Details of the Target
General Information of Target
| Target ID | LDTP00266 | |||||
|---|---|---|---|---|---|---|
| Target Name | Importin-5 (IPO5) | |||||
| Gene Name | IPO5 | |||||
| Gene ID | 3843 | |||||
| Synonyms |
KPNB3; RANBP5; Importin-5; Imp5; Importin subunit beta-3; Karyopherin beta-3; Ran-binding protein 5; RanBP5 |
|||||
| 3D Structure | ||||||
| Sequence |
MAAAAAEQQQFYLLLGNLLSPDNVVRKQAEETYENIPGQSKITFLLQAIRNTTAAEEARQ
MAAVLLRRLLSSAFDEVYPALPSDVQTAIKSELLMIIQMETQSSMRKKVCDIAAELARNL IDEDGNNQWPEGLKFLFDSVSSQNVGLREAALHIFWNFPGIFGNQQQHYLDVIKRMLVQC MQDQEHPSIRTLSARATAAFILANEHNVALFKHFADLLPGFLQAVNDSCYQNDDSVLKSL VEIADTVPKYLRPHLEATLQLSLKLCGDTSLNNMQRQLALEVIVTLSETAAAMLRKHTNI VAQTIPQMLAMMVDLEEDEDWANADELEDDDFDSNAVAGESALDRMACGLGGKLVLPMIK EHIMQMLQNPDWKYRHAGLMALSAIGEGCHQQMEGILNEIVNFVLLFLQDPHPRVRYAAC NAVGQMATDFAPGFQKKFHEKVIAALLQTMEDQGNQRVQAHAAAALINFTEDCPKSLLIP YLDNLVKHLHSIMVLKLQELIQKGTKLVLEQVVTSIASVADTAEEKFVPYYDLFMPSLKH IVENAVQKELRLLRGKTIECISLIGLAVGKEKFMQDASDVMQLLLKTQTDFNDMEDDDPQ ISYMISAWARMCKILGKEFQQYLPVVMGPLMKTASIKPEVALLDTQDMENMSDDDGWEFV NLGDQQSFGIKTAGLEEKSTACQMLVCYAKELKEGFVEYTEQVVKLMVPLLKFYFHDGVR VAAAESMPLLLECARVRGPEYLTQMWHFMCDALIKAIGTEPDSDVLSEIMHSFAKCIEVM GDGCLNNEHFEELGGILKAKLEEHFKNQELRQVKRQDEDYDEQVEESLQDEDDNDVYILT KVSDILHSIFSSYKEKVLPWFEQLLPLIVNLICPHRPWPDRQWGLCIFDDVIEHCSPASF KYAEYFLRPMLQYVCDNSPEVRQAAAYGLGVMAQYGGDNYRPFCTEALPLLVRVIQSADS KTKENVNATENCISAVGKIMKFKPDCVNVEEVLPHWLSWLPLHEDKEEAVQTFNYLCDLI ESNHPIVLGPNNTNLPKIFSIIAEGEMHEAIKHEDPCAKRLANVVRQVQTSGGLWTECIA QLSPEQQAAIQELLNSA |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Importin beta family, Importin beta-3 subfamily
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Functions in nuclear protein import as nuclear transport receptor. Serves as receptor for nuclear localization signals (NLS) in cargo substrates. Is thought to mediate docking of the importin/substrate complex to the nuclear pore complex (NPC) through binding to nucleoporin and the complex is subsequently translocated through the pore by an energy requiring, Ran-dependent mechanism. At the nucleoplasmic side of the NPC, Ran binds to the importin, the importin/substrate complex dissociates and importin is re-exported from the nucleus to the cytoplasm where GTP hydrolysis releases Ran. The directionality of nuclear import is thought to be conferred by an asymmetric distribution of the GTP- and GDP-bound forms of Ran between the cytoplasm and nucleus. Mediates the nuclear import of ribosomal proteins RPL23A, RPS7 and RPL5. In vitro, mediates nuclear import of H2A, H2B, H3 and H4 histones. Binds to CPEB3 and mediates its nuclear import following neuronal stimulation. In case of HIV-1 infection, binds and mediates the nuclear import of HIV-1 Rev.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AN3CA | SNV: p.K496R | DBIA Probe Info | |||
| CAL33 | SNV: p.L489M | DBIA Probe Info | |||
| COLO320 | SNV: p.S652G | DBIA Probe Info | |||
| LNCaP clone FGC | SNV: p.E319D | DBIA Probe Info | |||
| MEWO | SNV: p.Q808Ter | DBIA Probe Info | |||
| NCIH1155 | SNV: p.A338T | DBIA Probe Info | |||
| RKO | Deletion: p.I868SfsTer24 | DBIA Probe Info | |||
| RVH421 | SNV: p.G378R | DBIA Probe Info | |||
| SKMEL3 | SNV: p.M61T | DBIA Probe Info | |||
| SUDHL4 | Insertion: p.E549GfsTer11 | DBIA Probe Info | |||
| UACC257 | SNV: p.G555E | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
10.04 | LDD0402 | [1] | |
|
A-EBA Probe Info |
 |
4.51 | LDD0215 | [2] | |
|
CHEMBL5175495 Probe Info |
 |
7.58 | LDD0196 | [3] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [4] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [4] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [4] | |
|
TH211 Probe Info |
 |
Y940(10.99); Y714(7.30) | LDD0257 | [5] | |
|
TH214 Probe Info |
 |
Y853(15.15) | LDD0258 | [5] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [6] | |
|
ONAyne Probe Info |
 |
K108(0.50); K800(0.20); K961(0.59) | LDD0274 | [7] | |
|
STPyne Probe Info |
 |
K108(7.61); K436(10.00); K548(1.61); K613(4.74) | LDD0277 | [7] | |
|
Probe 1 Probe Info |
 |
Y699(26.87) | LDD3495 | [8] | |
|
JZ128-DTB Probe Info |
 |
C972(0.00); C682(0.00); C473(0.00); C1078(0.00) | LDD0462 | [9] | |
|
THZ1-DTB Probe Info |
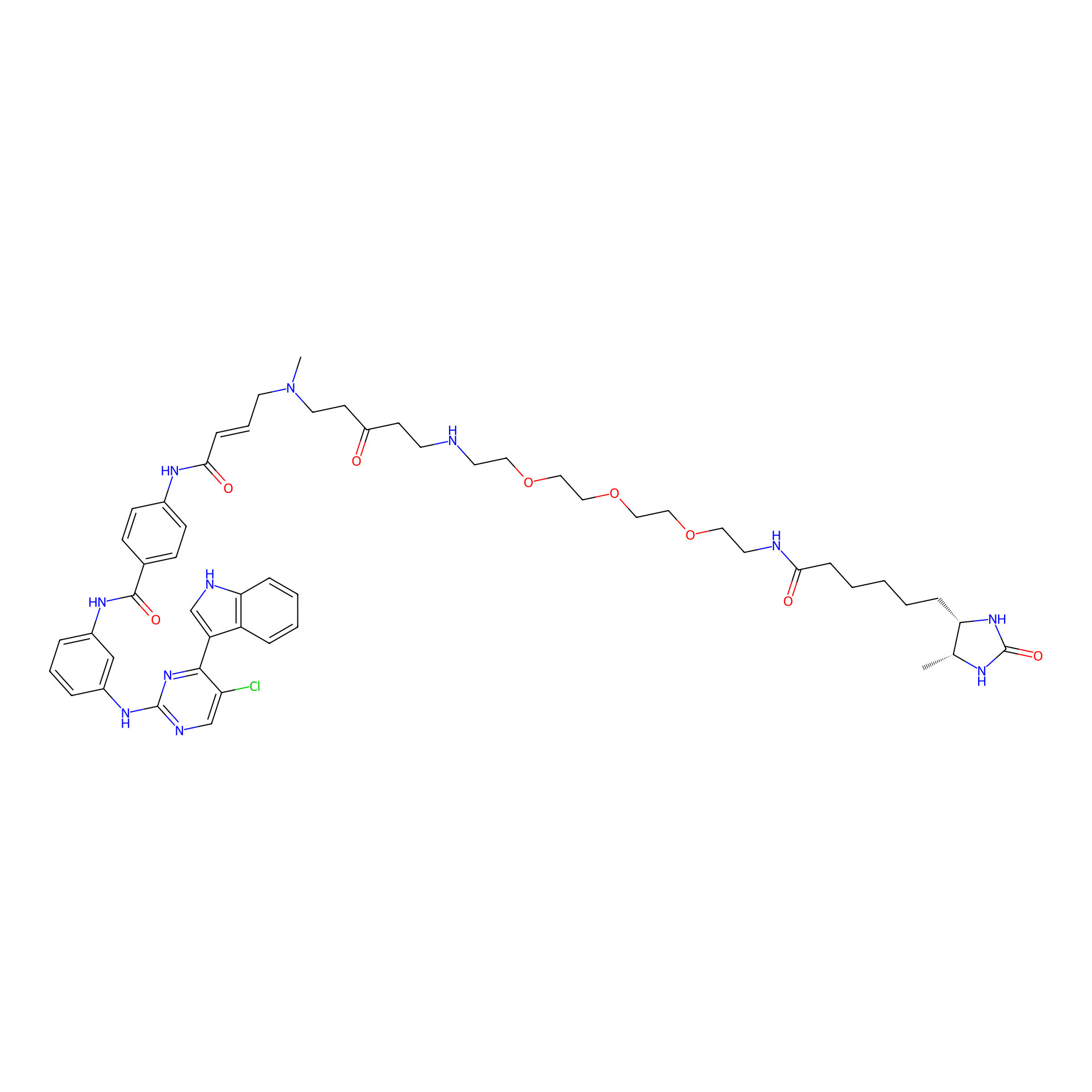 |
C733(1.08); C420(1.10); C944(1.02); C1078(0.98) | LDD0460 | [9] | |
|
Sulforaphane-probe2 Probe Info |
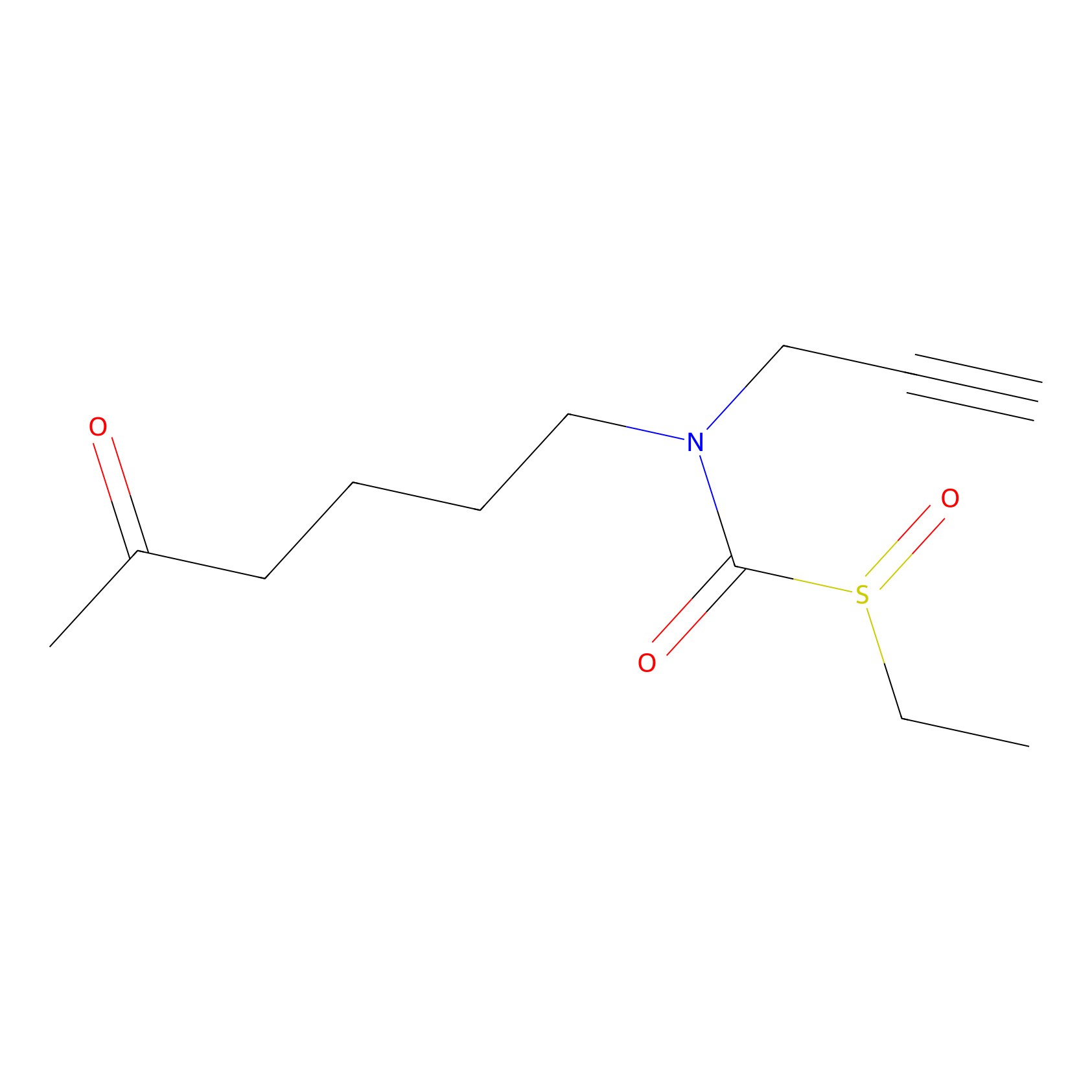 |
2.04 | LDD0042 | [10] | |
|
DA-P3 Probe Info |
 |
8.89 | LDD0183 | [11] | |
|
AHL-Pu-1 Probe Info |
 |
C784(4.71); C473(3.52); C229(20.00); C915(3.48) | LDD0169 | [12] | |
|
EA-probe Probe Info |
 |
C733(0.75) | LDD2210 | [13] | |
|
DBIA Probe Info |
 |
C682(4.36); C687(4.36); C560(4.44) | LDD0204 | [14] | |
|
HHS-475 Probe Info |
 |
Y905(0.82) | LDD0264 | [15] | |
|
HHS-465 Probe Info |
 |
Y622(4.97); Y714(2.86); Y905(10.00); Y935(2.82) | LDD2237 | [16] | |
|
5E-2FA Probe Info |
 |
H206(0.00); H186(0.00); H461(0.00) | LDD2235 | [17] | |
|
ATP probe Probe Info |
 |
K1052(0.00); K1059(0.00); K108(0.00); K806(0.00) | LDD0199 | [18] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C972(0.00); C750(0.00); C1057(0.00); C229(0.00) | LDD0038 | [19] | |
|
IA-alkyne Probe Info |
 |
C266(0.00); C944(0.00); C348(0.00); C473(0.00) | LDD0032 | [20] | |
|
IPIAA_H Probe Info |
 |
C110(0.00); C733(0.00); C944(0.00); C229(0.00) | LDD0030 | [21] | |
|
IPIAA_L Probe Info |
 |
C420(0.00); C944(0.00); C733(0.00); C750(0.00) | LDD0031 | [21] | |
|
Lodoacetamide azide Probe Info |
 |
C972(0.00); C986(0.00); C750(0.00); C229(0.00) | LDD0037 | [19] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [22] | |
|
JW-RF-010 Probe Info |
 |
C733(0.00); C180(0.00); C473(0.00); C110(0.00) | LDD0026 | [23] | |
|
NAIA_4 Probe Info |
 |
C180(0.00); C229(0.00); C420(0.00); C473(0.00) | LDD2226 | [24] | |
|
TFBX Probe Info |
 |
C110(0.00); C972(0.00); C420(0.00); C266(0.00) | LDD0027 | [23] | |
|
WYneN Probe Info |
 |
C972(0.00); C733(0.00); C110(0.00) | LDD0021 | [22] | |
|
WYneO Probe Info |
 |
C733(0.00); C110(0.00); C972(0.00); C266(0.00) | LDD0022 | [22] | |
|
aHNE Probe Info |
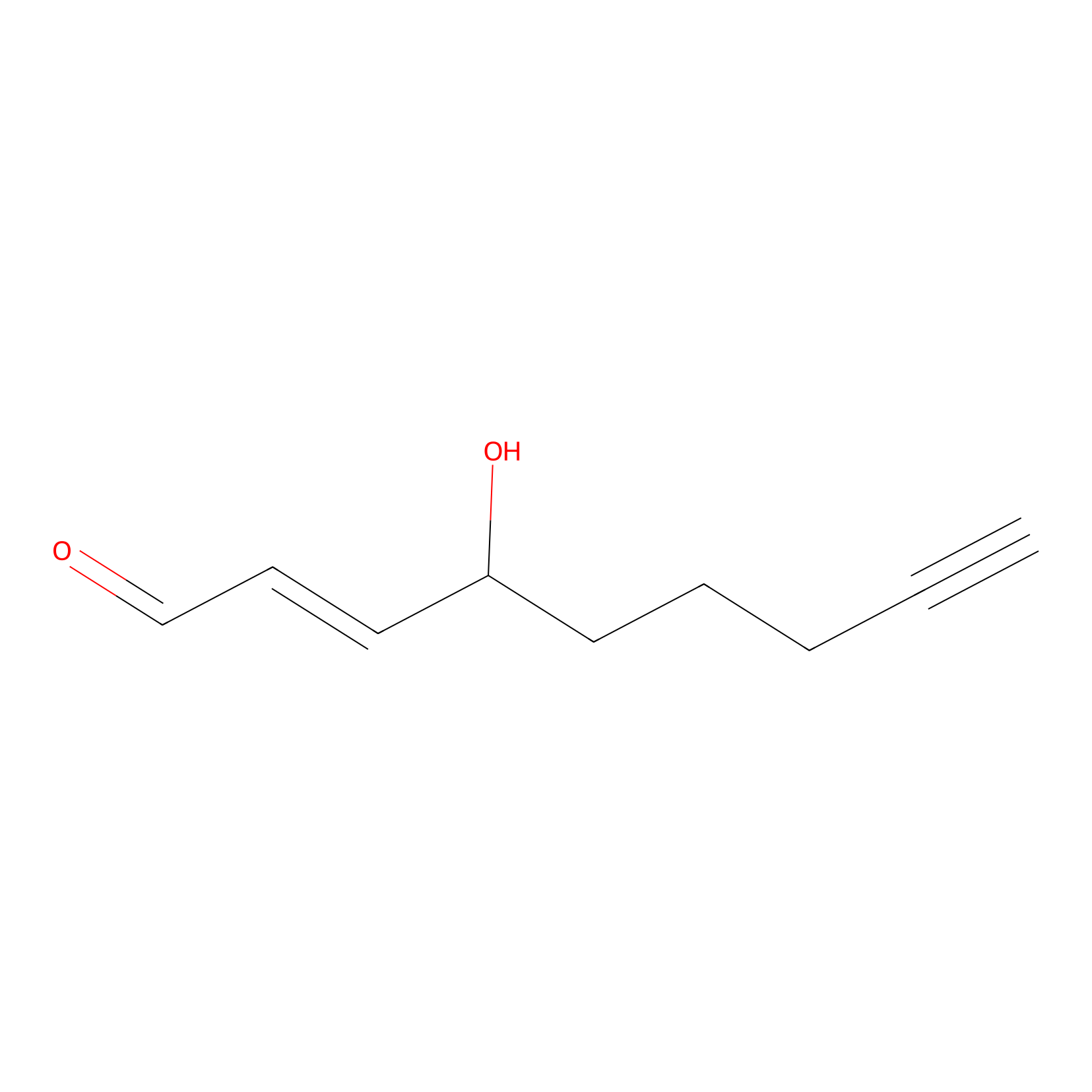 |
N.A. | LDD0001 | [22] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [25] | |
|
Compound 10 Probe Info |
 |
C1057(0.00); C110(0.00); C420(0.00); C473(0.00) | LDD2216 | [26] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [26] | |
|
ENE Probe Info |
 |
C348(0.00); C1057(0.00); C972(0.00); C110(0.00) | LDD0006 | [22] | |
|
IPM Probe Info |
 |
C733(0.00); C110(0.00); C972(0.00); C473(0.00) | LDD0005 | [22] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [27] | |
|
PPMS Probe Info |
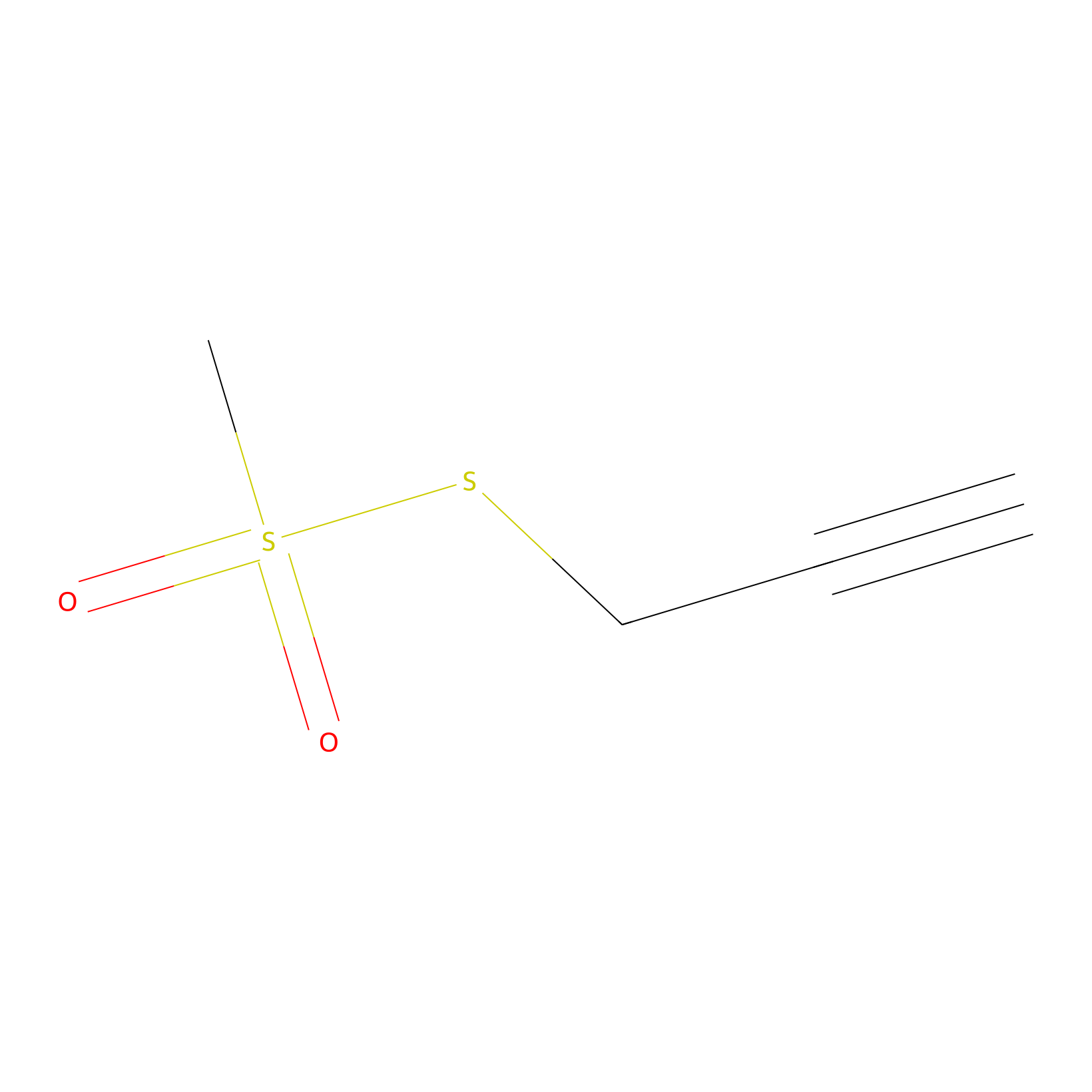 |
N.A. | LDD0008 | [22] | |
|
VSF Probe Info |
 |
C110(0.00); C972(0.00) | LDD0007 | [22] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [28] | |
|
1c-yne Probe Info |
 |
K693(0.00); K712(0.00); K441(0.00); K617(0.00) | LDD0228 | [25] | |
|
Acrolein Probe Info |
 |
C972(0.00); C473(0.00); C180(0.00); C733(0.00) | LDD0217 | [29] | |
|
Cinnamaldehyde Probe Info |
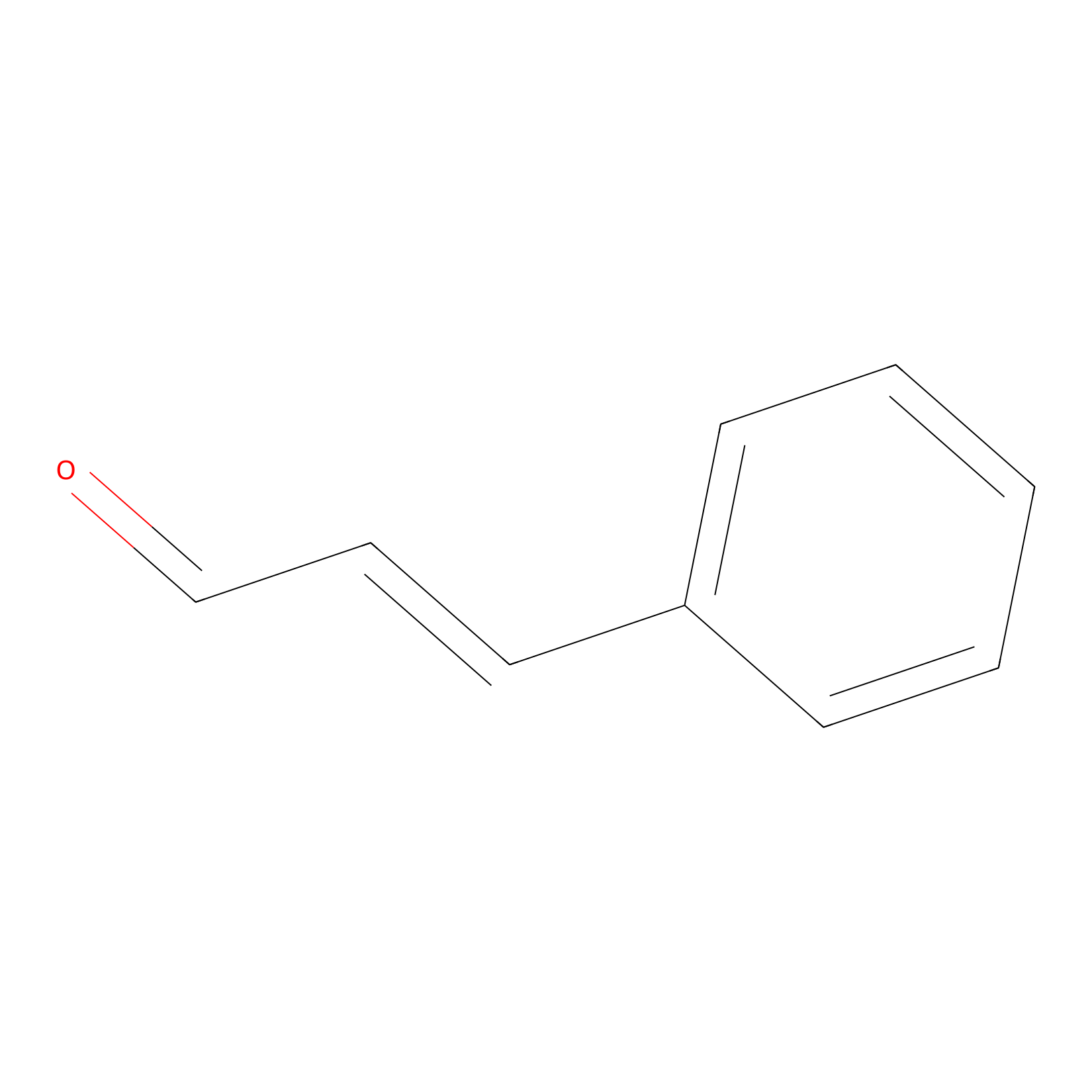 |
N.A. | LDD0220 | [29] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [29] | |
|
Methacrolein Probe Info |
 |
C473(0.00); C972(0.00); C420(0.00); C110(0.00) | LDD0218 | [29] | |
|
W1 Probe Info |
 |
C915(0.00); C733(0.00); C110(0.00); C972(0.00) | LDD0236 | [30] | |
|
AOyne Probe Info |
 |
9.70 | LDD0443 | [31] | |
|
NAIA_5 Probe Info |
 |
C972(0.00); C986(0.00); C560(0.00); C389(0.00) | LDD2223 | [24] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe13 Probe Info |
 |
14.43 | LDD0475 | [32] | |
|
FFF probe14 Probe Info |
 |
12.64 | LDD0477 | [32] | |
|
FFF probe2 Probe Info |
 |
5.54 | LDD0463 | [32] | |
|
FFF probe3 Probe Info |
 |
6.22 | LDD0465 | [32] | |
|
JN0003 Probe Info |
 |
12.07 | LDD0469 | [32] | |
|
STS-1 Probe Info |
 |
4.60 | LDD0136 | [33] | |
|
STS-2 Probe Info |
 |
8.20 | LDD0138 | [33] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [34] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [35] | |
|
OEA-DA Probe Info |
 |
4.98 | LDD0046 | [36] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C972(0.53); C733(0.58); C348(0.46); C180(0.44) | LDD2142 | [37] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C972(0.71); C733(0.85); C348(0.71); C180(0.80) | LDD2112 | [37] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C972(1.10); C733(0.71); C348(0.66); C180(0.64) | LDD2095 | [37] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C972(0.91); C733(0.90); C348(0.90); C180(1.01) | LDD2130 | [37] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C972(1.00); C733(1.32); C348(0.84); C180(1.11) | LDD2117 | [37] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C972(1.29); C733(1.47); C348(1.01); C180(1.20) | LDD2152 | [37] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C733(1.07); C348(1.07); C180(1.17); C110(1.12) | LDD2103 | [37] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C348(0.55); C180(0.50); C110(0.90); C420(0.40) | LDD2132 | [37] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C972(0.64); C733(0.68); C348(0.75); C180(0.83) | LDD2131 | [37] |
| LDCM0025 | 4SU-RNA | DM93 | C229(2.03); C915(2.33); C886(2.84) | LDD0170 | [12] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C784(4.71); C473(3.52); C229(20.00); C915(3.48) | LDD0169 | [12] |
| LDCM0214 | AC1 | HEK-293T | C1057(1.02); C180(1.01); C266(0.87); C348(0.98) | LDD1507 | [38] |
| LDCM0215 | AC10 | HEK-293T | C1057(1.00); C180(0.99); C266(0.94); C348(0.98) | LDD1508 | [38] |
| LDCM0226 | AC11 | HEK-293T | C1057(0.99); C180(1.06); C266(0.94); C348(0.92) | LDD1509 | [38] |
| LDCM0237 | AC12 | HEK-293T | C1057(1.06); C180(0.98); C266(0.90); C348(0.92) | LDD1510 | [38] |
| LDCM0259 | AC14 | HEK-293T | C1057(0.98); C180(1.02); C266(0.93); C348(0.93) | LDD1512 | [38] |
| LDCM0270 | AC15 | HEK-293T | C1057(0.94); C180(1.04); C266(0.91); C348(0.90) | LDD1513 | [38] |
| LDCM0276 | AC17 | HEK-293T | C1057(1.06); C180(1.04); C266(0.91); C348(1.04) | LDD1515 | [38] |
| LDCM0277 | AC18 | HEK-293T | C1057(1.00); C180(0.95); C266(0.97); C348(1.06) | LDD1516 | [38] |
| LDCM0278 | AC19 | HEK-293T | C1057(1.19); C180(0.88); C266(0.88); C348(1.26) | LDD1517 | [38] |
| LDCM0279 | AC2 | HEK-293T | C1057(0.96); C180(0.95); C266(0.96); C348(0.95) | LDD1518 | [38] |
| LDCM0280 | AC20 | HEK-293T | C1057(1.04); C180(1.02); C266(0.99); C348(0.99) | LDD1519 | [38] |
| LDCM0281 | AC21 | HEK-293T | C1057(1.00); C180(0.95); C266(0.93); C348(0.99) | LDD1520 | [38] |
| LDCM0282 | AC22 | HEK-293T | C1057(1.00); C180(1.02); C266(0.93); C348(1.00) | LDD1521 | [38] |
| LDCM0283 | AC23 | HEK-293T | C1057(0.94); C180(1.02); C266(0.97); C348(0.95) | LDD1522 | [38] |
| LDCM0284 | AC24 | HEK-293T | C1057(0.95); C266(1.00); C348(1.00); C972(1.01) | LDD1523 | [38] |
| LDCM0285 | AC25 | HEK-293T | C1057(1.02); C180(1.02); C266(0.93); C348(1.02) | LDD1524 | [38] |
| LDCM0286 | AC26 | HEK-293T | C1057(1.03); C180(0.98); C266(0.98); C348(0.99) | LDD1525 | [38] |
| LDCM0287 | AC27 | HEK-293T | C1057(1.04); C180(0.97); C266(1.04); C348(1.04) | LDD1526 | [38] |
| LDCM0288 | AC28 | HEK-293T | C1057(1.00); C180(1.04); C266(0.95); C348(0.97) | LDD1527 | [38] |
| LDCM0289 | AC29 | HEK-293T | C1057(0.99); C180(1.02); C266(0.95); C348(1.00) | LDD1528 | [38] |
| LDCM0290 | AC3 | HEK-293T | C1057(1.01); C180(0.96); C266(0.94); C348(0.95) | LDD1529 | [38] |
| LDCM0291 | AC30 | HEK-293T | C1057(1.01); C180(1.01); C266(1.02); C348(0.99) | LDD1530 | [38] |
| LDCM0292 | AC31 | HEK-293T | C1057(0.97); C180(0.98); C266(0.99); C348(0.96) | LDD1531 | [38] |
| LDCM0293 | AC32 | HEK-293T | C1057(0.94); C266(1.00); C348(1.03); C972(0.98) | LDD1532 | [38] |
| LDCM0294 | AC33 | HEK-293T | C1057(0.94); C180(1.02); C266(0.82); C348(1.00) | LDD1533 | [38] |
| LDCM0295 | AC34 | HEK-293T | C1057(0.97); C180(1.00); C266(0.97); C348(0.93) | LDD1534 | [38] |
| LDCM0296 | AC35 | HEK-293T | C1057(0.97); C180(1.03); C266(0.98); C348(0.90) | LDD1535 | [38] |
| LDCM0297 | AC36 | HEK-293T | C1057(1.02); C180(1.01); C266(0.89); C348(0.89) | LDD1536 | [38] |
| LDCM0298 | AC37 | HEK-293T | C1057(0.88); C180(1.07); C266(0.97); C348(0.87) | LDD1537 | [38] |
| LDCM0299 | AC38 | HEK-293T | C1057(0.90); C180(1.09); C266(0.96); C348(0.91) | LDD1538 | [38] |
| LDCM0300 | AC39 | HEK-293T | C1057(0.84); C180(0.99); C266(0.90); C348(0.87) | LDD1539 | [38] |
| LDCM0301 | AC4 | HEK-293T | C1057(1.07); C180(0.96); C266(0.94); C348(0.88) | LDD1540 | [38] |
| LDCM0302 | AC40 | HEK-293T | C1057(0.87); C266(0.97); C348(0.96); C972(0.90) | LDD1541 | [38] |
| LDCM0303 | AC41 | HEK-293T | C1057(0.91); C180(1.01); C266(0.98); C348(0.99) | LDD1542 | [38] |
| LDCM0304 | AC42 | HEK-293T | C1057(1.02); C180(1.03); C266(1.00); C348(1.01) | LDD1543 | [38] |
| LDCM0305 | AC43 | HEK-293T | C1057(0.98); C180(1.03); C266(0.96); C348(0.92) | LDD1544 | [38] |
| LDCM0306 | AC44 | HEK-293T | C1057(1.03); C180(0.98); C266(0.94); C348(0.93) | LDD1545 | [38] |
| LDCM0307 | AC45 | HEK-293T | C1057(0.92); C180(0.98); C266(0.96); C348(0.95) | LDD1546 | [38] |
| LDCM0308 | AC46 | HEK-293T | C1057(0.99); C180(1.08); C266(1.00); C348(0.96) | LDD1547 | [38] |
| LDCM0309 | AC47 | HEK-293T | C1057(0.87); C180(1.01); C266(0.92); C348(0.88) | LDD1548 | [38] |
| LDCM0310 | AC48 | HEK-293T | C1057(0.90); C266(1.03); C348(0.94); C972(0.88) | LDD1549 | [38] |
| LDCM0311 | AC49 | HEK-293T | C1057(0.98); C180(0.86); C266(0.90); C348(0.99) | LDD1550 | [38] |
| LDCM0312 | AC5 | HEK-293T | C1057(0.91); C180(1.02); C266(0.95); C348(0.94) | LDD1551 | [38] |
| LDCM0313 | AC50 | HEK-293T | C1057(0.93); C180(0.99); C266(1.01); C348(0.95) | LDD1552 | [38] |
| LDCM0314 | AC51 | HEK-293T | C1057(0.97); C180(1.04); C266(0.94); C348(0.95) | LDD1553 | [38] |
| LDCM0315 | AC52 | HEK-293T | C1057(1.07); C180(0.97); C266(0.95); C348(0.91) | LDD1554 | [38] |
| LDCM0316 | AC53 | HEK-293T | C1057(0.95); C180(0.94); C266(0.96); C348(0.93) | LDD1555 | [38] |
| LDCM0317 | AC54 | HEK-293T | C1057(0.96); C180(0.93); C266(0.98); C348(0.93) | LDD1556 | [38] |
| LDCM0318 | AC55 | HEK-293T | C1057(0.94); C180(0.93); C266(0.95); C348(0.89) | LDD1557 | [38] |
| LDCM0319 | AC56 | HEK-293T | C1057(0.90); C266(0.99); C348(0.98); C972(0.85) | LDD1558 | [38] |
| LDCM0320 | AC57 | HEK-293T | C1057(1.02); C180(0.92); C266(0.99); C348(1.04) | LDD1559 | [38] |
| LDCM0321 | AC58 | HEK-293T | C1057(0.94); C180(0.91); C266(0.94); C348(0.95) | LDD1560 | [38] |
| LDCM0322 | AC59 | HEK-293T | C1057(0.94); C180(0.98); C266(0.95); C348(0.94) | LDD1561 | [38] |
| LDCM0323 | AC6 | HEK-293T | C1057(0.99); C180(0.96); C266(0.98); C348(0.94) | LDD1562 | [38] |
| LDCM0324 | AC60 | HEK-293T | C1057(1.06); C180(0.97); C266(0.90); C348(0.99) | LDD1563 | [38] |
| LDCM0325 | AC61 | HEK-293T | C1057(0.96); C180(0.90); C266(0.94); C348(0.95) | LDD1564 | [38] |
| LDCM0326 | AC62 | HEK-293T | C1057(0.99); C180(1.02); C266(0.98); C348(1.00) | LDD1565 | [38] |
| LDCM0327 | AC63 | HEK-293T | C1057(0.93); C180(0.94); C266(0.98); C348(0.94) | LDD1566 | [38] |
| LDCM0328 | AC64 | HEK-293T | C1057(0.92); C266(0.99); C348(0.99); C972(0.94) | LDD1567 | [38] |
| LDCM0334 | AC7 | HEK-293T | C1057(0.95); C180(1.02); C266(0.94); C348(0.87) | LDD1568 | [38] |
| LDCM0345 | AC8 | HEK-293T | C1057(0.92); C266(0.98); C348(0.96); C972(0.88) | LDD1569 | [38] |
| LDCM0545 | Acetamide | MDA-MB-231 | C972(0.48); C733(1.04); C348(0.60); C180(0.72) | LDD2138 | [37] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C972(0.77); C733(0.80); C348(0.90); C180(0.86) | LDD2113 | [37] |
| LDCM0248 | AKOS034007472 | HEK-293T | C1057(0.99); C180(0.95); C266(0.97); C348(0.94) | LDD1511 | [38] |
| LDCM0356 | AKOS034007680 | HEK-293T | C1057(0.97); C180(0.94); C266(0.95); C348(0.99) | LDD1570 | [38] |
| LDCM0275 | AKOS034007705 | HEK-293T | C1057(0.97); C266(1.02); C348(0.99); C972(0.96) | LDD1514 | [38] |
| LDCM0156 | Aniline | NCI-H1299 | N.A. | LDD0405 | [1] |
| LDCM0102 | BDHI 8 | Jurkat | C682(4.36); C687(4.36); C560(4.44) | LDD0204 | [14] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C972(0.67); C733(0.85); C348(0.54); C180(0.99) | LDD2091 | [37] |
| LDCM0630 | CCW28-3 | 231MFP | C915(1.14) | LDD2214 | [39] |
| LDCM0108 | Chloroacetamide | HeLa | C473(0.00); C180(0.00); C972(0.00); C420(0.00) | LDD0222 | [29] |
| LDCM0632 | CL-Sc | Hep-G2 | C110(20.00); C733(1.08); C420(0.85); C682(0.79) | LDD2227 | [24] |
| LDCM0367 | CL1 | HEK-293T | C1057(0.99); C180(1.04); C266(0.93); C348(1.00) | LDD1571 | [38] |
| LDCM0368 | CL10 | HEK-293T | C1057(2.00); C180(1.17); C266(1.21); C348(1.74) | LDD1572 | [38] |
| LDCM0369 | CL100 | HEK-293T | C1057(1.06); C180(0.84); C266(0.82); C348(1.06) | LDD1573 | [38] |
| LDCM0370 | CL101 | HEK-293T | C1057(0.98); C180(1.03); C266(0.93); C348(0.98) | LDD1574 | [38] |
| LDCM0371 | CL102 | HEK-293T | C180(0.91); C266(0.88); C348(1.16); C972(0.98) | LDD1575 | [38] |
| LDCM0372 | CL103 | HEK-293T | C1057(0.99); C180(1.01); C266(0.89); C348(0.99) | LDD1576 | [38] |
| LDCM0373 | CL104 | HEK-293T | C1057(1.02); C180(0.92); C266(0.84); C348(0.98) | LDD1577 | [38] |
| LDCM0374 | CL105 | HEK-293T | C1057(1.03); C180(0.94); C266(0.99); C348(1.05) | LDD1578 | [38] |
| LDCM0375 | CL106 | HEK-293T | C180(0.91); C266(0.79); C348(1.05); C972(0.98) | LDD1579 | [38] |
| LDCM0376 | CL107 | HEK-293T | C1057(0.95); C180(0.96); C266(0.88); C348(1.01) | LDD1580 | [38] |
| LDCM0377 | CL108 | HEK-293T | C1057(0.95); C180(0.99); C266(0.76); C348(0.99) | LDD1581 | [38] |
| LDCM0378 | CL109 | HEK-293T | C1057(1.11); C180(1.03); C266(0.96); C348(1.13) | LDD1582 | [38] |
| LDCM0379 | CL11 | HEK-293T | C1057(0.81); C180(1.28); C266(1.05); C348(0.94) | LDD1583 | [38] |
| LDCM0380 | CL110 | HEK-293T | C180(0.96); C266(0.86); C348(1.38); C972(1.03) | LDD1584 | [38] |
| LDCM0381 | CL111 | HEK-293T | C1057(1.08); C180(0.86); C266(0.96); C348(1.11) | LDD1585 | [38] |
| LDCM0382 | CL112 | HEK-293T | C1057(0.98); C180(0.92); C266(0.82); C348(1.03) | LDD1586 | [38] |
| LDCM0383 | CL113 | HEK-293T | C1057(0.90); C180(1.05); C266(0.90); C348(0.90) | LDD1587 | [38] |
| LDCM0384 | CL114 | HEK-293T | C180(0.98); C266(0.89); C348(1.27); C972(1.01) | LDD1588 | [38] |
| LDCM0385 | CL115 | HEK-293T | C1057(0.91); C180(1.03); C266(0.91); C348(0.92) | LDD1589 | [38] |
| LDCM0386 | CL116 | HEK-293T | C1057(0.93); C180(0.89); C266(0.76); C348(0.94) | LDD1590 | [38] |
| LDCM0387 | CL117 | HEK-293T | C1057(0.91); C180(1.03); C266(0.92); C348(0.92) | LDD1591 | [38] |
| LDCM0388 | CL118 | HEK-293T | C180(0.92); C266(0.95); C348(0.96); C972(0.93) | LDD1592 | [38] |
| LDCM0389 | CL119 | HEK-293T | C1057(0.93); C180(0.95); C266(0.82); C348(0.93) | LDD1593 | [38] |
| LDCM0390 | CL12 | HEK-293T | C1057(0.81); C266(1.07); C348(0.95); C972(0.91) | LDD1594 | [38] |
| LDCM0391 | CL120 | HEK-293T | C1057(0.94); C180(0.91); C266(0.77); C348(0.92) | LDD1595 | [38] |
| LDCM0392 | CL121 | HEK-293T | C1057(0.95); C180(0.99); C266(0.93); C348(0.90) | LDD1596 | [38] |
| LDCM0393 | CL122 | HEK-293T | C180(0.89); C266(0.89); C348(1.00); C972(0.98) | LDD1597 | [38] |
| LDCM0394 | CL123 | HEK-293T | C1057(1.90); C180(0.98); C266(0.98); C348(1.45) | LDD1598 | [38] |
| LDCM0395 | CL124 | HEK-293T | C1057(1.26); C180(0.89); C266(0.77); C348(1.06) | LDD1599 | [38] |
| LDCM0396 | CL125 | HEK-293T | C1057(0.94); C180(0.92); C266(1.01); C348(1.00) | LDD1600 | [38] |
| LDCM0397 | CL126 | HEK-293T | C180(0.85); C266(0.93); C348(1.03); C972(0.99) | LDD1601 | [38] |
| LDCM0398 | CL127 | HEK-293T | C1057(0.96); C180(0.91); C266(0.92); C348(0.99) | LDD1602 | [38] |
| LDCM0399 | CL128 | HEK-293T | C1057(0.97); C180(0.79); C266(0.87); C348(0.97) | LDD1603 | [38] |
| LDCM0400 | CL13 | HEK-293T | C1057(0.97); C180(1.01); C266(0.87); C348(0.89) | LDD1604 | [38] |
| LDCM0401 | CL14 | HEK-293T | C180(0.99); C266(0.79); C348(0.84); C972(0.98) | LDD1605 | [38] |
| LDCM0402 | CL15 | HEK-293T | C1057(2.06); C180(0.98); C266(0.81); C348(1.48) | LDD1606 | [38] |
| LDCM0403 | CL16 | HEK-293T | C1057(0.90); C180(0.97); C266(0.72); C348(0.92) | LDD1607 | [38] |
| LDCM0404 | CL17 | HEK-293T | C1057(2.20); C180(0.99); C266(0.86); C348(1.50) | LDD1608 | [38] |
| LDCM0405 | CL18 | HEK-293T | C1057(1.02); C180(0.95); C266(0.97); C348(0.88) | LDD1609 | [38] |
| LDCM0406 | CL19 | HEK-293T | C1057(1.03); C180(1.03); C266(0.95); C348(0.91) | LDD1610 | [38] |
| LDCM0407 | CL2 | HEK-293T | C180(0.96); C266(0.77); C348(0.86); C972(1.03) | LDD1611 | [38] |
| LDCM0408 | CL20 | HEK-293T | C1057(0.91); C180(1.05); C266(0.92); C348(0.77) | LDD1612 | [38] |
| LDCM0409 | CL21 | HEK-293T | C1057(1.24); C180(1.02); C266(1.01); C348(1.34) | LDD1613 | [38] |
| LDCM0410 | CL22 | HEK-293T | C1057(0.91); C180(1.11); C266(1.02); C348(0.92) | LDD1614 | [38] |
| LDCM0411 | CL23 | HEK-293T | C1057(0.71); C180(1.46); C266(1.01); C348(0.81) | LDD1615 | [38] |
| LDCM0412 | CL24 | HEK-293T | C1057(0.69); C266(1.17); C348(0.90); C972(0.91) | LDD1616 | [38] |
| LDCM0413 | CL25 | HEK-293T | C1057(1.37); C180(1.05); C266(1.11); C348(1.30) | LDD1617 | [38] |
| LDCM0414 | CL26 | HEK-293T | C180(0.97); C266(0.89); C348(0.92); C972(0.95) | LDD1618 | [38] |
| LDCM0415 | CL27 | HEK-293T | C1057(0.98); C180(0.98); C266(0.93); C348(0.95) | LDD1619 | [38] |
| LDCM0416 | CL28 | HEK-293T | C1057(0.99); C180(0.95); C266(0.84); C348(1.02) | LDD1620 | [38] |
| LDCM0417 | CL29 | HEK-293T | C1057(0.97); C180(0.96); C266(0.92); C348(0.93) | LDD1621 | [38] |
| LDCM0418 | CL3 | HEK-293T | C1057(1.17); C180(1.04); C266(0.85); C348(1.00) | LDD1622 | [38] |
| LDCM0419 | CL30 | HEK-293T | C1057(0.99); C180(1.08); C266(0.90); C348(0.94) | LDD1623 | [38] |
| LDCM0420 | CL31 | HEK-293T | C1057(1.07); C180(0.93); C266(0.95); C348(1.02) | LDD1624 | [38] |
| LDCM0421 | CL32 | HEK-293T | C1057(0.96); C180(1.00); C266(0.89); C348(0.85) | LDD1625 | [38] |
| LDCM0422 | CL33 | HEK-293T | C1057(2.00); C180(1.01); C266(1.15); C348(2.36) | LDD1626 | [38] |
| LDCM0423 | CL34 | HEK-293T | C1057(0.97); C180(1.13); C266(1.08); C348(1.08) | LDD1627 | [38] |
| LDCM0424 | CL35 | HEK-293T | C1057(0.70); C180(1.33); C266(1.05); C348(0.89) | LDD1628 | [38] |
| LDCM0425 | CL36 | HEK-293T | C1057(0.66); C266(1.21); C348(1.00); C972(0.89) | LDD1629 | [38] |
| LDCM0426 | CL37 | HEK-293T | C1057(1.06); C180(0.99); C266(0.94); C348(1.00) | LDD1630 | [38] |
| LDCM0428 | CL39 | HEK-293T | C1057(1.00); C180(1.10); C266(0.87); C348(0.98) | LDD1632 | [38] |
| LDCM0429 | CL4 | HEK-293T | C1057(1.11); C180(1.01); C266(0.74); C348(1.05) | LDD1633 | [38] |
| LDCM0430 | CL40 | HEK-293T | C1057(1.00); C180(0.95); C266(0.93); C348(0.91) | LDD1634 | [38] |
| LDCM0431 | CL41 | HEK-293T | C1057(1.43); C180(1.02); C266(0.91); C348(1.11) | LDD1635 | [38] |
| LDCM0432 | CL42 | HEK-293T | C1057(1.04); C180(0.96); C266(0.90); C348(1.01) | LDD1636 | [38] |
| LDCM0433 | CL43 | HEK-293T | C1057(1.03); C180(1.12); C266(0.99); C348(0.95) | LDD1637 | [38] |
| LDCM0434 | CL44 | HEK-293T | C1057(0.96); C180(1.03); C266(0.96); C348(0.93) | LDD1638 | [38] |
| LDCM0435 | CL45 | HEK-293T | C1057(0.99); C180(0.96); C266(1.06); C348(1.10) | LDD1639 | [38] |
| LDCM0436 | CL46 | HEK-293T | C1057(0.93); C180(1.18); C266(1.11); C348(1.04) | LDD1640 | [38] |
| LDCM0437 | CL47 | HEK-293T | C1057(0.74); C180(1.30); C266(1.07); C348(0.94) | LDD1641 | [38] |
| LDCM0438 | CL48 | HEK-293T | C1057(0.68); C266(1.15); C348(1.03); C972(0.95) | LDD1642 | [38] |
| LDCM0439 | CL49 | HEK-293T | C1057(0.97); C180(1.01); C266(0.94); C348(0.87) | LDD1643 | [38] |
| LDCM0440 | CL5 | HEK-293T | C1057(1.00); C180(0.99); C266(0.88); C348(0.93) | LDD1644 | [38] |
| LDCM0441 | CL50 | HEK-293T | C180(0.96); C266(0.83); C348(0.98); C972(0.90) | LDD1645 | [38] |
| LDCM0443 | CL52 | HEK-293T | C1057(0.95); C180(1.00); C266(0.73); C348(0.91) | LDD1646 | [38] |
| LDCM0444 | CL53 | HEK-293T | C1057(1.96); C180(1.05); C266(0.87); C348(1.38) | LDD1647 | [38] |
| LDCM0445 | CL54 | HEK-293T | C1057(2.19); C180(0.97); C266(0.98); C348(1.75) | LDD1648 | [38] |
| LDCM0446 | CL55 | HEK-293T | C1057(1.01); C180(1.09); C266(0.98); C348(0.87) | LDD1649 | [38] |
| LDCM0447 | CL56 | HEK-293T | C1057(0.88); C180(1.04); C266(0.95); C348(0.78) | LDD1650 | [38] |
| LDCM0448 | CL57 | HEK-293T | C1057(0.97); C180(0.94); C266(0.91); C348(1.15) | LDD1651 | [38] |
| LDCM0449 | CL58 | HEK-293T | C1057(0.90); C180(1.09); C266(1.01); C348(0.95) | LDD1652 | [38] |
| LDCM0450 | CL59 | HEK-293T | C1057(0.71); C180(1.29); C266(1.11); C348(0.85) | LDD1653 | [38] |
| LDCM0451 | CL6 | HEK-293T | C1057(1.53); C180(1.01); C266(1.02); C348(1.22) | LDD1654 | [38] |
| LDCM0452 | CL60 | HEK-293T | C1057(0.63); C266(1.16); C348(0.97); C972(0.86) | LDD1655 | [38] |
| LDCM0453 | CL61 | HEK-293T | C1057(0.92); C180(1.03); C266(0.99); C348(0.88) | LDD1656 | [38] |
| LDCM0454 | CL62 | HEK-293T | C180(0.94); C266(0.88); C348(0.88); C972(0.93) | LDD1657 | [38] |
| LDCM0455 | CL63 | HEK-293T | C1057(0.94); C180(0.90); C266(0.95); C348(0.88) | LDD1658 | [38] |
| LDCM0456 | CL64 | HEK-293T | C1057(1.37); C180(0.90); C266(0.76); C348(1.19) | LDD1659 | [38] |
| LDCM0457 | CL65 | HEK-293T | C1057(1.00); C180(1.01); C266(0.89); C348(0.92) | LDD1660 | [38] |
| LDCM0458 | CL66 | HEK-293T | C1057(1.02); C180(0.97); C266(0.95); C348(0.96) | LDD1661 | [38] |
| LDCM0459 | CL67 | HEK-293T | C1057(1.01); C180(1.13); C266(0.96); C348(0.91) | LDD1662 | [38] |
| LDCM0460 | CL68 | HEK-293T | C1057(0.88); C180(1.07); C266(0.97); C348(0.90) | LDD1663 | [38] |
| LDCM0461 | CL69 | HEK-293T | C1057(0.95); C180(0.93); C266(1.00); C348(1.02) | LDD1664 | [38] |
| LDCM0462 | CL7 | HEK-293T | C1057(1.06); C180(1.02); C266(0.89); C348(0.95) | LDD1665 | [38] |
| LDCM0463 | CL70 | HEK-293T | C1057(1.00); C180(1.11); C266(1.02); C348(0.97) | LDD1666 | [38] |
| LDCM0464 | CL71 | HEK-293T | C1057(0.76); C180(1.31); C266(1.05); C348(0.85) | LDD1667 | [38] |
| LDCM0465 | CL72 | HEK-293T | C1057(0.72); C266(1.14); C348(0.94); C972(0.94) | LDD1668 | [38] |
| LDCM0466 | CL73 | HEK-293T | C1057(1.08); C180(0.92); C266(0.91); C348(0.99) | LDD1669 | [38] |
| LDCM0467 | CL74 | HEK-293T | C180(0.86); C266(0.85); C348(0.89); C972(0.98) | LDD1670 | [38] |
| LDCM0469 | CL76 | HEK-293T | C1057(0.95); C180(0.90); C266(0.82); C348(0.96) | LDD1672 | [38] |
| LDCM0470 | CL77 | HEK-293T | C1057(1.34); C180(0.98); C266(0.88); C348(1.19) | LDD1673 | [38] |
| LDCM0471 | CL78 | HEK-293T | C1057(0.94); C180(0.97); C266(0.94); C348(0.94) | LDD1674 | [38] |
| LDCM0472 | CL79 | HEK-293T | C1057(1.01); C180(1.01); C266(0.96); C348(0.93) | LDD1675 | [38] |
| LDCM0473 | CL8 | HEK-293T | C1057(10.76); C180(1.46); C266(1.39); C348(7.50) | LDD1676 | [38] |
| LDCM0474 | CL80 | HEK-293T | C1057(0.88); C180(1.02); C266(0.96); C348(0.87) | LDD1677 | [38] |
| LDCM0475 | CL81 | HEK-293T | C1057(0.87); C180(0.98); C266(1.01); C348(0.95) | LDD1678 | [38] |
| LDCM0476 | CL82 | HEK-293T | C1057(0.94); C180(1.12); C266(1.05); C348(0.95) | LDD1679 | [38] |
| LDCM0477 | CL83 | HEK-293T | C1057(0.95); C180(1.28); C266(1.02); C348(0.97) | LDD1680 | [38] |
| LDCM0478 | CL84 | HEK-293T | C1057(1.00); C266(1.12); C348(1.12); C972(0.96) | LDD1681 | [38] |
| LDCM0479 | CL85 | HEK-293T | C1057(0.91); C180(0.92); C266(0.92); C348(0.86) | LDD1682 | [38] |
| LDCM0480 | CL86 | HEK-293T | C180(0.77); C266(0.87); C348(0.92); C972(0.94) | LDD1683 | [38] |
| LDCM0481 | CL87 | HEK-293T | C1057(0.96); C180(0.88); C266(0.89); C348(0.93) | LDD1684 | [38] |
| LDCM0482 | CL88 | HEK-293T | C1057(0.94); C180(0.82); C266(0.78); C348(0.89) | LDD1685 | [38] |
| LDCM0483 | CL89 | HEK-293T | C1057(1.03); C180(0.85); C266(0.92); C348(0.95) | LDD1686 | [38] |
| LDCM0484 | CL9 | HEK-293T | C1057(0.86); C180(1.06); C266(1.00); C348(0.93) | LDD1687 | [38] |
| LDCM0485 | CL90 | HEK-293T | C1057(3.53); C180(0.92); C266(1.14); C348(2.65) | LDD1688 | [38] |
| LDCM0486 | CL91 | HEK-293T | C1057(0.96); C180(0.92); C266(1.01); C348(0.87) | LDD1689 | [38] |
| LDCM0487 | CL92 | HEK-293T | C1057(1.26); C180(0.95); C266(0.92); C348(1.03) | LDD1690 | [38] |
| LDCM0488 | CL93 | HEK-293T | C1057(1.14); C180(0.91); C266(1.00); C348(1.19) | LDD1691 | [38] |
| LDCM0489 | CL94 | HEK-293T | C1057(0.99); C180(1.05); C266(1.02); C348(0.93) | LDD1692 | [38] |
| LDCM0490 | CL95 | HEK-293T | C1057(2.92); C180(1.02); C266(1.24); C348(3.02) | LDD1693 | [38] |
| LDCM0491 | CL96 | HEK-293T | C1057(1.21); C266(1.06); C348(1.14); C972(0.97) | LDD1694 | [38] |
| LDCM0492 | CL97 | HEK-293T | C1057(1.22); C180(1.04); C266(0.95); C348(1.13) | LDD1695 | [38] |
| LDCM0493 | CL98 | HEK-293T | C180(1.00); C266(0.84); C348(1.04); C972(0.95) | LDD1696 | [38] |
| LDCM0494 | CL99 | HEK-293T | C1057(0.95); C180(0.95); C266(0.88); C348(0.94) | LDD1697 | [38] |
| LDCM0634 | CY-0357 | Hep-G2 | C1057(2.22) | LDD2228 | [24] |
| LDCM0495 | E2913 | HEK-293T | C1057(0.84); C180(0.99); C266(0.88); C348(0.84) | LDD1698 | [38] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C733(6.17); C972(4.06); C420(3.78); C110(3.55) | LDD1702 | [37] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 8.89 | LDD0183 | [11] |
| LDCM0175 | Ethacrynic acid | HeLa | C733(0.75) | LDD2210 | [13] |
| LDCM0625 | F8 | Ramos | C1057(1.73); C348(0.63); C110(1.10); C266(1.05) | LDD2187 | [40] |
| LDCM0572 | Fragment10 | Ramos | C1057(0.82); C348(1.40); C110(0.67); C266(0.58) | LDD2189 | [40] |
| LDCM0573 | Fragment11 | Ramos | C1057(0.10); C348(0.04); C110(0.01); C266(3.00) | LDD2190 | [40] |
| LDCM0574 | Fragment12 | Ramos | C1057(0.88); C348(1.54); C110(0.93); C266(0.58) | LDD2191 | [40] |
| LDCM0575 | Fragment13 | Ramos | C1057(0.95); C348(1.10); C110(0.95); C266(0.90) | LDD2192 | [40] |
| LDCM0576 | Fragment14 | Ramos | C1057(1.65); C348(1.07); C110(1.77); C266(1.74) | LDD2193 | [40] |
| LDCM0579 | Fragment20 | Ramos | C1057(1.10); C348(1.17); C110(0.87); C266(0.53) | LDD2194 | [40] |
| LDCM0580 | Fragment21 | Ramos | C1057(0.60); C348(0.63); C110(0.85); C266(0.62) | LDD2195 | [40] |
| LDCM0582 | Fragment23 | Ramos | C1057(1.35); C348(0.82); C110(1.31); C266(1.29) | LDD2196 | [40] |
| LDCM0578 | Fragment27 | Ramos | C1057(1.23); C348(0.81); C110(1.01); C266(0.81) | LDD2197 | [40] |
| LDCM0586 | Fragment28 | Ramos | C1057(0.52); C348(0.78); C110(0.95); C266(0.53) | LDD2198 | [40] |
| LDCM0588 | Fragment30 | Ramos | C1057(0.71); C348(1.12); C110(0.79); C266(0.78) | LDD2199 | [40] |
| LDCM0589 | Fragment31 | Ramos | C1057(1.26); C348(1.05); C110(1.02); C266(0.94) | LDD2200 | [40] |
| LDCM0590 | Fragment32 | Ramos | C1057(1.02); C348(1.26); C110(0.72); C266(0.53) | LDD2201 | [40] |
| LDCM0468 | Fragment33 | HEK-293T | C1057(0.98); C180(0.93); C266(0.90); C348(0.93) | LDD1671 | [38] |
| LDCM0596 | Fragment38 | Ramos | C1057(0.89); C348(1.34); C110(1.05); C266(1.39) | LDD2203 | [40] |
| LDCM0566 | Fragment4 | Ramos | C1057(1.26); C348(1.11); C110(1.63); C266(0.68) | LDD2184 | [40] |
| LDCM0427 | Fragment51 | HEK-293T | C180(0.97); C266(0.87); C348(0.94); C972(1.02) | LDD1631 | [38] |
| LDCM0610 | Fragment52 | Ramos | C1057(0.85); C348(1.30); C110(0.98); C266(0.96) | LDD2204 | [40] |
| LDCM0614 | Fragment56 | Ramos | C1057(1.17); C348(1.30); C110(0.86); C266(1.18) | LDD2205 | [40] |
| LDCM0569 | Fragment7 | Ramos | C1057(1.07); C348(1.39); C110(1.33); C266(0.78) | LDD2186 | [40] |
| LDCM0571 | Fragment9 | Ramos | C1057(0.83); C348(1.22); C110(1.01); C266(0.66) | LDD2188 | [40] |
| LDCM0116 | HHS-0101 | DM93 | Y905(0.82) | LDD0264 | [15] |
| LDCM0117 | HHS-0201 | DM93 | Y905(0.91) | LDD0265 | [15] |
| LDCM0118 | HHS-0301 | DM93 | Y169(0.05); Y905(1.05) | LDD0266 | [15] |
| LDCM0119 | HHS-0401 | DM93 | Y905(0.91) | LDD0267 | [15] |
| LDCM0120 | HHS-0701 | DM93 | Y905(0.91) | LDD0268 | [15] |
| LDCM0015 | HNE | MDA-MB-231 | C972(0.81) | LDD0346 | [40] |
| LDCM0107 | IAA | HeLa | H771(0.00); H540(0.00); C972(0.00); C180(0.00) | LDD0221 | [29] |
| LDCM0179 | JZ128 | PC-3 | C972(0.00); C682(0.00); C473(0.00); C1078(0.00) | LDD0462 | [9] |
| LDCM0022 | KB02 | HEK-293T | C1057(0.73); C348(0.85); C180(1.02); C266(0.96) | LDD1492 | [38] |
| LDCM0023 | KB03 | Jurkat | C473(8.60); C110(4.75) | LDD0209 | [14] |
| LDCM0024 | KB05 | COLO792 | C284(2.88); C491(2.21) | LDD3310 | [41] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C972(1.07); C348(1.39); C180(0.84); C110(1.01) | LDD2102 | [37] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C972(0.68); C733(0.94); C348(0.61); C180(1.15) | LDD2121 | [37] |
| LDCM0109 | NEM | HeLa | H461(0.00); H847(0.00); H540(0.00); H362(0.00) | LDD0223 | [29] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C972(0.81); C733(0.99); C348(0.64); C180(1.12) | LDD2089 | [37] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C972(0.83); C348(0.96); C180(1.07); C110(1.03) | LDD2090 | [37] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C972(1.00); C733(1.16); C348(1.13); C180(1.11) | LDD2092 | [37] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C972(1.01); C733(1.28); C348(1.09); C180(1.27) | LDD2093 | [37] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C972(2.13); C733(1.58); C348(1.57); C180(1.77) | LDD2094 | [37] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C972(0.17); C348(0.83); C180(0.06); C110(0.26) | LDD2096 | [37] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C972(1.05); C733(0.98); C348(0.84); C180(0.95) | LDD2097 | [37] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C972(1.10); C733(0.97); C348(0.72); C180(0.88) | LDD2098 | [37] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C972(0.90); C733(1.12); C348(0.93); C180(1.37) | LDD2099 | [37] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C972(0.56); C733(0.69); C348(0.65); C180(0.52) | LDD2100 | [37] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C972(0.94); C733(0.90); C348(1.22); C180(0.95) | LDD2101 | [37] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C972(0.62); C733(0.58); C348(0.62); C180(0.46) | LDD2104 | [37] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C972(2.71); C733(1.34); C348(1.32); C180(1.48) | LDD2105 | [37] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C972(0.66); C733(0.65); C180(0.42); C110(0.65) | LDD2106 | [37] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C972(0.79); C733(0.86); C348(0.68); C180(0.98) | LDD2107 | [37] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C972(0.67); C733(0.69); C348(0.46); C180(0.54) | LDD2108 | [37] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C972(0.70); C733(0.62); C348(0.56); C180(0.65) | LDD2109 | [37] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C972(0.76); C733(1.10); C180(0.58); C110(1.04) | LDD2110 | [37] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C972(1.04); C733(1.27); C348(0.85); C180(1.25) | LDD2111 | [37] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C733(0.65); C348(1.04); C180(0.58); C266(0.57) | LDD2114 | [37] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C972(0.46); C733(0.47); C348(0.51); C180(0.44) | LDD2115 | [37] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C972(0.73); C180(0.10); C110(0.60); C682(0.55) | LDD2116 | [37] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C972(0.49); C733(0.12); C348(0.32); C110(0.43) | LDD2118 | [37] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C972(2.03); C733(1.93); C348(1.96); C180(1.63) | LDD2119 | [37] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C972(0.96); C348(0.82); C180(0.57); C110(0.90) | LDD2120 | [37] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C733(0.11); C348(0.10); C180(0.12); C110(0.43) | LDD2122 | [37] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C348(0.75); C180(1.08); C266(1.16); C110(0.86) | LDD2123 | [37] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C110(0.44); C682(0.35); C687(0.85) | LDD2124 | [37] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C972(0.83); C733(0.93); C348(0.73); C180(1.03) | LDD2125 | [37] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C972(0.38); C348(0.08); C180(0.08); C110(0.31) | LDD2126 | [37] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C972(0.99); C733(1.20); C348(0.81); C180(1.23) | LDD2127 | [37] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C972(0.78); C348(0.93); C180(0.58); C110(1.47) | LDD2128 | [37] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C972(1.21); C733(1.38); C348(0.87); C180(1.22) | LDD2129 | [37] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C972(0.50); C733(0.54); C348(0.58); C180(0.52) | LDD2133 | [37] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C733(0.52); C180(0.47); C110(0.59); C1057(0.53) | LDD2134 | [37] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C972(0.98); C733(0.89); C348(0.85); C180(1.28) | LDD2135 | [37] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C972(1.09); C733(1.42); C348(0.94); C180(1.49) | LDD2136 | [37] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C972(1.23); C733(0.97); C348(0.72); C180(1.05) | LDD2137 | [37] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C180(3.88); C733(3.41); C972(2.48); C1057(2.28) | LDD1700 | [37] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C972(0.76); C733(1.02); C348(0.65); C180(1.08) | LDD2140 | [37] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C972(0.66); C733(0.61); C348(0.42); C180(0.53) | LDD2141 | [37] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C972(0.84); C348(0.91); C180(0.59); C110(0.98) | LDD2143 | [37] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C972(2.22); C733(2.02); C348(2.15); C180(1.86) | LDD2144 | [37] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C110(13.46) | LDD2145 | [37] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C972(0.86); C733(1.00); C348(0.79); C180(1.07) | LDD2146 | [37] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C972(2.60); C348(2.26); C110(2.77) | LDD2147 | [37] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C972(0.97); C733(0.42); C348(0.40); C180(0.59) | LDD2148 | [37] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C972(0.71); C180(0.07); C110(0.35); C687(0.90) | LDD2149 | [37] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C972(0.62); C733(0.74); C348(0.48); C180(0.84) | LDD2150 | [37] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C972(0.47); C733(0.24); C180(0.11); C110(0.44) | LDD2151 | [37] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C972(1.29); C733(2.08); C180(2.25); C110(2.60) | LDD2153 | [37] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C972(1.42); C110(1.31); C944(1.27) | LDD2206 | [42] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C348(1.13); C266(0.87); C972(0.86); C110(0.78) | LDD2207 | [42] |
| LDCM0131 | RA190 | MM1.R | C687(1.70); C110(1.66); C972(1.65) | LDD0304 | [43] |
| LDCM0003 | Sulforaphane | MCF-7 | 2.04 | LDD0042 | [10] |
| LDCM0021 | THZ1 | HeLa S3 | C733(1.08); C420(1.10); C944(1.02); C1078(0.98) | LDD0460 | [9] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Gamma-aminobutyric acid receptor-associated protein (GABARAP) | ATG8 family | O95166 | |||
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
Other
References
