Details of the Target
General Information of Target
| Target ID | LDTP02595 | |||||
|---|---|---|---|---|---|---|
| Target Name | Polyadenylate-binding protein 1 (PABPC1) | |||||
| Gene Name | PABPC1 | |||||
| Gene ID | 26986 | |||||
| Synonyms |
PAB1; PABP; PABP1; PABPC2; Polyadenylate-binding protein 1; PABP-1; Poly(A)-binding protein 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MNPSAPSYPMASLYVGDLHPDVTEAMLYEKFSPAGPILSIRVCRDMITRRSLGYAYVNFQ
QPADAERALDTMNFDVIKGKPVRIMWSQRDPSLRKSGVGNIFIKNLDKSIDNKALYDTFS AFGNILSCKVVCDENGSKGYGFVHFETQEAAERAIEKMNGMLLNDRKVFVGRFKSRKERE AELGARAKEFTNVYIKNFGEDMDDERLKDLFGKFGPALSVKVMTDESGKSKGFGFVSFER HEDAQKAVDEMNGKELNGKQIYVGRAQKKVERQTELKRKFEQMKQDRITRYQGVNLYVKN LDDGIDDERLRKEFSPFGTITSAKVMMEGGRSKGFGFVCFSSPEEATKAVTEMNGRIVAT KPLYVALAQRKEERQAHLTNQYMQRMASVRAVPNPVINPYQPAPPSGYFMAAIPQTQNRA AYYPPSQIAQLRPSPRWTAQGARPHPFQNMPGAIRPAAPRPPFSTMRPASSQVPRVMSTQ RVANTSTQTMGPRPAAAAAAATPAVRTVPQYKYAAGVRNPQQHLNAQPQVTMQQPAVHVQ GQEPLTASMLASAPPQEQKQMLGERLFPLIQAMHPTLAGKITGMLLEIDNSELLHMLESP ESLRSKVDEAVAVLQAHQAKEAAQKAVNSATGVPTV |
|||||
| Target Type |
Clinical trial
|
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Polyadenylate-binding protein type-1 family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Binds the poly(A) tail of mRNA, including that of its own transcript, and regulates processes of mRNA metabolism such as pre-mRNA splicing and mRNA stability. Its function in translational initiation regulation can either be enhanced by PAIP1 or repressed by PAIP2. Can probably bind to cytoplasmic RNA sequences other than poly(A) in vivo. Binds to N6-methyladenosine (m6A)-containing mRNAs and contributes to MYC stability by binding to m6A-containing MYC mRNAs. Involved in translationally coupled mRNA turnover. Implicated with other RNA-binding proteins in the cytoplasmic deadenylation/translational and decay interplay of the FOS mRNA mediated by the major coding-region determinant of instability (mCRD) domain. Involved in regulation of nonsense-mediated decay (NMD) of mRNAs containing premature stop codons; for the recognition of premature termination codons (PTC) and initiation of NMD a competitive interaction between UPF1 and PABPC1 with the ribosome-bound release factors is proposed. By binding to long poly(A) tails, may protect them from uridylation by ZCCHC6/ZCCHC11 and hence contribute to mRNA stability.; (Microbial infection) Positively regulates the replication of dengue virus (DENV).
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P1 Probe Info |
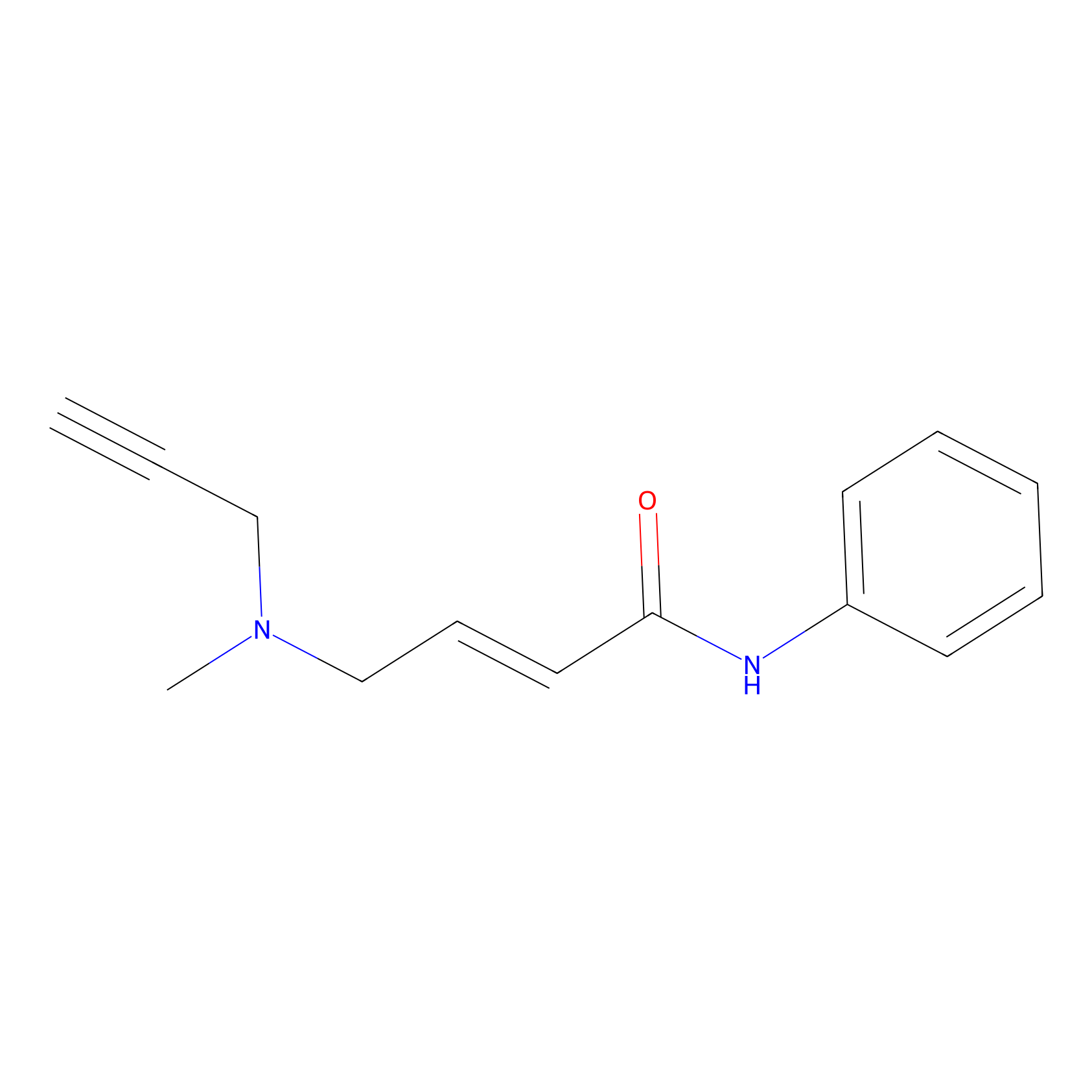 |
10.00 | LDD0452 | [1] | |
|
P2 Probe Info |
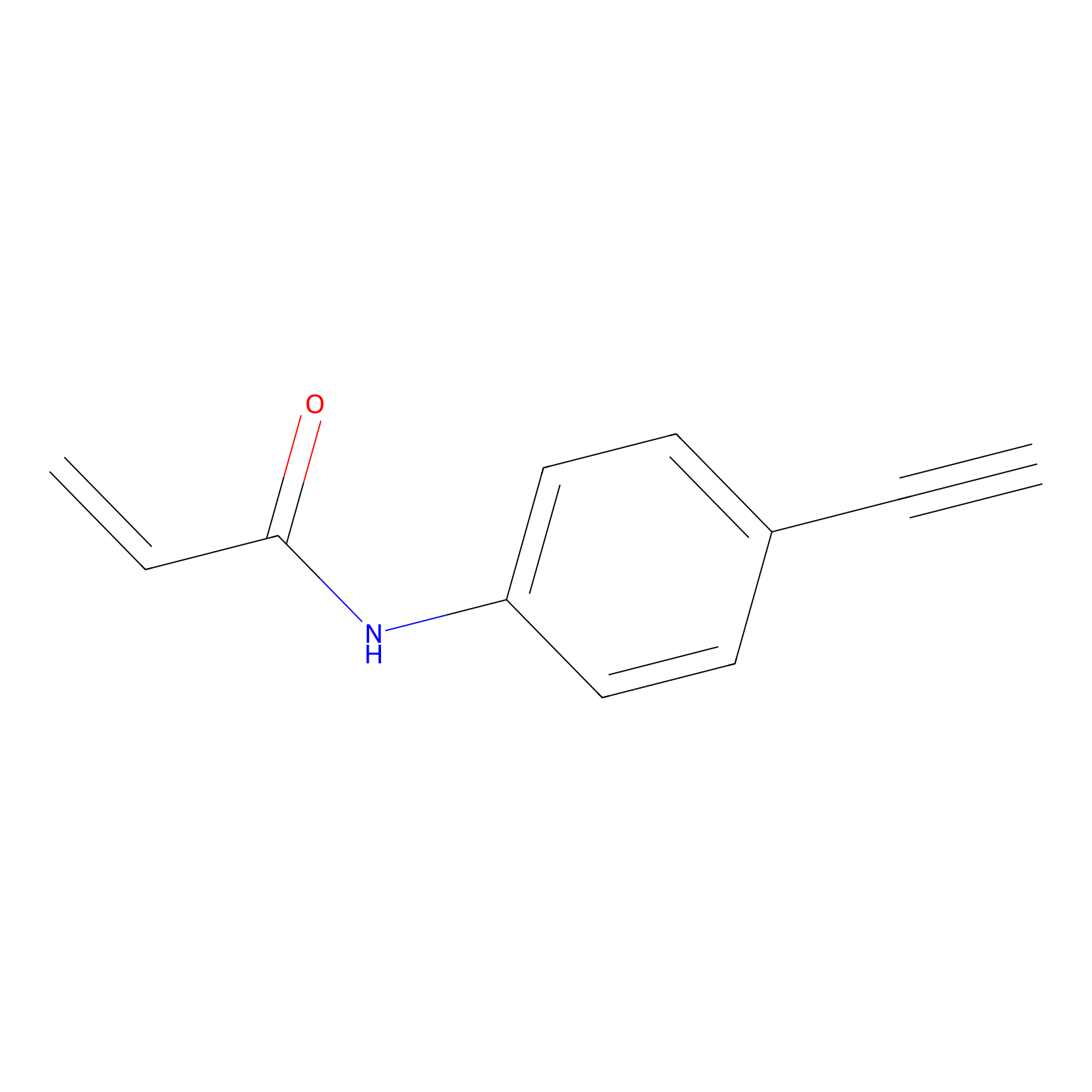 |
10.00 | LDD0449 | [1] | |
|
P3 Probe Info |
 |
10.00 | LDD0450 | [1] | |
|
P8 Probe Info |
 |
3.64 | LDD0451 | [1] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [2] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [2] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
TH211 Probe Info |
 |
Y364(17.49); Y297(11.61); Y291(9.30); Y382(6.91) | LDD0257 | [3] | |
|
TH214 Probe Info |
 |
Y364(19.01); Y291(5.92) | LDD0258 | [3] | |
|
TH216 Probe Info |
 |
Y364(20.00); Y140(13.17); Y382(9.17) | LDD0259 | [3] | |
|
AZ-9 Probe Info |
 |
E66(0.94); E239(10.00); D165(10.00); Q61(0.94) | LDD2208 | [4] | |
|
ONAyne Probe Info |
 |
K512(0.00); K620(0.00); K312(0.00); K606(0.00) | LDD0273 | [5] | |
|
OPA-S-S-alkyne Probe Info |
 |
K157(1.38); K231(1.77); K512(3.06) | LDD3494 | [6] | |
|
Probe 1 Probe Info |
 |
Y56(6.46); Y140(9.07); Y194(12.40); Y291(42.43) | LDD3495 | [7] | |
|
THZ1-DTB Probe Info |
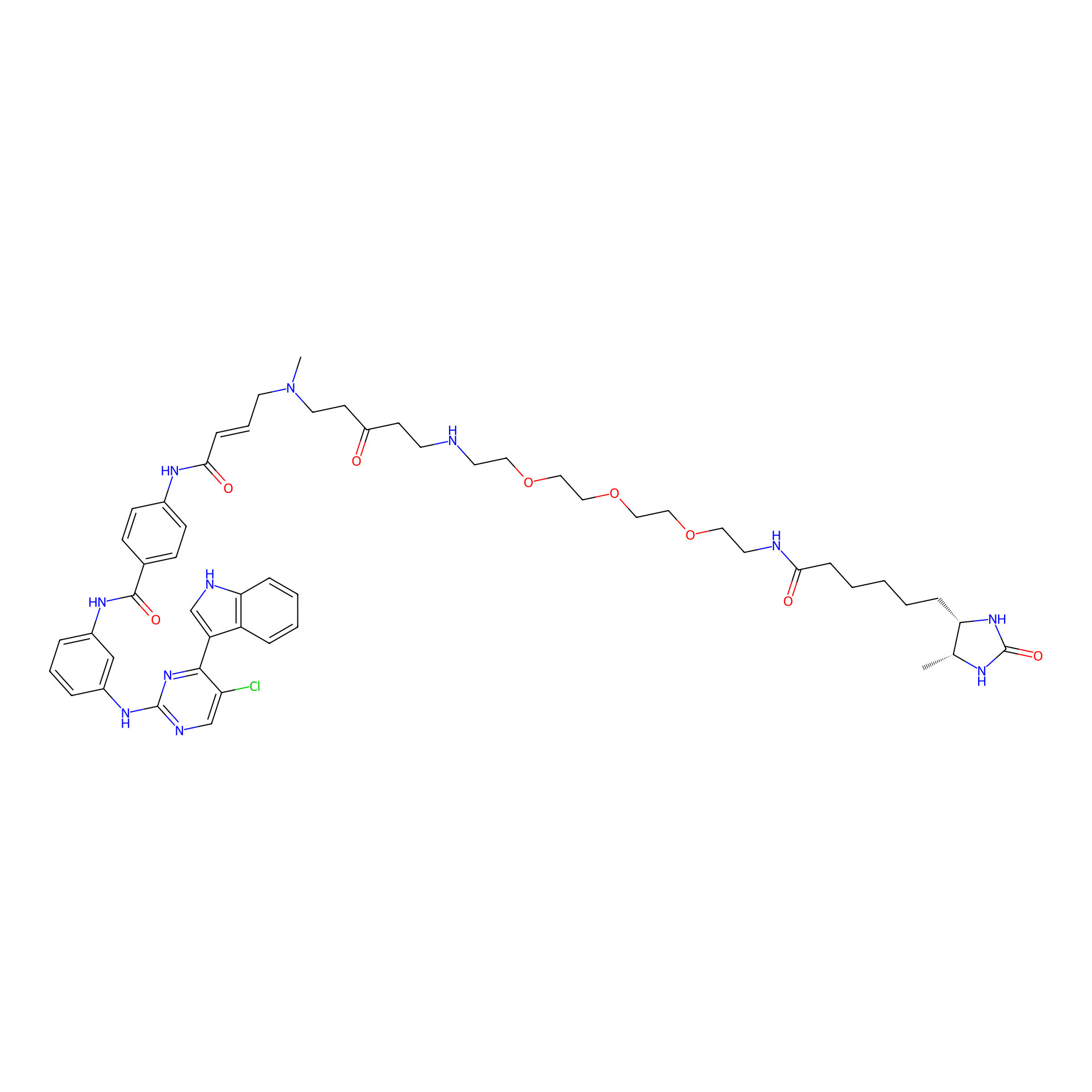 |
C128(1.08) | LDD0460 | [8] | |
|
BTD Probe Info |
 |
C132(1.88) | LDD1700 | [9] | |
|
AHL-Pu-1 Probe Info |
 |
C339(2.44) | LDD0169 | [10] | |
|
HHS-482 Probe Info |
 |
Y291(0.89); Y297(0.94); Y364(0.93); Y382(0.88) | LDD0285 | [11] | |
|
HHS-475 Probe Info |
 |
Y297(0.70); Y291(0.78); Y140(0.81); Y511(0.84) | LDD0264 | [12] | |
|
HHS-465 Probe Info |
 |
Y140(6.93); Y291(8.36); Y297(8.02); Y382(7.90) | LDD2237 | [13] | |
|
DBIA Probe Info |
 |
C132(0.83) | LDD1507 | [14] | |
|
5E-2FA Probe Info |
 |
H241(0.00); H595(0.00); H574(0.00); H377(0.00) | LDD2235 | [15] | |
|
ATP probe Probe Info |
 |
K188(0.00); K259(0.00); K138(0.00); K108(0.00) | LDD0199 | [16] | |
|
m-APA Probe Info |
 |
H595(0.00); H574(0.00); H377(0.00); H617(0.00) | LDD2231 | [15] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C128(0.00); C339(0.00); C132(0.00) | LDD0038 | [17] | |
|
IA-alkyne Probe Info |
 |
C132(0.00); C128(0.00); C339(0.00) | LDD0032 | [18] | |
|
IPIAA_H Probe Info |
 |
C339(0.00); C132(0.00) | LDD0030 | [19] | |
|
IPIAA_L Probe Info |
 |
C132(0.00); C339(0.00) | LDD0031 | [19] | |
|
Lodoacetamide azide Probe Info |
 |
C128(0.00); C339(0.00); C132(0.00) | LDD0037 | [17] | |
|
ATP probe Probe Info |
 |
K324(0.00); K606(0.00); K246(0.00); K312(0.00) | LDD0035 | [20] | |
|
IPM Probe Info |
 |
C339(0.00); C132(0.00) | LDD0025 | [21] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [21] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [22] | |
|
Compound 10 Probe Info |
 |
C132(0.00); C339(0.00) | LDD2216 | [23] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [23] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [22] | |
|
NHS Probe Info |
 |
K284(0.00); K512(0.00); K188(0.00); K259(0.00) | LDD0010 | [22] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0152 | [24] | |
|
SF Probe Info |
 |
K104(0.00); K312(0.00); K196(0.00); Y56(0.00) | LDD0028 | [25] | |
|
STPyne Probe Info |
 |
K620(0.00); K512(0.00) | LDD0009 | [22] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [22] | |
|
1c-yne Probe Info |
 |
K620(0.00); K312(0.00); K606(0.00); K361(0.00) | LDD0228 | [26] | |
|
Acrolein Probe Info |
 |
C128(0.00); H144(0.00) | LDD0217 | [27] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [27] | |
|
W1 Probe Info |
 |
C132(0.00); C339(0.00) | LDD0236 | [28] | |
|
NAIA_5 Probe Info |
 |
C128(0.00); C339(0.00); C43(0.00); C132(0.00) | LDD2223 | [29] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
7.19 | LDD0471 | [30] | |
|
FFF probe13 Probe Info |
 |
11.31 | LDD0475 | [30] | |
|
FFF probe14 Probe Info |
 |
9.00 | LDD0477 | [30] | |
|
FFF probe2 Probe Info |
 |
6.54 | LDD0463 | [30] | |
|
FFF probe3 Probe Info |
 |
6.17 | LDD0464 | [30] | |
|
JN0003 Probe Info |
 |
7.10 | LDD0469 | [30] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [31] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [32] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [33] | |
|
OEA-DA Probe Info |
 |
4.29 | LDD0046 | [34] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0068 | [35] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C132(0.73); C339(0.35) | LDD2142 | [9] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C132(0.96); C339(0.34) | LDD2112 | [9] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C132(0.85) | LDD2095 | [9] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C132(0.75) | LDD2130 | [9] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C132(1.04) | LDD2117 | [9] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C132(1.70) | LDD2152 | [9] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C132(0.63) | LDD2131 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C339(2.44) | LDD0169 | [10] |
| LDCM0562 | Abegg_cp(-)-11 | HeLa | C128(2.01) | LDD0313 | [21] |
| LDCM0214 | AC1 | HEK-293T | C132(0.83) | LDD1507 | [14] |
| LDCM0215 | AC10 | HEK-293T | C132(0.84) | LDD1508 | [14] |
| LDCM0226 | AC11 | HEK-293T | C132(0.84) | LDD1509 | [14] |
| LDCM0237 | AC12 | HEK-293T | C132(0.99) | LDD1510 | [14] |
| LDCM0259 | AC14 | HEK-293T | C132(0.89) | LDD1512 | [14] |
| LDCM0270 | AC15 | HEK-293T | C132(0.89) | LDD1513 | [14] |
| LDCM0276 | AC17 | HEK-293T | C132(0.92) | LDD1515 | [14] |
| LDCM0277 | AC18 | HEK-293T | C132(0.93) | LDD1516 | [14] |
| LDCM0278 | AC19 | HEK-293T | C132(0.78) | LDD1517 | [14] |
| LDCM0279 | AC2 | HEK-293T | C132(0.87) | LDD1518 | [14] |
| LDCM0280 | AC20 | HEK-293T | C132(0.97) | LDD1519 | [14] |
| LDCM0281 | AC21 | HEK-293T | C132(0.97) | LDD1520 | [14] |
| LDCM0282 | AC22 | HEK-293T | C132(0.96) | LDD1521 | [14] |
| LDCM0283 | AC23 | HEK-293T | C132(0.98) | LDD1522 | [14] |
| LDCM0284 | AC24 | HEK-293T | C132(0.93) | LDD1523 | [14] |
| LDCM0285 | AC25 | HEK-293T | C132(0.94) | LDD1524 | [14] |
| LDCM0286 | AC26 | HEK-293T | C132(0.95) | LDD1525 | [14] |
| LDCM0287 | AC27 | HEK-293T | C132(0.90) | LDD1526 | [14] |
| LDCM0288 | AC28 | HEK-293T | C132(0.95) | LDD1527 | [14] |
| LDCM0289 | AC29 | HEK-293T | C132(0.97) | LDD1528 | [14] |
| LDCM0290 | AC3 | HEK-293T | C132(0.83) | LDD1529 | [14] |
| LDCM0291 | AC30 | HEK-293T | C132(0.96) | LDD1530 | [14] |
| LDCM0292 | AC31 | HEK-293T | C132(0.98) | LDD1531 | [14] |
| LDCM0293 | AC32 | HEK-293T | C132(0.93) | LDD1532 | [14] |
| LDCM0294 | AC33 | HEK-293T | C132(0.81) | LDD1533 | [14] |
| LDCM0295 | AC34 | HEK-293T | C132(0.82) | LDD1534 | [14] |
| LDCM0296 | AC35 | HEK-293T | C132(0.80) | LDD1535 | [14] |
| LDCM0297 | AC36 | HEK-293T | C132(0.94) | LDD1536 | [14] |
| LDCM0298 | AC37 | HEK-293T | C132(0.93) | LDD1537 | [14] |
| LDCM0299 | AC38 | HEK-293T | C132(0.86) | LDD1538 | [14] |
| LDCM0300 | AC39 | HEK-293T | C132(0.85) | LDD1539 | [14] |
| LDCM0301 | AC4 | HEK-293T | C132(0.96) | LDD1540 | [14] |
| LDCM0302 | AC40 | HEK-293T | C132(0.77) | LDD1541 | [14] |
| LDCM0303 | AC41 | HEK-293T | C132(0.83) | LDD1542 | [14] |
| LDCM0304 | AC42 | HEK-293T | C132(0.82) | LDD1543 | [14] |
| LDCM0305 | AC43 | HEK-293T | C132(0.79) | LDD1544 | [14] |
| LDCM0306 | AC44 | HEK-293T | C132(0.98) | LDD1545 | [14] |
| LDCM0307 | AC45 | HEK-293T | C132(0.92) | LDD1546 | [14] |
| LDCM0308 | AC46 | HEK-293T | C132(0.86) | LDD1547 | [14] |
| LDCM0309 | AC47 | HEK-293T | C132(0.88) | LDD1548 | [14] |
| LDCM0310 | AC48 | HEK-293T | C132(0.79) | LDD1549 | [14] |
| LDCM0311 | AC49 | HEK-293T | C132(0.84) | LDD1550 | [14] |
| LDCM0312 | AC5 | HEK-293T | C132(0.95) | LDD1551 | [14] |
| LDCM0313 | AC50 | HEK-293T | C132(0.87) | LDD1552 | [14] |
| LDCM0314 | AC51 | HEK-293T | C132(0.90) | LDD1553 | [14] |
| LDCM0315 | AC52 | HEK-293T | C132(0.97) | LDD1554 | [14] |
| LDCM0316 | AC53 | HEK-293T | C132(0.97) | LDD1555 | [14] |
| LDCM0317 | AC54 | HEK-293T | C132(0.90) | LDD1556 | [14] |
| LDCM0318 | AC55 | HEK-293T | C132(0.85) | LDD1557 | [14] |
| LDCM0319 | AC56 | HEK-293T | C132(0.84) | LDD1558 | [14] |
| LDCM0320 | AC57 | HEK-293T | C132(0.86) | LDD1559 | [14] |
| LDCM0321 | AC58 | HEK-293T | C132(0.88) | LDD1560 | [14] |
| LDCM0322 | AC59 | HEK-293T | C132(0.89) | LDD1561 | [14] |
| LDCM0323 | AC6 | HEK-293T | C132(0.87) | LDD1562 | [14] |
| LDCM0324 | AC60 | HEK-293T | C132(0.96) | LDD1563 | [14] |
| LDCM0325 | AC61 | HEK-293T | C132(0.95) | LDD1564 | [14] |
| LDCM0326 | AC62 | HEK-293T | C132(0.93) | LDD1565 | [14] |
| LDCM0327 | AC63 | HEK-293T | C132(0.90) | LDD1566 | [14] |
| LDCM0328 | AC64 | HEK-293T | C132(0.84) | LDD1567 | [14] |
| LDCM0334 | AC7 | HEK-293T | C132(0.92) | LDD1568 | [14] |
| LDCM0345 | AC8 | HEK-293T | C132(0.80) | LDD1569 | [14] |
| LDCM0545 | Acetamide | MDA-MB-231 | C132(0.87) | LDD2138 | [9] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C132(0.63) | LDD2113 | [9] |
| LDCM0248 | AKOS034007472 | HEK-293T | C132(0.92) | LDD1511 | [14] |
| LDCM0356 | AKOS034007680 | HEK-293T | C132(0.86) | LDD1570 | [14] |
| LDCM0275 | AKOS034007705 | HEK-293T | C132(0.79) | LDD1514 | [14] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C132(1.01) | LDD2091 | [9] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [27] |
| LDCM0632 | CL-Sc | Hep-G2 | C339(3.54); C132(1.53); C132(1.30); C339(0.85) | LDD2227 | [29] |
| LDCM0367 | CL1 | HEK-293T | C132(0.79) | LDD1571 | [14] |
| LDCM0368 | CL10 | HEK-293T | C132(0.67) | LDD1572 | [14] |
| LDCM0369 | CL100 | HEK-293T | C132(0.79) | LDD1573 | [14] |
| LDCM0370 | CL101 | HEK-293T | C132(0.89) | LDD1574 | [14] |
| LDCM0371 | CL102 | HEK-293T | C132(0.80) | LDD1575 | [14] |
| LDCM0372 | CL103 | HEK-293T | C132(0.88) | LDD1576 | [14] |
| LDCM0373 | CL104 | HEK-293T | C132(0.80) | LDD1577 | [14] |
| LDCM0374 | CL105 | HEK-293T | C132(0.97) | LDD1578 | [14] |
| LDCM0375 | CL106 | HEK-293T | C132(0.87) | LDD1579 | [14] |
| LDCM0376 | CL107 | HEK-293T | C132(0.95) | LDD1580 | [14] |
| LDCM0377 | CL108 | HEK-293T | C132(0.88) | LDD1581 | [14] |
| LDCM0378 | CL109 | HEK-293T | C132(0.92) | LDD1582 | [14] |
| LDCM0379 | CL11 | HEK-293T | C132(0.81) | LDD1583 | [14] |
| LDCM0380 | CL110 | HEK-293T | C132(0.88) | LDD1584 | [14] |
| LDCM0381 | CL111 | HEK-293T | C132(0.93) | LDD1585 | [14] |
| LDCM0382 | CL112 | HEK-293T | C132(1.29) | LDD1586 | [14] |
| LDCM0383 | CL113 | HEK-293T | C132(0.88) | LDD1587 | [14] |
| LDCM0384 | CL114 | HEK-293T | C132(0.80) | LDD1588 | [14] |
| LDCM0385 | CL115 | HEK-293T | C132(0.87) | LDD1589 | [14] |
| LDCM0386 | CL116 | HEK-293T | C132(0.75) | LDD1590 | [14] |
| LDCM0387 | CL117 | HEK-293T | C132(0.89) | LDD1591 | [14] |
| LDCM0388 | CL118 | HEK-293T | C132(0.80) | LDD1592 | [14] |
| LDCM0389 | CL119 | HEK-293T | C132(0.90) | LDD1593 | [14] |
| LDCM0390 | CL12 | HEK-293T | C132(0.71) | LDD1594 | [14] |
| LDCM0391 | CL120 | HEK-293T | C132(0.77) | LDD1595 | [14] |
| LDCM0392 | CL121 | HEK-293T | C132(0.88) | LDD1596 | [14] |
| LDCM0393 | CL122 | HEK-293T | C132(0.83) | LDD1597 | [14] |
| LDCM0394 | CL123 | HEK-293T | C132(0.76) | LDD1598 | [14] |
| LDCM0395 | CL124 | HEK-293T | C132(0.72) | LDD1599 | [14] |
| LDCM0396 | CL125 | HEK-293T | C132(0.98) | LDD1600 | [14] |
| LDCM0397 | CL126 | HEK-293T | C132(0.85) | LDD1601 | [14] |
| LDCM0398 | CL127 | HEK-293T | C132(0.93) | LDD1602 | [14] |
| LDCM0399 | CL128 | HEK-293T | C132(0.82) | LDD1603 | [14] |
| LDCM0400 | CL13 | HEK-293T | C132(0.77) | LDD1604 | [14] |
| LDCM0401 | CL14 | HEK-293T | C132(0.75) | LDD1605 | [14] |
| LDCM0402 | CL15 | HEK-293T | C132(0.67) | LDD1606 | [14] |
| LDCM0403 | CL16 | HEK-293T | C132(0.67) | LDD1607 | [14] |
| LDCM0404 | CL17 | HEK-293T | C132(0.66) | LDD1608 | [14] |
| LDCM0405 | CL18 | HEK-293T | C132(0.78) | LDD1609 | [14] |
| LDCM0406 | CL19 | HEK-293T | C132(0.81) | LDD1610 | [14] |
| LDCM0407 | CL2 | HEK-293T | C132(0.77) | LDD1611 | [14] |
| LDCM0408 | CL20 | HEK-293T | C132(0.92) | LDD1612 | [14] |
| LDCM0409 | CL21 | HEK-293T | C132(0.77) | LDD1613 | [14] |
| LDCM0410 | CL22 | HEK-293T | C132(0.82) | LDD1614 | [14] |
| LDCM0411 | CL23 | HEK-293T | C132(0.82) | LDD1615 | [14] |
| LDCM0412 | CL24 | HEK-293T | C132(0.76) | LDD1616 | [14] |
| LDCM0413 | CL25 | HEK-293T | C132(0.83) | LDD1617 | [14] |
| LDCM0414 | CL26 | HEK-293T | C132(0.84) | LDD1618 | [14] |
| LDCM0415 | CL27 | HEK-293T | C132(0.89) | LDD1619 | [14] |
| LDCM0416 | CL28 | HEK-293T | C132(0.75) | LDD1620 | [14] |
| LDCM0417 | CL29 | HEK-293T | C132(0.84) | LDD1621 | [14] |
| LDCM0418 | CL3 | HEK-293T | C132(0.79) | LDD1622 | [14] |
| LDCM0419 | CL30 | HEK-293T | C132(0.87) | LDD1623 | [14] |
| LDCM0420 | CL31 | HEK-293T | C132(0.84) | LDD1624 | [14] |
| LDCM0421 | CL32 | HEK-293T | C132(0.92) | LDD1625 | [14] |
| LDCM0422 | CL33 | HEK-293T | C132(0.73) | LDD1626 | [14] |
| LDCM0423 | CL34 | HEK-293T | C132(0.85) | LDD1627 | [14] |
| LDCM0424 | CL35 | HEK-293T | C132(0.93) | LDD1628 | [14] |
| LDCM0425 | CL36 | HEK-293T | C132(0.82) | LDD1629 | [14] |
| LDCM0426 | CL37 | HEK-293T | C132(0.94) | LDD1630 | [14] |
| LDCM0428 | CL39 | HEK-293T | C132(0.83) | LDD1632 | [14] |
| LDCM0429 | CL4 | HEK-293T | C132(0.63) | LDD1633 | [14] |
| LDCM0430 | CL40 | HEK-293T | C132(0.81) | LDD1634 | [14] |
| LDCM0431 | CL41 | HEK-293T | C132(0.81) | LDD1635 | [14] |
| LDCM0432 | CL42 | HEK-293T | C132(0.86) | LDD1636 | [14] |
| LDCM0433 | CL43 | HEK-293T | C132(0.80) | LDD1637 | [14] |
| LDCM0434 | CL44 | HEK-293T | C132(0.95) | LDD1638 | [14] |
| LDCM0435 | CL45 | HEK-293T | C132(0.86) | LDD1639 | [14] |
| LDCM0436 | CL46 | HEK-293T | C132(0.93) | LDD1640 | [14] |
| LDCM0437 | CL47 | HEK-293T | C132(0.87) | LDD1641 | [14] |
| LDCM0438 | CL48 | HEK-293T | C132(0.83) | LDD1642 | [14] |
| LDCM0439 | CL49 | HEK-293T | C132(0.83) | LDD1643 | [14] |
| LDCM0440 | CL5 | HEK-293T | C132(0.77) | LDD1644 | [14] |
| LDCM0441 | CL50 | HEK-293T | C132(0.71) | LDD1645 | [14] |
| LDCM0443 | CL52 | HEK-293T | C132(0.63) | LDD1646 | [14] |
| LDCM0444 | CL53 | HEK-293T | C132(0.66) | LDD1647 | [14] |
| LDCM0445 | CL54 | HEK-293T | C132(0.64) | LDD1648 | [14] |
| LDCM0446 | CL55 | HEK-293T | C132(0.73) | LDD1649 | [14] |
| LDCM0447 | CL56 | HEK-293T | C132(0.90) | LDD1650 | [14] |
| LDCM0448 | CL57 | HEK-293T | C132(0.75) | LDD1651 | [14] |
| LDCM0449 | CL58 | HEK-293T | C132(0.76) | LDD1652 | [14] |
| LDCM0450 | CL59 | HEK-293T | C132(0.80) | LDD1653 | [14] |
| LDCM0451 | CL6 | HEK-293T | C132(0.73) | LDD1654 | [14] |
| LDCM0452 | CL60 | HEK-293T | C132(0.73) | LDD1655 | [14] |
| LDCM0453 | CL61 | HEK-293T | C132(0.92) | LDD1656 | [14] |
| LDCM0454 | CL62 | HEK-293T | C132(0.80) | LDD1657 | [14] |
| LDCM0455 | CL63 | HEK-293T | C132(0.82) | LDD1658 | [14] |
| LDCM0456 | CL64 | HEK-293T | C132(0.59) | LDD1659 | [14] |
| LDCM0457 | CL65 | HEK-293T | C132(0.77) | LDD1660 | [14] |
| LDCM0458 | CL66 | HEK-293T | C132(0.80) | LDD1661 | [14] |
| LDCM0459 | CL67 | HEK-293T | C132(0.78) | LDD1662 | [14] |
| LDCM0460 | CL68 | HEK-293T | C132(0.91) | LDD1663 | [14] |
| LDCM0461 | CL69 | HEK-293T | C132(0.84) | LDD1664 | [14] |
| LDCM0462 | CL7 | HEK-293T | C132(0.84) | LDD1665 | [14] |
| LDCM0463 | CL70 | HEK-293T | C132(0.82) | LDD1666 | [14] |
| LDCM0464 | CL71 | HEK-293T | C132(0.86) | LDD1667 | [14] |
| LDCM0465 | CL72 | HEK-293T | C132(0.75) | LDD1668 | [14] |
| LDCM0466 | CL73 | HEK-293T | C132(0.85) | LDD1669 | [14] |
| LDCM0467 | CL74 | HEK-293T | C132(0.80) | LDD1670 | [14] |
| LDCM0469 | CL76 | HEK-293T | C132(0.71) | LDD1672 | [14] |
| LDCM0470 | CL77 | HEK-293T | C132(0.72) | LDD1673 | [14] |
| LDCM0471 | CL78 | HEK-293T | C132(0.83) | LDD1674 | [14] |
| LDCM0472 | CL79 | HEK-293T | C132(0.81) | LDD1675 | [14] |
| LDCM0473 | CL8 | HEK-293T | C132(0.63) | LDD1676 | [14] |
| LDCM0474 | CL80 | HEK-293T | C132(0.97) | LDD1677 | [14] |
| LDCM0475 | CL81 | HEK-293T | C132(0.92) | LDD1678 | [14] |
| LDCM0476 | CL82 | HEK-293T | C132(0.84) | LDD1679 | [14] |
| LDCM0477 | CL83 | HEK-293T | C132(0.84) | LDD1680 | [14] |
| LDCM0478 | CL84 | HEK-293T | C132(0.76) | LDD1681 | [14] |
| LDCM0479 | CL85 | HEK-293T | C132(0.89) | LDD1682 | [14] |
| LDCM0480 | CL86 | HEK-293T | C132(0.81) | LDD1683 | [14] |
| LDCM0481 | CL87 | HEK-293T | C132(0.91) | LDD1684 | [14] |
| LDCM0482 | CL88 | HEK-293T | C132(0.68) | LDD1685 | [14] |
| LDCM0483 | CL89 | HEK-293T | C132(0.80) | LDD1686 | [14] |
| LDCM0484 | CL9 | HEK-293T | C132(0.89) | LDD1687 | [14] |
| LDCM0485 | CL90 | HEK-293T | C132(0.58) | LDD1688 | [14] |
| LDCM0486 | CL91 | HEK-293T | C132(0.76) | LDD1689 | [14] |
| LDCM0487 | CL92 | HEK-293T | C132(0.90) | LDD1690 | [14] |
| LDCM0488 | CL93 | HEK-293T | C132(0.84) | LDD1691 | [14] |
| LDCM0489 | CL94 | HEK-293T | C132(0.83) | LDD1692 | [14] |
| LDCM0490 | CL95 | HEK-293T | C132(0.84) | LDD1693 | [14] |
| LDCM0491 | CL96 | HEK-293T | C132(0.74) | LDD1694 | [14] |
| LDCM0492 | CL97 | HEK-293T | C132(0.85) | LDD1695 | [14] |
| LDCM0493 | CL98 | HEK-293T | C132(0.82) | LDD1696 | [14] |
| LDCM0494 | CL99 | HEK-293T | C132(0.89) | LDD1697 | [14] |
| LDCM0634 | CY-0357 | Hep-G2 | C132(3.35); C339(0.92) | LDD2228 | [29] |
| LDCM0495 | E2913 | HEK-293T | C132(0.76) | LDD1698 | [14] |
| LDCM0625 | F8 | Ramos | C339(0.63); C132(1.73); C128(0.39) | LDD2187 | [36] |
| LDCM0572 | Fragment10 | Ramos | C339(0.85); C132(1.07); C128(0.72) | LDD2189 | [36] |
| LDCM0573 | Fragment11 | Ramos | C339(0.06); C128(0.63) | LDD2190 | [36] |
| LDCM0574 | Fragment12 | Ramos | C339(0.87); C132(0.76); C128(4.03) | LDD2191 | [36] |
| LDCM0575 | Fragment13 | Ramos | C339(0.93); C132(1.27); C128(1.80) | LDD2192 | [36] |
| LDCM0576 | Fragment14 | Ramos | C339(0.66); C132(0.87) | LDD2193 | [36] |
| LDCM0579 | Fragment20 | Ramos | C339(0.93); C132(0.91); C128(0.93) | LDD2194 | [36] |
| LDCM0580 | Fragment21 | Ramos | C339(1.25); C132(1.17); C128(7.84) | LDD2195 | [36] |
| LDCM0582 | Fragment23 | Ramos | C339(1.36); C132(1.12); C128(4.76) | LDD2196 | [36] |
| LDCM0578 | Fragment27 | Ramos | C339(1.21); C132(1.25) | LDD2197 | [36] |
| LDCM0586 | Fragment28 | Ramos | C339(1.07); C132(0.64) | LDD2198 | [36] |
| LDCM0588 | Fragment30 | Ramos | C339(0.92); C132(1.24); C128(4.60) | LDD2199 | [36] |
| LDCM0589 | Fragment31 | Ramos | C339(1.35); C132(1.04); C128(2.87) | LDD2200 | [36] |
| LDCM0590 | Fragment32 | Ramos | C339(0.65); C132(0.85); C128(2.61) | LDD2201 | [36] |
| LDCM0468 | Fragment33 | HEK-293T | C132(0.82) | LDD1671 | [14] |
| LDCM0596 | Fragment38 | Ramos | C339(1.45); C132(0.83); C128(14.13) | LDD2203 | [36] |
| LDCM0566 | Fragment4 | Ramos | C339(0.75); C132(0.86); C128(0.65) | LDD2184 | [36] |
| LDCM0427 | Fragment51 | HEK-293T | C132(0.88) | LDD1631 | [14] |
| LDCM0610 | Fragment52 | Ramos | C339(1.15); C132(1.45); C128(2.85) | LDD2204 | [36] |
| LDCM0614 | Fragment56 | Ramos | C339(0.91); C132(1.68); C128(2.71) | LDD2205 | [36] |
| LDCM0569 | Fragment7 | Ramos | C339(0.67); C132(0.77); C128(0.51) | LDD2186 | [36] |
| LDCM0571 | Fragment9 | Ramos | C339(0.69); C132(0.95); C128(0.91) | LDD2188 | [36] |
| LDCM0116 | HHS-0101 | DM93 | Y297(0.70); Y291(0.78); Y140(0.81); Y511(0.84) | LDD0264 | [12] |
| LDCM0117 | HHS-0201 | DM93 | Y297(0.62); Y291(0.67); Y140(0.75); Y56(0.78) | LDD0265 | [12] |
| LDCM0118 | HHS-0301 | DM93 | Y291(0.73); Y194(0.76); Y513(0.76); Y297(0.80) | LDD0266 | [12] |
| LDCM0119 | HHS-0401 | DM93 | Y297(0.72); Y291(0.77); Y364(0.86); Y382(0.87) | LDD0267 | [12] |
| LDCM0120 | HHS-0701 | DM93 | Y364(0.75); Y291(0.81); Y140(0.82); Y382(0.84) | LDD0268 | [12] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [27] |
| LDCM0123 | JWB131 | DM93 | Y291(0.89); Y297(0.94); Y364(0.93); Y382(0.88) | LDD0285 | [11] |
| LDCM0124 | JWB142 | DM93 | Y140(0.88); Y291(0.65); Y297(0.68); Y364(0.69) | LDD0286 | [11] |
| LDCM0125 | JWB146 | DM93 | Y140(1.56); Y291(1.09); Y297(0.95); Y364(1.06) | LDD0287 | [11] |
| LDCM0126 | JWB150 | DM93 | Y140(3.52); Y291(2.17); Y297(3.02); Y364(2.66) | LDD0288 | [11] |
| LDCM0127 | JWB152 | DM93 | Y140(3.55); Y291(1.62); Y297(1.86); Y364(1.77) | LDD0289 | [11] |
| LDCM0128 | JWB198 | DM93 | Y291(0.98); Y297(1.13); Y364(1.10); Y382(0.94) | LDD0290 | [11] |
| LDCM0129 | JWB202 | DM93 | Y140(1.06); Y291(0.40); Y297(0.63); Y364(0.44) | LDD0291 | [11] |
| LDCM0130 | JWB211 | DM93 | Y140(1.34); Y291(0.88); Y297(0.88); Y364(0.98) | LDD0292 | [11] |
| LDCM0022 | KB02 | Ramos | C339(0.48); C132(0.92); C128(0.43) | LDD2182 | [36] |
| LDCM0023 | KB03 | Ramos | C339(0.70); C132(0.76); C128(0.96) | LDD2183 | [36] |
| LDCM0024 | KB05 | Ramos | C339(0.51); C132(0.76); C128(0.73) | LDD2185 | [36] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C132(0.94) | LDD2102 | [9] |
| LDCM0109 | NEM | HeLa | H144(0.00); H377(0.00); C128(0.00) | LDD0223 | [27] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C132(0.64) | LDD2089 | [9] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C132(1.16) | LDD2090 | [9] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C132(0.77) | LDD2092 | [9] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C132(1.32) | LDD2094 | [9] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C132(0.66) | LDD2097 | [9] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C132(0.68) | LDD2098 | [9] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C132(1.32) | LDD2099 | [9] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C132(0.72) | LDD2100 | [9] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C132(0.63) | LDD2101 | [9] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C132(0.93) | LDD2104 | [9] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C132(1.01) | LDD2105 | [9] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C339(0.36) | LDD2106 | [9] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C132(0.88); C339(0.50) | LDD2108 | [9] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C132(0.84) | LDD2109 | [9] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C132(1.63) | LDD2111 | [9] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C132(0.71) | LDD2114 | [9] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C132(0.36) | LDD2115 | [9] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C132(0.46) | LDD2116 | [9] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C132(2.03) | LDD2119 | [9] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C132(1.15) | LDD2120 | [9] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C132(0.37) | LDD2122 | [9] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C132(0.25) | LDD2124 | [9] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C132(0.99) | LDD2125 | [9] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C132(0.18) | LDD2126 | [9] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C132(1.00) | LDD2127 | [9] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C132(0.97) | LDD2128 | [9] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C132(0.27) | LDD2133 | [9] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C132(0.30) | LDD2134 | [9] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C132(0.80) | LDD2135 | [9] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C132(1.29) | LDD2136 | [9] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C132(1.10) | LDD2137 | [9] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C132(1.88) | LDD1700 | [9] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C132(0.96) | LDD2140 | [9] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C132(0.80) | LDD2141 | [9] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C132(1.03) | LDD2143 | [9] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C132(3.31) | LDD2144 | [9] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C132(7.04); C339(0.35) | LDD2145 | [9] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C132(1.07) | LDD2146 | [9] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C132(0.65) | LDD2148 | [9] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C132(1.38) | LDD2153 | [9] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C339(1.04); C132(0.92) | LDD2206 | [37] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C132(0.71); C339(0.63) | LDD2207 | [37] |
| LDCM0021 | THZ1 | HeLa S3 | C128(1.08) | LDD0460 | [8] |
| LDCM0113 | W17 | Hep-G2 | C132(11.84) | LDD0240 | [28] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Amyloid-beta precursor protein (APP) | APP family | P05067 | |||
| Nuclear cap-binding protein subunit 1 (NCBP1) | NCBP1 family | Q09161 | |||
Other
References
