Details of the Target
General Information of Target
| Target ID | LDTP05238 | |||||
|---|---|---|---|---|---|---|
| Target Name | Solute carrier family 25 member 3 (SLC25A3) | |||||
| Gene Name | SLC25A3 | |||||
| Gene ID | 5250 | |||||
| Synonyms |
PHC; Solute carrier family 25 member 3; Phosphate carrier protein, mitochondrial; Phosphate transport protein; PTP |
|||||
| 3D Structure | ||||||
| Sequence |
MFSSVAHLARANPFNTPHLQLVHDGLGDLRSSSPGPTGQPRRPRNLAAAAVEEQYSCDYG
SGRFFILCGLGGIISCGTTHTALVPLDLVKCRMQVDPQKYKGIFNGFSVTLKEDGVRGLA KGWAPTFLGYSMQGLCKFGFYEVFKVLYSNMLGEENTYLWRTSLYLAASASAEFFADIAL APMEAAKVRIQTQPGYANTLRDAAPKMYKEEGLKAFYKGVAPLWMRQIPYTMMKFACFER TVEALYKFVVPKPRSECSKPEQLVVTFVAGYIAGVFCAIVSHPADSVVSVLNKEKGSSAS LVLKRLGFKGVWKGLFARIIMIGTLTALQWFIYDSVKVYFRLPRPPPPEMPESLKKKLGL TQ |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Mitochondrial carrier (TC 2.A.29) family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Inorganic ion transporter that transports phosphate or copper ions across the mitochondrial inner membrane into the matrix compartment. Mediates proton-coupled symport of phosphate ions necessary for mitochondrial oxidative phosphorylation of ADP to ATP. Transports copper ions probably in the form of anionic copper(I) complexes to maintain mitochondrial matrix copper pool and to supply copper for cytochrome C oxidase complex assembly. May also play a role in regulation of the mitochondrial permeability transition pore (mPTP).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
9.95 | LDD0402 | [2] | |
|
Alkylaryl probe 3 Probe Info |
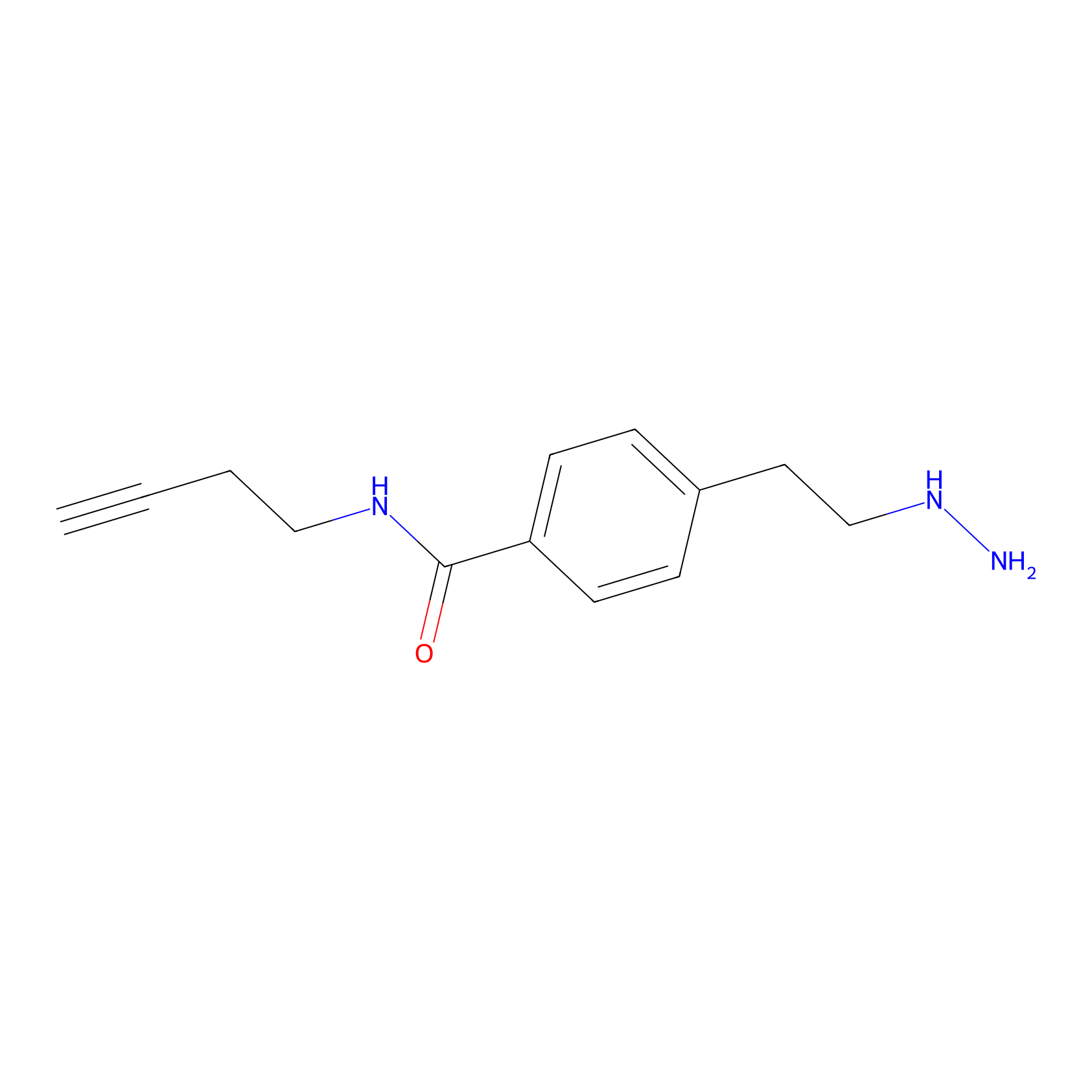 |
6.74 | LDD0381 | [3] | |
|
CHEMBL5175495 Probe Info |
 |
8.08 | LDD0196 | [4] | |
|
CY-1 Probe Info |
 |
22.66 | LDD0243 | [5] | |
|
CY4 Probe Info |
 |
8.24 | LDD0244 | [5] | |
|
N1 Probe Info |
 |
18.79 | LDD0242 | [5] | |
|
W1 Probe Info |
 |
27.57 | LDD0235 | [6] | |
|
TH211 Probe Info |
 |
Y196(20.00); Y208(10.44); Y100(5.42) | LDD0260 | [7] | |
|
TH214 Probe Info |
 |
Y208(11.01) | LDD0258 | [7] | |
|
TH216 Probe Info |
 |
Y208(10.83); Y100(8.39); Y196(6.72) | LDD0259 | [7] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [8] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [8] | |
|
ONAyne Probe Info |
 |
K206(1.41); K209(0.40); K252(0.28); K295(0.98) | LDD0274 | [9] | |
|
BTD Probe Info |
 |
C237(6.08) | LDD1699 | [10] | |
|
OPA-S-S-alkyne Probe Info |
 |
K218(0.82); K234(1.80); K252(3.68); K313(4.20) | LDD3494 | [11] | |
|
Probe 1 Probe Info |
 |
Y196(188.66); Y246(37.16) | LDD3495 | [12] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [13] | |
|
DBIA Probe Info |
 |
C136(1.93) | LDD0080 | [14] | |
|
AMP probe Probe Info |
 |
K99(0.00); K101(0.00) | LDD0200 | [15] | |
|
ATP probe Probe Info |
 |
K99(0.00); K101(0.00); K295(0.00); K304(0.00) | LDD0199 | [15] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [16] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [17] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [18] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [18] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [16] | |
|
Alkyne tyramide Probe Info |
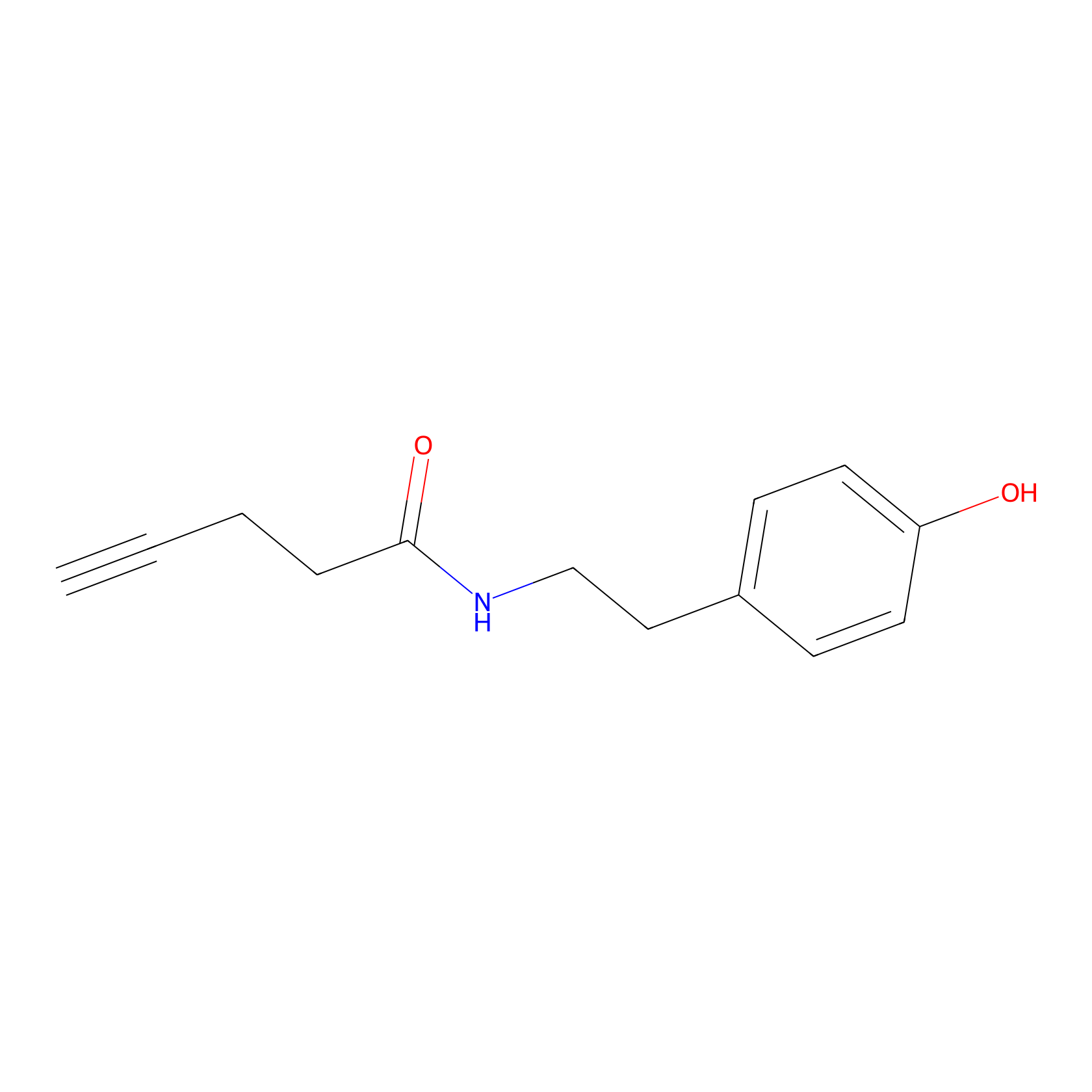 |
Y196(0.00); Y208(0.00) | LDD0003 | [19] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [20] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [21] | |
|
TFBX Probe Info |
 |
C237(0.00); C136(0.00) | LDD0027 | [21] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [19] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [22] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [19] | |
|
IPM Probe Info |
 |
C237(0.00); C136(0.00) | LDD0005 | [19] | |
|
NHS Probe Info |
 |
K218(0.00); K99(0.00); K214(0.00); K304(0.00) | LDD0010 | [19] | |
|
SF Probe Info |
 |
Y141(0.00); Y196(0.00); K355(0.00) | LDD0028 | [23] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [19] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [24] | |
|
1c-yne Probe Info |
 |
K247(0.00); K218(0.00); K101(0.00) | LDD0228 | [25] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [26] | |
|
Cinnamaldehyde Probe Info |
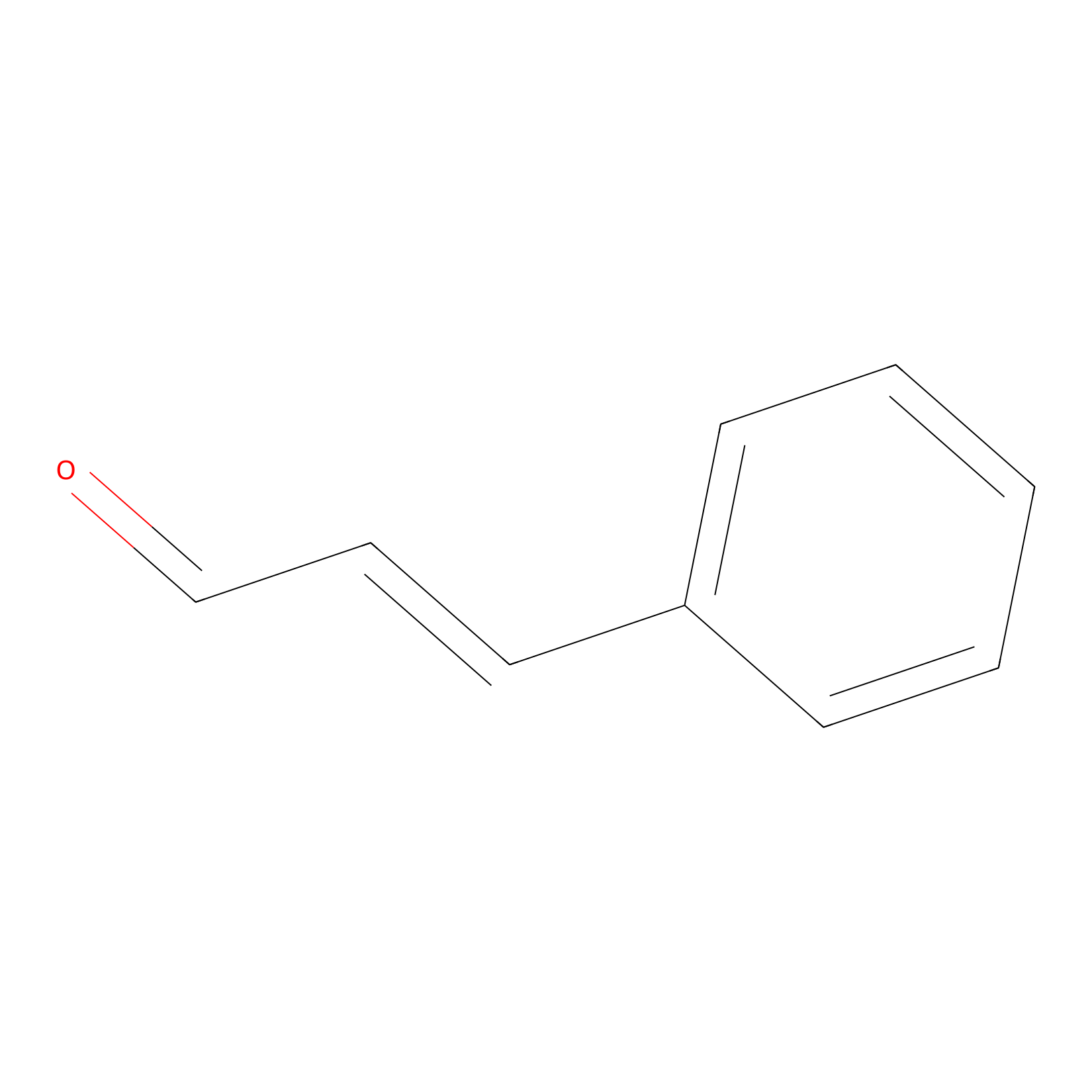 |
N.A. | LDD0220 | [26] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [26] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [26] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [27] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [28] | |
|
NAIA_5 Probe Info |
 |
C91(0.00); C136(0.00); C277(0.00); C257(0.00) | LDD2223 | [29] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [28] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
IMP2070 Probe Info |
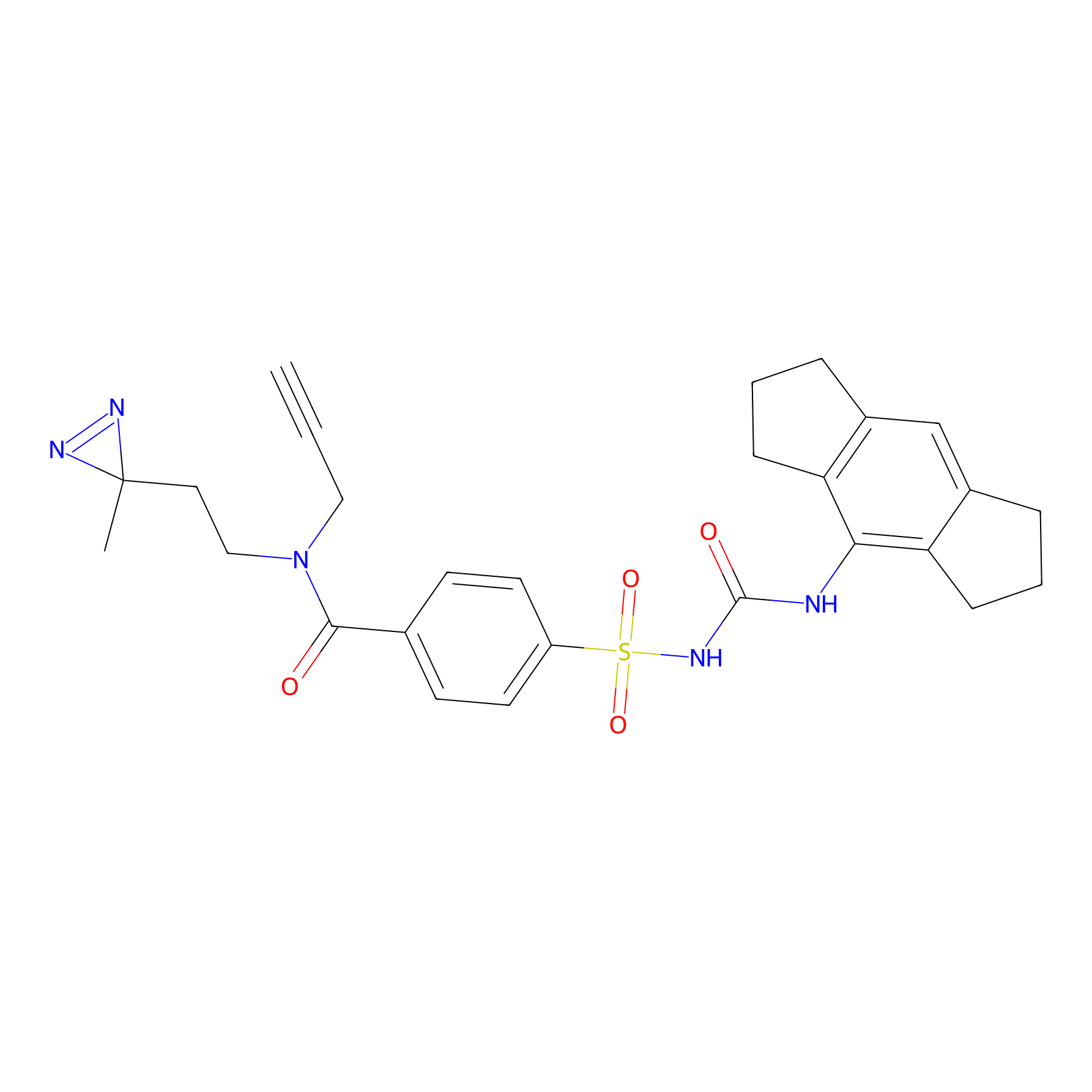 |
1.53 | LDD0367 | [30] | |
|
C004 Probe Info |
 |
6.28 | LDD1714 | [31] | |
|
C196 Probe Info |
 |
11.08 | LDD1872 | [31] | |
|
C220 Probe Info |
 |
15.24 | LDD1894 | [31] | |
|
C362 Probe Info |
 |
30.70 | LDD2023 | [31] | |
|
FFF probe11 Probe Info |
 |
10.90 | LDD0471 | [32] | |
|
FFF probe13 Probe Info |
 |
19.18 | LDD0475 | [32] | |
|
FFF probe14 Probe Info |
 |
13.10 | LDD0477 | [32] | |
|
FFF probe2 Probe Info |
 |
13.50 | LDD0463 | [32] | |
|
FFF probe3 Probe Info |
 |
10.99 | LDD0464 | [32] | |
|
FFF probe4 Probe Info |
 |
12.32 | LDD0466 | [32] | |
|
FFF probe6 Probe Info |
 |
12.94 | LDD0467 | [32] | |
|
JN0003 Probe Info |
 |
15.98 | LDD0469 | [32] | |
|
STS-2 Probe Info |
 |
2.36 | LDD0139 | [33] | |
|
BD-F Probe Info |
 |
T162(0.00); E173(0.00); I332(0.00); S171(0.00) | LDD0024 | [34] | |
|
LD-F Probe Info |
 |
S169(0.00); E173(0.00); M207(0.00); A168(0.00) | LDD0015 | [34] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [35] | |
|
OEA-DA Probe Info |
 |
3.35 | LDD0046 | [36] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C237(0.82) | LDD2142 | [10] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C237(1.09) | LDD2112 | [10] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C237(0.70) | LDD2095 | [10] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C237(0.91) | LDD2130 | [10] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C237(1.25) | LDD2117 | [10] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C237(1.29) | LDD2152 | [10] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C237(0.97) | LDD2103 | [10] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C237(0.65) | LDD2132 | [10] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C237(0.70) | LDD2131 | [10] |
| LDCM0545 | Acetamide | MDA-MB-231 | C237(0.63) | LDD2138 | [10] |
| LDCM0156 | Aniline | NCI-H1299 | 12.14 | LDD0403 | [2] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C237(1.08) | LDD2091 | [10] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [26] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C237(47.55) | LDD1702 | [10] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [13] |
| LDCM0625 | F8 | Ramos | C136(1.24) | LDD2187 | [37] |
| LDCM0572 | Fragment10 | Ramos | C136(1.86) | LDD2189 | [37] |
| LDCM0573 | Fragment11 | Ramos | C136(0.11) | LDD2190 | [37] |
| LDCM0574 | Fragment12 | Ramos | C136(3.30) | LDD2191 | [37] |
| LDCM0575 | Fragment13 | Ramos | C136(0.95) | LDD2192 | [37] |
| LDCM0579 | Fragment20 | Ramos | C136(2.87) | LDD2194 | [37] |
| LDCM0580 | Fragment21 | Ramos | C136(1.01) | LDD2195 | [37] |
| LDCM0582 | Fragment23 | Ramos | C136(0.48) | LDD2196 | [37] |
| LDCM0578 | Fragment27 | Ramos | C136(0.71) | LDD2197 | [37] |
| LDCM0586 | Fragment28 | Ramos | C136(0.73) | LDD2198 | [37] |
| LDCM0588 | Fragment30 | Ramos | C136(1.39) | LDD2199 | [37] |
| LDCM0589 | Fragment31 | Ramos | C136(0.96) | LDD2200 | [37] |
| LDCM0590 | Fragment32 | Ramos | C136(2.15) | LDD2201 | [37] |
| LDCM0468 | Fragment33 | Ramos | C136(1.66) | LDD2202 | [37] |
| LDCM0596 | Fragment38 | Ramos | C136(1.44) | LDD2203 | [37] |
| LDCM0566 | Fragment4 | Ramos | C136(3.25) | LDD2184 | [37] |
| LDCM0610 | Fragment52 | Ramos | C136(2.34) | LDD2204 | [37] |
| LDCM0614 | Fragment56 | Ramos | C136(1.90) | LDD2205 | [37] |
| LDCM0569 | Fragment7 | Ramos | C136(3.00) | LDD2186 | [37] |
| LDCM0571 | Fragment9 | Ramos | C136(4.20) | LDD2188 | [37] |
| LDCM0015 | HNE | MDA-MB-231 | C136(1.14) | LDD0346 | [37] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [26] |
| LDCM0022 | KB02 | HCT 116 | C136(1.93) | LDD0080 | [14] |
| LDCM0023 | KB03 | HCT 116 | C136(2.95) | LDD0081 | [14] |
| LDCM0024 | KB05 | HCT 116 | C136(2.92) | LDD0082 | [14] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C237(1.41) | LDD2102 | [10] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C237(0.75) | LDD2121 | [10] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [26] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C237(0.96) | LDD2089 | [10] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C237(1.59) | LDD2090 | [10] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C237(1.10) | LDD2092 | [10] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C237(1.00) | LDD2093 | [10] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C237(1.48) | LDD2094 | [10] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C237(0.85) | LDD2097 | [10] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C237(1.64) | LDD2098 | [10] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C237(0.97) | LDD2099 | [10] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C237(1.90) | LDD2100 | [10] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C237(0.77) | LDD2101 | [10] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C237(0.87) | LDD2104 | [10] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C237(1.14) | LDD2105 | [10] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C237(1.17) | LDD2106 | [10] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C237(1.07) | LDD2107 | [10] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C237(0.75) | LDD2108 | [10] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C237(0.92) | LDD2109 | [10] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C237(4.02) | LDD2110 | [10] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C237(1.19) | LDD2111 | [10] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C237(1.02) | LDD2114 | [10] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C237(0.57) | LDD2115 | [10] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C237(0.22) | LDD2118 | [10] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C237(1.62) | LDD2119 | [10] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C237(0.95) | LDD2120 | [10] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C237(0.05) | LDD2122 | [10] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C237(1.03) | LDD2123 | [10] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C237(1.04) | LDD2125 | [10] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C237(1.65) | LDD2127 | [10] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C237(1.01) | LDD2128 | [10] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C237(1.06) | LDD2129 | [10] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C237(0.69) | LDD2133 | [10] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C237(0.58) | LDD2134 | [10] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C237(1.25) | LDD2135 | [10] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C237(0.97) | LDD2136 | [10] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C237(0.86) | LDD2137 | [10] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C237(2.14) | LDD1700 | [10] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C237(0.99) | LDD2140 | [10] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C237(0.66) | LDD2141 | [10] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C237(0.79) | LDD2143 | [10] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C237(1.71) | LDD2144 | [10] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C237(8.90) | LDD2145 | [10] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C237(1.12) | LDD2146 | [10] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C237(1.20) | LDD2147 | [10] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C237(0.60) | LDD2148 | [10] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C237(0.66) | LDD2150 | [10] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C237(0.06) | LDD2151 | [10] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C136(1.04); C237(0.89) | LDD2206 | [38] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C237(0.81); C136(0.76) | LDD2207 | [38] |
| LDCM0131 | RA190 | MM1.R | C136(1.64); C237(1.54) | LDD0304 | [39] |
The Interaction Atlas With This Target
References
