Details of the Target
General Information of Target
| Target ID | LDTP04284 | |||||
|---|---|---|---|---|---|---|
| Target Name | Proteasome subunit beta type-3 (PSMB3) | |||||
| Gene Name | PSMB3 | |||||
| Gene ID | 5691 | |||||
| Synonyms |
Proteasome subunit beta type-3; Proteasome chain 13; Proteasome component C10-II; Proteasome theta chain |
|||||
| 3D Structure | ||||||
| Sequence |
MSIMSYNGGAVMAMKGKNCVAIAADRRFGIQAQMVTTDFQKIFPMGDRLYIGLAGLATDV
QTVAQRLKFRLNLYELKEGRQIKPYTLMSMVANLLYEKRFGPYYTEPVIAGLDPKTFKPF ICSLDLIGCPMVTDDFVVSGTCAEQMYGMCESLWEPNMDPDHLFETISQAMLNAVDRDAV SGMGVIVHIIEKDKITTRTLKARMD |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Peptidase T1B family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Non-catalytic component of the 20S core proteasome complex involved in the proteolytic degradation of most intracellular proteins. This complex plays numerous essential roles within the cell by associating with different regulatory particles. Associated with two 19S regulatory particles, forms the 26S proteasome and thus participates in the ATP-dependent degradation of ubiquitinated proteins. The 26S proteasome plays a key role in the maintenance of protein homeostasis by removing misfolded or damaged proteins that could impair cellular functions, and by removing proteins whose functions are no longer required. Associated with the PA200 or PA28, the 20S proteasome mediates ubiquitin-independent protein degradation. This type of proteolysis is required in several pathways including spermatogenesis (20S-PA200 complex) or generation of a subset of MHC class I-presented antigenic peptides (20S-PA28 complex).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
6.11 | LDD0402 | [1] | |
|
TH211 Probe Info |
 |
Y104(12.39) | LDD0257 | [2] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [3] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [4] | |
|
OPA-S-S-alkyne Probe Info |
 |
K77(2.88) | LDD3494 | [5] | |
|
Probe 1 Probe Info |
 |
Y104(9.31) | LDD3495 | [6] | |
|
BTD Probe Info |
 |
C19(1.86) | LDD1700 | [7] | |
|
DBIA Probe Info |
 |
C19(4.08) | LDD0204 | [8] | |
|
HHS-475 Probe Info |
 |
Y104(0.82) | LDD0264 | [9] | |
|
HHS-465 Probe Info |
 |
Y104(6.70) | LDD2237 | [10] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [11] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [12] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [12] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [13] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [13] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [12] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [14] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [15] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [14] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [16] | |
|
SF Probe Info |
 |
K77(0.00); Y104(0.00) | LDD0028 | [17] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [16] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [18] | |
|
AOyne Probe Info |
 |
11.70 | LDD0443 | [19] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [15] | |
|
HHS-482 Probe Info |
 |
Y104(1.26) | LDD2239 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C108 Probe Info |
 |
9.51 | LDD1795 | [20] | |
|
C178 Probe Info |
 |
29.45 | LDD1857 | [20] | |
|
C226 Probe Info |
 |
10.63 | LDD1899 | [20] | |
|
C252 Probe Info |
 |
9.00 | LDD1925 | [20] | |
|
C277 Probe Info |
 |
7.57 | LDD1947 | [20] | |
|
C282 Probe Info |
 |
16.45 | LDD1952 | [20] | |
|
C387 Probe Info |
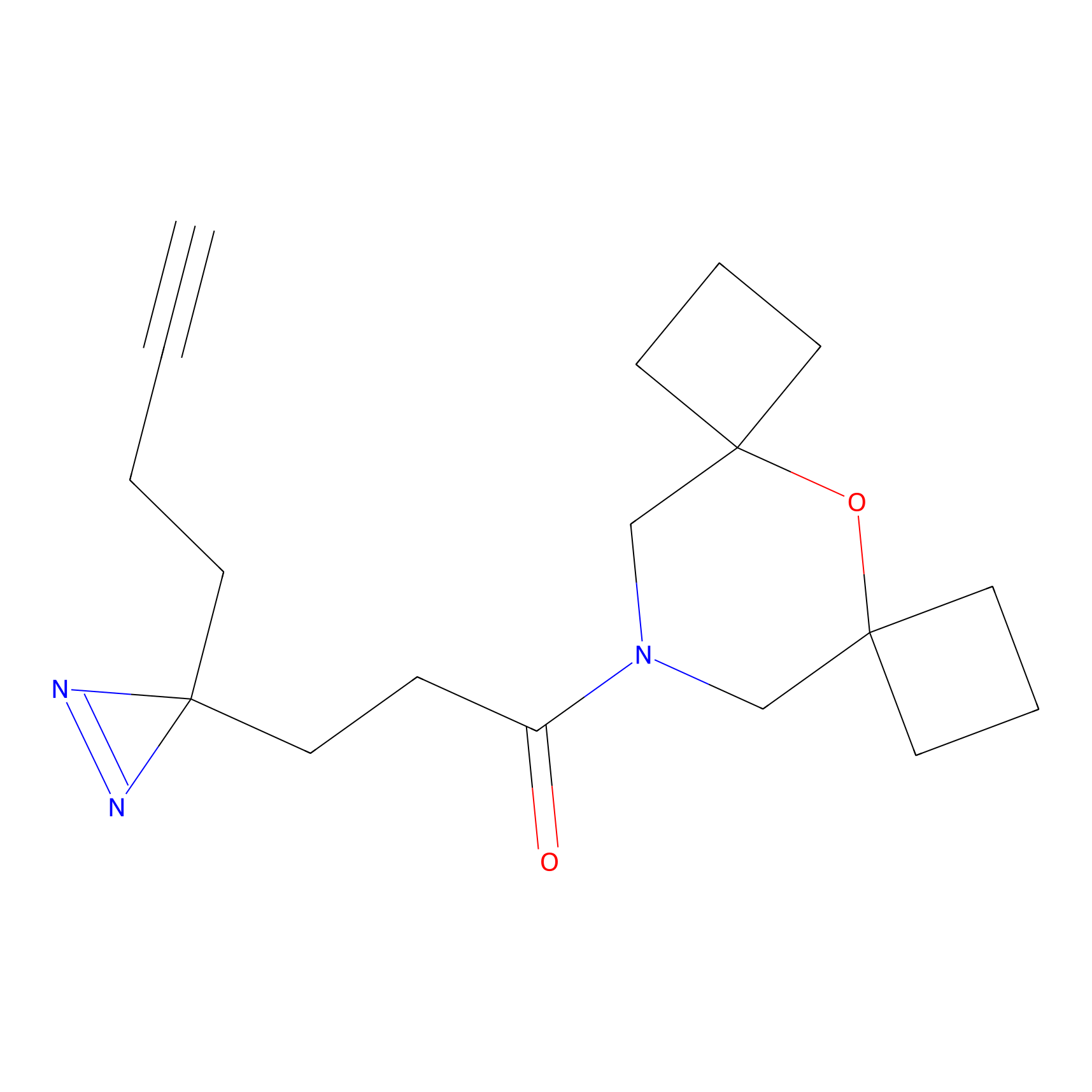 |
5.70 | LDD2046 | [20] | |
|
C431 Probe Info |
 |
28.64 | LDD2086 | [20] | |
|
FFF probe12 Probe Info |
 |
5.25 | LDD0473 | [21] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [21] | |
|
FFF probe14 Probe Info |
 |
12.09 | LDD0477 | [21] | |
|
FFF probe15 Probe Info |
 |
6.60 | LDD0478 | [21] | |
|
FFF probe2 Probe Info |
 |
12.84 | LDD0463 | [21] | |
|
FFF probe3 Probe Info |
 |
9.42 | LDD0465 | [21] | |
|
FFF probe6 Probe Info |
 |
12.40 | LDD0467 | [21] | |
|
Diazir Probe Info |
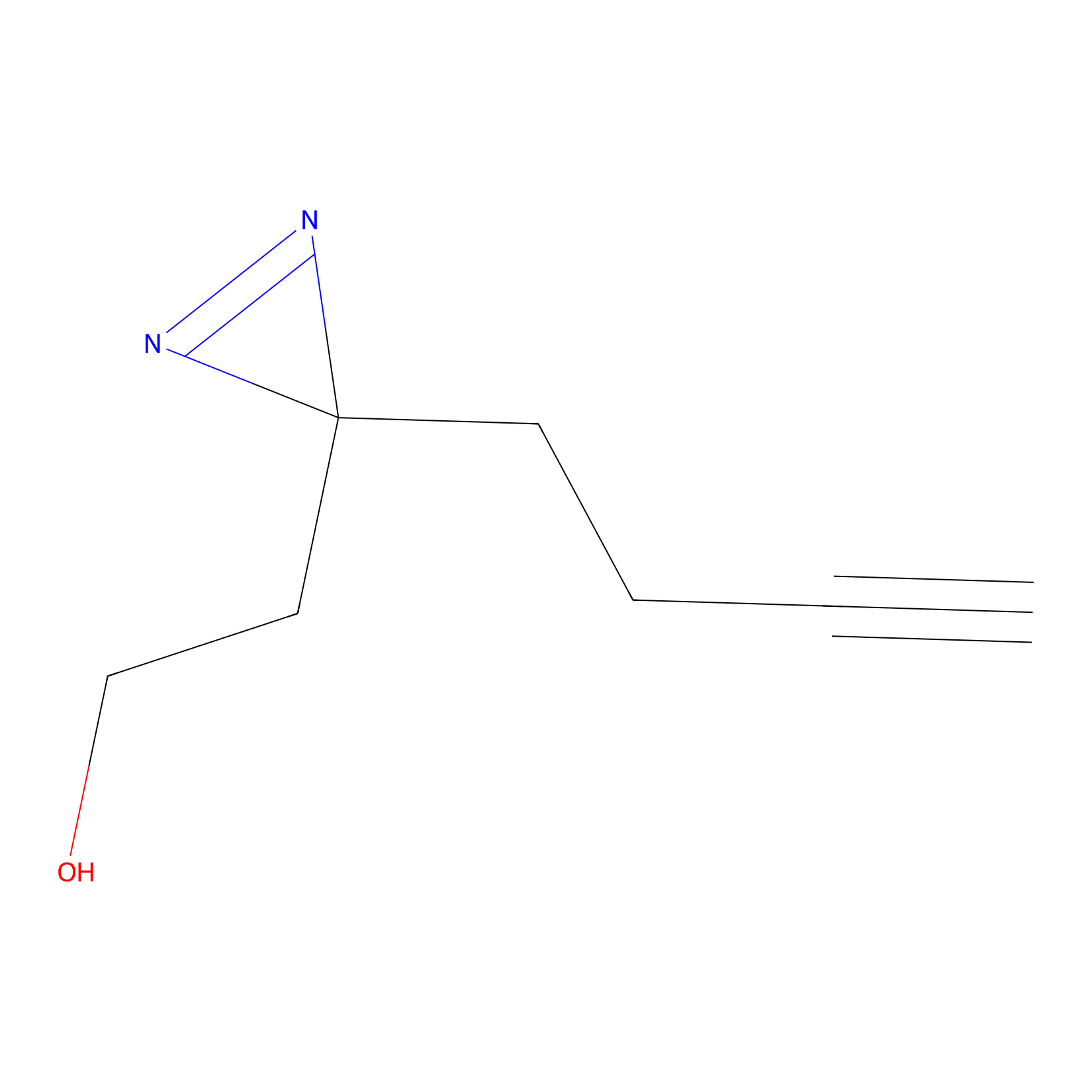 |
N.A. | LDD0011 | [16] | |
|
Photonaproxen Probe Info |
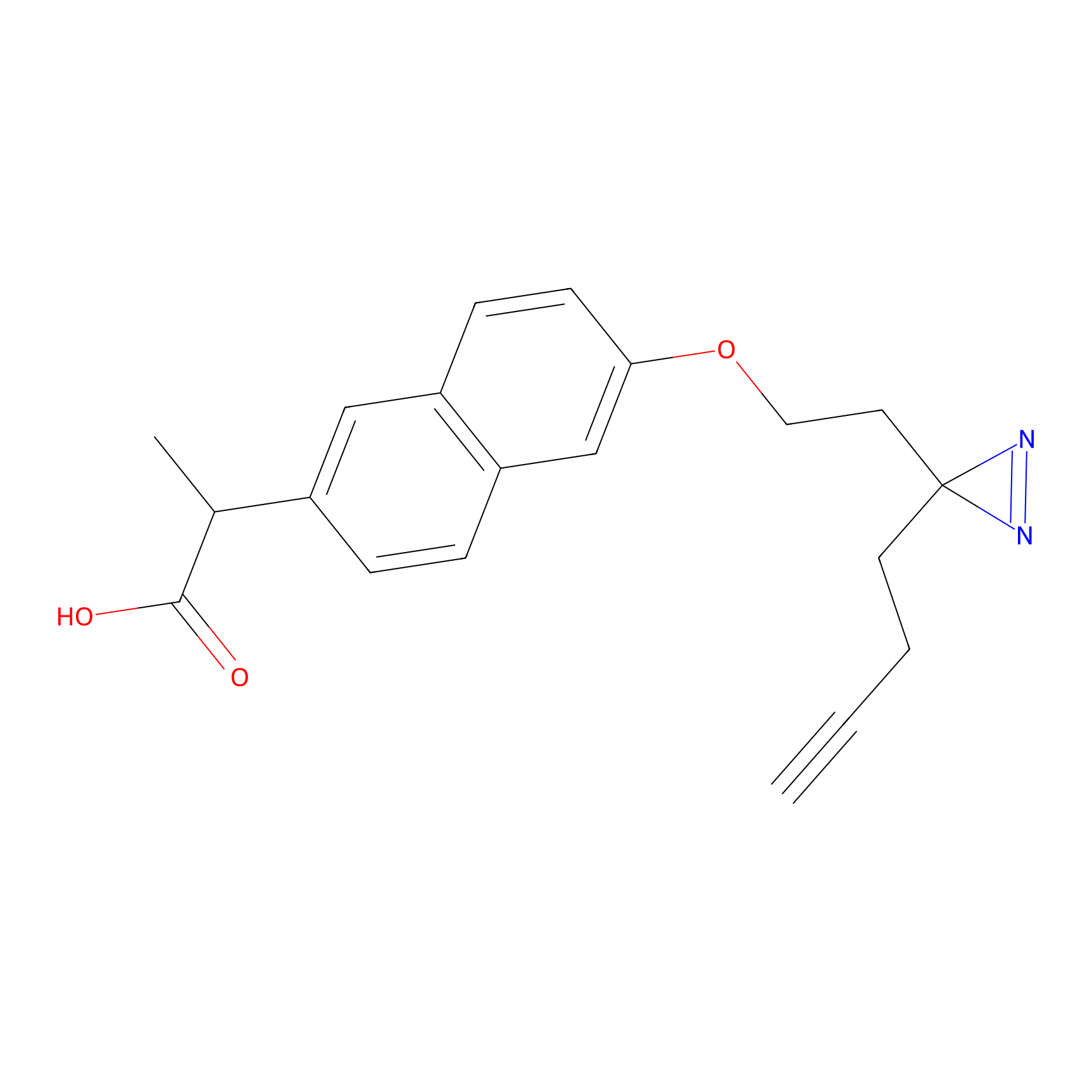 |
E106(0.00); Y104(0.00) | LDD0157 | [22] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [23] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [24] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C19(0.36) | LDD2142 | [7] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C19(0.43) | LDD2112 | [7] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C19(0.54) | LDD2095 | [7] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C19(0.80) | LDD2130 | [7] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C19(0.85) | LDD2117 | [7] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C19(1.18) | LDD2152 | [7] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C19(0.68) | LDD2132 | [7] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C19(0.45) | LDD2131 | [7] |
| LDCM0237 | AC12 | HEK-293T | C19(1.05) | LDD1510 | [25] |
| LDCM0259 | AC14 | HEK-293T | C19(0.89) | LDD1512 | [25] |
| LDCM0270 | AC15 | HEK-293T | C19(0.95) | LDD1513 | [25] |
| LDCM0280 | AC20 | HEK-293T | C19(0.95) | LDD1519 | [25] |
| LDCM0282 | AC22 | HEK-293T | C19(1.04) | LDD1521 | [25] |
| LDCM0283 | AC23 | HEK-293T | C19(0.99) | LDD1522 | [25] |
| LDCM0284 | AC24 | HEK-293T | C19(1.06) | LDD1523 | [25] |
| LDCM0285 | AC25 | HCT 116 | C19(1.05) | LDD0602 | [26] |
| LDCM0286 | AC26 | HCT 116 | C19(1.13) | LDD0603 | [26] |
| LDCM0287 | AC27 | HCT 116 | C19(1.02) | LDD0604 | [26] |
| LDCM0288 | AC28 | HCT 116 | C19(1.01) | LDD0605 | [26] |
| LDCM0289 | AC29 | HCT 116 | C19(1.27) | LDD0606 | [26] |
| LDCM0291 | AC30 | HCT 116 | C19(1.23) | LDD0608 | [26] |
| LDCM0292 | AC31 | HCT 116 | C19(1.20) | LDD0609 | [26] |
| LDCM0293 | AC32 | HCT 116 | C19(1.29) | LDD0610 | [26] |
| LDCM0294 | AC33 | HCT 116 | C19(1.06) | LDD0611 | [26] |
| LDCM0295 | AC34 | HCT 116 | C19(1.07) | LDD0612 | [26] |
| LDCM0296 | AC35 | HCT 116 | C19(0.93) | LDD0613 | [26] |
| LDCM0297 | AC36 | HCT 116 | C19(0.89) | LDD0614 | [26] |
| LDCM0298 | AC37 | HCT 116 | C19(0.91) | LDD0615 | [26] |
| LDCM0299 | AC38 | HCT 116 | C19(1.02) | LDD0616 | [26] |
| LDCM0300 | AC39 | HCT 116 | C19(0.97) | LDD0617 | [26] |
| LDCM0301 | AC4 | HEK-293T | C19(1.24) | LDD1540 | [25] |
| LDCM0302 | AC40 | HCT 116 | C19(1.01) | LDD0619 | [26] |
| LDCM0303 | AC41 | HCT 116 | C19(1.01) | LDD0620 | [26] |
| LDCM0304 | AC42 | HCT 116 | C19(1.03) | LDD0621 | [26] |
| LDCM0305 | AC43 | HCT 116 | C19(1.38) | LDD0622 | [26] |
| LDCM0306 | AC44 | HCT 116 | C19(1.06) | LDD0623 | [26] |
| LDCM0307 | AC45 | HCT 116 | C19(0.92) | LDD0624 | [26] |
| LDCM0308 | AC46 | HCT 116 | C19(0.83) | LDD0625 | [26] |
| LDCM0309 | AC47 | HCT 116 | C19(0.91) | LDD0626 | [26] |
| LDCM0310 | AC48 | HCT 116 | C19(1.20) | LDD0627 | [26] |
| LDCM0311 | AC49 | HCT 116 | C19(0.93) | LDD0628 | [26] |
| LDCM0313 | AC50 | HCT 116 | C19(1.05) | LDD0630 | [26] |
| LDCM0314 | AC51 | HCT 116 | C19(0.99) | LDD0631 | [26] |
| LDCM0315 | AC52 | HCT 116 | C19(0.94) | LDD0632 | [26] |
| LDCM0316 | AC53 | HCT 116 | C19(1.06) | LDD0633 | [26] |
| LDCM0317 | AC54 | HCT 116 | C19(0.98) | LDD0634 | [26] |
| LDCM0318 | AC55 | HCT 116 | C19(0.79) | LDD0635 | [26] |
| LDCM0319 | AC56 | HCT 116 | C19(0.99) | LDD0636 | [26] |
| LDCM0323 | AC6 | HEK-293T | C19(1.07) | LDD1562 | [25] |
| LDCM0324 | AC60 | HEK-293T | C19(1.07) | LDD1563 | [25] |
| LDCM0326 | AC62 | HEK-293T | C19(1.20) | LDD1565 | [25] |
| LDCM0327 | AC63 | HEK-293T | C19(0.99) | LDD1566 | [25] |
| LDCM0328 | AC64 | HEK-293T | C19(1.00) | LDD1567 | [25] |
| LDCM0334 | AC7 | HEK-293T | C19(0.98) | LDD1568 | [25] |
| LDCM0345 | AC8 | HEK-293T | C19(0.98) | LDD1569 | [25] |
| LDCM0545 | Acetamide | MDA-MB-231 | C19(1.01) | LDD2138 | [7] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C19(0.50) | LDD2113 | [7] |
| LDCM0275 | AKOS034007705 | HEK-293T | C19(1.11) | LDD1514 | [25] |
| LDCM0020 | ARS-1620 | HCC44 | C19(1.23) | LDD2171 | [26] |
| LDCM0103 | BDHI 10 | Jurkat | C19(4.03) | LDD0205 | [8] |
| LDCM0102 | BDHI 8 | Jurkat | C19(4.08) | LDD0204 | [8] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C19(0.50) | LDD2091 | [7] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [18] |
| LDCM0367 | CL1 | HEK-293T | C19(0.96) | LDD1571 | [25] |
| LDCM0368 | CL10 | HEK-293T | C19(0.56) | LDD1572 | [25] |
| LDCM0370 | CL101 | HEK-293T | C19(1.06) | LDD1574 | [25] |
| LDCM0374 | CL105 | HEK-293T | C19(1.13) | LDD1578 | [25] |
| LDCM0378 | CL109 | HEK-293T | C19(1.09) | LDD1582 | [25] |
| LDCM0379 | CL11 | HEK-293T | C19(0.57) | LDD1583 | [25] |
| LDCM0382 | CL112 | HCT 116 | C19(1.10) | LDD0699 | [26] |
| LDCM0383 | CL113 | HCT 116 | C19(1.17) | LDD0700 | [26] |
| LDCM0384 | CL114 | HCT 116 | C19(1.30) | LDD0701 | [26] |
| LDCM0385 | CL115 | HCT 116 | C19(1.03) | LDD0702 | [26] |
| LDCM0386 | CL116 | HCT 116 | C19(1.04) | LDD0703 | [26] |
| LDCM0387 | CL117 | HCT 116 | C19(1.04) | LDD0704 | [26] |
| LDCM0388 | CL118 | HCT 116 | C19(1.14) | LDD0705 | [26] |
| LDCM0389 | CL119 | HCT 116 | C19(1.30) | LDD0706 | [26] |
| LDCM0390 | CL12 | HEK-293T | C19(1.03) | LDD1594 | [25] |
| LDCM0391 | CL120 | HCT 116 | C19(1.25) | LDD0708 | [26] |
| LDCM0392 | CL121 | HCT 116 | C19(0.92) | LDD0709 | [26] |
| LDCM0393 | CL122 | HCT 116 | C19(0.77) | LDD0710 | [26] |
| LDCM0394 | CL123 | HCT 116 | C19(0.73) | LDD0711 | [26] |
| LDCM0395 | CL124 | HCT 116 | C19(0.82) | LDD0712 | [26] |
| LDCM0396 | CL125 | HEK-293T | C19(0.96) | LDD1600 | [25] |
| LDCM0400 | CL13 | HEK-293T | C19(0.88) | LDD1604 | [25] |
| LDCM0408 | CL20 | HEK-293T | C19(0.84) | LDD1612 | [25] |
| LDCM0410 | CL22 | HEK-293T | C19(0.67) | LDD1614 | [25] |
| LDCM0411 | CL23 | HEK-293T | C19(0.71) | LDD1615 | [25] |
| LDCM0412 | CL24 | HEK-293T | C19(1.18) | LDD1616 | [25] |
| LDCM0413 | CL25 | HEK-293T | C19(1.08) | LDD1617 | [25] |
| LDCM0421 | CL32 | HEK-293T | C19(0.96) | LDD1625 | [25] |
| LDCM0423 | CL34 | HEK-293T | C19(0.78) | LDD1627 | [25] |
| LDCM0424 | CL35 | HEK-293T | C19(0.64) | LDD1628 | [25] |
| LDCM0425 | CL36 | HEK-293T | C19(1.21) | LDD1629 | [25] |
| LDCM0426 | CL37 | HEK-293T | C19(0.98) | LDD1630 | [25] |
| LDCM0434 | CL44 | HEK-293T | C19(0.96) | LDD1638 | [25] |
| LDCM0436 | CL46 | HCT 116 | C19(0.52) | LDD0753 | [26] |
| LDCM0437 | CL47 | HCT 116 | C19(0.71) | LDD0754 | [26] |
| LDCM0438 | CL48 | HCT 116 | C19(0.64) | LDD0755 | [26] |
| LDCM0439 | CL49 | HCT 116 | C19(0.62) | LDD0756 | [26] |
| LDCM0441 | CL50 | HCT 116 | C19(0.56) | LDD0758 | [26] |
| LDCM0442 | CL51 | HCT 116 | C19(0.58) | LDD0759 | [26] |
| LDCM0443 | CL52 | HCT 116 | C19(0.61) | LDD0760 | [26] |
| LDCM0444 | CL53 | HCT 116 | C19(0.95) | LDD0761 | [26] |
| LDCM0445 | CL54 | HCT 116 | C19(0.49) | LDD0762 | [26] |
| LDCM0446 | CL55 | HCT 116 | C19(0.57) | LDD0763 | [26] |
| LDCM0447 | CL56 | HCT 116 | C19(0.76) | LDD0764 | [26] |
| LDCM0448 | CL57 | HCT 116 | C19(0.76) | LDD0765 | [26] |
| LDCM0449 | CL58 | HCT 116 | C19(0.61) | LDD0766 | [26] |
| LDCM0450 | CL59 | HCT 116 | C19(0.74) | LDD0767 | [26] |
| LDCM0452 | CL60 | HCT 116 | C19(0.39) | LDD0769 | [26] |
| LDCM0453 | CL61 | HEK-293T | C19(0.96) | LDD1656 | [25] |
| LDCM0460 | CL68 | HEK-293T | C19(1.03) | LDD1663 | [25] |
| LDCM0463 | CL70 | HEK-293T | C19(0.73) | LDD1666 | [25] |
| LDCM0464 | CL71 | HEK-293T | C19(0.73) | LDD1667 | [25] |
| LDCM0465 | CL72 | HEK-293T | C19(1.14) | LDD1668 | [25] |
| LDCM0466 | CL73 | HEK-293T | C19(0.91) | LDD1669 | [25] |
| LDCM0473 | CL8 | HEK-293T | C19(0.97) | LDD1676 | [25] |
| LDCM0474 | CL80 | HEK-293T | C19(1.03) | LDD1677 | [25] |
| LDCM0476 | CL82 | HEK-293T | C19(0.82) | LDD1679 | [25] |
| LDCM0477 | CL83 | HEK-293T | C19(0.73) | LDD1680 | [25] |
| LDCM0478 | CL84 | HEK-293T | C19(1.10) | LDD1681 | [25] |
| LDCM0479 | CL85 | HEK-293T | C19(0.88) | LDD1682 | [25] |
| LDCM0487 | CL92 | HEK-293T | C19(1.08) | LDD1690 | [25] |
| LDCM0489 | CL94 | HEK-293T | C19(0.83) | LDD1692 | [25] |
| LDCM0490 | CL95 | HEK-293T | C19(0.87) | LDD1693 | [25] |
| LDCM0491 | CL96 | HEK-293T | C19(1.04) | LDD1694 | [25] |
| LDCM0492 | CL97 | HEK-293T | C19(1.03) | LDD1695 | [25] |
| LDCM0625 | F8 | Ramos | C19(0.98) | LDD2187 | [27] |
| LDCM0572 | Fragment10 | Ramos | C19(0.49) | LDD2189 | [27] |
| LDCM0573 | Fragment11 | Ramos | C19(0.64) | LDD2190 | [27] |
| LDCM0574 | Fragment12 | Ramos | C19(0.45) | LDD2191 | [27] |
| LDCM0575 | Fragment13 | Ramos | C19(1.16) | LDD2192 | [27] |
| LDCM0576 | Fragment14 | Ramos | C19(0.83) | LDD2193 | [27] |
| LDCM0579 | Fragment20 | Ramos | C19(0.41) | LDD2194 | [27] |
| LDCM0580 | Fragment21 | Ramos | C19(0.71) | LDD2195 | [27] |
| LDCM0582 | Fragment23 | Ramos | C19(1.49) | LDD2196 | [27] |
| LDCM0578 | Fragment27 | Ramos | C19(0.91) | LDD2197 | [27] |
| LDCM0586 | Fragment28 | Ramos | C19(1.20) | LDD2198 | [27] |
| LDCM0588 | Fragment30 | Ramos | C19(1.33) | LDD2199 | [27] |
| LDCM0589 | Fragment31 | Ramos | C19(0.88) | LDD2200 | [27] |
| LDCM0590 | Fragment32 | Ramos | C19(0.52) | LDD2201 | [27] |
| LDCM0468 | Fragment33 | Ramos | C19(0.72) | LDD2202 | [27] |
| LDCM0596 | Fragment38 | Ramos | C19(1.53) | LDD2203 | [27] |
| LDCM0566 | Fragment4 | Ramos | C19(0.64) | LDD2184 | [27] |
| LDCM0610 | Fragment52 | Ramos | C19(0.90) | LDD2204 | [27] |
| LDCM0614 | Fragment56 | Ramos | C19(1.11) | LDD2205 | [27] |
| LDCM0569 | Fragment7 | Ramos | C19(0.58) | LDD2186 | [27] |
| LDCM0571 | Fragment9 | Ramos | C19(0.28) | LDD2188 | [27] |
| LDCM0116 | HHS-0101 | DM93 | Y104(0.82) | LDD0264 | [9] |
| LDCM0117 | HHS-0201 | DM93 | Y104(0.78) | LDD0265 | [9] |
| LDCM0118 | HHS-0301 | DM93 | Y104(0.88) | LDD0266 | [9] |
| LDCM0119 | HHS-0401 | DM93 | Y104(1.14) | LDD0267 | [9] |
| LDCM0120 | HHS-0701 | DM93 | Y104(1.45) | LDD0268 | [9] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [18] |
| LDCM0022 | KB02 | Ramos | C19(0.54) | LDD2182 | [27] |
| LDCM0023 | KB03 | Ramos | C19(0.71) | LDD2183 | [27] |
| LDCM0024 | KB05 | COLO792 | C19(2.00) | LDD3310 | [28] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C19(1.40) | LDD2102 | [7] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C19(0.59) | LDD2121 | [7] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [18] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C19(0.54) | LDD2089 | [7] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C19(1.26) | LDD2090 | [7] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C19(1.34) | LDD2092 | [7] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C19(0.98) | LDD2093 | [7] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C19(2.60) | LDD2094 | [7] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C19(0.20) | LDD2096 | [7] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C19(0.55) | LDD2097 | [7] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C19(0.98) | LDD2099 | [7] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C19(0.47) | LDD2100 | [7] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C19(0.71) | LDD2101 | [7] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C19(0.34) | LDD2104 | [7] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C19(2.37) | LDD2105 | [7] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C19(0.30) | LDD2106 | [7] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C19(1.04) | LDD2107 | [7] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C19(0.26) | LDD2108 | [7] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C19(0.44) | LDD2109 | [7] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C19(0.60) | LDD2110 | [7] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C19(0.99) | LDD2111 | [7] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C19(0.77) | LDD2114 | [7] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C19(0.38) | LDD2115 | [7] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C19(0.43) | LDD2116 | [7] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C19(0.36) | LDD2118 | [7] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C19(1.48) | LDD2119 | [7] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C19(0.55) | LDD2120 | [7] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C19(0.82) | LDD2123 | [7] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C19(0.76) | LDD2125 | [7] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C19(0.25) | LDD2126 | [7] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C19(0.88) | LDD2127 | [7] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C19(0.53) | LDD2128 | [7] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C19(0.95) | LDD2129 | [7] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C19(0.51) | LDD2133 | [7] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C19(0.48) | LDD2134 | [7] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C19(1.41) | LDD2135 | [7] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C19(1.05) | LDD2136 | [7] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C19(0.74) | LDD2137 | [7] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C19(1.86) | LDD1700 | [7] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C19(0.66) | LDD2141 | [7] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C19(0.37) | LDD2143 | [7] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C19(2.05) | LDD2144 | [7] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C19(0.16) | LDD2145 | [7] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C19(0.92) | LDD2146 | [7] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C19(1.80) | LDD2147 | [7] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C19(0.46) | LDD2148 | [7] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C19(0.26) | LDD2149 | [7] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C19(0.54) | LDD2150 | [7] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C19(0.27) | LDD2151 | [7] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C19(2.61) | LDD2153 | [7] |
| LDCM0021 | THZ1 | HCT 116 | C19(1.23) | LDD2173 | [26] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
| Syntenin-1 (SDCBP) | . | O00560 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Proteasome maturation protein (POMP) | POMP/UMP1 family | Q9Y244 | |||
References
