Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AZ-9 Probe Info |
 |
E383(10.00); D386(10.00) | LDD2208 | [1] | |
|
ONAyne Probe Info |
 |
K314(0.00); K102(0.00) | LDD0273 | [2] | |
|
HHS-465 Probe Info |
 |
Y150(8.76); Y401(0.63) | LDD2237 | [3] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0358 | [4] | |
|
NHS Probe Info |
 |
K871(0.00); K102(0.00); K298(0.00); K314(0.00) | LDD0010 | [5] | |
|
SF Probe Info |
 |
Y115(0.00); Y14(0.00); Y401(0.00); Y384(0.00) | LDD0028 | [6] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [5] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [7] | |
|
1c-yne Probe Info |
 |
K127(0.00); K223(0.00) | LDD0228 | [4] | |
|
AOyne Probe Info |
 |
14.90 | LDD0443 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C003 Probe Info |
 |
13.45 | LDD1713 | [9] | |
|
C052 Probe Info |
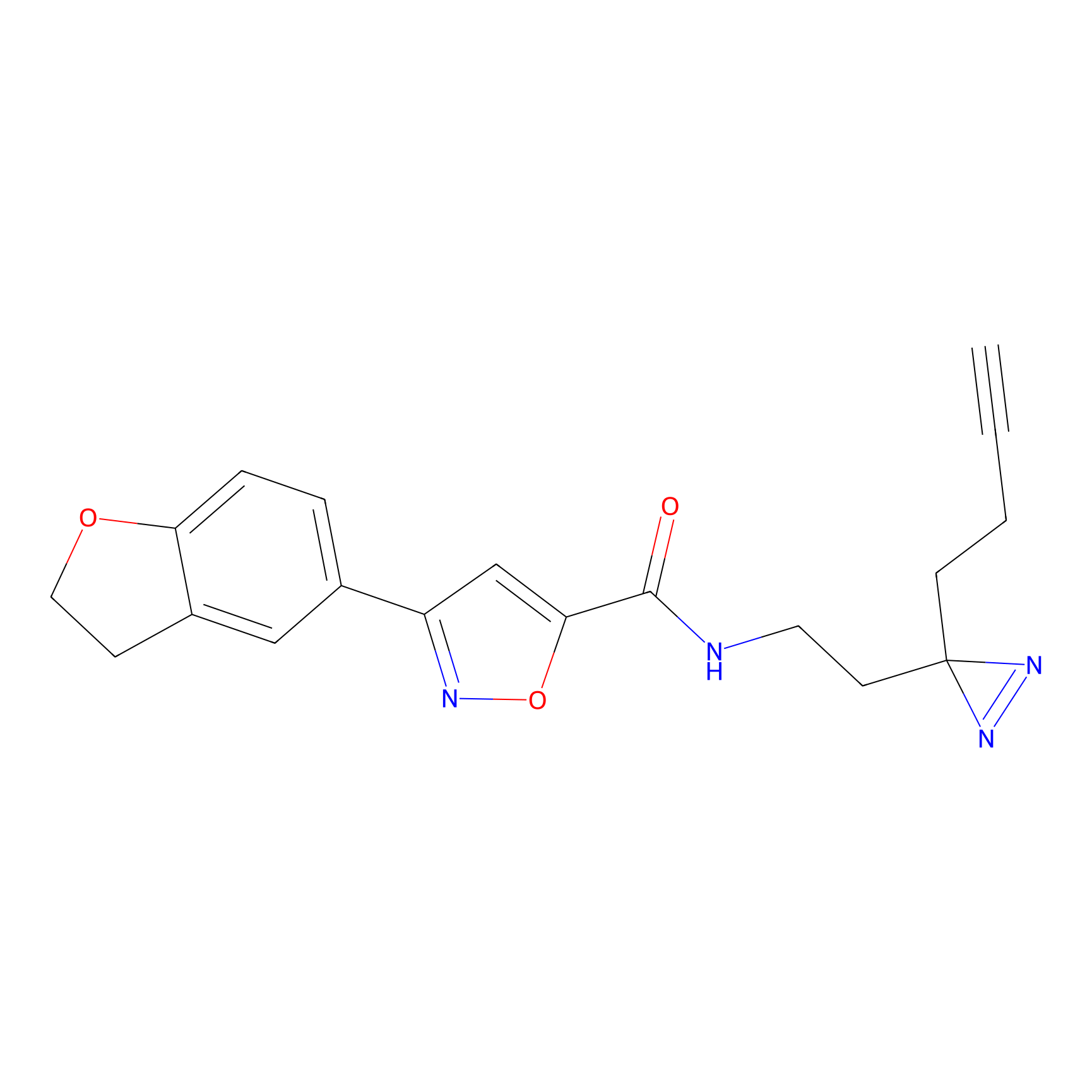 |
6.28 | LDD1750 | [9] | |
|
C053 Probe Info |
 |
5.50 | LDD1751 | [9] | |
|
C055 Probe Info |
 |
12.55 | LDD1752 | [9] | |
|
C056 Probe Info |
 |
16.45 | LDD1753 | [9] | |
|
C087 Probe Info |
 |
20.25 | LDD1779 | [9] | |
|
C106 Probe Info |
 |
23.75 | LDD1793 | [9] | |
|
C108 Probe Info |
 |
10.27 | LDD1795 | [9] | |
|
C112 Probe Info |
 |
23.43 | LDD1799 | [9] | |
|
C158 Probe Info |
 |
9.78 | LDD1838 | [9] | |
|
C159 Probe Info |
 |
7.73 | LDD1839 | [9] | |
|
C160 Probe Info |
 |
7.73 | LDD1840 | [9] | |
|
C161 Probe Info |
 |
13.74 | LDD1841 | [9] | |
|
C187 Probe Info |
 |
20.39 | LDD1865 | [9] | |
|
C191 Probe Info |
 |
13.09 | LDD1868 | [9] | |
|
C193 Probe Info |
 |
4.99 | LDD1869 | [9] | |
|
C201 Probe Info |
 |
29.24 | LDD1877 | [9] | |
|
C206 Probe Info |
 |
17.27 | LDD1881 | [9] | |
|
C210 Probe Info |
 |
67.65 | LDD1884 | [9] | |
|
C214 Probe Info |
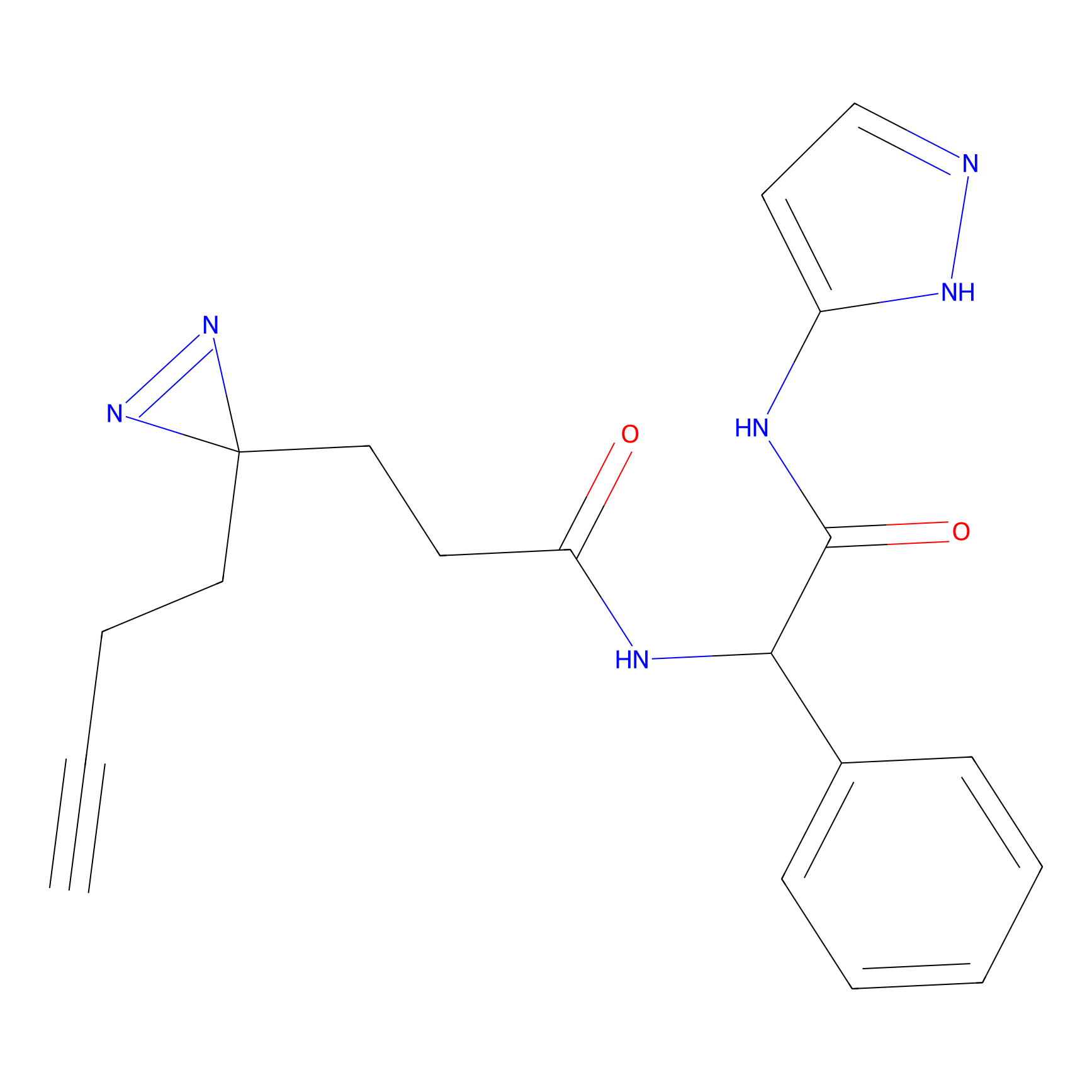 |
5.62 | LDD1888 | [9] | |
|
C284 Probe Info |
 |
22.01 | LDD1954 | [9] | |
|
C313 Probe Info |
 |
23.43 | LDD1980 | [9] | |
|
C338 Probe Info |
 |
19.70 | LDD2001 | [9] | |
|
C355 Probe Info |
 |
25.81 | LDD2016 | [9] | |
|
C361 Probe Info |
 |
17.51 | LDD2022 | [9] | |
|
C362 Probe Info |
 |
35.02 | LDD2023 | [9] | |
|
C364 Probe Info |
 |
15.35 | LDD2025 | [9] | |
|
C391 Probe Info |
 |
18.38 | LDD2050 | [9] | |
|
C407 Probe Info |
 |
12.21 | LDD2064 | [9] | |
|
FFF probe11 Probe Info |
 |
16.59 | LDD0471 | [10] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [10] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [10] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0464 | [10] | |
|
OEA-DA Probe Info |
 |
16.52 | LDD0046 | [11] | |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transcription factor Jun (JUN) | BZIP family | P05412 | |||
| Mothers against decapentaplegic homolog 2 (SMAD2) | Dwarfin/SMAD family | Q15796 | |||
Other
References
