM0254 | AC135 | HCT 116 | C584(0.74); C166(0.85); C247(0.91); C277(0.90) | LDD0571 | [16] |
| LDCM0255 | AC136 | HCT 116 | C584(0.67); C166(0.78); C247(1.06); C277(1.05) | LDD0572 | [16] |
| LDCM0256 | AC137 | HCT 116 | C584(0.56); C166(0.87); C247(1.06); C277(1.08) | LDD0573 | [16] |
| LDCM0257 | AC138 | HCT 116 | C584(0.66); C166(0.84); C247(1.11); C277(1.01) | LDD0574 | [16] |
| LDCM0258 | AC139 | HCT 116 | C584(0.71); C166(0.74); C247(1.04); C277(0.90) | LDD0575 | [16] |
| LDCM0259 | AC14 | HCT 116 | C584(0.71); C166(1.99); C247(1.08); C277(0.88) | LDD0576 | [16] |
| LDCM0260 | AC140 | HCT 116 | C584(0.71); C166(0.69); C247(1.06); C277(0.92) | LDD0577 | [16] |
| LDCM0261 | AC141 | HCT 116 | C584(0.67); C166(0.68); C247(1.07); C277(0.97) | LDD0578 | [16] |
| LDCM0262 | AC142 | HCT 116 | C584(0.63); C166(0.77); C247(1.04); C277(0.91) | LDD0579 | [16] |
| LDCM0263 | AC143 | HCT 116 | C584(1.01); C166(0.58); C247(0.93); C277(0.93) | LDD0580 | [16] |
| LDCM0264 | AC144 | HCT 116 | C298(0.68); C247(0.71); C277(0.78); C319(0.89) | LDD0581 | [16] |
| LDCM0265 | AC145 | HCT 116 | C298(0.78); C166(0.95); C247(0.95); C447(0.95) | LDD0582 | [16] |
| LDCM0266 | AC146 | HCT 116 | C298(0.60); C166(0.67); C277(0.75); C247(0.83) | LDD0583 | [16] |
| LDCM0267 | AC147 | HCT 116 | C298(0.71); C277(0.76); C247(0.76); C319(0.88) | LDD0584 | [16] |
| LDCM0268 | AC148 | HCT 116 | C298(0.54); C166(0.59); C319(0.63); C277(0.65) | LDD0585 | [16] |
| LDCM0269 | AC149 | HCT 116 | C298(0.57); C166(0.59); C277(0.68); C319(0.71) | LDD0586 | [16] |
| LDCM0270 | AC15 | HCT 116 | C584(0.84); C277(0.86); C447(1.00); C298(1.04) | LDD0587 | [16] |
| LDCM0271 | AC150 | HCT 116 | C166(0.74); C298(0.76); C319(0.81); C247(0.90) | LDD0588 | [16] |
| LDCM0272 | AC151 | HCT 116 | C298(0.77); C166(0.83); C277(0.84); C447(0.89) | LDD0589 | [16] |
| LDCM0273 | AC152 | HCT 116 | C298(0.56); C319(0.62); C166(0.67); C277(0.68) | LDD0590 | [16] |
| LDCM0274 | AC153 | HCT 116 | C166(0.34); C298(0.54); C277(0.61); C319(0.62) | LDD0591 | [16] |
| LDCM0621 | AC154 | HCT 116 | C584(1.39); C166(0.75); C247(0.85); C277(0.78) | LDD2158 | [16] |
| LDCM0622 | AC155 | HCT 116 | C584(1.28); C166(0.44); C247(0.92); C277(0.82) | LDD2159 | [16] |
| LDCM0623 | AC156 | HCT 116 | C584(1.11); C166(0.66); C247(1.01); C277(0.95) | LDD2160 | [16] |
| LDCM0624 | AC157 | HCT 116 | C584(1.26); C166(0.46); C247(1.08); C277(0.91) | LDD2161 | [16] |
| LDCM0276 | AC17 | HCT 116 | C166(0.89); C447(0.93); C298(0.95); C247(1.00) | LDD0593 | [16] |
| LDCM0277 | AC18 | HCT 116 | C584(0.73); C247(0.89); C447(0.98); C277(1.06) | LDD0594 | [16] |
| LDCM0278 | AC19 | HCT 116 | C447(0.85); C277(0.92); C247(0.92); C584(0.99) | LDD0595 | [16] |
| LDCM0279 | AC2 | HCT 116 | C584(0.74); C166(0.92); C298(0.92); C277(0.93) | LDD0596 | [16] |
| LDCM0280 | AC20 | HCT 116 | C166(0.89); C298(0.92); C447(0.93); C247(0.97) | LDD0597 | [16] |
| LDCM0281 | AC21 | HCT 116 | C447(0.87); C247(0.93); C277(1.16); C298(1.30) | LDD0598 | [16] |
| LDCM0282 | AC22 | HCT 116 | C277(0.91); C447(0.94); C247(0.96); C166(0.98) | LDD0599 | [16] |
| LDCM0283 | AC23 | HCT 116 | C447(0.83); C247(0.92); C298(0.99); C277(1.07) | LDD0600 | [16] |
| LDCM0284 | AC24 | HCT 116 | C298(0.86); C447(0.88); C277(0.92); C166(0.98) | LDD0601 | [16] |
| LDCM0285 | AC25 | HCT 116 | C319(0.73); C166(0.91); C298(0.92); C247(0.98) | LDD0602 | [16] |
| LDCM0286 | AC26 | HCT 116 | C319(0.67); C298(0.68); C166(0.72); C247(0.81) | LDD0603 | [16] |
| LDCM0287 | AC27 | HCT 116 | C319(0.64); C166(0.79); C298(0.83); C247(0.87) | LDD0604 | [16] |
| LDCM0288 | AC28 | HCT 116 | C584(0.59); C166(0.81); C319(0.82); C247(0.85) | LDD0605 | [16] |
| LDCM0289 | AC29 | HCT 116 | C166(0.59); C298(0.66); C247(0.75); C319(0.79) | LDD0606 | [16] |
| LDCM0290 | AC3 | HCT 116 | C584(0.70); C447(1.00); C298(1.04); C277(1.06) | LDD0607 | [16] |
| LDCM0291 | AC30 | HCT 116 | C319(0.55); C166(0.56); C298(0.63); C247(0.79) | LDD0608 | [16] |
| LDCM0292 | AC31 | HCT 116 | C319(0.63); C166(0.65); C298(0.68); C247(0.75) | LDD0609 | [16] |
| LDCM0293 | AC32 | HCT 116 | C166(0.49); C319(0.53); C298(0.55); C247(0.74) | LDD0610 | [16] |
| LDCM0294 | AC33 | HCT 116 | C166(0.62); C298(0.65); C319(0.65); C247(0.68) | LDD0611 | [16] |
| LDCM0295 | AC34 | HCT 116 | C584(0.47); C319(0.57); C166(0.67); C298(0.71) | LDD0612 | [16] |
| LDCM0296 | AC35 | HCT 116 | C447(0.95); C247(1.12); C319(1.20); C584(1.23) | LDD0613 | [16] |
| LDCM0297 | AC36 | HCT 116 | C166(0.93); C447(0.96); C298(0.99); C247(1.01) | LDD0614 | [16] |
| LDCM0298 | AC37 | HCT 116 | C447(0.98); C247(1.08); C584(1.26); C319(1.29) | LDD0615 | [16] |
| LDCM0299 | AC38 | HCT 116 | C166(0.77); C447(0.98); C247(1.00); C298(1.04) | LDD0616 | [16] |
| LDCM0300 | AC39 | HCT 116 | C166(0.79); C447(0.88); C247(0.98); C319(1.02) | LDD0617 | [16] |
| LDCM0301 | AC4 | HCT 116 | C584(0.71); C277(1.00); C447(1.04); C298(1.11) | LDD0618 | [16] |
| LDCM0302 | AC40 | HCT 116 | C166(0.87); C447(0.90); C247(0.98); C584(0.98) | LDD0619 | [16] |
| LDCM0303 | AC41 | HCT 116 | C247(0.86); C298(0.88); C447(0.91); C166(1.06) | LDD0620 | [16] |
| LDCM0304 | AC42 | HCT 116 | C247(0.74); C447(0.84); C298(0.85); C166(0.91) | LDD0621 | [16] |
| LDCM0305 | AC43 | HCT 116 | C166(0.77); C298(0.82); C447(0.83); C247(0.87) | LDD0622 | [16] |
| LDCM0306 | AC44 | HCT 116 | C247(0.83); C584(0.91); C447(0.97); C319(1.09) | LDD0623 | [16] |
| LDCM0307 | AC45 | HCT 116 | C247(0.71); C447(0.91); C277(0.97); C584(1.04) | LDD0624 | [16] |
| LDCM0308 | AC46 | HCT 116 | C319(0.81); C298(0.83); C277(0.87); C247(0.90) | LDD0625 | [16] |
| LDCM0309 | AC47 | HCT 116 | C319(0.82); C584(0.90); C247(0.93); C298(0.99) | LDD0626 | [16] |
| LDCM0310 | AC48 | HCT 116 | C584(0.71); C298(0.88); C277(0.93); C247(0.95) | LDD0627 | [16] |
| LDCM0311 | AC49 | HCT 116 | C584(0.64); C298(0.67); C247(0.69); C277(0.76) | LDD0628 | [16] |
| LDCM0312 | AC5 | HCT 116 | C584(0.55); C298(1.04); C277(1.11); C447(1.12) | LDD0629 | [16] |
| LDCM0313 | AC50 | HCT 116 | C298(0.75); C319(0.78); C277(0.81); C584(0.83) | LDD0630 | [16] |
| LDCM0314 | AC51 | HCT 116 | C447(0.90); C277(1.02); C247(1.10); C584(1.19) | LDD0631 | [16] |
| LDCM0315 | AC52 | HCT 116 | C584(0.93); C447(0.96); C247(1.08); C298(1.09) | LDD0632 | [16] |
| LDCM0316 | AC53 | HCT 116 | C277(0.75); C298(0.76); C247(0.76); C447(0.81) | LDD0633 | [16] |
| LDCM0317 | AC54 | HCT 116 | C584(0.80); C247(0.81); C277(0.83); C447(1.01) | LDD0634 | [16] |
| LDCM0318 | AC55 | HCT 116 | C298(0.66); C247(0.71); C447(0.76); C584(0.76) | LDD0635 | [16] |
| LDCM0319 | AC56 | HCT 116 | C298(0.71); C247(0.78); C277(0.81); C584(0.82) | LDD0636 | [16] |
| LDCM0320 | AC57 | HCT 116 | C584(0.76); C277(0.80); C247(0.85); C447(0.95) | LDD0637 | [16] |
| LDCM0321 | AC58 | HCT 116 | C319(0.69); C584(0.70); C277(0.81); C247(0.84) | LDD0638 | [16] |
| LDCM0322 | AC59 | HCT 116 | C584(0.53); C166(0.68); C277(0.75); C247(0.75) | LDD0639 | [16] |
| LDCM0323 | AC6 | HCT 116 | C277(0.80); C319(0.81); C584(0.86); C298(0.86) | LDD0640 | [16] |
| LDCM0324 | AC60 | HCT 116 | C166(0.55); C247(0.66); C584(0.68); C277(0.76) | LDD0641 | [16] |
| LDCM0325 | AC61 | HCT 116 | C277(0.64); C584(0.66); C247(0.74); C319(0.84) | LDD0642 | [16] |
| LDCM0326 | AC62 | HCT 116 | C584(0.54); C277(0.59); C247(0.73); C166(0.75) | LDD0643 | [16] |
| LDCM0327 | AC63 | HCT 116 | C584(0.67); C247(0.70); C166(0.70); C277(0.86) | LDD0644 | [16] |
| LDCM0328 | AC64 | HCT 116 | C584(0.57); C247(0.67); C166(0.70); C277(0.77) | LDD0645 | [16] |
| LDCM0329 | AC65 | HCT 116 | C584(0.60); C277(0.68); C247(0.76); C319(0.76) | LDD0646 | [16] |
| LDCM0330 | AC66 | HCT 116 | C584(0.53); C277(0.69); C247(0.75); C298(0.78) | LDD0647 | [16] |
| LDCM0331 | AC67 | HCT 116 | C584(0.43); C166(0.59); C277(0.59); C247(0.63) | LDD0648 | [16] |
| LDCM0332 | AC68 | HCT 116 | C247(0.88); C319(0.89); C277(0.89); C298(0.90) | LDD0649 | [16] |
| LDCM0333 | AC69 | HCT 116 | C298(0.93); C319(0.96); C447(0.98); C277(1.03) | LDD0650 | [16] |
| LDCM0334 | AC7 | HCT 116 | C584(0.76); C298(0.93); C277(0.95); C247(0.99) | LDD0651 | [16] |
| LDCM0335 | AC70 | HCT 116 | C298(0.66); C247(0.74); C277(0.87); C447(0.88) | LDD0652 | [16] |
| LDCM0336 | AC71 | HCT 116 | C247(0.96); C447(0.97); C298(1.09); C277(1.10) | LDD0653 | [16] |
| LDCM0337 | AC72 | HCT 116 | C298(0.89); C247(0.90); C447(0.94); C319(1.00) | LDD0654 | [16] |
| LDCM0338 | AC73 | HCT 116 | C298(0.71); C247(0.73); C277(0.79); C447(0.85) | LDD0655 | [16] |
| LDCM0339 | AC74 | HCT 116 | C247(0.85); C447(0.86); C298(0.89); C277(0.91) | LDD0656 | [16] |
| LDCM0340 | AC75 | HCT 116 | C247(0.78); C298(0.82); C277(0.88); C447(0.91) | LDD0657 | [16] |
| LDCM0341 | AC76 | HCT 116 | C447(0.87); C247(0.93); C298(0.95); C277(0.98) | LDD0658 | [16] |
| LDCM0342 | AC77 | HCT 116 | C277(0.66); C298(0.75); C247(0.81); C447(0.88) | LDD0659 | [16] |
| LDCM0343 | AC78 | HCT 116 | C298(0.86); C447(0.91); C247(0.95); C277(0.96) | LDD0660 | [16] |
| LDCM0344 | AC79 | HCT 116 | C277(0.90); C298(0.94); C447(1.00); C319(1.20) | LDD0661 | [16] |
| LDCM0345 | AC8 | HCT 116 | C584(0.74); C298(0.79); C319(0.89); C277(0.96) | LDD0662 | [16] |
| LDCM0346 | AC80 | HCT 116 | C277(0.83); C247(0.89); C447(0.92); C298(0.96) | LDD0663 | [16] |
| LDCM0347 | AC81 | HCT 116 | C247(0.80); C277(0.89); C447(0.94); C298(1.03) | LDD0664 | [16] |
| LDCM0348 | AC82 | HCT 116 | C277(0.86); C298(0.90); C247(0.90); C319(0.98) | LDD0665 | [16] |
| LDCM0349 | AC83 | HCT 116 | C277(0.59); C319(0.60); C247(0.66); C447(0.90) | LDD0666 | [16] |
| LDCM0350 | AC84 | HCT 116 | C319(0.60); C277(0.65); C247(0.72); C298(0.78) | LDD0667 | [16] |
| LDCM0351 | AC85 | HCT 116 | C319(0.70); C277(0.72); C247(0.73); C298(0.79) | LDD0668 | [16] |
| LDCM0352 | AC86 | HCT 116 | C319(0.80); C277(0.82); C247(0.84); C298(0.90) | LDD0669 | [16] |
| LDCM0353 | AC87 | HCT 116 | C277(0.82); C298(0.85); C247(0.88); C319(0.89) | LDD0670 | [16] |
| LDCM0354 | AC88 | HCT 116 | C319(0.77); C277(0.81); C447(0.94); C298(0.94) | LDD0671 | [16] |
| LDCM0355 | AC89 | HCT 116 | C277(0.63); C319(0.66); C247(0.80); C447(0.86) | LDD0672 | [16] |
| LDCM0357 | AC90 | HCT 116 | C447(0.87); C584(0.96); C298(1.00); C247(1.08) | LDD0674 | [16] |
| LDCM0358 | AC91 | HCT 116 | C277(0.64); C319(0.68); C247(0.73); C447(1.03) | LDD0675 | [16] |
| LDCM0359 | AC92 | HCT 116 | C277(0.58); C319(0.64); C247(0.71); C298(0.85) | LDD0676 | [16] |
| LDCM0360 | AC93 | HCT 116 | C319(0.70); C277(0.83); C247(0.90); C447(0.93) | LDD0677 | [16] |
| LDCM0361 | AC94 | HCT 116 | C319(0.79); C277(0.79); C247(0.91); C447(0.91) | LDD0678 | [16] |
| LDCM0362 | AC95 | HCT 116 | C298(0.66); C277(0.67); C447(0.77); C247(0.77) | LDD0679 | [16] |
| LDCM0363 | AC96 | HCT 116 | C277(0.75); C319(0.78); C247(0.80); C298(0.89) | LDD0680 | [16] |
| LDCM0364 | AC97 | HCT 116 | C277(0.62); C319(0.62); C298(0.68); C447(0.72) | LDD0681 | [16] |
| LDCM0365 | AC98 | HCT 116 | C247(0.44); C277(0.45); C447(0.75); C166(0.80) | LDD0682 | [16] |
| LDCM0366 | AC99 | HCT 116 | C247(0.63); C277(0.72); C584(0.76); C447(0.84) | LDD0683 | [16] |
| LDCM0545 | Acetamide | MDA-MB-231 | C431(0.47) | LDD2138 | [9] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C166(0.67); C431(0.87) | LDD2113 | [9] |
| LDCM0248 | AKOS034007472 | HCT 116 | C584(0.92); C166(1.66); C247(1.02); C277(0.90) | LDD0565 | [16] |
| LDCM0356 | AKOS034007680 | HCT 116 | C298(0.86); C447(0.90); C277(0.93); C247(0.95) | LDD0673 | [16] |
| LDCM0275 | AKOS034007705 | HCT 116 | C277(0.72); C584(0.73); C319(0.82); C247(0.90) | LDD0592 | [16] |
| LDCM0156 | Aniline | NCI-H1299 | N.A. | LDD0404 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C319(1.07); C584(1.15); C247(1.33); C447(0.92) | LDD2171 | [16] |
| LDCM0151 | AZ-11 | HeLa | 10.00 | LDD2154 | [12] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C166(0.65); C584(0.49); C431(0.86); C270(0.46) | LDD2091 | [9] |
| LDCM0630 | CCW28-3 | 231MFP | C584(1.40) | LDD2214 | [36] |
| LDCM0108 | Chloroacetamide | HeLa | C584(0.00); C247(0.00); C298(0.00); C277(0.00) | LDD0222 | [30] |
| LDCM0632 | CL-Sc | Hep-G2 | C584(20.00); C277(2.01); C247(1.33); C584(1.07) | LDD2227 | [23] |
| LDCM0367 | CL1 | HCT 116 | C166(0.77); C319(0.85); C447(0.89); C298(1.05) | LDD0684 | [16] |
| LDCM0368 | CL10 | HCT 116 | C277(0.66); C166(0.72); C298(0.83); C447(0.86) | LDD0685 | [16] |
| LDCM0369 | CL100 | HCT 116 | C584(0.68); C166(0.85); C298(0.90); C447(0.95) | LDD0686 | [16] |
| LDCM0370 | CL101 | HCT 116 | C298(0.76); C277(0.80); C584(0.82); C447(0.92) | LDD0687 | [16] |
| LDCM0371 | CL102 | HCT 116 | C447(0.86); C298(0.89); C584(0.95); C277(0.95) | LDD0688 | [16] |
| LDCM0372 | CL103 | HCT 116 | C447(0.94); C277(0.99); C298(1.06); C584(1.12) | LDD0689 | [16] |
| LDCM0373 | CL104 | HCT 116 | C447(0.90); C277(0.94); C584(1.01); C247(1.07) | LDD0690 | [16] |
| LDCM0374 | CL105 | HCT 116 | C584(0.84); C247(0.85); C277(0.90); C447(0.95) | LDD0691 | [16] |
| LDCM0375 | CL106 | HCT 116 | C247(0.77); C584(0.80); C277(0.91); C447(1.03) | LDD0692 | [16] |
| LDCM0376 | CL107 | HCT 116 | C584(0.81); C247(0.86); C298(0.87); C447(0.93) | LDD0693 | [16] |
| LDCM0377 | CL108 | HCT 116 | C584(0.62); C298(0.77); C277(0.78); C247(0.86) | LDD0694 | [16] |
| LDCM0378 | CL109 | HCT 116 | C277(0.82); C247(0.84); C447(0.87); C298(0.89) | LDD0695 | [16] |
| LDCM0379 | CL11 | HCT 116 | C277(0.73); C298(0.82); C584(0.83); C447(0.86) | LDD0696 | [16] |
| LDCM0380 | CL110 | HCT 116 | C277(0.69); C247(0.74); C584(0.79); C447(0.82) | LDD0697 | [16] |
| LDCM0381 | CL111 | HCT 116 | C447(0.80); C247(0.82); C277(0.83); C584(0.89) | LDD0698 | [16] |
| LDCM0382 | CL112 | HCT 116 | C319(0.78); C166(0.83); C447(0.88); C247(0.89) | LDD0699 | [16] |
| LDCM0383 | CL113 | HCT 116 | C166(0.50); C298(0.56); C319(0.59); C247(0.71) | LDD0700 | [16] |
| LDCM0384 | CL114 | HCT 116 | C166(0.52); C298(0.57); C319(0.66); C247(0.79) | LDD0701 | [16] |
| LDCM0385 | CL115 | HCT 116 | C166(0.49); C298(0.61); C319(0.61); C247(0.75) | LDD0702 | [16] |
| LDCM0386 | CL116 | HCT 116 | C319(0.62); C298(0.62); C166(0.68); C247(0.78) | LDD0703 | [16] |
| LDCM0387 | CL117 | HCT 116 | C166(0.79); C447(0.84); C247(0.89); C319(0.89) | LDD0704 | [16] |
| LDCM0388 | CL118 | HCT 116 | C166(0.85); C447(0.92); C247(0.93); C298(0.94) | LDD0705 | [16] |
| LDCM0389 | CL119 | HCT 116 | C447(0.82); C247(0.86); C166(0.90); C298(0.95) | LDD0706 | [16] |
| LDCM0390 | CL12 | HCT 116 | C277(0.69); C298(0.79); C584(0.79); C247(0.88) | LDD0707 | [16] |
| LDCM0391 | CL120 | HCT 116 | C166(0.82); C298(0.85); C247(0.86); C447(0.87) | LDD0708 | [16] |
| LDCM0392 | CL121 | HCT 116 | C277(0.65); C319(0.68); C584(0.85); C298(0.93) | LDD0709 | [16] |
| LDCM0393 | CL122 | HCT 116 | C584(0.59); C298(0.84); C247(0.88); C319(0.90) | LDD0710 | [16] |
| LDCM0394 | CL123 | HCT 116 | C277(0.72); C298(0.74); C247(0.79); C584(0.81) | LDD0711 | [16] |
| LDCM0395 | CL124 | HCT 116 | C247(0.71); C584(0.72); C298(0.73); C277(0.80) | LDD0712 | [16] |
| LDCM0396 | CL125 | HCT 116 | C584(0.70); C166(0.73); C298(0.86); C447(0.97) | LDD0713 | [16] |
| LDCM0397 | CL126 | HCT 116 | C166(0.77); C277(0.80); C298(0.81); C584(0.82) | LDD0714 | [16] |
| LDCM0398 | CL127 | HCT 116 | C584(0.75); C166(0.78); C319(0.82); C277(0.85) | LDD0715 | [16] |
| LDCM0399 | CL128 | HCT 116 | C584(0.71); C247(0.78); C277(0.82); C298(0.91) | LDD0716 | [16] |
| LDCM0400 | CL13 | HCT 116 | C584(0.72); C277(0.73); C298(0.80); C447(0.89) | LDD0717 | [16] |
| LDCM0401 | CL14 | HCT 116 | C166(0.68); C277(0.80); C447(0.88); C298(0.92) | LDD0718 | [16] |
| LDCM0402 | CL15 | HCT 116 | C277(0.76); C247(0.85); C298(0.86); C584(0.92) | LDD0719 | [16] |
| LDCM0403 | CL16 | HCT 116 | C166(0.71); C277(0.73); C298(0.78); C584(0.84) | LDD0720 | [16] |
| LDCM0404 | CL17 | HCT 116 | C277(0.77); C166(0.78); C247(0.85); C584(0.87) | LDD0721 | [16] |
| LDCM0405 | CL18 | HCT 116 | C298(0.66); C277(0.66); C247(0.74); C166(0.86) | LDD0722 | [16] |
| LDCM0406 | CL19 | HCT 116 | C277(0.64); C584(0.82); C298(0.82); C247(0.84) | LDD0723 | [16] |
| LDCM0407 | CL2 | HCT 116 | C166(0.91); C319(0.92); C447(1.00); C298(1.03) | LDD0724 | [16] |
| LDCM0408 | CL20 | HCT 116 | C166(0.54); C277(0.65); C298(0.71); C247(0.84) | LDD0725 | [16] |
| LDCM0409 | CL21 | HCT 116 | C277(0.62); C298(0.67); C247(0.71); C584(0.83) | LDD0726 | [16] |
| LDCM0410 | CL22 | HCT 116 | C277(0.65); C298(0.68); C247(0.68); C584(0.80) | LDD0727 | [16] |
| LDCM0411 | CL23 | HCT 116 | C298(0.70); C277(0.74); C247(0.83); C447(0.91) | LDD0728 | [16] |
| LDCM0412 | CL24 | HCT 116 | C277(0.62); C298(0.68); C247(0.74); C584(0.82) | LDD0729 | [16] |
| LDCM0413 | CL25 | HCT 116 | C584(0.86); C166(1.05); C247(0.65); C277(0.60) | LDD0730 | [16] |
| LDCM0414 | CL26 | HCT 116 | C584(0.94); C166(0.89); C247(0.73); C277(0.75) | LDD0731 | [16] |
| LDCM0415 | CL27 | HCT 116 | C584(1.05); C166(1.01); C247(0.92); C277(0.70) | LDD0732 | [16] |
| LDCM0416 | CL28 | HCT 116 | C584(0.91); C166(0.93); C247(0.77); C277(0.54) | LDD0733 | [16] |
| LDCM0417 | CL29 | HCT 116 | C584(0.84); C166(0.98); C247(0.76); C277(0.56) | LDD0734 | [16] |
| LDCM0418 | CL3 | HCT 116 | C584(0.99); C166(0.80); C247(1.27); C277(1.16) | LDD0735 | [16] |
| LDCM0419 | CL30 | HCT 116 | C584(0.89); C166(1.72); C247(0.88); C277(0.63) | LDD0736 | [16] |
| LDCM0420 | CL31 | HCT 116 | C584(0.72); C166(0.96); C247(0.86); C277(0.88) | LDD0737 | [16] |
| LDCM0421 | CL32 | HCT 116 | C584(0.66); C166(0.74); C247(0.77); C277(0.62) | LDD0738 | [16] |
| LDCM0422 | CL33 | HCT 116 | C584(0.82); C166(0.78); C247(0.76); C277(0.62) | LDD0739 | [16] |
| LDCM0423 | CL34 | HCT 116 | C584(0.66); C166(0.67); C247(0.74); C277(0.65) | LDD0740 | [16] |
| LDCM0424 | CL35 | HCT 116 | C584(0.64); C166(0.72); C247(0.85); C277(0.62) | LDD0741 | [16] |
| LDCM0425 | CL36 | HCT 116 | C584(0.64); C166(0.76); C247(0.88); C277(0.73) | LDD0742 | [16] |
| LDCM0426 | CL37 | HCT 116 | C584(0.57); C166(0.98); C247(0.79); C277(1.00) | LDD0743 | [16] |
| LDCM0428 | CL39 | HCT 116 | C584(0.74); C166(0.99); C247(0.77); C277(0.59) | LDD0745 | [16] |
| LDCM0429 | CL4 | HCT 116 | C584(1.05); C166(1.02); C247(1.05); C277(0.82) | LDD0746 | [16] |
| LDCM0430 | CL40 | HCT 116 | C584(0.71); C166(0.90); C247(0.81); C277(0.77) | LDD0747 | [16] |
| LDCM0431 | CL41 | HCT 116 | C584(0.70); C166(0.82); C247(0.92); C277(0.75) | LDD0748 | [16] |
| LDCM0432 | CL42 | HCT 116 | C584(0.63); C166(1.15); C247(0.82); C277(0.85) | LDD0749 | [16] |
| LDCM0433 | CL43 | HCT 116 | C584(0.60); C166(0.92); C247(0.77); C277(0.58) | LDD0750 | [16] |
| LDCM0434 | CL44 | HCT 116 | C584(0.73); C166(0.98); C247(0.84); C277(0.67) | LDD0751 | [16] |
| LDCM0435 | CL45 | HCT 116 | C584(0.65); C166(0.90); C247(0.76); C277(0.63) | LDD0752 | [16] |
| LDCM0436 | CL46 | HCT 116 | C584(1.27); C166(1.02); C247(1.10); C277(1.49) | LDD0753 | [16] |
| LDCM0437 | CL47 | HCT 116 | C584(1.24); C166(1.09); C247(1.14); C277(1.52) | LDD0754 | [16] |
| LDCM0438 | CL48 | HCT 116 | C584(1.08); C166(0.93); C247(1.14); C277(1.30) | LDD0755 | [16] |
| LDCM0439 | CL49 | HCT 116 | C584(0.96); C166(1.27); C247(1.27); C277(1.21) | LDD0756 | [16] |
| LDCM0440 | CL5 | HCT 116 | C584(1.03); C166(0.91); C247(1.14); C277(1.02) | LDD0757 | [16] |
| LDCM0441 | CL50 | HCT 116 | C584(1.14); C166(1.06); C247(1.50); C277(1.46) | LDD0758 | [16] |
| LDCM0442 | CL51 | HCT 116 | C584(1.22); C166(1.24); C247(1.42); C277(1.38) | LDD0759 | [16] |
| LDCM0443 | CL52 | HCT 116 | C584(1.24); C166(0.72); C247(1.06); C277(1.25) | LDD0760 | [16] |
| LDCM0444 | CL53 | HCT 116 | C584(1.26); C166(1.04); C247(1.31); C277(1.43) | LDD0761 | [16] |
| LDCM0445 | CL54 | HCT 116 | C584(1.07); C166(1.38); C247(1.23); C277(1.42) | LDD0762 | [16] |
| LDCM0446 | CL55 | HCT 116 | C584(1.15); C166(1.33); C247(1.33); C277(1.29) | LDD0763 | [16] |
| LDCM0447 | CL56 | HCT 116 | C584(1.07); C166(1.24); C247(1.12); C277(1.36) | LDD0764 | [16] |
| LDCM0448 | CL57 | HCT 116 | C584(1.20); C166(1.57); C247(1.25); C277(1.15) | LDD0765 | [16] |
| LDCM0449 | CL58 | HCT 116 | C584(1.24); C166(0.83); C247(1.35); C277(1.34) | LDD0766 | [16] |
| LDCM0450 | CL59 | HCT 116 | C584(1.07); C166(1.45); C247(1.23); C277(1.25) | LDD0767 | [16] |
| LDCM0451 | CL6 | HCT 116 | C584(0.97); C166(0.71); C247(0.90); C277(0.75) | LDD0768 | [16] |
| LDCM0452 | CL60 | HCT 116 | C584(0.93); C166(0.95); C247(1.21); C277(1.28) | LDD0769 | [16] |
| LDCM0453 | CL61 | HCT 116 | C584(1.09); C166(0.88); C247(0.93); C277(0.98) | LDD0770 | [16] |
| LDCM0454 | CL62 | HCT 116 | C584(1.08); C166(0.81); C247(0.92); C277(0.94) | LDD0771 | [16] |
| LDCM0455 | CL63 | HCT 116 | C584(0.91); C166(0.79); C247(0.83); C277(0.94) | LDD0772 | [16] |
| LDCM0456 | CL64 | HCT 116 | C584(0.76); C166(0.89); C247(0.83); C277(0.83) | LDD0773 | [16] |
| LDCM0457 | CL65 | HCT 116 | C584(0.99); C166(0.96); C247(0.96); C277(1.02) | LDD0774 | [16] |
| LDCM0458 | CL66 | HCT 116 | C584(0.86); C166(0.81); C247(0.77); C277(0.72) | LDD0775 | [16] |
| LDCM0459 | CL67 | HCT 116 | C584(0.96); C166(1.00); C247(0.82); C277(0.85) | LDD0776 | [16] |
| LDCM0460 | CL68 | HCT 116 | C584(1.07); C166(1.19); C247(0.66); C277(0.80) | LDD0777 | [16] |
| LDCM0461 | CL69 | HCT 116 | C584(1.09); C166(0.59); C247(0.74); C277(0.68) | LDD0778 | [16] |
| LDCM0462 | CL7 | HCT 116 | C584(0.80); C166(0.95); C247(0.93); C277(0.80) | LDD0779 | [16] |
| LDCM0463 | CL70 | HCT 116 | C584(0.75); C166(0.74); C247(0.79); C277(0.68) | LDD0780 | [16] |
| LDCM0464 | CL71 | HCT 116 | C584(0.94); C166(0.83); C247(0.78); C277(0.85) | LDD0781 | [16] |
| LDCM0465 | CL72 | HCT 116 | C584(1.12); C166(0.73); C247(0.86); C277(0.93) | LDD0782 | [16] |
| LDCM0466 | CL73 | HCT 116 | C584(0.91); C166(0.94); C247(0.77); C277(0.93) | LDD0783 | [16] |
| LDCM0467 | CL74 | HCT 116 | C584(1.14); C166(0.70); C247(0.74); C277(0.83) | LDD0784 | [16] |
| LDCM0469 | CL76 | HCT 116 | C584(0.87); C247(0.72); C277(0.82); C298(1.01) | LDD0786 | [16] |
| LDCM0470 | CL77 | HCT 116 | C584(0.73); C247(0.97); C277(1.11); C298(0.99) | LDD0787 | [16] |
| LDCM0471 | CL78 | HCT 116 | C584(0.87); C247(1.01); C277(1.01); C298(1.00) | LDD0788 | [16] |
| LDCM0472 | CL79 | HCT 116 | C584(0.69); C247(0.77); C277(0.76); C298(1.09) | LDD0789 | [16] |
| LDCM0473 | CL8 | HCT 116 | C584(1.08); C166(0.55); C247(0.73); C277(0.72) | LDD0790 | [16] |
| LDCM0474 | CL80 | HCT 116 | C584(0.81); C247(0.79); C277(0.81); C298(0.99) | LDD0791 | [16] |
| LDCM0475 | CL81 | HCT 116 | C584(0.68); C247(0.97); C277(1.03); C298(1.10) | LDD0792 | [16] |
| LDCM0476 | CL82 | HCT 116 | C584(0.66); C247(0.77); C277(0.81); C298(1.13) | LDD0793 | [16] |
| LDCM0477 | CL83 | HCT 116 | C584(0.63); C247(0.73); C277(0.82); C298(0.98) | LDD0794 | [16] |
| LDCM0478 | CL84 | HCT 116 | C584(0.59); C247(0.67); C277(0.69); C298(1.00) | LDD0795 | [16] |
| LDCM0479 | CL85 | HCT 116 | C584(0.63); C247(0.90); C277(0.82); C298(0.90) | LDD0796 | [16] |
| LDCM0480 | CL86 | HCT 116 | C584(0.62); C247(1.15); C277(1.08); C298(1.26) | LDD0797 | [16] |
| LDCM0481 | CL87 | HCT 116 | C584(0.59); C247(0.90); C277(0.88); C298(1.09) | LDD0798 | [16] |
| LDCM0482 | CL88 | HCT 116 | C584(0.57); C247(0.72); C277(0.78); C298(1.07) | LDD0799 | [16] |
| LDCM0483 | CL89 | HCT 116 | C584(0.57); C247(0.79); C277(0.68); C298(0.86) | LDD0800 | [16] |
| LDCM0484 | CL9 | HCT 116 | C584(0.97); C166(0.84); C247(1.05); C277(0.92) | LDD0801 | [16] |
| LDCM0485 | CL90 | HCT 116 | C584(0.73); C247(1.19); C277(1.13); C298(1.10) | LDD0802 | [16] |
| LDCM0486 | CL91 | HCT 116 | C584(0.90); C166(1.50); C247(1.15); C277(1.02) | LDD0803 | [16] |
| LDCM0487 | CL92 | HCT 116 | C584(0.89); C166(1.67); C247(1.23); C277(1.14) | LDD0804 | [16] |
| LDCM0488 | CL93 | HCT 116 | C584(0.82); C166(1.73); C247(1.27); C277(1.25) | LDD0805 | [16] |
| LDCM0489 | CL94 | HCT 116 | C584(0.92); C166(0.89); C247(1.24); C277(0.97) | LDD0806 | [16] |
| LDCM0490 | CL95 | HCT 116 | C584(1.01); C166(1.18); C247(1.12); C277(0.95) | LDD0807 | [16] |
| LDCM0491 | CL96 | HCT 116 | C584(0.86); C166(1.14); C247(1.18); C277(0.92) | LDD0808 | [16] |
| LDCM0492 | CL97 | HCT 116 | C584(0.61); C166(0.85); C247(1.07); C277(0.80) | LDD0809 | [16] |
| LDCM0493 | CL98 | HCT 116 | C584(0.69); C166(0.81); C247(0.93); C277(0.96) | LDD0810 | [16] |
| LDCM0494 | CL99 | HCT 116 | C584(0.69); C166(1.06); C247(1.04); C277(1.02) | LDD0811 | [16] |
| LDCM0495 | E2913 | HEK-293T | C584(1.01); C166(0.94); C277(1.00); C447(1.03) | LDD1698 | [37] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C166(1.18); C584(1.10) | LDD1702 | [9] |
| LDCM0625 | F8 | Ramos | C298(0.89); C247(0.47); C270(1.71); C584(1.63) | LDD2187 | [38] |
| LDCM0572 | Fragment10 | Ramos | C298(0.75); C247(0.57); C270(0.68); C584(1.11) | LDD2189 | [38] |
| LDCM0573 | Fragment11 | Ramos | C298(2.01); C247(1.50); C270(1.76); C584(0.13) | LDD2190 | [38] |
| LDCM0574 | Fragment12 | Ramos | C298(0.80); C247(0.60); C270(0.56); C584(0.71) | LDD2191 | [38] |
| LDCM0575 | Fragment13 | Ramos | C298(1.42); C247(0.95); C270(0.69); C584(0.82) | LDD2192 | [38] |
| LDCM0576 | Fragment14 | Ramos | C298(0.99); C247(0.66); C270(0.90); C584(0.86) | LDD2193 | [38] |
| LDCM0579 | Fragment20 | Ramos | C298(0.59); C247(0.53); C270(0.37); C584(0.60) | LDD2194 | [38] |
| LDCM0580 | Fragment21 | Ramos | C298(1.86); C247(0.99); C270(0.79); C584(0.83) | LDD2195 | [38] |
| LDCM0582 | Fragment23 | Ramos | C298(2.49); C247(1.25); C270(0.87); C584(0.83) | LDD2196 | [38] |
| LDCM0578 | Fragment27 | Ramos | C298(1.96); C247(1.16); C270(0.63); C584(0.92) | LDD2197 | [38] |
| LDCM0586 | Fragment28 | Ramos | C298(0.91); C247(0.72); C270(0.99); C584(0.77) | LDD2198 | [38] |
| LDCM0588 | Fragment30 | Ramos | C298(1.21); C247(1.02); C270(0.89); C584(1.21) | LDD2199 | [38] |
| LDCM0589 | Fragment31 | Ramos | C298(2.60); C247(1.28); C270(0.87); C584(0.86) | LDD2200 | [38] |
| LDCM0590 | Fragment32 | Ramos | C298(0.92); C247(0.48); C270(0.58); C584(1.27) | LDD2201 | [38] |
| LDCM0468 | Fragment33 | HCT 116 | C584(1.03); C166(1.17); C247(0.79); C277(0.90) | LDD0785 | [16] |
| LDCM0596 | Fragment38 | Ramos | C298(1.01); C247(0.99); C270(0.74); C584(0.90) | LDD2203 | [38] |
| LDCM0566 | Fragment4 | Ramos | C298(0.79); C247(0.66); C270(0.89); C584(1.11) | LDD2184 | [38] |
| LDCM0427 | Fragment51 | HCT 116 | C584(0.70); C166(0.81); C247(0.71); C277(0.87) | LDD0744 | [16] |
| LDCM0610 | Fragment52 | Ramos | C298(1.25); C247(1.14); C270(0.80); C584(0.96) | LDD2204 | [38] |
| LDCM0614 | Fragment56 | Ramos | C298(2.30); C247(1.19); C270(1.29); C584(1.13) | LDD2205 | [38] |
| LDCM0569 | Fragment7 | Ramos | C298(0.50); C247(0.71); C270(0.59); C584(1.21) | LDD2186 | [38] |
| LDCM0571 | Fragment9 | Ramos | C298(0.79); C247(0.51); C270(0.59); C584(0.79) | LDD2188 | [38] |
| LDCM0116 | HHS-0101 | DM93 | Y75(0.90) | LDD0264 | [14] |
| LDCM0117 | HHS-0201 | DM93 | Y75(0.81) | LDD0265 | [14] |
| LDCM0118 | HHS-0301 | DM93 | Y75(0.71) | LDD0266 | [14] |
| LDCM0119 | HHS-0401 | DM93 | Y75(0.89) | LDD0267 | [14] |
| LDCM0120 | HHS-0701 | DM93 | Y75(1.05) | LDD0268 | [14] |
| LDCM0107 | IAA | HeLa | C247(0.00); C298(0.00); H554(0.00); H504(0.00) | LDD0221 | [30] |
| LDCM0022 | KB02 | HCT 116 | C277(3.18); C166(1.70); C584(1.93) | LDD0080 | [16] |
| LDCM0023 | KB03 | HCT 116 | C277(3.65); C166(2.75); C584(1.27) | LDD0081 | [16] |
| LDCM0024 | KB05 | HCT 116 | C277(3.28); C166(2.06); C584(1.62) | LDD0082 | [16] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C166(0.79); C584(0.90); C270(1.09) | LDD2102 | [9] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C166(0.51); C431(1.02) | LDD2121 | [9] |
| LDCM0109 | NEM | HeLa | H504(0.00); H138(0.00); H554(0.00) | LDD0223 | [30] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C166(0.86); C584(0.75); C431(0.78) | LDD2089 | [9] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C166(1.13); C584(1.26); C270(0.89) | LDD2090 | [9] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C166(1.00); C584(0.81); C431(0.94); C270(0.86) | LDD2092 | [9] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C166(0.92); C431(0.79) | LDD2093 | [9] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C584(1.11); C431(1.62) | LDD2094 | [9] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C584(0.95); C431(0.36); C270(0.52) | LDD2096 | [9] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C166(0.57); C431(0.71) | LDD2097 | [9] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C584(0.96); C431(0.98); C270(0.87) | LDD2098 | [9] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C166(0.92); C584(0.88) | LDD2099 | [9] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C166(0.80); C584(1.08); C431(0.89) | LDD2100 | [9] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C166(1.01); C431(1.47) | LDD2101 | [9] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C166(0.78); C584(0.96); C431(0.75); C270(0.85) | LDD2104 | [9] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C584(1.30); C431(1.55); C270(2.21) | LDD2105 | [9] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C584(0.68); C431(0.71); C270(0.54) | LDD2106 | [9] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C166(0.92); C584(0.99); C431(0.96) | LDD2107 | [9] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C166(0.85); C584(0.91); C431(1.10); C270(0.86) | LDD2108 | [9] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C166(0.72); C584(0.84); C431(0.66); C270(1.05) | LDD2109 | [9] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C166(0.74); C584(2.21); C431(1.18) | LDD2110 | [9] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C166(0.91); C431(1.02); C270(1.04) | LDD2111 | [9] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C166(0.42); C584(0.79); C431(0.83); C270(0.64) | LDD2114 | [9] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C431(0.52) | LDD2115 | [9] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C431(0.54) | LDD2116 | [9] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C166(1.19); C584(1.25); C431(0.85); C270(1.02) | LDD2118 | [9] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C166(2.27); C584(2.10); C431(1.49) | LDD2119 | [9] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C166(0.94); C584(0.97); C431(0.62) | LDD2120 | [9] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C166(0.80); C431(0.55); C270(0.68) | LDD2122 | [9] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C166(0.85); C584(0.94); C431(0.80); C270(0.81) | LDD2123 | [9] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C166(1.33); C584(0.97); C431(0.42) | LDD2124 | [9] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C166(0.73); C584(0.94); C431(0.60) | LDD2125 | [9] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C166(1.65); C584(1.27); C270(0.73) | LDD2126 | [9] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C166(0.88); C431(1.00); C270(1.03) | LDD2127 | [9] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C166(1.09); C584(1.13); C431(0.62) | LDD2128 | [9] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C166(0.96); C584(1.10); C431(0.89); C270(0.80) | LDD2129 | [9] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C166(0.78); C431(0.55) | LDD2133 | [9] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C166(0.41) | LDD2134 | [9] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C166(0.96); C584(0.72); C431(1.11) | LDD2135 | [9] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C166(0.91); C584(0.94); C431(0.86); C270(1.01) | LDD2136 | [9] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C166(0.87); C431(0.72); C270(1.04) | LDD2137 | [9] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C431(1.53); C166(1.34) | LDD1700 | [9] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C166(0.76); C584(0.78); C431(0.66) | LDD2140 | [9] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C166(0.92); C584(0.87); C431(1.02); C270(0.65) | LDD2141 | [9] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C166(1.29); C431(1.27) | LDD2143 | [9] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C166(2.77); C584(1.14) | LDD2144 | [9] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C166(0.79); C431(0.90) | LDD2146 | [9] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C166(4.09) | LDD2147 | [9] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C166(0.55); C270(0.52) | LDD2148 | [9] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C166(1.41); C431(0.53) | LDD2149 | [9] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C584(1.28); C431(0.66); C270(0.41) | LDD2150 | [9] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C584(1.22); C584(0.95); C584(0.78) | LDD2206 | [39] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C584(1.01); C584(0.96); C584(0.64); C584(0.38) | LDD2207 | [39] |
| LDCM0131 | RA190 | MM1.R | C584(1.47); C319(1.32) | LDD0304 | [40] |
| LDCM0021 | THZ1 | HCT 116 | C319(1.07); C584(1.15); C247(1.33); C447(0.92) | LDD2173 | [16] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Cellular tumor antigen p53 (TP53) | P53 family | P04637 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nuclear factor of activated T-cells 5 (NFAT5) | . | O94916 | |||
Other
References
