Details of the Target
General Information of Target
| Target ID | LDTP02750 | |||||
|---|---|---|---|---|---|---|
| Target Name | Heterogeneous nuclear ribonucleoprotein L (HNRNPL) | |||||
| Gene Name | HNRNPL | |||||
| Gene ID | 3191 | |||||
| Synonyms |
HNRPL; Heterogeneous nuclear ribonucleoprotein L; hnRNP L |
|||||
| 3D Structure | ||||||
| Sequence |
MSRRLLPRAEKRRRRLEQRQQPDEQRRRSGAMVKMAAAGGGGGGGRYYGGGSEGGRAPKR
LKTDNAGDQHGGGGGGGGGAGAAGGGGGGENYDDPHKTPASPVVHIRGLIDGVVEADLVE ALQEFGPISYVVVMPKKRQALVEFEDVLGACNAVNYAADNQIYIAGHPAFVNYSTSQKIS RPGDSDDSRSVNSVLLFTILNPIYSITTDVLYTICNPCGPVQRIVIFRKNGVQAMVEFDS VQSAQRAKASLNGADIYSGCCTLKIEYAKPTRLNVFKNDQDTWDYTNPNLSGQGDPGSNP NKRQRQPPLLGDHPAEYGGPHGGYHSHYHDEGYGPPPPHYEGRRMGPPVGGHRRGPSRYG PQYGHPPPPPPPPEYGPHADSPVLMVYGLDQSKMNCDRVFNVFCLYGNVEKVKFMKSKPG AAMVEMADGYAVDRAITHLNNNFMFGQKLNVCVSKQPAIMPGQSYGLEDGSCSYKDFSES RNNRFSTPEQAAKNRIQHPSNVLHFFNAPLEVTEENFFEICDELGVKRPSSVKVFSGKSE RSSSGLLEWESKSDALETLGFLNHYQMKNPNGPYPYTLKLCFSTAQHAS |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Nucleus, nucleoplasm
|
|||||
| Function |
Splicing factor binding to exonic or intronic sites and acting as either an activator or repressor of exon inclusion. Exhibits a binding preference for CA-rich elements. Component of the heterogeneous nuclear ribonucleoprotein (hnRNP) complexes and associated with most nascent transcripts. Associates, together with APEX1, to the negative calcium responsive element (nCaRE) B2 of the APEX2 promoter. As part of a ribonucleoprotein complex composed at least of ZNF827, HNRNPK and the circular RNA circZNF827 that nucleates the complex on chromatin, may negatively regulate the transcription of genes involved in neuronal differentiation. Regulates alternative splicing of a core group of genes involved in neuronal differentiation, likely by mediating H3K36me3-coupled transcription elongation and co-transcriptional RNA processing via interaction with CHD8.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 42MGBA | SNV: p.I436V | DBIA Probe Info | |||
| EFO27 | Deletion: p.P367HfsTer28 | DBIA Probe Info | |||
| ETK1 | SNV: p.Q20E | DBIA Probe Info | |||
| HUH28 | SNV: p.Q463Ter | DBIA Probe Info | |||
| IGROV1 | Deletion: p.P367HfsTer28 | DBIA Probe Info | |||
| Ishikawa (Heraklio) 02 ER | SNV: p.S190N | DBIA Probe Info | |||
| JURKAT | SNV: p.P335A; p.S551T | Compound 10 Probe Info | |||
| LNCaP clone FGC | SNV: p.H105R | DBIA Probe Info | |||
| MDAMB231 | SNV: p.G323A | . | |||
| MEWO | SNV: p.P419L | DBIA Probe Info | |||
| MKN45 | SNV: p.M567V | DBIA Probe Info | |||
| OVK18 | Insertion: p.P338HfsTer58 | DBIA Probe Info | |||
| SNU1 | Deletion: p.P337HfsTer58 | DBIA Probe Info | |||
| SSP25 | SNV: p.Q20E | DBIA Probe Info | |||
| TOV21G | SNV: p.H378N | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
14.54 | LDD0402 | [1] | |
|
A-EBA Probe Info |
 |
3.87 | LDD0215 | [2] | |
|
CY-1 Probe Info |
 |
33.86 | LDD0243 | [3] | |
|
CY4 Probe Info |
 |
12.51 | LDD0244 | [3] | |
|
N1 Probe Info |
 |
8.10 | LDD0242 | [3] | |
|
TH211 Probe Info |
 |
Y465(20.00); Y474(20.00); Y375(16.13); Y48(15.63) | LDD0257 | [4] | |
|
TH216 Probe Info |
 |
Y47(15.29); Y430(6.93) | LDD0259 | [4] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [5] | |
|
ONAyne Probe Info |
 |
N.A. | LDD0273 | [6] | |
|
OPA-S-S-alkyne Probe Info |
 |
K538(1.29); K302(1.66); K493(1.94); K269(2.23) | LDD3494 | [7] | |
|
Probe 1 Probe Info |
 |
Y48(50.26); Y92(9.22); Y285(29.86); Y430(11.72) | LDD3495 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C151(10.75) | LDD0168 | [9] | |
|
HHS-475 Probe Info |
 |
Y47(0.26) | LDD0264 | [10] | |
|
DBIA Probe Info |
 |
C452(1.10); C472(0.94) | LDD0078 | [11] | |
|
5E-2FA Probe Info |
 |
H564(0.00); H365(0.00); H378(0.00) | LDD2235 | [12] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [13] | |
|
ATP probe Probe Info |
 |
K493(0.00); K455(0.00); K418(0.00); K533(0.00) | LDD0199 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C260(0.00); C261(0.00); C521(0.00); C581(0.00) | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
C452(0.00); C472(0.00); C581(0.00); C260(0.00) | LDD0032 | [15] | |
|
IPIAA_H Probe Info |
 |
C404(0.00); C472(0.00); C521(0.00) | LDD0030 | [16] | |
|
IPIAA_L Probe Info |
 |
C472(0.00); C151(0.00); C260(0.00); C452(0.00) | LDD0031 | [16] | |
|
Lodoacetamide azide Probe Info |
 |
C260(0.00); C261(0.00); C521(0.00); C581(0.00) | LDD0037 | [14] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [17] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [18] | |
|
JW-RF-010 Probe Info |
 |
C260(0.00); C472(0.00); C581(0.00) | LDD0026 | [19] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [20] | |
|
TFBX Probe Info |
 |
C261(0.00); C472(0.00); C581(0.00); C260(0.00) | LDD0027 | [19] | |
|
WYneN Probe Info |
 |
C581(0.00); C452(0.00); C472(0.00) | LDD0021 | [18] | |
|
WYneO Probe Info |
 |
C581(0.00); C472(0.00); C452(0.00); C260(0.00) | LDD0022 | [18] | |
|
aHNE Probe Info |
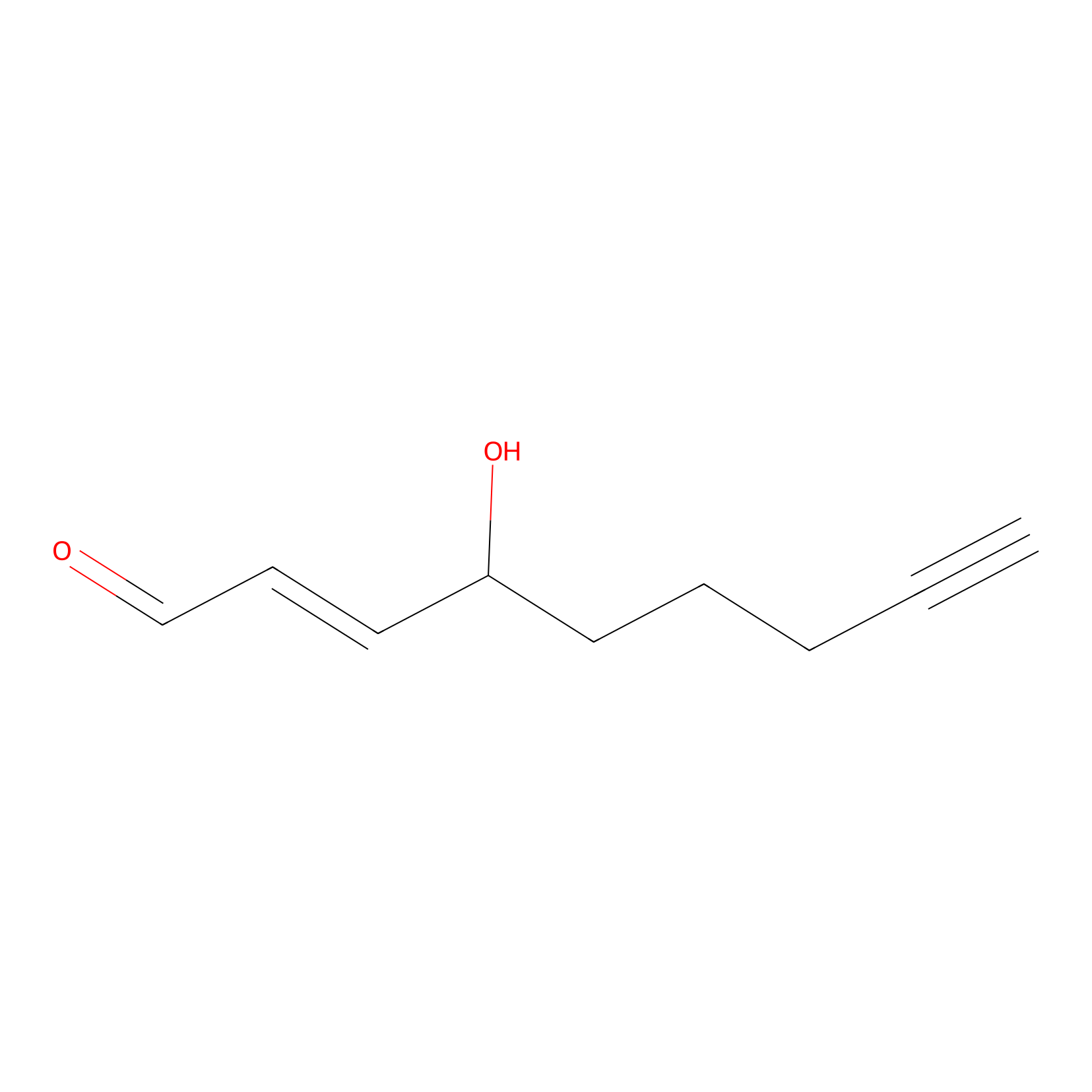 |
N.A. | LDD0001 | [18] | |
|
1d-yne Probe Info |
 |
K97(0.00); K277(0.00); K568(0.00); K418(0.00) | LDD0356 | [21] | |
|
Compound 10 Probe Info |
 |
C260(0.00); C261(0.00); C452(0.00); C472(0.00) | LDD2216 | [22] | |
|
Compound 11 Probe Info |
 |
C260(0.00); C261(0.00); C472(0.00) | LDD2213 | [22] | |
|
ENE Probe Info |
 |
C581(0.00); C260(0.00); C472(0.00) | LDD0006 | [18] | |
|
IPM Probe Info |
 |
C452(0.00); C472(0.00); C581(0.00) | LDD0005 | [18] | |
|
NHS Probe Info |
 |
K302(0.00); K493(0.00); K62(0.00) | LDD0010 | [18] | |
|
OSF Probe Info |
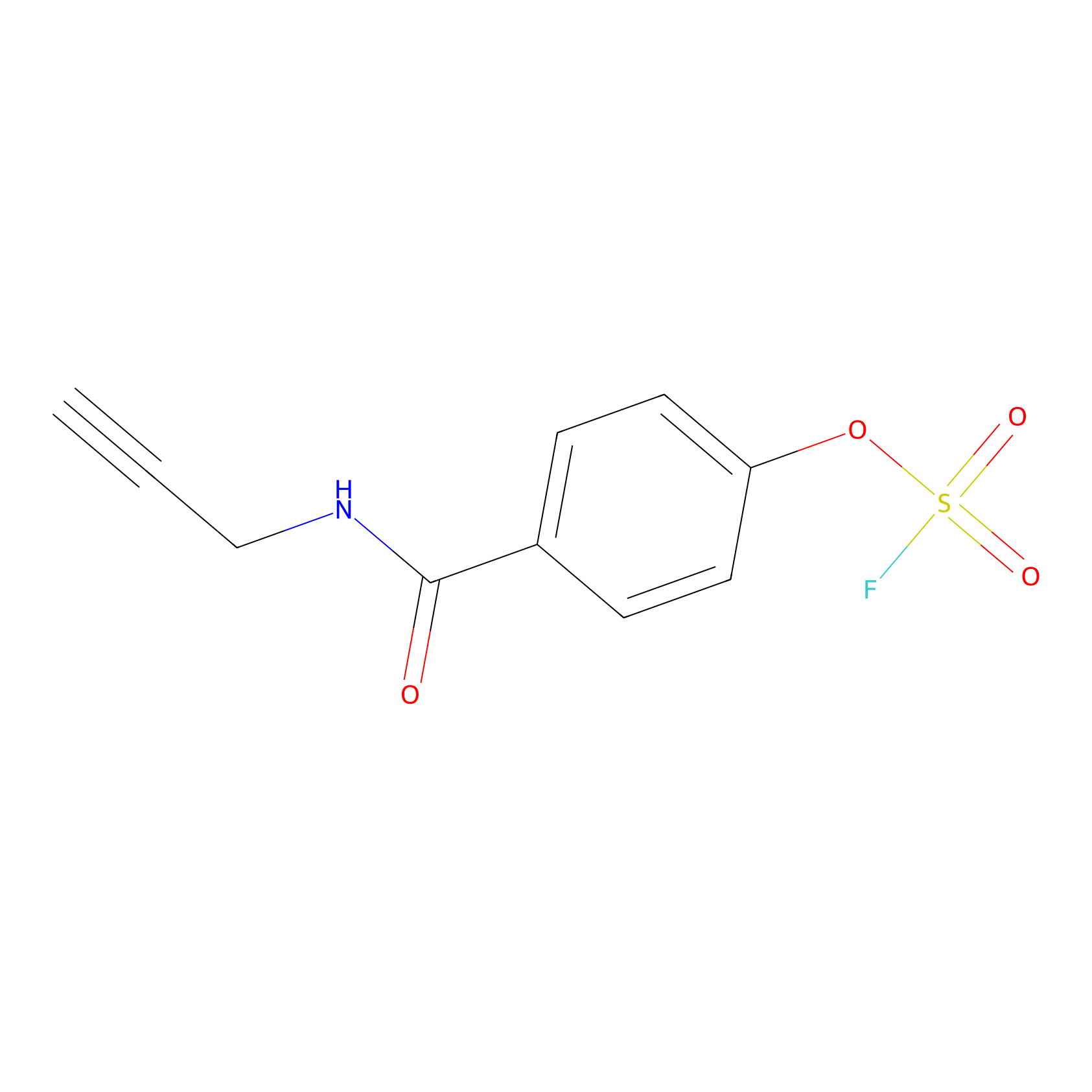 |
Y47(0.00); H438(0.00) | LDD0029 | [23] | |
|
PF-06672131 Probe Info |
 |
C452(0.00); C472(0.00) | LDD0017 | [24] | |
|
PPMS Probe Info |
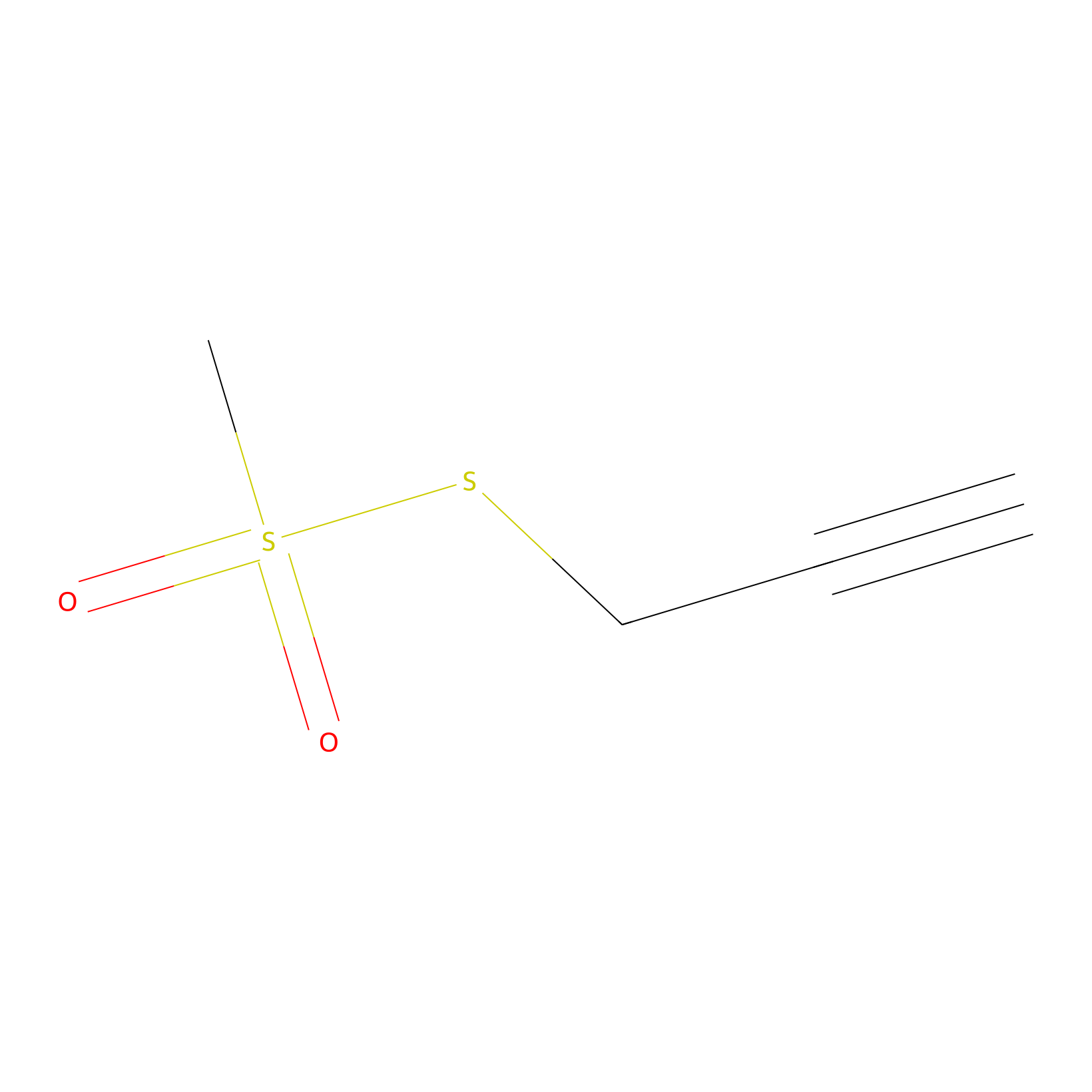 |
N.A. | LDD0008 | [18] | |
|
SF Probe Info |
 |
Y48(0.00); Y430(0.00); K493(0.00); Y359(0.00) | LDD0028 | [23] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [18] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [18] | |
|
YN-1 Probe Info |
 |
N.A. | LDD0446 | [5] | |
|
Phosphinate-6 Probe Info |
 |
C452(0.00); C581(0.00); C472(0.00); C151(0.00) | LDD0018 | [25] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [26] | |
|
1c-yne Probe Info |
 |
K62(0.00); K97(0.00); K229(0.00); K302(0.00) | LDD0228 | [21] | |
|
Acrolein Probe Info |
 |
H70(0.00); C404(0.00); C260(0.00); C472(0.00) | LDD0217 | [27] | |
|
Cinnamaldehyde Probe Info |
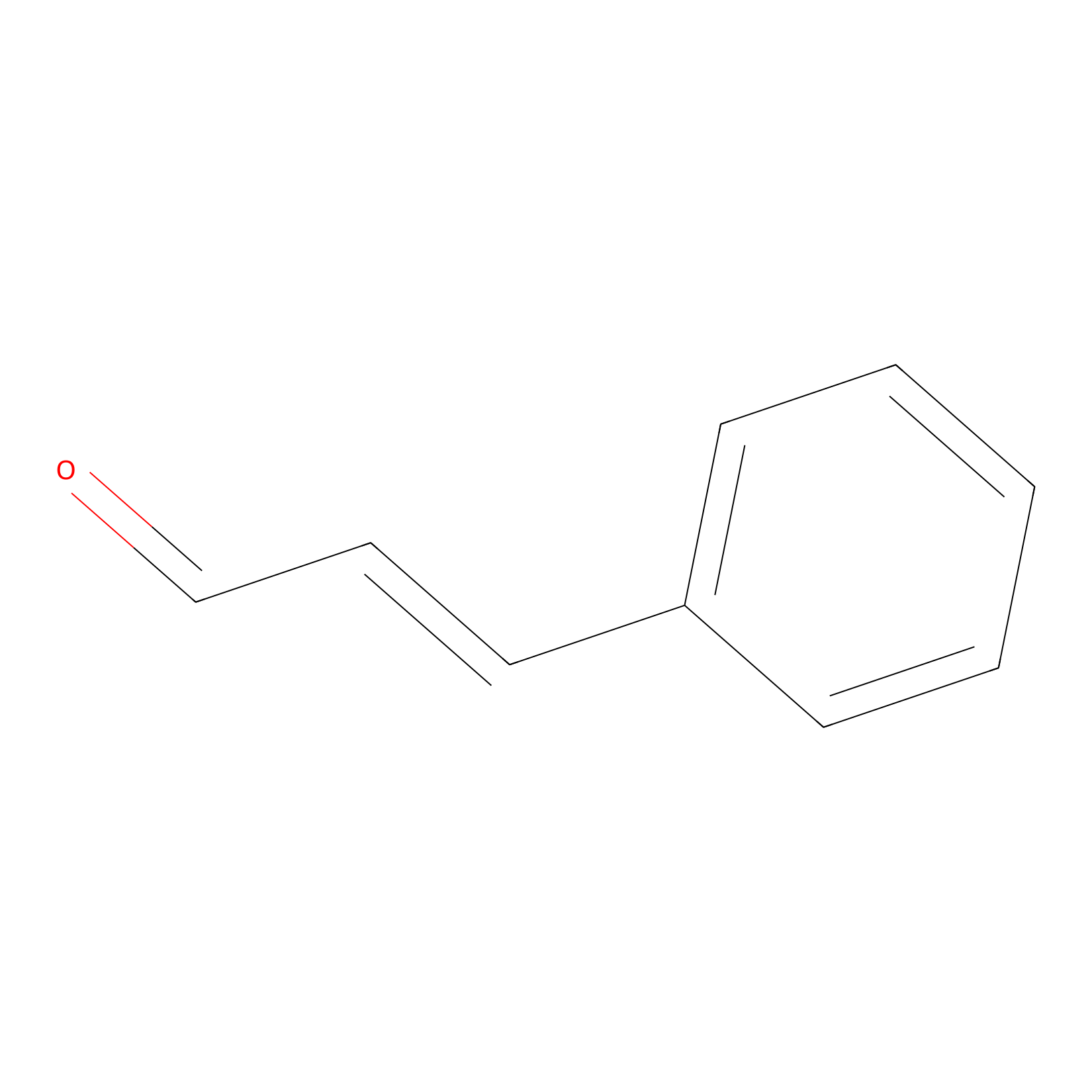 |
N.A. | LDD0220 | [27] | |
|
Crotonaldehyde Probe Info |
 |
C260(0.00); H105(0.00); C404(0.00); C472(0.00) | LDD0219 | [27] | |
|
Methacrolein Probe Info |
 |
C260(0.00); C472(0.00); C581(0.00) | LDD0218 | [27] | |
|
W1 Probe Info |
 |
C452(0.00); C472(0.00); C404(0.00); C260(0.00) | LDD0236 | [28] | |
|
AOyne Probe Info |
 |
10.50 | LDD0443 | [29] | |
|
NAIA_5 Probe Info |
 |
C452(0.00); C260(0.00); C521(0.00); C581(0.00) | LDD2223 | [20] | |
|
HHS-465 Probe Info |
 |
Y47(0.00); Y48(0.00) | LDD2240 | [30] | |
|
HHS-482 Probe Info |
 |
Y359(0.95); Y430(0.86); Y47(1.10); Y48(1.07) | LDD2239 | [31] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
5.00 | LDD0471 | [32] | |
|
FFF probe13 Probe Info |
 |
6.96 | LDD0475 | [32] | |
|
FFF probe14 Probe Info |
 |
6.35 | LDD0477 | [32] | |
|
FFF probe2 Probe Info |
 |
5.79 | LDD0463 | [32] | |
|
FFF probe3 Probe Info |
 |
6.61 | LDD0464 | [32] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [33] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [34] | |
|
Diazir Probe Info |
 |
N.A. | LDD0011 | [18] | |
|
BD-F Probe Info |
 |
H365(0.00); Y363(0.00); P371(0.00) | LDD0024 | [35] | |
|
LD-F Probe Info |
 |
P370(0.00); E120(0.00); S129(0.00); P369(0.00) | LDD0015 | [35] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [36] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [37] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C472(0.96); C452(0.56); C581(0.98) | LDD2142 | [38] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C472(1.02); C452(0.93); C581(1.13) | LDD2112 | [38] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C452(0.76) | LDD2130 | [38] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C472(1.01); C581(1.47); C261(1.21); C260(1.20) | LDD2117 | [38] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C452(1.05); C581(1.43) | LDD2152 | [38] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C452(1.02); C581(1.40) | LDD2103 | [38] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C472(0.76) | LDD2132 | [38] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C472(0.71); C452(0.62); C581(0.58) | LDD2131 | [38] |
| LDCM0025 | 4SU-RNA | HEK-293T | C151(10.75) | LDD0168 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C151(6.04) | LDD0169 | [9] |
| LDCM0214 | AC1 | HCT 116 | C260(1.11); C261(1.10); C452(1.52); C472(1.38) | LDD0531 | [11] |
| LDCM0215 | AC10 | HCT 116 | C260(0.96); C261(0.92); C404(0.74); C452(1.10) | LDD0532 | [11] |
| LDCM0216 | AC100 | HCT 116 | C260(0.58); C261(0.60); C452(1.22); C472(1.05) | LDD0533 | [11] |
| LDCM0217 | AC101 | HCT 116 | C260(0.53); C261(0.57); C452(1.11); C472(1.07) | LDD0534 | [11] |
| LDCM0218 | AC102 | HCT 116 | C260(0.56); C261(0.63); C452(1.10); C472(1.09) | LDD0535 | [11] |
| LDCM0219 | AC103 | HCT 116 | C260(0.45); C261(0.47); C452(1.14); C472(1.20) | LDD0536 | [11] |
| LDCM0220 | AC104 | HCT 116 | C260(0.55); C261(0.55); C452(1.16); C472(1.16) | LDD0537 | [11] |
| LDCM0221 | AC105 | HCT 116 | C260(0.47); C261(0.51); C452(0.99); C472(1.12) | LDD0538 | [11] |
| LDCM0222 | AC106 | HCT 116 | C260(0.45); C261(0.49); C452(1.13); C472(1.15) | LDD0539 | [11] |
| LDCM0223 | AC107 | HCT 116 | C260(0.51); C261(0.54); C452(1.11); C472(1.12) | LDD0540 | [11] |
| LDCM0224 | AC108 | HCT 116 | C260(0.55); C261(0.69); C452(1.34); C472(1.05) | LDD0541 | [11] |
| LDCM0225 | AC109 | HCT 116 | C260(0.61); C261(0.69); C452(1.15); C472(0.89) | LDD0542 | [11] |
| LDCM0226 | AC11 | HCT 116 | C260(1.16); C261(1.22); C404(0.76); C452(1.88) | LDD0543 | [11] |
| LDCM0227 | AC110 | HCT 116 | C260(0.51); C261(0.64); C452(1.15); C472(0.97) | LDD0544 | [11] |
| LDCM0228 | AC111 | HCT 116 | C260(0.50); C261(0.62); C452(1.09); C472(1.01) | LDD0545 | [11] |
| LDCM0229 | AC112 | HCT 116 | C260(0.55); C261(0.56); C452(1.27); C472(1.13) | LDD0546 | [11] |
| LDCM0230 | AC113 | HCT 116 | C260(1.08); C261(1.08); C404(0.79); C452(1.04) | LDD0547 | [11] |
| LDCM0231 | AC114 | HCT 116 | C260(0.90); C261(0.93); C404(0.51); C452(1.00) | LDD0548 | [11] |
| LDCM0232 | AC115 | HCT 116 | C260(0.82); C261(0.89); C404(0.31); C452(0.88) | LDD0549 | [11] |
| LDCM0233 | AC116 | HCT 116 | C260(0.79); C261(0.84); C404(0.27); C452(0.83) | LDD0550 | [11] |
| LDCM0234 | AC117 | HCT 116 | C260(0.88); C261(0.89); C404(0.67); C452(0.98) | LDD0551 | [11] |
| LDCM0235 | AC118 | HCT 116 | C260(0.97); C261(0.95); C404(0.59); C452(1.08) | LDD0552 | [11] |
| LDCM0236 | AC119 | HCT 116 | C260(0.88); C261(0.93); C404(0.38); C452(0.99) | LDD0553 | [11] |
| LDCM0237 | AC12 | HCT 116 | C260(1.25); C261(1.17); C404(1.06); C452(1.63) | LDD0554 | [11] |
| LDCM0238 | AC120 | HCT 116 | C260(1.01); C261(0.94); C404(0.60); C452(1.03) | LDD0555 | [11] |
| LDCM0239 | AC121 | HCT 116 | C260(0.98); C261(0.92); C404(0.53); C452(1.01) | LDD0556 | [11] |
| LDCM0240 | AC122 | HCT 116 | C260(0.91); C261(1.03); C404(0.60); C452(1.15) | LDD0557 | [11] |
| LDCM0241 | AC123 | HCT 116 | C260(1.01); C261(0.98); C404(0.95); C452(1.12) | LDD0558 | [11] |
| LDCM0242 | AC124 | HCT 116 | C260(1.07); C261(1.00); C404(0.90); C452(1.00) | LDD0559 | [11] |
| LDCM0243 | AC125 | HCT 116 | C260(1.00); C261(1.06); C404(0.66); C452(0.99) | LDD0560 | [11] |
| LDCM0244 | AC126 | HCT 116 | C260(0.82); C261(0.88); C404(0.22); C452(0.85) | LDD0561 | [11] |
| LDCM0245 | AC127 | HCT 116 | C260(0.78); C261(0.81); C404(0.31); C452(0.89) | LDD0562 | [11] |
| LDCM0246 | AC128 | HCT 116 | C218(0.57); C260(1.04); C261(0.87); C452(0.64) | LDD0563 | [11] |
| LDCM0247 | AC129 | HCT 116 | C218(1.11); C260(1.16); C261(1.08); C452(1.19) | LDD0564 | [11] |
| LDCM0249 | AC130 | HCT 116 | C218(0.64); C260(0.97); C261(0.90); C452(1.02) | LDD0566 | [11] |
| LDCM0250 | AC131 | HCT 116 | C218(1.40); C260(1.05); C261(0.99); C452(0.94) | LDD0567 | [11] |
| LDCM0251 | AC132 | HCT 116 | C218(1.08); C260(0.93); C261(1.02); C452(1.31) | LDD0568 | [11] |
| LDCM0252 | AC133 | HCT 116 | C218(0.83); C260(0.97); C261(1.00); C452(1.05) | LDD0569 | [11] |
| LDCM0253 | AC134 | HCT 116 | C218(0.78); C260(0.86); C261(0.90); C452(0.90) | LDD0570 | [11] |
| LDCM0254 | AC135 | HCT 116 | C218(0.73); C260(0.86); C261(0.91); C452(0.92) | LDD0571 | [11] |
| LDCM0255 | AC136 | HCT 116 | C218(0.87); C260(0.85); C261(0.91); C452(1.18) | LDD0572 | [11] |
| LDCM0256 | AC137 | HCT 116 | C218(1.05); C260(0.94); C261(1.01); C452(1.29) | LDD0573 | [11] |
| LDCM0257 | AC138 | HCT 116 | C218(0.73); C260(0.88); C261(0.94); C452(1.30) | LDD0574 | [11] |
| LDCM0258 | AC139 | HCT 116 | C218(1.21); C260(0.82); C261(0.89); C452(1.20) | LDD0575 | [11] |
| LDCM0259 | AC14 | HCT 116 | C260(1.32); C261(1.24); C404(0.97); C452(1.05) | LDD0576 | [11] |
| LDCM0260 | AC140 | HCT 116 | C218(0.83); C260(0.76); C261(0.84); C452(1.12) | LDD0577 | [11] |
| LDCM0261 | AC141 | HCT 116 | C218(1.02); C260(0.79); C261(0.89); C452(1.20) | LDD0578 | [11] |
| LDCM0262 | AC142 | HCT 116 | C218(1.59); C260(1.09); C261(1.21); C452(1.68) | LDD0579 | [11] |
| LDCM0263 | AC143 | HCT 116 | C260(0.82); C261(0.87); C404(0.68); C452(1.41) | LDD0580 | [11] |
| LDCM0264 | AC144 | HCT 116 | C404(0.31); C260(0.75); C261(0.79); C581(0.81) | LDD0581 | [11] |
| LDCM0265 | AC145 | HCT 116 | C404(0.65); C581(0.72); C260(0.87); C261(0.92) | LDD0582 | [11] |
| LDCM0266 | AC146 | HCT 116 | C404(0.45); C261(0.73); C260(0.75); C581(0.81) | LDD0583 | [11] |
| LDCM0267 | AC147 | HCT 116 | C404(0.20); C452(0.66); C260(0.69); C261(0.74) | LDD0584 | [11] |
| LDCM0268 | AC148 | HCT 116 | C404(0.15); C260(0.60); C261(0.61); C452(0.66) | LDD0585 | [11] |
| LDCM0269 | AC149 | HCT 116 | C404(0.18); C260(0.64); C261(0.64); C452(0.72) | LDD0586 | [11] |
| LDCM0270 | AC15 | HCT 116 | C404(0.74); C261(1.01); C260(1.07); C472(1.13) | LDD0587 | [11] |
| LDCM0271 | AC150 | HCT 116 | C404(0.75); C581(0.92); C260(0.95); C261(0.98) | LDD0588 | [11] |
| LDCM0272 | AC151 | HCT 116 | C404(0.85); C260(0.86); C261(0.97); C472(1.05) | LDD0589 | [11] |
| LDCM0273 | AC152 | HCT 116 | C404(0.21); C452(0.77); C261(0.84); C260(0.86) | LDD0590 | [11] |
| LDCM0274 | AC153 | HCT 116 | C404(0.12); C260(0.72); C261(0.73); C452(0.78) | LDD0591 | [11] |
| LDCM0621 | AC154 | HCT 116 | C260(0.88); C261(0.99); C404(0.40); C452(0.85) | LDD2158 | [11] |
| LDCM0622 | AC155 | HCT 116 | C260(0.86); C261(0.95); C404(0.30); C452(0.96) | LDD2159 | [11] |
| LDCM0623 | AC156 | HCT 116 | C260(0.91); C261(0.94); C404(0.63); C452(1.00) | LDD2160 | [11] |
| LDCM0624 | AC157 | HCT 116 | C260(1.10); C261(1.15); C404(0.89); C452(0.99) | LDD2161 | [11] |
| LDCM0276 | AC17 | HCT 116 | C261(0.93); C404(0.95); C260(0.97); C452(0.99) | LDD0593 | [11] |
| LDCM0277 | AC18 | HCT 116 | C404(0.51); C260(0.88); C261(0.93); C452(0.96) | LDD0594 | [11] |
| LDCM0278 | AC19 | HCT 116 | C404(0.76); C581(0.85); C452(0.87); C261(0.91) | LDD0595 | [11] |
| LDCM0279 | AC2 | HCT 116 | C581(0.85); C261(1.01); C260(1.17); C472(1.24) | LDD0596 | [11] |
| LDCM0280 | AC20 | HCT 116 | C404(0.82); C261(0.88); C452(0.89); C260(0.96) | LDD0597 | [11] |
| LDCM0281 | AC21 | HCT 116 | C404(0.79); C452(0.88); C260(0.92); C261(0.96) | LDD0598 | [11] |
| LDCM0282 | AC22 | HCT 116 | C581(0.73); C404(0.84); C472(0.94); C261(1.02) | LDD0599 | [11] |
| LDCM0283 | AC23 | HCT 116 | C404(0.68); C261(0.94); C472(1.01); C260(1.03) | LDD0600 | [11] |
| LDCM0284 | AC24 | HCT 116 | C452(0.90); C404(0.90); C261(0.91); C581(0.93) | LDD0601 | [11] |
| LDCM0285 | AC25 | HCT 116 | C581(0.51); C261(0.85); C260(0.88); C452(1.01) | LDD0602 | [11] |
| LDCM0286 | AC26 | HCT 116 | C581(0.56); C261(0.80); C260(0.83); C452(1.15) | LDD0603 | [11] |
| LDCM0287 | AC27 | HCT 116 | C581(0.61); C261(0.78); C260(0.78); C452(1.01) | LDD0604 | [11] |
| LDCM0288 | AC28 | HCT 116 | C581(0.57); C260(0.79); C261(0.79); C452(1.01) | LDD0605 | [11] |
| LDCM0289 | AC29 | HCT 116 | C260(0.73); C261(0.73); C581(0.94); C452(0.98) | LDD0606 | [11] |
| LDCM0290 | AC3 | HCT 116 | C260(1.11); C261(1.12); C472(1.25); C581(1.42) | LDD0607 | [11] |
| LDCM0291 | AC30 | HCT 116 | C261(0.79); C260(0.83); C581(0.94); C452(1.19) | LDD0608 | [11] |
| LDCM0292 | AC31 | HCT 116 | C581(0.70); C260(0.79); C261(0.82); C452(1.10) | LDD0609 | [11] |
| LDCM0293 | AC32 | HCT 116 | C261(0.73); C260(0.73); C581(0.82); C452(1.09) | LDD0610 | [11] |
| LDCM0294 | AC33 | HCT 116 | C261(0.75); C260(0.79); C581(0.79); C452(1.06) | LDD0611 | [11] |
| LDCM0295 | AC34 | HCT 116 | C260(0.69); C261(0.73); C452(1.02); C472(1.58) | LDD0612 | [11] |
| LDCM0296 | AC35 | HCT 116 | C581(0.66); C260(1.07); C261(1.12); C472(1.70) | LDD0613 | [11] |
| LDCM0297 | AC36 | HCT 116 | C581(0.65); C261(0.94); C260(0.94); C472(1.00) | LDD0614 | [11] |
| LDCM0298 | AC37 | HCT 116 | C581(0.69); C472(0.85); C260(0.90); C261(0.98) | LDD0615 | [11] |
| LDCM0299 | AC38 | HCT 116 | C581(0.85); C472(0.91); C452(0.93); C261(1.01) | LDD0616 | [11] |
| LDCM0300 | AC39 | HCT 116 | C404(0.60); C260(0.92); C261(0.98); C472(1.07) | LDD0617 | [11] |
| LDCM0301 | AC4 | HCT 116 | C260(1.18); C261(1.25); C581(1.25); C472(1.41) | LDD0618 | [11] |
| LDCM0302 | AC40 | HCT 116 | C404(0.52); C260(0.69); C452(0.80); C261(0.87) | LDD0619 | [11] |
| LDCM0303 | AC41 | HCT 116 | C581(0.81); C261(0.91); C260(0.95); C472(1.00) | LDD0620 | [11] |
| LDCM0304 | AC42 | HCT 116 | C581(0.67); C260(0.82); C452(0.89); C261(0.95) | LDD0621 | [11] |
| LDCM0305 | AC43 | HCT 116 | C581(0.57); C261(0.92); C260(0.92); C472(1.08) | LDD0622 | [11] |
| LDCM0306 | AC44 | HCT 116 | C581(0.71); C404(0.76); C260(0.91); C472(0.97) | LDD0623 | [11] |
| LDCM0307 | AC45 | HCT 116 | C404(0.51); C260(0.82); C452(0.88); C261(0.92) | LDD0624 | [11] |
| LDCM0308 | AC46 | HCT 116 | C260(0.76); C581(0.79); C261(1.00); C452(1.01) | LDD0625 | [11] |
| LDCM0309 | AC47 | HCT 116 | C404(0.64); C260(0.71); C581(0.72); C261(0.93) | LDD0626 | [11] |
| LDCM0310 | AC48 | HCT 116 | C260(0.64); C581(0.78); C261(0.81); C404(0.81) | LDD0627 | [11] |
| LDCM0311 | AC49 | HCT 116 | C404(0.39); C260(0.60); C261(0.74); C452(1.07) | LDD0628 | [11] |
| LDCM0312 | AC5 | HCT 116 | C261(1.27); C581(1.44); C260(1.44); C472(1.53) | LDD0629 | [11] |
| LDCM0313 | AC50 | HCT 116 | C404(0.36); C260(0.62); C261(0.79); C452(0.83) | LDD0630 | [11] |
| LDCM0314 | AC51 | HCT 116 | C581(0.67); C260(0.71); C261(0.99); C472(1.15) | LDD0631 | [11] |
| LDCM0315 | AC52 | HCT 116 | C404(0.80); C260(0.81); C452(0.96); C261(0.97) | LDD0632 | [11] |
| LDCM0316 | AC53 | HCT 116 | C404(0.44); C260(0.74); C581(0.78); C261(0.88) | LDD0633 | [11] |
| LDCM0317 | AC54 | HCT 116 | C404(0.54); C260(0.80); C261(0.90); C581(0.95) | LDD0634 | [11] |
| LDCM0318 | AC55 | HCT 116 | C404(0.37); C260(0.65); C581(0.70); C261(0.79) | LDD0635 | [11] |
| LDCM0319 | AC56 | HCT 116 | C404(0.44); C260(0.78); C261(0.89); C452(1.00) | LDD0636 | [11] |
| LDCM0320 | AC57 | HCT 116 | C404(0.45); C581(0.70); C260(0.87); C261(0.87) | LDD0637 | [11] |
| LDCM0321 | AC58 | HCT 116 | C404(0.56); C261(0.77); C260(0.79); C452(0.82) | LDD0638 | [11] |
| LDCM0322 | AC59 | HCT 116 | C404(0.28); C261(0.77); C260(0.80); C452(0.82) | LDD0639 | [11] |
| LDCM0323 | AC6 | HCT 116 | C404(0.66); C581(0.75); C261(0.92); C452(0.93) | LDD0640 | [11] |
| LDCM0324 | AC60 | HCT 116 | C404(0.57); C261(0.80); C260(0.83); C581(0.90) | LDD0641 | [11] |
| LDCM0325 | AC61 | HCT 116 | C581(0.75); C261(0.81); C260(0.81); C404(1.10) | LDD0642 | [11] |
| LDCM0326 | AC62 | HCT 116 | C404(0.26); C581(0.66); C260(0.73); C261(0.73) | LDD0643 | [11] |
| LDCM0327 | AC63 | HCT 116 | C404(0.58); C581(0.62); C261(0.87); C260(0.95) | LDD0644 | [11] |
| LDCM0328 | AC64 | HCT 116 | C404(0.26); C581(0.65); C452(0.79); C260(0.83) | LDD0645 | [11] |
| LDCM0329 | AC65 | HCT 116 | C404(0.17); C261(0.76); C260(0.76); C581(0.78) | LDD0646 | [11] |
| LDCM0330 | AC66 | HCT 116 | C404(0.25); C260(0.79); C261(0.81); C581(0.84) | LDD0647 | [11] |
| LDCM0331 | AC67 | HCT 116 | C404(0.12); C261(0.63); C260(0.66); C581(0.77) | LDD0648 | [11] |
| LDCM0332 | AC68 | HCT 116 | C218(0.60); C452(0.71); C261(0.87); C472(0.87) | LDD0649 | [11] |
| LDCM0333 | AC69 | HCT 116 | C218(0.47); C452(0.65); C261(0.82); C260(0.87) | LDD0650 | [11] |
| LDCM0334 | AC7 | HCT 116 | C581(0.70); C404(0.84); C261(1.03); C260(1.06) | LDD0651 | [11] |
| LDCM0335 | AC70 | HCT 116 | C218(0.48); C452(0.53); C260(0.65); C261(0.66) | LDD0652 | [11] |
| LDCM0336 | AC71 | HCT 116 | C452(0.69); C472(0.84); C581(0.86); C261(0.97) | LDD0653 | [11] |
| LDCM0337 | AC72 | HCT 116 | C452(0.69); C260(0.79); C261(0.80); C218(0.80) | LDD0654 | [11] |
| LDCM0338 | AC73 | HCT 116 | C218(0.39); C452(0.58); C260(0.65); C261(0.65) | LDD0655 | [11] |
| LDCM0339 | AC74 | HCT 116 | C218(0.57); C452(0.57); C261(0.72); C260(0.74) | LDD0656 | [11] |
| LDCM0340 | AC75 | HCT 116 | C218(0.39); C452(0.62); C260(0.66); C261(0.66) | LDD0657 | [11] |
| LDCM0341 | AC76 | HCT 116 | C218(0.48); C452(0.52); C261(0.71); C260(0.74) | LDD0658 | [11] |
| LDCM0342 | AC77 | HCT 116 | C218(0.45); C452(0.52); C261(0.75); C260(0.80) | LDD0659 | [11] |
| LDCM0343 | AC78 | HCT 116 | C218(0.65); C261(0.83); C260(0.86); C452(0.97) | LDD0660 | [11] |
| LDCM0344 | AC79 | HCT 116 | C452(0.71); C260(0.86); C472(0.89); C261(0.89) | LDD0661 | [11] |
| LDCM0345 | AC8 | HCT 116 | C404(0.77); C261(1.10); C260(1.12); C581(1.43) | LDD0662 | [11] |
| LDCM0346 | AC80 | HCT 116 | C452(0.62); C218(0.76); C261(0.83); C472(0.84) | LDD0663 | [11] |
| LDCM0347 | AC81 | HCT 116 | C218(0.42); C452(0.96); C472(0.98); C581(1.12) | LDD0664 | [11] |
| LDCM0348 | AC82 | HCT 116 | C218(0.43); C452(0.75); C261(0.78); C260(0.80) | LDD0665 | [11] |
| LDCM0349 | AC83 | HCT 116 | C261(0.46); C260(0.54); C404(0.67); C452(0.67) | LDD0666 | [11] |
| LDCM0350 | AC84 | HCT 116 | C261(0.49); C260(0.57); C404(0.73); C452(0.96) | LDD0667 | [11] |
| LDCM0351 | AC85 | HCT 116 | C261(0.58); C260(0.62); C404(0.92); C581(1.06) | LDD0668 | [11] |
| LDCM0352 | AC86 | HCT 116 | C261(0.65); C260(0.66); C404(0.85); C581(0.95) | LDD0669 | [11] |
| LDCM0353 | AC87 | HCT 116 | C261(0.78); C581(0.89); C260(0.92); C452(1.07) | LDD0670 | [11] |
| LDCM0354 | AC88 | HCT 116 | C261(0.63); C260(0.71); C404(0.83); C452(0.91) | LDD0671 | [11] |
| LDCM0355 | AC89 | HCT 116 | C261(0.53); C260(0.61); C404(0.77); C452(0.85) | LDD0672 | [11] |
| LDCM0357 | AC90 | HCT 116 | C581(0.59); C261(0.75); C260(0.91); C472(1.02) | LDD0674 | [11] |
| LDCM0358 | AC91 | HCT 116 | C260(0.50); C261(0.52); C452(0.59); C404(0.60) | LDD0675 | [11] |
| LDCM0359 | AC92 | HCT 116 | C261(0.44); C260(0.51); C404(0.61); C452(0.66) | LDD0676 | [11] |
| LDCM0360 | AC93 | HCT 116 | C261(0.66); C260(0.76); C404(0.90); C581(1.00) | LDD0677 | [11] |
| LDCM0361 | AC94 | HCT 116 | C260(0.78); C581(0.84); C261(0.94); C404(0.97) | LDD0678 | [11] |
| LDCM0362 | AC95 | HCT 116 | C261(0.63); C260(0.72); C404(0.91); C581(1.05) | LDD0679 | [11] |
| LDCM0363 | AC96 | HCT 116 | C260(0.69); C261(0.80); C452(0.83); C404(0.83) | LDD0680 | [11] |
| LDCM0364 | AC97 | HCT 116 | C261(0.47); C260(0.52); C452(0.77); C404(0.81) | LDD0681 | [11] |
| LDCM0365 | AC98 | HCT 116 | C260(0.37); C261(0.44); C581(0.76); C452(1.20) | LDD0682 | [11] |
| LDCM0366 | AC99 | HCT 116 | C260(0.64); C261(0.68); C472(1.09); C581(1.16) | LDD0683 | [11] |
| LDCM0545 | Acetamide | MDA-MB-231 | C472(0.65); C452(0.32); C581(0.63) | LDD2138 | [38] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C452(0.83); C581(0.80); C261(1.14) | LDD2113 | [38] |
| LDCM0248 | AKOS034007472 | HCT 116 | C260(1.13); C261(1.15); C404(0.97); C452(1.20) | LDD0565 | [11] |
| LDCM0356 | AKOS034007680 | HCT 116 | C404(0.84); C581(0.86); C260(1.02); C452(1.06) | LDD0673 | [11] |
| LDCM0275 | AKOS034007705 | HCT 116 | C404(0.20); C261(0.72); C260(0.84); C452(0.88) | LDD0592 | [11] |
| LDCM0156 | Aniline | NCI-H1299 | C151(0.00); C472(0.00) | LDD0405 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C452(1.10); C472(0.94) | LDD0078 | [11] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C452(0.35); C581(0.68) | LDD2091 | [38] |
| LDCM0108 | Chloroacetamide | HeLa | H96(0.00); H70(0.00); C260(0.00); C472(0.00) | LDD0222 | [27] |
| LDCM0632 | CL-Sc | Hep-G2 | C581(1.07); C452(0.42) | LDD2227 | [20] |
| LDCM0367 | CL1 | HCT 116 | C581(0.86); C260(1.01); C261(1.01); C404(1.03) | LDD0684 | [11] |
| LDCM0368 | CL10 | HCT 116 | C404(0.50); C260(0.73); C218(0.74); C261(0.74) | LDD0685 | [11] |
| LDCM0369 | CL100 | HCT 116 | C581(0.83); C261(1.03); C260(1.04); C472(1.33) | LDD0686 | [11] |
| LDCM0370 | CL101 | HCT 116 | C404(0.62); C581(0.75); C452(0.77); C261(1.02) | LDD0687 | [11] |
| LDCM0371 | CL102 | HCT 116 | C581(0.66); C404(0.85); C452(0.91); C472(0.98) | LDD0688 | [11] |
| LDCM0372 | CL103 | HCT 116 | C581(0.78); C404(0.99); C472(1.00); C452(1.03) | LDD0689 | [11] |
| LDCM0373 | CL104 | HCT 116 | C581(0.92); C404(1.01); C261(1.05); C472(1.13) | LDD0690 | [11] |
| LDCM0374 | CL105 | HCT 116 | C404(0.63); C581(0.93); C260(0.96); C261(1.12) | LDD0691 | [11] |
| LDCM0375 | CL106 | HCT 116 | C404(0.40); C260(0.90); C261(1.02); C452(1.17) | LDD0692 | [11] |
| LDCM0376 | CL107 | HCT 116 | C404(0.29); C260(0.92); C261(1.03); C452(1.12) | LDD0693 | [11] |
| LDCM0377 | CL108 | HCT 116 | C404(0.27); C260(0.88); C452(0.92); C261(0.92) | LDD0694 | [11] |
| LDCM0378 | CL109 | HCT 116 | C404(0.62); C260(0.86); C261(0.89); C452(0.99) | LDD0695 | [11] |
| LDCM0379 | CL11 | HCT 116 | C404(0.34); C261(0.71); C260(0.75); C218(0.78) | LDD0696 | [11] |
| LDCM0380 | CL110 | HCT 116 | C404(0.46); C261(0.80); C260(0.98); C452(1.09) | LDD0697 | [11] |
| LDCM0381 | CL111 | HCT 116 | C404(0.64); C260(0.91); C261(0.93); C472(1.22) | LDD0698 | [11] |
| LDCM0382 | CL112 | HCT 116 | C581(0.66); C261(0.92); C260(1.00); C452(1.00) | LDD0699 | [11] |
| LDCM0383 | CL113 | HCT 116 | C260(0.74); C261(0.76); C581(0.87); C452(1.14) | LDD0700 | [11] |
| LDCM0384 | CL114 | HCT 116 | C261(0.76); C260(0.83); C452(1.19); C472(1.28) | LDD0701 | [11] |
| LDCM0385 | CL115 | HCT 116 | C260(0.79); C261(0.81); C452(1.01); C581(1.22) | LDD0702 | [11] |
| LDCM0386 | CL116 | HCT 116 | C261(0.82); C260(0.84); C472(1.20); C581(1.20) | LDD0703 | [11] |
| LDCM0387 | CL117 | HCT 116 | C404(0.57); C260(0.70); C261(0.75); C452(0.81) | LDD0704 | [11] |
| LDCM0388 | CL118 | HCT 116 | C581(0.59); C260(0.80); C261(0.89); C452(0.97) | LDD0705 | [11] |
| LDCM0389 | CL119 | HCT 116 | C581(0.55); C404(0.81); C260(0.88); C261(0.91) | LDD0706 | [11] |
| LDCM0390 | CL12 | HCT 116 | C404(0.30); C260(0.63); C218(0.63); C261(0.67) | LDD0707 | [11] |
| LDCM0391 | CL120 | HCT 116 | C581(0.50); C404(0.85); C260(0.85); C261(0.94) | LDD0708 | [11] |
| LDCM0392 | CL121 | HCT 116 | C260(0.59); C261(0.68); C404(0.79); C581(0.91) | LDD0709 | [11] |
| LDCM0393 | CL122 | HCT 116 | C404(0.89); C261(0.98); C260(1.06); C581(1.07) | LDD0710 | [11] |
| LDCM0394 | CL123 | HCT 116 | C404(0.36); C260(0.71); C581(0.77); C452(0.79) | LDD0711 | [11] |
| LDCM0395 | CL124 | HCT 116 | C404(0.36); C260(0.69); C261(0.84); C581(0.85) | LDD0712 | [11] |
| LDCM0396 | CL125 | HCT 116 | C581(0.72); C261(1.00); C404(1.04); C260(1.05) | LDD0713 | [11] |
| LDCM0397 | CL126 | HCT 116 | C404(0.99); C261(1.04); C472(1.06); C260(1.08) | LDD0714 | [11] |
| LDCM0398 | CL127 | HCT 116 | C581(0.92); C404(0.95); C260(0.98); C261(0.98) | LDD0715 | [11] |
| LDCM0399 | CL128 | HCT 116 | C404(0.29); C581(0.63); C260(0.83); C261(0.83) | LDD0716 | [11] |
| LDCM0400 | CL13 | HCT 116 | C404(0.62); C261(0.73); C260(0.90); C218(0.91) | LDD0717 | [11] |
| LDCM0401 | CL14 | HCT 116 | C404(0.75); C581(0.77); C261(0.78); C260(0.85) | LDD0718 | [11] |
| LDCM0402 | CL15 | HCT 116 | C581(0.58); C218(0.68); C261(0.70); C404(0.73) | LDD0719 | [11] |
| LDCM0403 | CL16 | HCT 116 | C404(0.81); C581(0.92); C472(1.07); C260(1.16) | LDD0720 | [11] |
| LDCM0404 | CL17 | HCT 116 | C581(0.72); C260(0.87); C261(0.92); C472(1.09) | LDD0721 | [11] |
| LDCM0405 | CL18 | HCT 116 | C581(0.78); C260(0.80); C261(0.84); C472(1.05) | LDD0722 | [11] |
| LDCM0406 | CL19 | HCT 116 | C261(0.85); C404(0.86); C581(0.86); C260(0.89) | LDD0723 | [11] |
| LDCM0407 | CL2 | HCT 116 | C581(0.64); C261(0.99); C472(0.99); C452(1.05) | LDD0724 | [11] |
| LDCM0408 | CL20 | HCT 116 | C404(0.43); C452(0.73); C260(0.80); C261(0.80) | LDD0725 | [11] |
| LDCM0409 | CL21 | HCT 116 | C404(0.31); C260(0.69); C452(0.70); C261(0.74) | LDD0726 | [11] |
| LDCM0410 | CL22 | HCT 116 | C404(0.22); C261(0.86); C260(0.88); C452(0.89) | LDD0727 | [11] |
| LDCM0411 | CL23 | HCT 116 | C404(0.53); C261(0.90); C260(0.91); C472(1.03) | LDD0728 | [11] |
| LDCM0412 | CL24 | HCT 116 | C404(0.31); C261(0.71); C260(0.76); C452(0.83) | LDD0729 | [11] |
| LDCM0413 | CL25 | HCT 116 | C260(0.84); C261(0.82); C404(0.26); C452(0.95) | LDD0730 | [11] |
| LDCM0414 | CL26 | HCT 116 | C260(0.95); C261(0.93); C404(0.64); C452(1.20) | LDD0731 | [11] |
| LDCM0415 | CL27 | HCT 116 | C260(0.89); C261(0.89); C404(0.44); C452(1.06) | LDD0732 | [11] |
| LDCM0416 | CL28 | HCT 116 | C260(0.91); C261(0.75); C404(0.56); C452(0.92) | LDD0733 | [11] |
| LDCM0417 | CL29 | HCT 116 | C260(0.80); C261(0.81); C404(0.46); C452(0.85) | LDD0734 | [11] |
| LDCM0418 | CL3 | HCT 116 | C218(1.28); C260(0.97); C261(0.96); C404(0.98) | LDD0735 | [11] |
| LDCM0419 | CL30 | HCT 116 | C260(0.98); C261(0.89); C404(0.95); C452(1.06) | LDD0736 | [11] |
| LDCM0420 | CL31 | HCT 116 | C218(0.59); C260(0.85); C261(0.78); C404(1.09) | LDD0737 | [11] |
| LDCM0421 | CL32 | HCT 116 | C218(0.65); C260(0.89); C261(0.81); C404(1.48) | LDD0738 | [11] |
| LDCM0422 | CL33 | HCT 116 | C218(0.45); C260(0.83); C261(0.79); C404(1.17) | LDD0739 | [11] |
| LDCM0423 | CL34 | HCT 116 | C218(0.33); C260(0.73); C261(0.73); C404(0.44) | LDD0740 | [11] |
| LDCM0424 | CL35 | HCT 116 | C218(0.42); C260(0.75); C261(0.79); C404(0.54) | LDD0741 | [11] |
| LDCM0425 | CL36 | HCT 116 | C218(0.39); C260(0.92); C261(0.94); C404(0.50) | LDD0742 | [11] |
| LDCM0426 | CL37 | HCT 116 | C218(0.31); C260(0.71); C261(0.74); C404(0.44) | LDD0743 | [11] |
| LDCM0428 | CL39 | HCT 116 | C218(0.40); C260(0.72); C261(0.70); C404(0.35) | LDD0745 | [11] |
| LDCM0429 | CL4 | HCT 116 | C218(0.65); C260(0.91); C261(0.85); C404(0.88) | LDD0746 | [11] |
| LDCM0430 | CL40 | HCT 116 | C218(0.38); C260(0.90); C261(0.84); C404(0.58) | LDD0747 | [11] |
| LDCM0431 | CL41 | HCT 116 | C218(0.39); C260(0.84); C261(0.88); C404(0.64) | LDD0748 | [11] |
| LDCM0432 | CL42 | HCT 116 | C218(0.33); C260(0.86); C261(0.85); C404(0.45) | LDD0749 | [11] |
| LDCM0433 | CL43 | HCT 116 | C218(0.32); C260(0.64); C261(0.75); C404(0.47) | LDD0750 | [11] |
| LDCM0434 | CL44 | HCT 116 | C218(0.35); C260(0.82); C261(0.90); C404(0.76) | LDD0751 | [11] |
| LDCM0435 | CL45 | HCT 116 | C218(0.35); C260(0.75); C261(0.81); C404(0.38) | LDD0752 | [11] |
| LDCM0436 | CL46 | HCT 116 | C260(0.96); C261(1.04); C452(1.16); C472(1.09) | LDD0753 | [11] |
| LDCM0437 | CL47 | HCT 116 | C260(1.37); C261(1.16); C452(1.26); C472(1.13) | LDD0754 | [11] |
| LDCM0438 | CL48 | HCT 116 | C260(1.11); C261(1.10); C452(0.92); C472(0.77) | LDD0755 | [11] |
| LDCM0439 | CL49 | HCT 116 | C260(0.90); C261(0.94); C452(1.00); C472(0.93) | LDD0756 | [11] |
| LDCM0440 | CL5 | HCT 116 | C218(0.98); C260(0.96); C261(0.87); C404(0.77) | LDD0757 | [11] |
| LDCM0441 | CL50 | HCT 116 | C260(0.97); C261(0.99); C452(1.00); C472(0.96) | LDD0758 | [11] |
| LDCM0442 | CL51 | HCT 116 | C260(0.91); C261(0.96); C452(1.04); C472(0.99) | LDD0759 | [11] |
| LDCM0443 | CL52 | HCT 116 | C260(1.01); C261(0.94); C452(1.12); C472(1.09) | LDD0760 | [11] |
| LDCM0444 | CL53 | HCT 116 | C260(0.99); C261(0.96); C452(0.95); C472(0.96) | LDD0761 | [11] |
| LDCM0445 | CL54 | HCT 116 | C260(0.94); C261(1.02); C452(1.09); C472(1.07) | LDD0762 | [11] |
| LDCM0446 | CL55 | HCT 116 | C260(1.01); C261(1.06); C452(1.10); C472(1.00) | LDD0763 | [11] |
| LDCM0447 | CL56 | HCT 116 | C260(1.11); C261(1.11); C452(1.36); C472(1.21) | LDD0764 | [11] |
| LDCM0448 | CL57 | HCT 116 | C260(1.16); C261(0.99); C452(1.09); C472(0.99) | LDD0765 | [11] |
| LDCM0449 | CL58 | HCT 116 | C260(1.16); C261(0.97); C452(1.12); C472(0.99) | LDD0766 | [11] |
| LDCM0450 | CL59 | HCT 116 | C260(1.04); C261(1.02); C452(1.16); C472(1.06) | LDD0767 | [11] |
| LDCM0451 | CL6 | HCT 116 | C218(0.75); C260(0.80); C261(0.72); C404(0.82) | LDD0768 | [11] |
| LDCM0452 | CL60 | HCT 116 | C260(1.07); C261(0.98); C452(1.24); C472(1.12) | LDD0769 | [11] |
| LDCM0453 | CL61 | HCT 116 | C260(0.94); C261(0.97); C404(0.69); C452(0.93) | LDD0770 | [11] |
| LDCM0454 | CL62 | HCT 116 | C260(0.96); C261(0.98); C404(0.70); C452(1.06) | LDD0771 | [11] |
| LDCM0455 | CL63 | HCT 116 | C260(1.01); C261(1.02); C404(0.45); C452(1.06) | LDD0772 | [11] |
| LDCM0456 | CL64 | HCT 116 | C260(0.92); C261(0.94); C404(0.33); C452(0.91) | LDD0773 | [11] |
| LDCM0457 | CL65 | HCT 116 | C260(1.02); C261(1.03); C404(0.59); C452(1.08) | LDD0774 | [11] |
| LDCM0458 | CL66 | HCT 116 | C260(0.86); C261(0.88); C404(0.15); C452(0.83) | LDD0775 | [11] |
| LDCM0459 | CL67 | HCT 116 | C260(0.84); C261(0.85); C404(0.31); C452(0.89) | LDD0776 | [11] |
| LDCM0460 | CL68 | HCT 116 | C260(0.91); C261(0.92); C404(0.63); C452(1.00) | LDD0777 | [11] |
| LDCM0461 | CL69 | HCT 116 | C260(0.88); C261(0.89); C404(1.52); C452(0.95) | LDD0778 | [11] |
| LDCM0462 | CL7 | HCT 116 | C218(0.87); C260(0.81); C261(0.71); C404(0.34) | LDD0779 | [11] |
| LDCM0463 | CL70 | HCT 116 | C260(0.87); C261(0.88); C404(0.69); C452(1.04) | LDD0780 | [11] |
| LDCM0464 | CL71 | HCT 116 | C260(1.08); C261(1.09); C404(0.46); C452(1.03) | LDD0781 | [11] |
| LDCM0465 | CL72 | HCT 116 | C260(0.98); C261(0.99); C404(0.88); C452(1.04) | LDD0782 | [11] |
| LDCM0466 | CL73 | HCT 116 | C260(0.98); C261(1.02); C404(0.89); C452(1.05) | LDD0783 | [11] |
| LDCM0467 | CL74 | HCT 116 | C260(0.87); C261(0.86); C404(0.61); C452(0.90) | LDD0784 | [11] |
| LDCM0469 | CL76 | HCT 116 | C260(0.96); C261(1.05); C404(1.06); C452(1.05) | LDD0786 | [11] |
| LDCM0470 | CL77 | HCT 116 | C260(1.16); C261(1.11); C404(1.17); C452(1.11) | LDD0787 | [11] |
| LDCM0471 | CL78 | HCT 116 | C260(1.08); C261(0.94); C404(0.72); C452(1.16) | LDD0788 | [11] |
| LDCM0472 | CL79 | HCT 116 | C260(0.97); C261(0.96); C404(0.34); C452(0.85) | LDD0789 | [11] |
| LDCM0473 | CL8 | HCT 116 | C218(0.57); C260(0.67); C261(0.65); C404(0.31) | LDD0790 | [11] |
| LDCM0474 | CL80 | HCT 116 | C260(0.93); C261(0.98); C404(1.35); C452(1.02) | LDD0791 | [11] |
| LDCM0475 | CL81 | HCT 116 | C260(1.00); C261(0.94); C404(0.60); C452(1.05) | LDD0792 | [11] |
| LDCM0476 | CL82 | HCT 116 | C260(0.99); C261(0.95); C404(0.24); C452(0.92) | LDD0793 | [11] |
| LDCM0477 | CL83 | HCT 116 | C260(0.86); C261(0.91); C404(0.24); C452(0.85) | LDD0794 | [11] |
| LDCM0478 | CL84 | HCT 116 | C260(0.80); C261(0.83); C404(0.24); C452(0.86) | LDD0795 | [11] |
| LDCM0479 | CL85 | HCT 116 | C260(0.96); C261(0.97); C404(0.48); C452(1.16) | LDD0796 | [11] |
| LDCM0480 | CL86 | HCT 116 | C260(1.17); C261(1.09); C404(0.88); C452(1.64) | LDD0797 | [11] |
| LDCM0481 | CL87 | HCT 116 | C260(1.01); C261(1.14); C404(0.45); C452(1.08) | LDD0798 | [11] |
| LDCM0482 | CL88 | HCT 116 | C260(0.89); C261(0.91); C404(0.24); C452(0.89) | LDD0799 | [11] |
| LDCM0483 | CL89 | HCT 116 | C260(0.86); C261(0.82); C404(0.28); C452(1.09) | LDD0800 | [11] |
| LDCM0484 | CL9 | HCT 116 | C218(1.30); C260(0.84); C261(0.75); C404(0.59) | LDD0801 | [11] |
| LDCM0485 | CL90 | HCT 116 | C260(0.93); C261(0.93); C404(0.86); C452(0.91) | LDD0802 | [11] |
| LDCM0486 | CL91 | HCT 116 | C260(1.28); C261(1.00); C452(1.23); C472(1.27) | LDD0803 | [11] |
| LDCM0487 | CL92 | HCT 116 | C260(1.17); C261(0.97); C452(1.13); C472(1.12) | LDD0804 | [11] |
| LDCM0488 | CL93 | HCT 116 | C260(1.15); C261(1.08); C452(1.15); C472(1.14) | LDD0805 | [11] |
| LDCM0489 | CL94 | HCT 116 | C260(1.12); C261(0.90); C452(0.99); C472(1.20) | LDD0806 | [11] |
| LDCM0490 | CL95 | HCT 116 | C260(1.04); C261(1.04); C452(1.16); C472(1.33) | LDD0807 | [11] |
| LDCM0491 | CL96 | HCT 116 | C260(1.08); C261(1.09); C452(1.21); C472(1.26) | LDD0808 | [11] |
| LDCM0492 | CL97 | HCT 116 | C260(0.96); C261(0.93); C452(1.30); C472(1.33) | LDD0809 | [11] |
| LDCM0493 | CL98 | HCT 116 | C260(0.97); C261(0.95); C452(1.26); C472(1.40) | LDD0810 | [11] |
| LDCM0494 | CL99 | HCT 116 | C260(1.29); C261(1.11); C452(1.38); C472(1.38) | LDD0811 | [11] |
| LDCM0634 | CY-0357 | Hep-G2 | C581(2.15); C472(0.52) | LDD2228 | [20] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C472(0.94); C581(0.90) | LDD1702 | [38] |
| LDCM0625 | F8 | Ramos | C452(1.01); 1.19; C581(0.85); C404(0.32) | LDD2187 | [39] |
| LDCM0572 | Fragment10 | Ramos | C452(0.89); 0.94; C581(0.54); C404(0.38) | LDD2189 | [39] |
| LDCM0573 | Fragment11 | Ramos | C452(1.18); 20.00; C581(0.02); C404(20.00) | LDD2190 | [39] |
| LDCM0574 | Fragment12 | Ramos | C452(0.74); 1.09; C581(0.63); C404(0.46) | LDD2191 | [39] |
| LDCM0575 | Fragment13 | Ramos | C452(1.12); 1.39; C581(3.16); C404(0.97) | LDD2192 | [39] |
| LDCM0576 | Fragment14 | Ramos | C452(0.74); 0.77; C581(0.75); C404(0.58) | LDD2193 | [39] |
| LDCM0579 | Fragment20 | Ramos | C452(0.69); 1.03; C581(0.31); C404(0.55) | LDD2194 | [39] |
| LDCM0580 | Fragment21 | Ramos | C452(1.05); 1.40; C581(0.81); C404(0.78) | LDD2195 | [39] |
| LDCM0582 | Fragment23 | Ramos | C452(1.25); 0.31; C581(0.61); C404(0.86) | LDD2196 | [39] |
| LDCM0578 | Fragment27 | Ramos | C452(1.14); 1.72; C581(0.79); C404(0.67) | LDD2197 | [39] |
| LDCM0586 | Fragment28 | Ramos | C452(0.63); 0.34; C581(0.80); C472(0.98) | LDD2198 | [39] |
| LDCM0588 | Fragment30 | Ramos | C452(1.47); 1.36; C581(0.90); C404(0.59) | LDD2199 | [39] |
| LDCM0589 | Fragment31 | Ramos | C452(1.23); 1.76; C581(0.82); C472(0.37) | LDD2200 | [39] |
| LDCM0590 | Fragment32 | Ramos | C452(0.71); 0.72; C581(0.57); C404(0.67) | LDD2201 | [39] |
| LDCM0468 | Fragment33 | HCT 116 | C260(1.05); C261(1.07); C404(0.52); C452(0.98) | LDD0785 | [11] |
| LDCM0596 | Fragment38 | Ramos | C452(0.98); 1.08; C581(0.52); C404(1.15) | LDD2203 | [39] |
| LDCM0566 | Fragment4 | Ramos | C452(0.89); 0.81; C581(0.75); C404(0.59) | LDD2184 | [39] |
| LDCM0427 | Fragment51 | HCT 116 | C218(0.29); C260(0.67); C261(0.75); C404(0.36) | LDD0744 | [11] |
| LDCM0610 | Fragment52 | Ramos | C452(1.72); 1.83; C581(1.89); C404(0.46) | LDD2204 | [39] |
| LDCM0614 | Fragment56 | Ramos | C452(1.57); 1.90; C581(1.20); C404(0.39) | LDD2205 | [39] |
| LDCM0569 | Fragment7 | Ramos | C452(0.64); 0.89; C581(0.45); C404(0.42) | LDD2186 | [39] |
| LDCM0571 | Fragment9 | Ramos | C452(0.67); 1.26; C581(0.42); C404(0.45) | LDD2188 | [39] |
| LDCM0116 | HHS-0101 | DM93 | Y47(0.26) | LDD0264 | [10] |
| LDCM0118 | HHS-0301 | DM93 | Y47(0.05) | LDD0266 | [10] |
| LDCM0119 | HHS-0401 | DM93 | Y47(0.05) | LDD0267 | [10] |
| LDCM0120 | HHS-0701 | DM93 | Y47(0.06) | LDD0268 | [10] |
| LDCM0107 | IAA | HeLa | H96(0.00); H70(0.00); H564(0.00); C404(0.00) | LDD0221 | [27] |
| LDCM0022 | KB02 | HCT 116 | C261(1.60); C260(1.55); C472(1.73); C581(1.06) | LDD0080 | [11] |
| LDCM0023 | KB03 | HCT 116 | C261(1.98); C260(1.93); C472(1.12); C581(0.86) | LDD0081 | [11] |
| LDCM0024 | KB05 | HCT 116 | C261(3.27); C260(3.26); C472(1.12); C581(1.07) | LDD0082 | [11] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C472(1.12); C581(1.06) | LDD2102 | [38] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C472(0.79); C452(0.38); C581(0.81) | LDD2121 | [38] |
| LDCM0109 | NEM | HeLa | H96(0.00); H564(0.00); H70(0.00); H105(0.00) | LDD0223 | [27] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C472(0.92); C452(1.18); C581(0.97) | LDD2089 | [38] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C472(1.08) | LDD2090 | [38] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C472(1.09); C581(0.91) | LDD2092 | [38] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C452(0.84); C581(1.74) | LDD2093 | [38] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C581(1.23) | LDD2094 | [38] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C452(0.33) | LDD2096 | [38] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C581(0.79) | LDD2097 | [38] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C472(0.98); C581(0.98) | LDD2098 | [38] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C472(0.99); C261(1.81) | LDD2099 | [38] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C472(1.03); C452(0.23); C581(0.45) | LDD2100 | [38] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C581(0.97); C261(1.31) | LDD2101 | [38] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C472(1.03); C452(1.05); C581(0.70) | LDD2104 | [38] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C472(1.34); C581(1.05) | LDD2105 | [38] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C472(0.85) | LDD2106 | [38] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C472(0.89); C452(1.07); C581(1.11); C260(1.17) | LDD2107 | [38] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C472(0.92); C581(1.10) | LDD2108 | [38] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C452(0.50); C581(0.97); C261(0.87) | LDD2109 | [38] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C472(2.28) | LDD2110 | [38] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C452(0.71) | LDD2111 | [38] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C472(0.75); C581(0.52) | LDD2114 | [38] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C261(0.62); C260(0.87) | LDD2115 | [38] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C452(0.57); C581(0.74) | LDD2116 | [38] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C452(0.49) | LDD2118 | [38] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C452(2.48); C581(1.21) | LDD2119 | [38] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C452(0.84); C581(0.79) | LDD2120 | [38] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C472(0.97); C452(0.60); C581(1.05); C261(1.11) | LDD2123 | [38] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C472(0.97); C581(1.27); C261(1.11) | LDD2125 | [38] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C452(0.34) | LDD2126 | [38] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C452(0.68); C581(1.08) | LDD2127 | [38] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C472(1.14); C452(0.86) | LDD2129 | [38] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C452(1.14); C581(0.65) | LDD2133 | [38] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C452(0.57); C581(0.59); C261(0.77) | LDD2134 | [38] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C472(1.14); C581(1.39) | LDD2135 | [38] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C472(1.24); C452(0.88); C581(1.60); C261(1.39) | LDD2136 | [38] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C472(1.02); C452(0.80); C581(0.90); C261(1.00) | LDD2137 | [38] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C581(3.33); C452(1.35) | LDD1700 | [38] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C472(1.02); C452(0.53); C581(1.38); C261(1.22) | LDD2140 | [38] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C472(0.86); C452(0.37); C581(1.06) | LDD2141 | [38] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C452(0.90) | LDD2143 | [38] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C452(2.84); C581(1.86) | LDD2144 | [38] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C472(1.97); C581(1.35) | LDD2146 | [38] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C452(0.49); C581(0.57); C261(0.67) | LDD2148 | [38] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C452(0.37) | LDD2149 | [38] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C472(0.58); C581(0.54) | LDD2150 | [38] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C472(1.71); C452(0.88) | LDD2153 | [38] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C472(1.33); C472(0.88) | LDD2206 | [40] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C472(0.88); C581(0.71); C472(0.56) | LDD2207 | [40] |
| LDCM0131 | RA190 | MM1.R | C404(1.44); C581(1.26); C260(1.25); C261(1.25) | LDD0304 | [41] |
| LDCM0021 | THZ1 | HCT 116 | C404(1.38); C452(1.14); C472(1.07); C260(1.33) | LDD2173 | [11] |
| LDCM0111 | W14 | Hep-G2 | C452(0.89) | LDD0238 | [28] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| von Hippel-Lindau disease tumor suppressor (VHL) | VHL family | P40337 | |||
Other
References
