Details of the Target
General Information of Target
| Target ID | LDTP05228 | |||||
|---|---|---|---|---|---|---|
| Target Name | Calcium-transporting ATPase type 2C member 1 (ATP2C1) | |||||
| Gene Name | ATP2C1 | |||||
| Gene ID | 27032 | |||||
| Synonyms |
KIAA1347; PMR1L; Calcium-transporting ATPase type 2C member 1; ATPase 2C1; EC 7.2.2.10; ATP-dependent Ca(2+) pump PMR1; Ca(2+)/Mn(2+)-ATPase 2C1; Secretory pathway Ca(2+)-transporting ATPase type 1; SPCA1
|
|||||
| 3D Structure | ||||||
| Sequence |
MKVARFQKIPNGENETMIPVLTSKKASELPVSEVASILQADLQNGLNKCEVSHRRAFHGW
NEFDISEDEPLWKKYISQFKNPLIMLLLASAVISVLMHQFDDAVSITVAILIVVTVAFVQ EYRSEKSLEELSKLVPPECHCVREGKLEHTLARDLVPGDTVCLSVGDRVPADLRLFEAVD LSIDESSLTGETTPCSKVTAPQPAATNGDLASRSNIAFMGTLVRCGKAKGVVIGTGENSE FGEVFKMMQAEEAPKTPLQKSMDLLGKQLSFYSFGIIGIIMLVGWLLGKDILEMFTISVS LAVAAIPEGLPIVVTVTLALGVMRMVKKRAIVKKLPIVETLGCCNVICSDKTGTLTKNEM TVTHIFTSDGLHAEVTGVGYNQFGEVIVDGDVVHGFYNPAVSRIVEAGCVCNDAVIRNNT LMGKPTEGALIALAMKMGLDGLQQDYIRKAEYPFSSEQKWMAVKCVHRTQQDRPEICFMK GAYEQVIKYCTTYQSKGQTLTLTQQQRDVYQQEKARMGSAGLRVLALASGPELGQLTFLG LVGIIDPPRTGVKEAVTTLIASGVSIKMITGDSQETAVAIASRLGLYSKTSQSVSGEEID AMDVQQLSQIVPKVAVFYRASPRHKMKIIKSLQKNGSVVAMTGDGVNDAVALKAADIGVA MGQTGTDVCKEAADMILVDDDFQTIMSAIEEGKGIYNNIKNFVRFQLSTSIAALTLISLA TLMNFPNPLNAMQILWINIIMDGPPAQSLGVEPVDKDVIRKPPRNWKDSILTKNLILKIL VSSIIIVCGTLFVFWRELRDNVITPRDTTMTFTCFVFFDMFNALSSRSQTKSVFEIGLCS NRMFCYAVLGSIMGQLLVIYFPPLQKVFQTESLSILDLLFLLGLTSSVCIVAEIIKKVER SREKIQKHVSSTSSSFLEV |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Cation transport ATPase (P-type) (TC 3.A.3) family, Type IIA subfamily
|
|||||
| Subcellular location |
Golgi apparatus, trans-Golgi network membrane
|
|||||
| Function |
ATP-driven pump that supplies the Golgi apparatus with Ca(2+) and Mn(2+) ions, both essential cofactors for processing and trafficking of newly synthesized proteins in the secretory pathway. Within a catalytic cycle, acquires Ca(2+) or Mn(2+) ions on the cytoplasmic side of the membrane and delivers them to the lumenal side. The transfer of ions across the membrane is coupled to ATP hydrolysis and is associated with a transient phosphorylation that shifts the pump conformation from inward-facing to outward-facing state. Plays a primary role in the maintenance of Ca(2+) homeostasis in the trans-Golgi compartment with a functional impact on Golgi and post-Golgi protein sorting as well as a structural impact on cisternae morphology. Responsible for loading the Golgi stores with Ca(2+) ions in keratinocytes, contributing to keratinocyte differentiation and epidermis integrity. Participates in Ca(2+) and Mn(2+) ions uptake into the Golgi store of hippocampal neurons and regulates protein trafficking required for neural polarity. May also play a role in the maintenance of Ca(2+) and Mn(2+) homeostasis and signaling in the cytosol while preventing cytotoxicity.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A498 | SNV: p.K831T | . | |||
| CCK81 | Deletion: p.A26QfsTer21 | DBIA Probe Info | |||
| DU145 | SNV: p.A579T | DBIA Probe Info | |||
| EFO27 | SNV: p.A892T | DBIA Probe Info | |||
| IPC298 | SNV: p.M741R | DBIA Probe Info | |||
| KYM1 | SNV: p.E240Ter | DBIA Probe Info | |||
| LNCaP clone FGC | SNV: p.S298I | DBIA Probe Info | |||
| MDAMB231 | SNV: p.S588F | . | |||
| MDAMB453 | SNV: p.E598K | DBIA Probe Info | |||
| NALM6 | Insertion: p.D877_L878insK | . | |||
| RPMI7951 | SNV: p.A450S | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
HDSF-alk Probe Info |
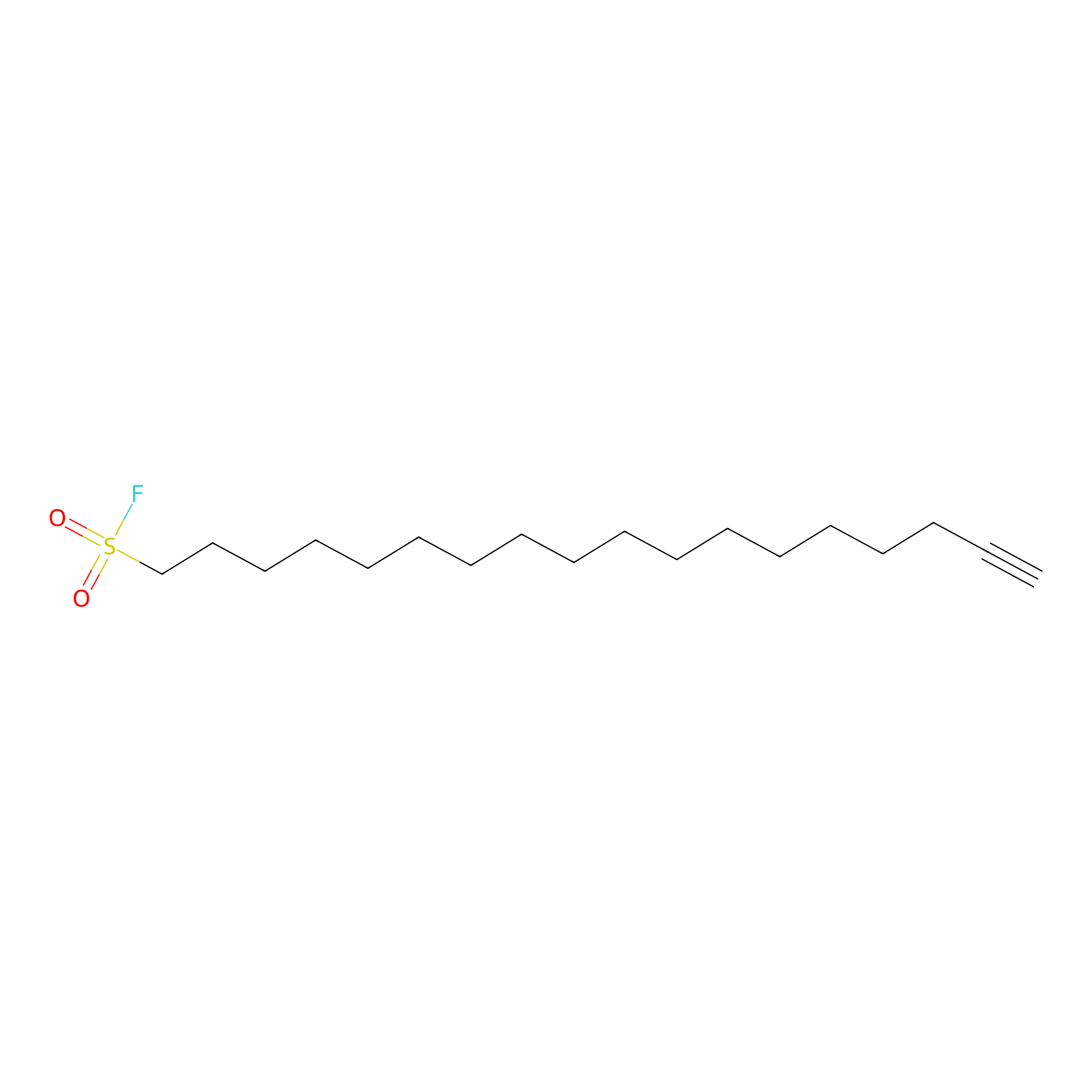 |
1.66 | LDD0197 | [2] | |
|
FBPP2 Probe Info |
 |
2.73 | LDD0318 | [3] | |
|
TG42 Probe Info |
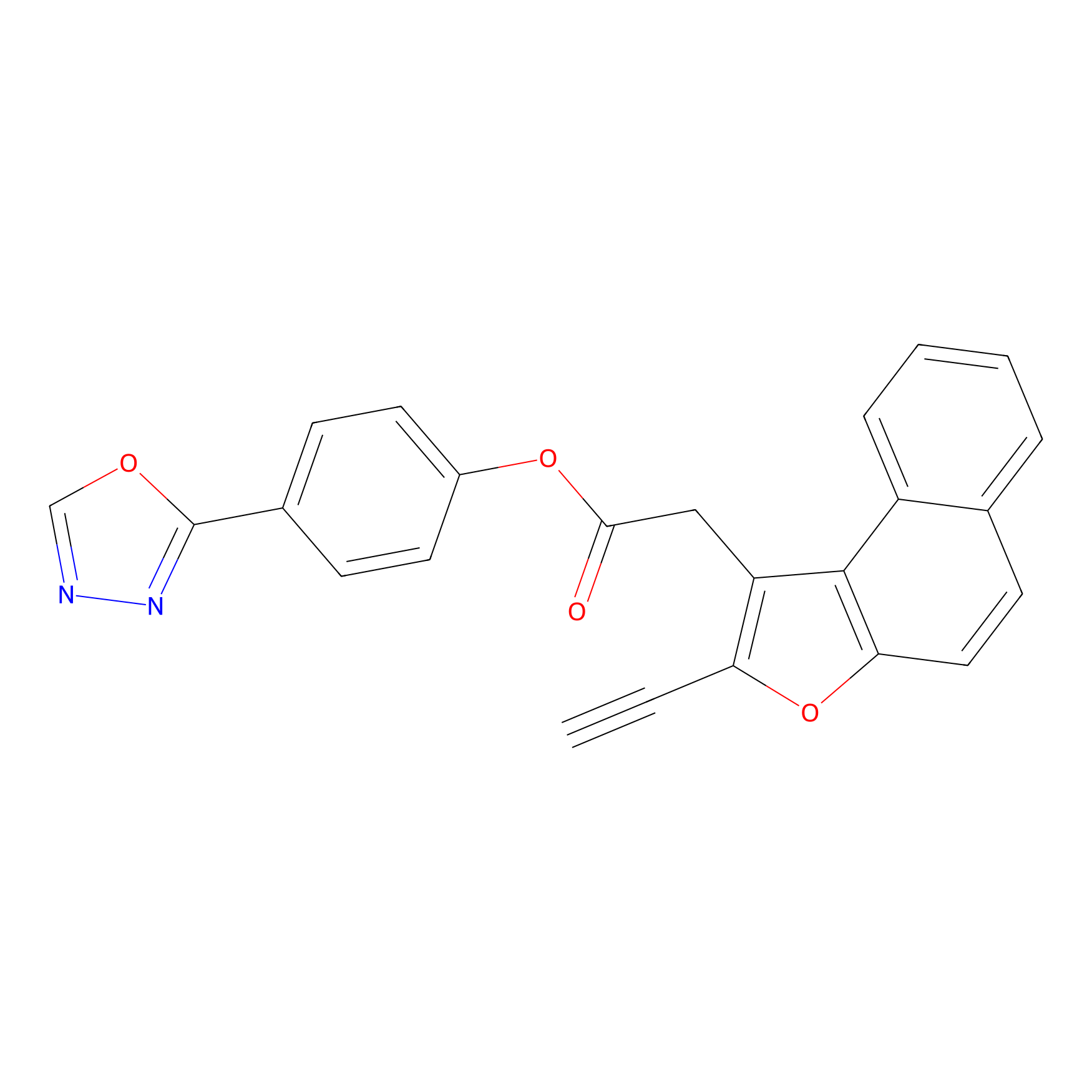 |
4.97 | LDD0326 | [4] | |
|
BTD Probe Info |
 |
C141(0.41) | LDD2104 | [5] | |
|
DA-P3 Probe Info |
 |
13.42 | LDD0179 | [6] | |
|
DBIA Probe Info |
 |
C49(70.64) | LDD0209 | [7] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [8] | |
|
Lodoacetamide azide Probe Info |
 |
C669(0.00); C477(0.00); C490(0.00); C162(0.00) | LDD0037 | [8] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [9] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [10] | |
|
IPM Probe Info |
 |
C49(0.00); C162(0.00) | LDD0147 | [9] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [10] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [11] | |
|
Methacrolein Probe Info |
 |
C669(0.00); C49(0.00) | LDD0218 | [11] | |
|
AOyne Probe Info |
 |
10.90 | LDD0443 | [12] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
8.11 | LDD1740 | [14] | |
|
C056 Probe Info |
 |
17.27 | LDD1753 | [14] | |
|
C091 Probe Info |
 |
11.55 | LDD1782 | [14] | |
|
C092 Probe Info |
 |
19.16 | LDD1783 | [14] | |
|
C134 Probe Info |
 |
24.93 | LDD1816 | [14] | |
|
C135 Probe Info |
 |
12.13 | LDD1817 | [14] | |
|
C177 Probe Info |
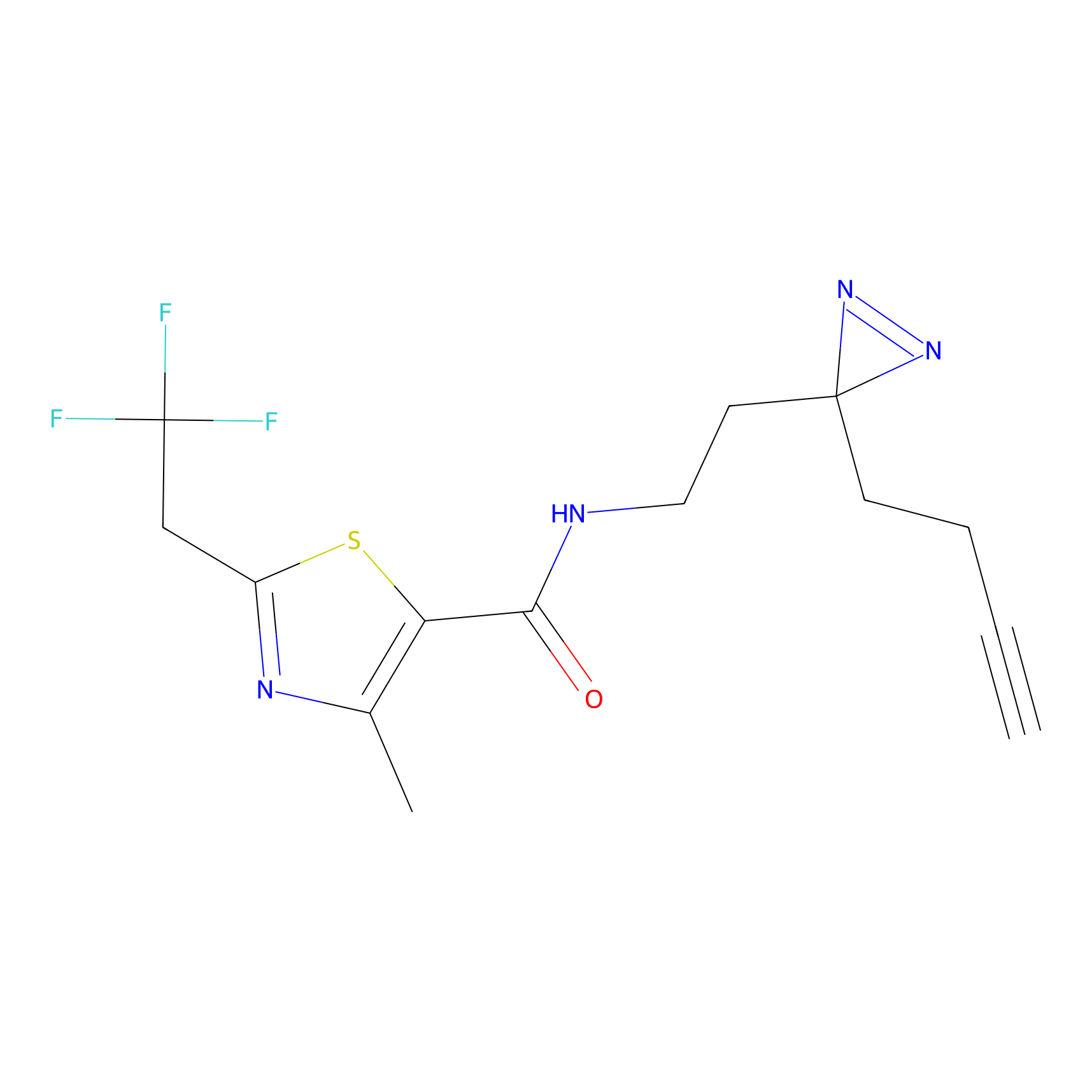 |
11.24 | LDD1856 | [14] | |
|
C201 Probe Info |
 |
29.04 | LDD1877 | [14] | |
|
C210 Probe Info |
 |
43.11 | LDD1884 | [14] | |
|
C226 Probe Info |
 |
6.02 | LDD1899 | [14] | |
|
C228 Probe Info |
 |
16.11 | LDD1901 | [14] | |
|
C264 Probe Info |
 |
18.25 | LDD1935 | [14] | |
|
C287 Probe Info |
 |
11.55 | LDD1957 | [14] | |
|
C289 Probe Info |
 |
36.25 | LDD1959 | [14] | |
|
C355 Probe Info |
 |
32.00 | LDD2016 | [14] | |
|
C356 Probe Info |
 |
9.32 | LDD2017 | [14] | |
|
C362 Probe Info |
 |
26.54 | LDD2023 | [14] | |
|
C390 Probe Info |
 |
25.63 | LDD2049 | [14] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [15] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [15] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [16] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C141(0.72) | LDD2112 | [5] |
| LDCM0156 | Aniline | NCI-H1299 | 12.17 | LDD0403 | [1] |
| LDCM0087 | Capsaicin | HEK-293T | 4.20 | LDD0185 | [6] |
| LDCM0028 | Dobutamine | HEK-293T | 4.05 | LDD0180 | [6] |
| LDCM0027 | Dopamine | HEK-293T | 13.42 | LDD0179 | [6] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C49(7.19); C839(1.02); C141(0.95) | LDD1702 | [5] |
| LDCM0022 | KB02 | HEK-293T | C490(0.98); C669(0.89); C162(1.08); C839(0.86) | LDD1492 | [17] |
| LDCM0023 | KB03 | Jurkat | C49(70.64) | LDD0209 | [7] |
| LDCM0024 | KB05 | HMCB | C703(3.21); C196(1.13) | LDD3312 | [18] |
| LDCM0030 | Luteolin | HEK-293T | 11.27 | LDD0182 | [6] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C141(0.41) | LDD2104 | [5] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C477(0.81); C141(0.97) | LDD2108 | [5] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C141(0.44) | LDD2110 | [5] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C477(0.93); C141(0.33) | LDD2114 | [5] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C477(1.09) | LDD2120 | [5] |
The Interaction Atlas With This Target
The Drug(s) Related To This Target
Approved
Investigative
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Calcium | . | DB01373 | |||
References
