Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alkylaryl probe 2 Probe Info |
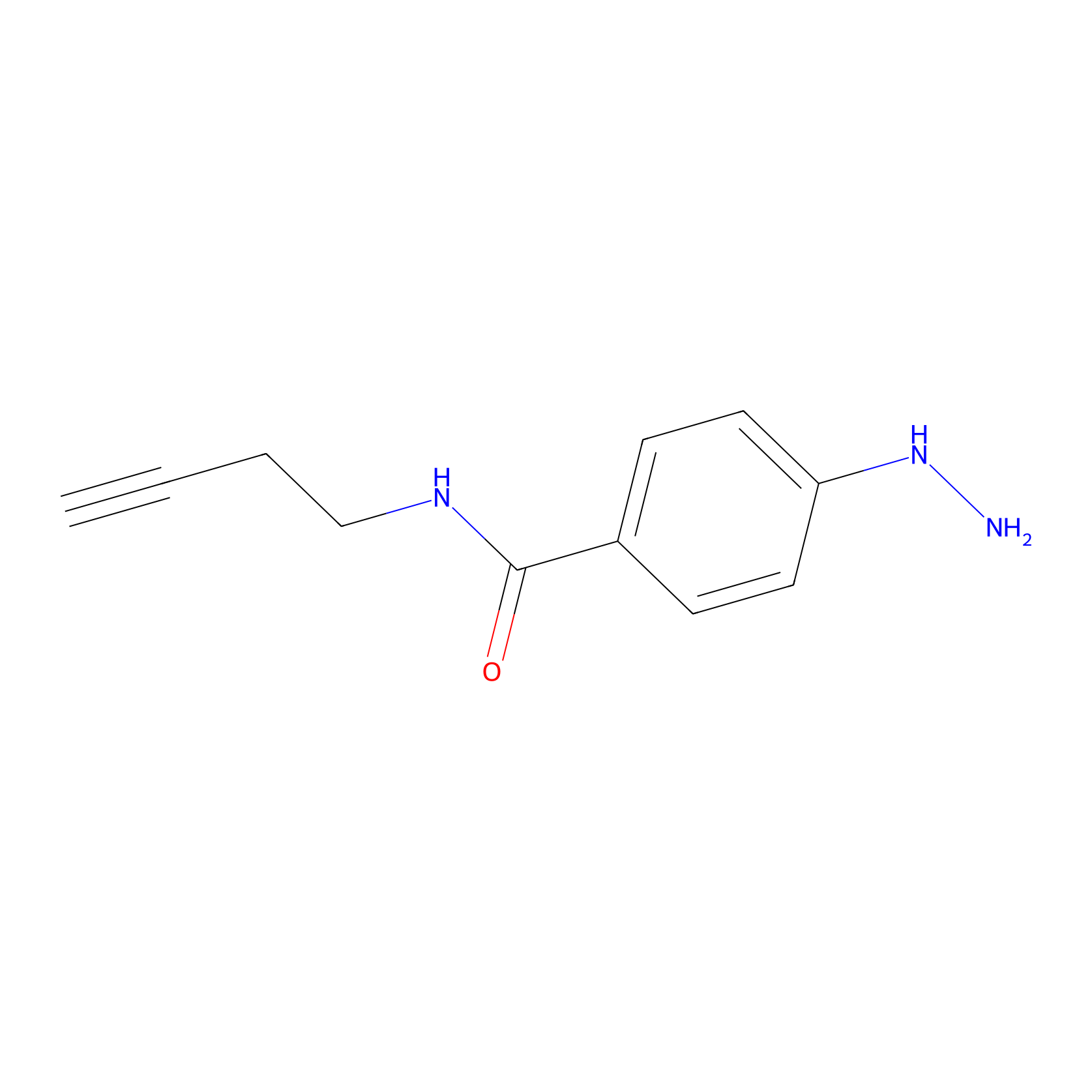 |
20.00 | LDD0389 | [1] | |
|
TH211 Probe Info |
 |
Y356(20.00) | LDD0260 | [2] | |
|
C-Sul Probe Info |
 |
6.61 | LDD0066 | [3] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [4] | |
|
STPyne Probe Info |
 |
K189(1.85); K194(6.48); K341(5.06) | LDD0277 | [5] | |
|
m-APA Probe Info |
 |
11.79 | LDD0403 | [6] | |
|
Alkylaryl probe 3 Probe Info |
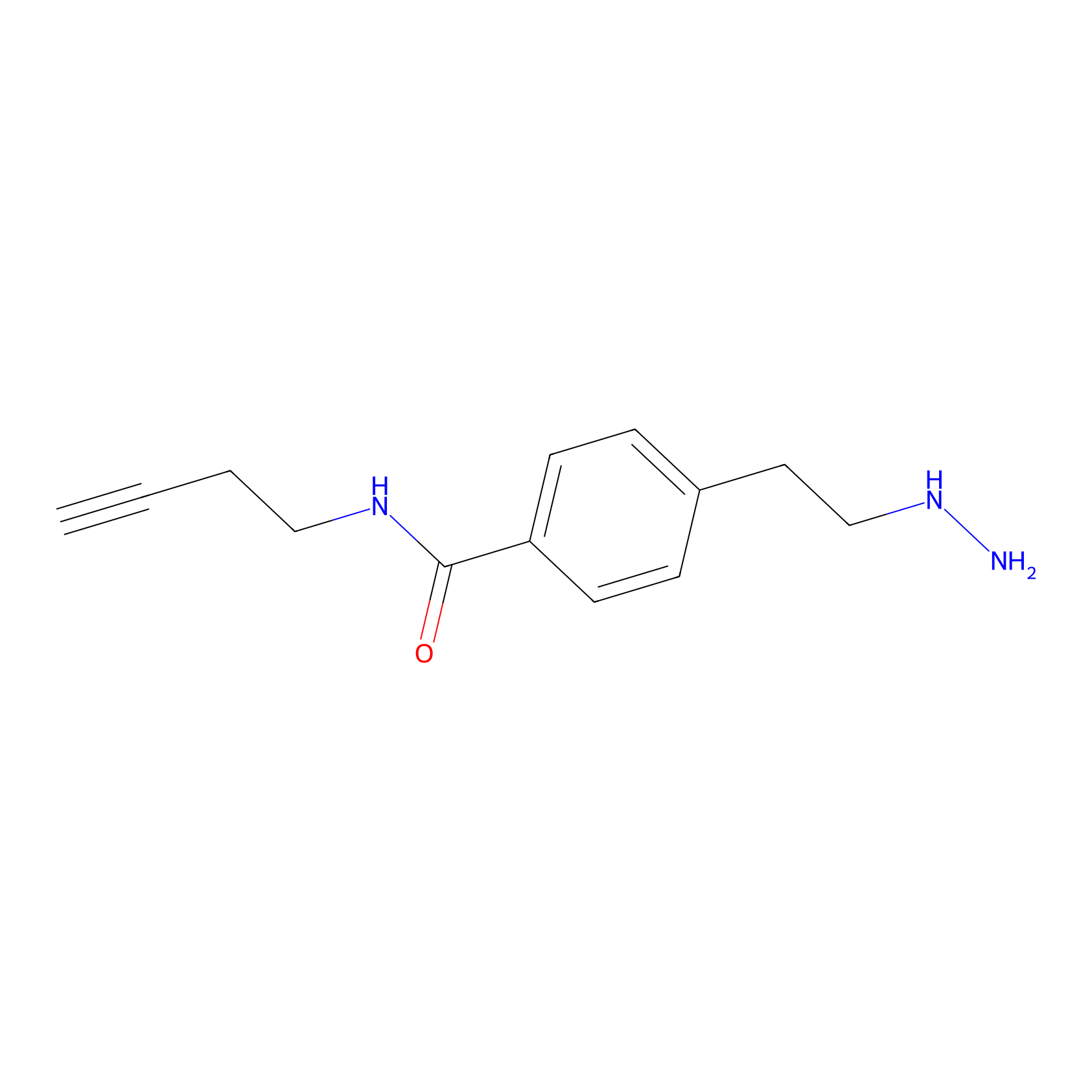 |
20.00 | LDD0282 | [7] | |
|
HPAP Probe Info |
 |
4.05 | LDD0062 | [8] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [9] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [10] | |
|
Crotonaldehyde Probe Info |
 |
H171(0.00); H167(0.00) | LDD0219 | [9] | |
|
AOyne Probe Info |
 |
10.80 | LDD0443 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C027 Probe Info |
 |
7.11 | LDD1733 | [12] | |
|
C040 Probe Info |
 |
14.22 | LDD1740 | [12] | |
|
C092 Probe Info |
 |
32.90 | LDD1783 | [12] | |
|
C094 Probe Info |
 |
25.81 | LDD1785 | [12] | |
|
C112 Probe Info |
 |
19.16 | LDD1799 | [12] | |
|
C166 Probe Info |
 |
7.84 | LDD1846 | [12] | |
|
C218 Probe Info |
 |
14.42 | LDD1892 | [12] | |
|
C219 Probe Info |
 |
4.92 | LDD1893 | [12] | |
|
C220 Probe Info |
 |
14.32 | LDD1894 | [12] | |
|
C231 Probe Info |
 |
13.36 | LDD1904 | [12] | |
|
C235 Probe Info |
 |
25.46 | LDD1908 | [12] | |
|
C289 Probe Info |
 |
44.02 | LDD1959 | [12] | |
|
C349 Probe Info |
 |
9.99 | LDD2010 | [12] | |
|
C350 Probe Info |
 |
23.92 | LDD2011 | [12] | |
|
C362 Probe Info |
 |
38.05 | LDD2023 | [12] | |
|
C366 Probe Info |
 |
7.01 | LDD2027 | [12] | |
|
C407 Probe Info |
 |
13.93 | LDD2064 | [12] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0472 | [13] | |
|
FFF probe15 Probe Info |
 |
20.00 | LDD0478 | [13] | |
|
FFF probe3 Probe Info |
 |
15.71 | LDD0465 | [13] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [14] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [15] | |
|
OEA-DA Probe Info |
 |
5.77 | LDD0046 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 11.79 | LDD0403 | [6] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [9] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [9] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [9] |
| LDCM0014 | Panhematin | HEK-293T | 4.05 | LDD0062 | [8] |
| LDCM0099 | Phenelzine | HEK-293T | 20.00 | LDD0282 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Cyclic AMP-responsive element-binding protein 3-like protein 1 (CREB3L1) | BZIP family | Q96BA8 | |||
GPCR
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Myeloid cell surface antigen CD33 (CD33) | SIGLEC (sialic acid binding Ig-like lectin) family | P20138 | |||
| Poliovirus receptor (PVR) | Nectin family | P15151 | |||
Other
References
