Details of the Target
General Information of Target
| Target ID | LDTP05991 | |||||
|---|---|---|---|---|---|---|
| Target Name | Spliceosome RNA helicase DDX39B (DDX39B) | |||||
| Gene Name | DDX39B | |||||
| Gene ID | 7919 | |||||
| Synonyms |
BAT1; UAP56; Spliceosome RNA helicase DDX39B; EC 3.6.4.13; 56 kDa U2AF65-associated protein; ATP-dependent RNA helicase p47; DEAD box protein UAP56; HLA-B-associated transcript 1 protein |
|||||
| 3D Structure | ||||||
| Sequence |
MAENDVDNELLDYEDDEVETAAGGDGAEAPAKKDVKGSYVSIHSSGFRDFLLKPELLRAI
VDCGFEHPSEVQHECIPQAILGMDVLCQAKSGMGKTAVFVLATLQQLEPVTGQVSVLVMC HTRELAFQISKEYERFSKYMPNVKVAVFFGGLSIKKDEEVLKKNCPHIVVGTPGRILALA RNKSLNLKHIKHFILDECDKMLEQLDMRRDVQEIFRMTPHEKQVMMFSATLSKEIRPVCR KFMQDPMEIFVDDETKLTLHGLQQYYVKLKDNEKNRKLFDLLDVLEFNQVVIFVKSVQRC IALAQLLVEQNFPAIAIHRGMPQEERLSRYQQFKDFQRRILVATNLFGRGMDIERVNIAF NYDMPEDSDTYLHRVARAGRFGTKGLAITFVSDENDAKILNDVQDRFEVNISELPDEIDI SSYIEQTR |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
DEAD box helicase family, DECD subfamily
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Involved in nuclear export of spliced and unspliced mRNA. Assembling component of the TREX complex which is thought to couple mRNA transcription, processing and nuclear export, and specifically associates with spliced mRNA and not with unspliced pre-mRNA. TREX is recruited to spliced mRNAs by a transcription-independent mechanism, binds to mRNA upstream of the exon-junction complex (EJC) and is recruited in a splicing- and cap-dependent manner to a region near the 5' end of the mRNA where it functions in mRNA export to the cytoplasm via the TAP/NFX1 pathway. May undergo several rounds of ATP hydrolysis during assembly of TREX to drive subsequent loading of components such as ALYREF/THOC and CHTOP onto mRNA. Also associates with pre-mRNA independent of ALYREF/THOC4 and the THO complex. Involved in the nuclear export of intronless mRNA; the ATP-bound form is proposed to recruit export adapter ALYREF/THOC4 to intronless mRNA; its ATPase activity is cooperatively stimulated by RNA and ALYREF/THOC4 and ATP hydrolysis is thought to trigger the dissociation from RNA to allow the association of ALYREF/THOC4 and the NXF1-NXT1 heterodimer. Involved in transcription elongation and genome stability.; Splice factor that is required for the first ATP-dependent step in spliceosome assembly and for the interaction of U2 snRNP with the branchpoint. Has both RNA-stimulated ATP binding/hydrolysis activity and ATP-dependent RNA unwinding activity. Even with the stimulation of RNA, the ATPase activity is weak. Can only hydrolyze ATP but not other NTPs. The RNA stimulation of ATPase activity does not have a strong preference for the sequence and length of the RNA. However, ssRNA stimulates the ATPase activity much more strongly than dsRNA. Can unwind 5' or 3' overhangs or blunt end RNA duplexes in vitro. The ATPase and helicase activities are not influenced by U2AF2; the effect of ALYREF/THOC4 is reported conflictingly with [] reporting a stimulatory effect.; (Microbial infection) The TREX complex is essential for the export of Kaposi's sarcoma-associated herpesvirus (KSHV) intronless mRNAs and infectious virus production.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AN3CA | Insertion: p.H220PfsTer34 | DBIA Probe Info | |||
| HCT116 | Deletion: p.F149LfsTer13 | . | |||
| HCT15 | SNV: p.R181Q | DBIA Probe Info | |||
| HT | SNV: p.E134G | DBIA Probe Info | |||
| IM95 | SNV: p.M321K | DBIA Probe Info | |||
| JURKAT | SNV: p.Q128Ter | Compound 10 Probe Info | |||
| MOLT4 | SNV: p.R135H; p.M140I; p.Q244Ter | IA-alkyne Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
8.53 | LDD0402 | [1] | |
|
AZ-9 Probe Info |
 |
7.03 | LDD0393 | [2] | |
|
CHEMBL5175495 Probe Info |
 |
7.18 | LDD0196 | [3] | |
|
CY4 Probe Info |
 |
2.16 | LDD0244 | [4] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [4] | |
|
TH211 Probe Info |
 |
Y362(20.00); Y39(12.10); Y266(10.55); Y265(10.22) | LDD0257 | [5] | |
|
TH214 Probe Info |
 |
Y39(6.58); Y266(6.45) | LDD0258 | [5] | |
|
TH216 Probe Info |
 |
Y265(11.41); Y39(9.90); Y266(7.89) | LDD0259 | [5] | |
|
STPyne Probe Info |
 |
K163(5.16); K183(4.74); K188(9.80); K241(9.40) | LDD0277 | [6] | |
|
ONAyne Probe Info |
 |
K334(0.00); K53(0.00) | LDD0273 | [6] | |
|
Probe 1 Probe Info |
 |
Y139(16.64); Y362(8.63) | LDD3495 | [7] | |
|
DBIA Probe Info |
 |
C315(1.53); C180(2.01) | LDD3311 | [8] | |
|
THZ1-DTB Probe Info |
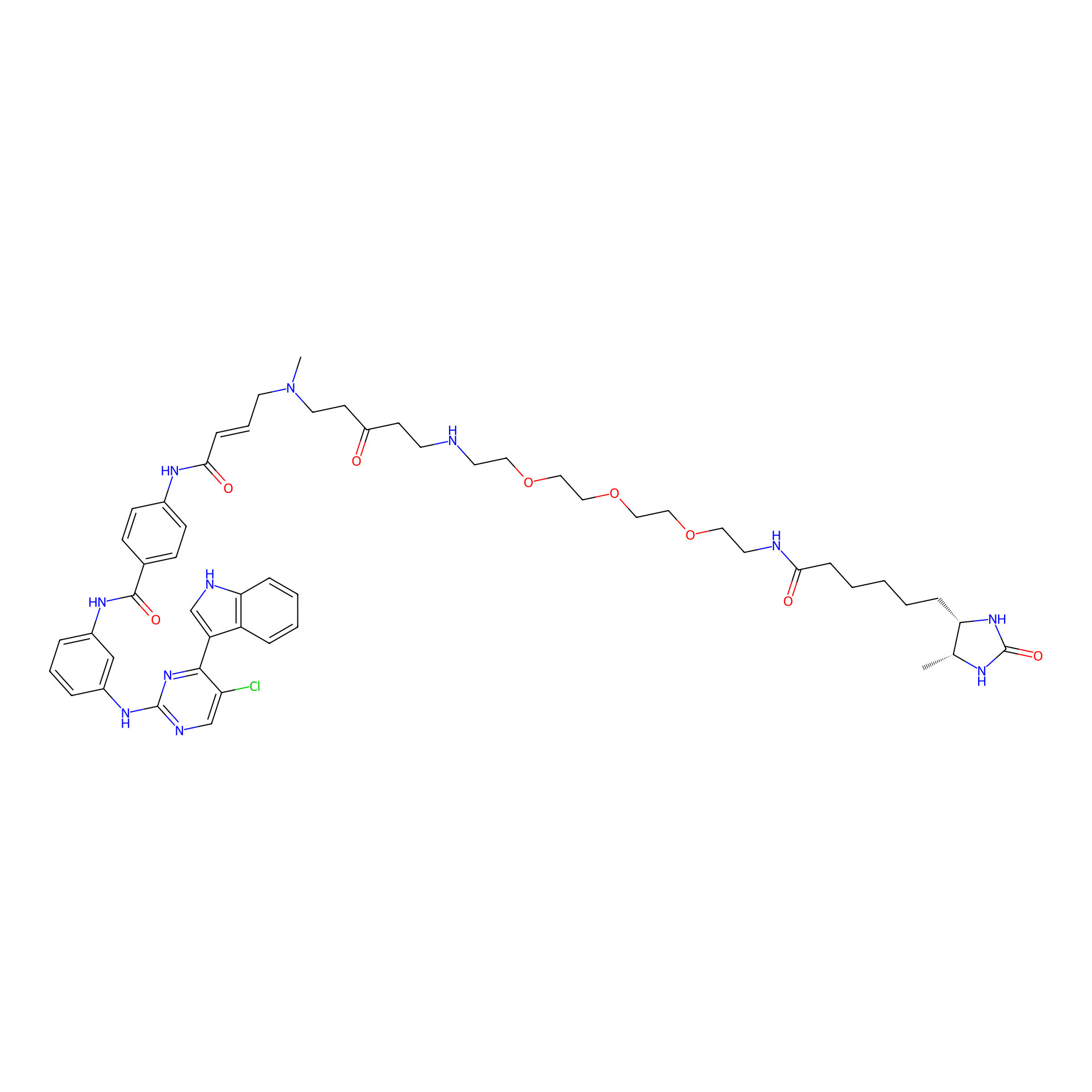 |
C75(1.11) | LDD0460 | [9] | |
|
AHL-Pu-1 Probe Info |
 |
C198(2.56) | LDD0171 | [10] | |
|
HHS-482 Probe Info |
 |
Y39(0.61) | LDD0285 | [11] | |
|
HHS-475 Probe Info |
 |
Y39(0.75); Y266(1.15); Y265(1.70) | LDD0264 | [12] | |
|
HHS-465 Probe Info |
 |
Y265(10.00); Y266(10.00); Y39(10.00) | LDD2237 | [13] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [14] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [15] | |
|
ATP probe Probe Info |
 |
K36(0.00); K163(0.00); K241(0.00); K156(0.00) | LDD0199 | [15] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0358 | [16] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C300(0.00); C198(0.00); C165(0.00); C120(0.00) | LDD0038 | [17] | |
|
IA-alkyne Probe Info |
 |
C198(0.00); C300(0.00); C165(0.00); C63(0.00) | LDD0032 | [18] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [19] | |
|
IPIAA_L Probe Info |
 |
C198(0.00); C63(0.00) | LDD0031 | [19] | |
|
Lodoacetamide azide Probe Info |
 |
C300(0.00); C239(0.00); C198(0.00); C165(0.00) | LDD0037 | [17] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [20] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [21] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [21] | |
|
NAIA_4 Probe Info |
 |
C75(0.00); C63(0.00); C198(0.00) | LDD2226 | [22] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [21] | |
|
WYneN Probe Info |
 |
C165(0.00); C239(0.00) | LDD0021 | [20] | |
|
Compound 10 Probe Info |
 |
C165(0.00); C198(0.00) | LDD2216 | [23] | |
|
NHS Probe Info |
 |
K268(0.00); K36(0.00); K53(0.00); K241(0.00) | LDD0010 | [20] | |
|
OSF Probe Info |
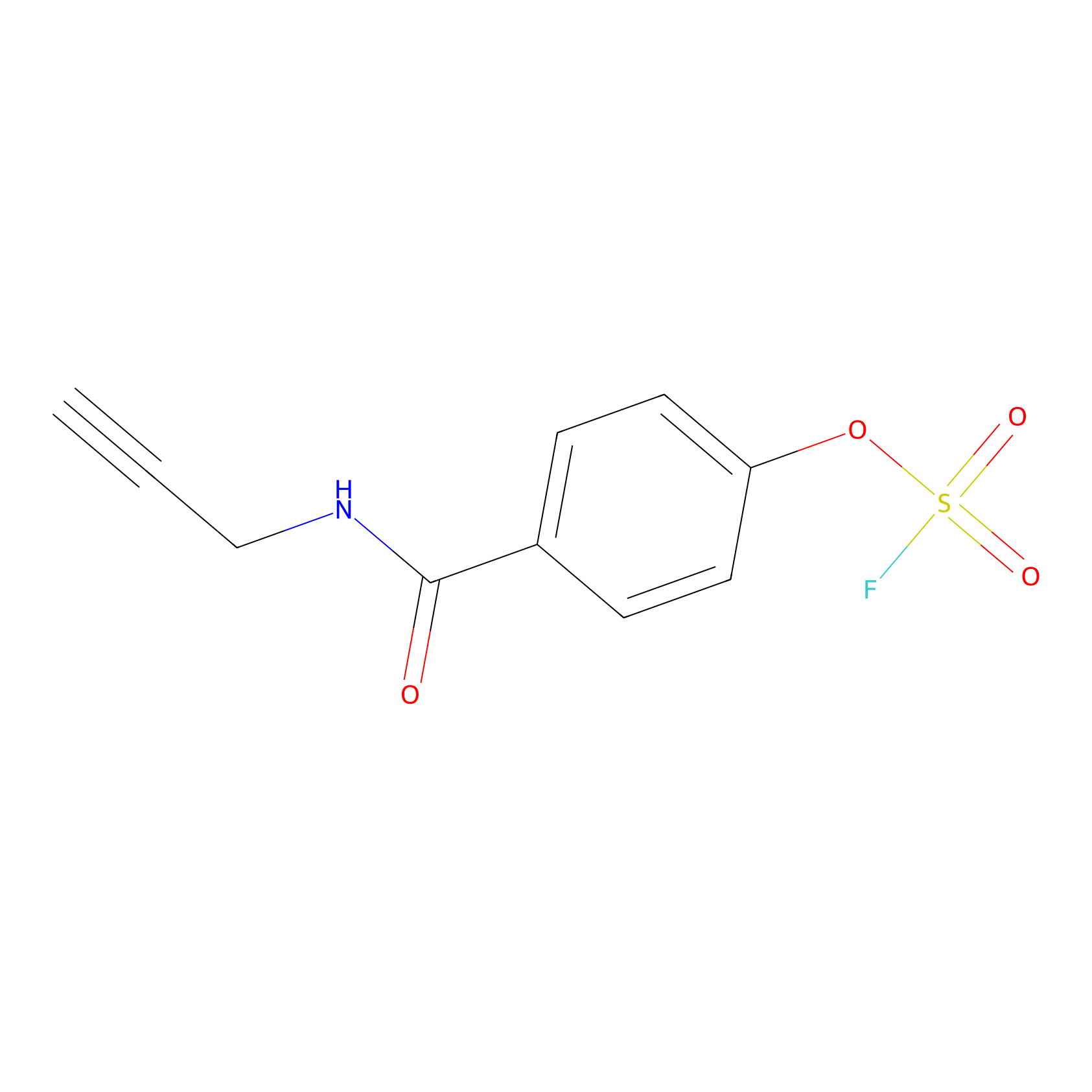 |
Y39(0.00); H373(0.00) | LDD0029 | [24] | |
|
SF Probe Info |
 |
K188(0.00); K156(0.00); K334(0.00); Y362(0.00) | LDD0028 | [24] | |
|
Phosphinate-6 Probe Info |
 |
C63(0.00); C87(0.00) | LDD0018 | [25] | |
|
1c-yne Probe Info |
 |
K384(0.00); K241(0.00) | LDD0228 | [16] | |
|
Acrolein Probe Info |
 |
C165(0.00); C198(0.00); H192(0.00) | LDD0217 | [26] | |
|
Cinnamaldehyde Probe Info |
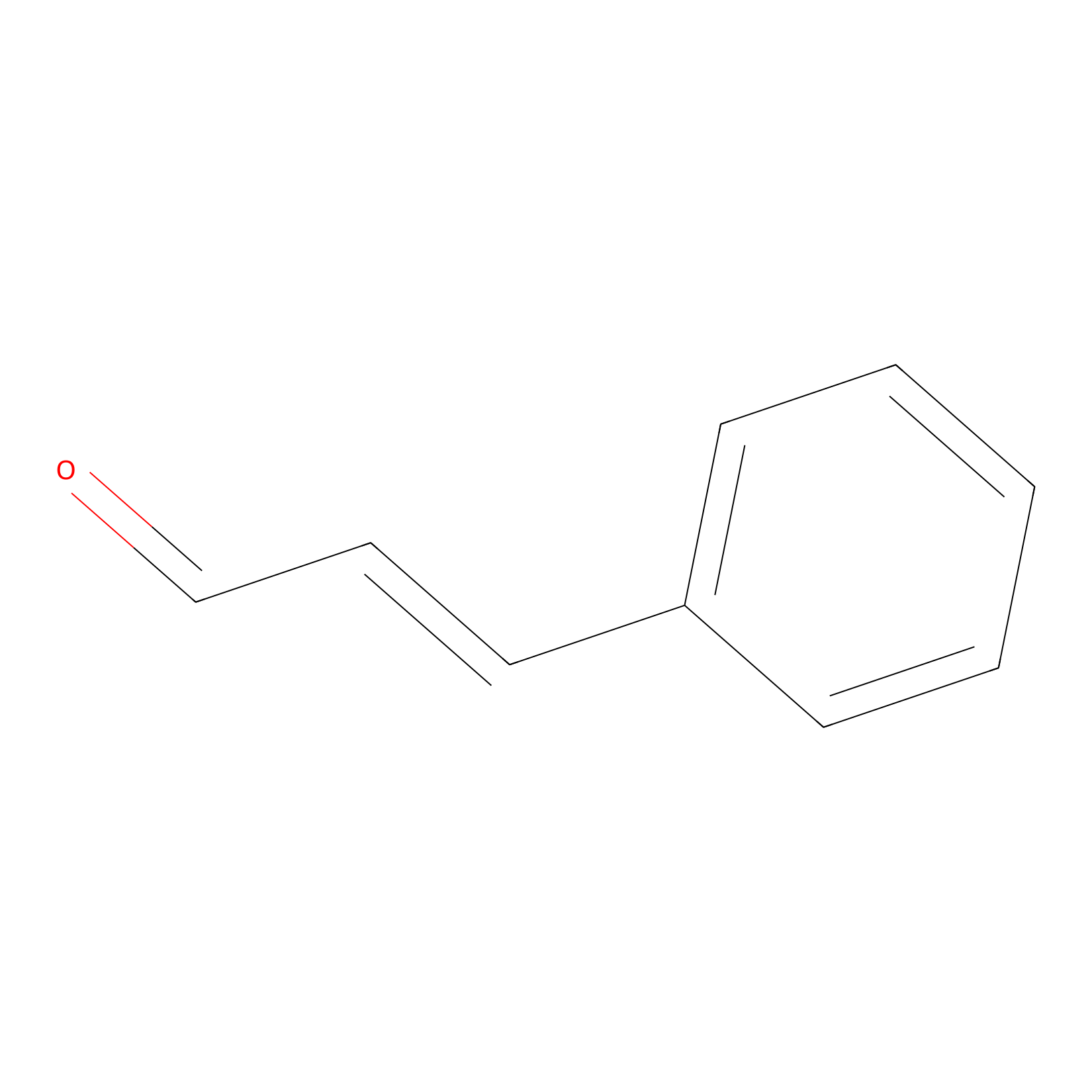 |
N.A. | LDD0220 | [26] | |
|
Crotonaldehyde Probe Info |
 |
C198(0.00); C165(0.00); C239(0.00); H167(0.00) | LDD0219 | [26] | |
|
Methacrolein Probe Info |
 |
C165(0.00); C198(0.00); H167(0.00) | LDD0218 | [26] | |
|
W1 Probe Info |
 |
C239(0.00); C198(0.00); C165(0.00); K53(0.00) | LDD0236 | [27] | |
|
AOyne Probe Info |
 |
8.50 | LDD0443 | [28] | |
|
NAIA_5 Probe Info |
 |
C300(0.00); C165(0.00); C239(0.00); C198(0.00) | LDD2223 | [22] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [29] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe13 Probe Info |
 |
6.50 | LDD0475 | [30] | |
|
STS-2 Probe Info |
 |
3.86 | LDD0138 | [31] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [32] | |
|
Staurosporine capture compound Probe Info |
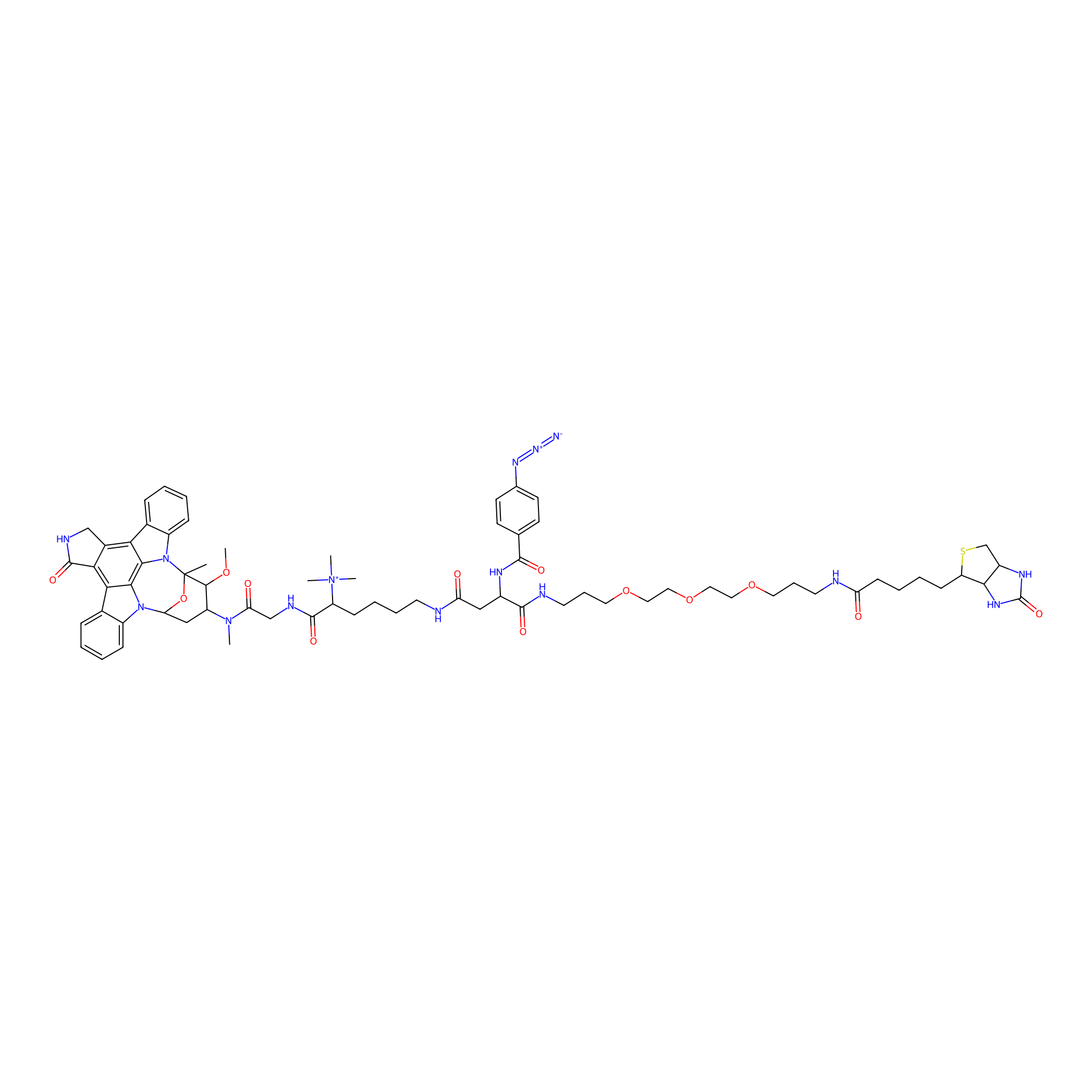 |
N.A. | LDD0083 | [33] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [34] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0069 | [35] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C165(0.68); C239(0.83) | LDD2142 | [36] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C165(0.90); C239(0.86) | LDD2112 | [36] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C165(0.82); C239(0.58) | LDD2095 | [36] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C165(1.02); C239(0.85) | LDD2130 | [36] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C165(0.98); C239(1.07) | LDD2117 | [36] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C165(1.33); C239(1.36); C198(2.75) | LDD2152 | [36] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C165(0.71); C239(0.88) | LDD2103 | [36] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C165(0.49); C239(0.63) | LDD2132 | [36] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C165(0.60); C239(0.87) | LDD2131 | [36] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C198(2.56) | LDD0171 | [10] |
| LDCM0214 | AC1 | HEK-293T | C300(1.06) | LDD0812 | [37] |
| LDCM0215 | AC10 | HEK-293T | C300(1.03) | LDD0813 | [37] |
| LDCM0216 | AC100 | HEK-293T | C300(1.17) | LDD0814 | [37] |
| LDCM0217 | AC101 | HEK-293T | C300(1.17) | LDD0815 | [37] |
| LDCM0218 | AC102 | HEK-293T | C300(1.09) | LDD0816 | [37] |
| LDCM0219 | AC103 | HEK-293T | C300(1.31) | LDD0817 | [37] |
| LDCM0220 | AC104 | HEK-293T | C300(1.30) | LDD0818 | [37] |
| LDCM0221 | AC105 | HEK-293T | C300(0.78) | LDD0819 | [37] |
| LDCM0222 | AC106 | HEK-293T | C300(0.89) | LDD0820 | [37] |
| LDCM0223 | AC107 | HEK-293T | C300(1.08) | LDD0821 | [37] |
| LDCM0224 | AC108 | HEK-293T | C300(1.01) | LDD0822 | [37] |
| LDCM0225 | AC109 | HEK-293T | C300(0.92) | LDD0823 | [37] |
| LDCM0226 | AC11 | HEK-293T | C300(0.91) | LDD0824 | [37] |
| LDCM0227 | AC110 | HEK-293T | C300(1.30) | LDD0825 | [37] |
| LDCM0228 | AC111 | HEK-293T | C300(1.29) | LDD0826 | [37] |
| LDCM0229 | AC112 | HEK-293T | C300(1.47) | LDD0827 | [37] |
| LDCM0230 | AC113 | HEK-293T | C300(1.04) | LDD0828 | [37] |
| LDCM0231 | AC114 | HEK-293T | C300(1.01) | LDD0829 | [37] |
| LDCM0232 | AC115 | HEK-293T | C300(1.09) | LDD0830 | [37] |
| LDCM0233 | AC116 | HEK-293T | C300(0.94) | LDD0831 | [37] |
| LDCM0234 | AC117 | HEK-293T | C300(1.07) | LDD0832 | [37] |
| LDCM0235 | AC118 | HEK-293T | C300(1.19) | LDD0833 | [37] |
| LDCM0236 | AC119 | HEK-293T | C300(1.15) | LDD0834 | [37] |
| LDCM0237 | AC12 | HEK-293T | C300(1.07) | LDD0835 | [37] |
| LDCM0238 | AC120 | HEK-293T | C300(1.14) | LDD0836 | [37] |
| LDCM0239 | AC121 | HEK-293T | C300(1.35) | LDD0837 | [37] |
| LDCM0240 | AC122 | HEK-293T | C300(1.42) | LDD0838 | [37] |
| LDCM0241 | AC123 | HEK-293T | C300(1.45) | LDD0839 | [37] |
| LDCM0242 | AC124 | HEK-293T | C300(1.28) | LDD0840 | [37] |
| LDCM0243 | AC125 | HEK-293T | C300(1.37) | LDD0841 | [37] |
| LDCM0244 | AC126 | HEK-293T | C300(1.64) | LDD0842 | [37] |
| LDCM0245 | AC127 | HEK-293T | C300(1.76) | LDD0843 | [37] |
| LDCM0246 | AC128 | HEK-293T | C300(1.11) | LDD0844 | [37] |
| LDCM0247 | AC129 | HEK-293T | C300(1.27) | LDD0845 | [37] |
| LDCM0249 | AC130 | HEK-293T | C300(1.07) | LDD0847 | [37] |
| LDCM0250 | AC131 | HEK-293T | C300(0.97) | LDD0848 | [37] |
| LDCM0251 | AC132 | HEK-293T | C300(1.39) | LDD0849 | [37] |
| LDCM0252 | AC133 | HEK-293T | C300(1.68) | LDD0850 | [37] |
| LDCM0253 | AC134 | HEK-293T | C300(1.26) | LDD0851 | [37] |
| LDCM0254 | AC135 | HEK-293T | C300(1.24) | LDD0852 | [37] |
| LDCM0255 | AC136 | HEK-293T | C300(1.68) | LDD0853 | [37] |
| LDCM0256 | AC137 | HEK-293T | C300(1.59) | LDD0854 | [37] |
| LDCM0257 | AC138 | HEK-293T | C300(1.77) | LDD0855 | [37] |
| LDCM0258 | AC139 | HEK-293T | C300(1.43) | LDD0856 | [37] |
| LDCM0259 | AC14 | HEK-293T | C300(1.12) | LDD0857 | [37] |
| LDCM0260 | AC140 | HEK-293T | C300(1.26) | LDD0858 | [37] |
| LDCM0261 | AC141 | HEK-293T | C300(1.54) | LDD0859 | [37] |
| LDCM0262 | AC142 | HEK-293T | C300(1.23) | LDD0860 | [37] |
| LDCM0263 | AC143 | HEK-293T | C300(1.07) | LDD0861 | [37] |
| LDCM0264 | AC144 | HEK-293T | C300(1.18) | LDD0862 | [37] |
| LDCM0265 | AC145 | HEK-293T | C300(1.44) | LDD0863 | [37] |
| LDCM0266 | AC146 | HEK-293T | C300(1.37) | LDD0864 | [37] |
| LDCM0267 | AC147 | HEK-293T | C300(1.47) | LDD0865 | [37] |
| LDCM0268 | AC148 | HEK-293T | C300(1.44) | LDD0866 | [37] |
| LDCM0269 | AC149 | HEK-293T | C300(0.99) | LDD0867 | [37] |
| LDCM0270 | AC15 | HEK-293T | C300(1.32) | LDD0868 | [37] |
| LDCM0271 | AC150 | HEK-293T | C300(1.28) | LDD0869 | [37] |
| LDCM0272 | AC151 | HEK-293T | C300(1.12) | LDD0870 | [37] |
| LDCM0273 | AC152 | HEK-293T | C300(1.14) | LDD0871 | [37] |
| LDCM0274 | AC153 | HEK-293T | C300(1.20) | LDD0872 | [37] |
| LDCM0621 | AC154 | HEK-293T | C300(1.02) | LDD2162 | [37] |
| LDCM0622 | AC155 | HEK-293T | C300(1.02) | LDD2163 | [37] |
| LDCM0623 | AC156 | HEK-293T | C300(1.47) | LDD2164 | [37] |
| LDCM0624 | AC157 | HEK-293T | C300(1.75) | LDD2165 | [37] |
| LDCM0276 | AC17 | HEK-293T | C300(2.25) | LDD0874 | [37] |
| LDCM0277 | AC18 | HEK-293T | C300(1.37) | LDD0875 | [37] |
| LDCM0278 | AC19 | HEK-293T | C300(1.46) | LDD0876 | [37] |
| LDCM0279 | AC2 | HEK-293T | C300(1.29) | LDD0877 | [37] |
| LDCM0280 | AC20 | HEK-293T | C300(2.25) | LDD0878 | [37] |
| LDCM0281 | AC21 | HEK-293T | C300(1.78) | LDD0879 | [37] |
| LDCM0282 | AC22 | HEK-293T | C300(1.66) | LDD0880 | [37] |
| LDCM0283 | AC23 | HEK-293T | C300(1.37) | LDD0881 | [37] |
| LDCM0284 | AC24 | HEK-293T | C300(1.31) | LDD0882 | [37] |
| LDCM0285 | AC25 | HEK-293T | C300(1.13) | LDD0883 | [37] |
| LDCM0286 | AC26 | HEK-293T | C300(1.22) | LDD0884 | [37] |
| LDCM0287 | AC27 | HEK-293T | C300(1.07) | LDD0885 | [37] |
| LDCM0288 | AC28 | HEK-293T | C300(1.41) | LDD0886 | [37] |
| LDCM0289 | AC29 | HEK-293T | C300(1.18) | LDD0887 | [37] |
| LDCM0290 | AC3 | HEK-293T | C300(1.18) | LDD0888 | [37] |
| LDCM0291 | AC30 | HEK-293T | C300(1.02) | LDD0889 | [37] |
| LDCM0292 | AC31 | HEK-293T | C300(0.95) | LDD0890 | [37] |
| LDCM0293 | AC32 | HEK-293T | C300(0.97) | LDD0891 | [37] |
| LDCM0294 | AC33 | HEK-293T | C300(1.17) | LDD0892 | [37] |
| LDCM0295 | AC34 | HEK-293T | C300(0.98) | LDD0893 | [37] |
| LDCM0296 | AC35 | HEK-293T | C300(1.29) | LDD0894 | [37] |
| LDCM0297 | AC36 | HEK-293T | C300(1.40) | LDD0895 | [37] |
| LDCM0298 | AC37 | HEK-293T | C300(1.42) | LDD0896 | [37] |
| LDCM0299 | AC38 | HEK-293T | C300(1.25) | LDD0897 | [37] |
| LDCM0300 | AC39 | HEK-293T | C300(1.39) | LDD0898 | [37] |
| LDCM0301 | AC4 | HEK-293T | C300(1.32) | LDD0899 | [37] |
| LDCM0302 | AC40 | HEK-293T | C300(1.23) | LDD0900 | [37] |
| LDCM0303 | AC41 | HEK-293T | C300(1.26) | LDD0901 | [37] |
| LDCM0304 | AC42 | HEK-293T | C300(1.10) | LDD0902 | [37] |
| LDCM0305 | AC43 | HEK-293T | C300(1.25) | LDD0903 | [37] |
| LDCM0306 | AC44 | HEK-293T | C300(1.35) | LDD0904 | [37] |
| LDCM0307 | AC45 | HEK-293T | C300(1.36) | LDD0905 | [37] |
| LDCM0308 | AC46 | HEK-293T | C300(1.22) | LDD0906 | [37] |
| LDCM0309 | AC47 | HEK-293T | C300(1.29) | LDD0907 | [37] |
| LDCM0310 | AC48 | HEK-293T | C300(1.20) | LDD0908 | [37] |
| LDCM0311 | AC49 | HEK-293T | C300(1.14) | LDD0909 | [37] |
| LDCM0312 | AC5 | HEK-293T | C300(1.22) | LDD0910 | [37] |
| LDCM0313 | AC50 | HEK-293T | C300(1.13) | LDD0911 | [37] |
| LDCM0314 | AC51 | HEK-293T | C300(1.26) | LDD0912 | [37] |
| LDCM0315 | AC52 | HEK-293T | C300(1.24) | LDD0913 | [37] |
| LDCM0316 | AC53 | HEK-293T | C300(1.10) | LDD0914 | [37] |
| LDCM0317 | AC54 | HEK-293T | C300(1.46) | LDD0915 | [37] |
| LDCM0318 | AC55 | HEK-293T | C300(1.29) | LDD0916 | [37] |
| LDCM0319 | AC56 | HEK-293T | C300(1.39) | LDD0917 | [37] |
| LDCM0320 | AC57 | HEK-293T | C300(1.02) | LDD0918 | [37] |
| LDCM0321 | AC58 | HEK-293T | C300(1.16) | LDD0919 | [37] |
| LDCM0322 | AC59 | HEK-293T | C300(1.07) | LDD0920 | [37] |
| LDCM0323 | AC6 | HEK-293T | C300(0.92) | LDD0921 | [37] |
| LDCM0324 | AC60 | HEK-293T | C300(1.09) | LDD0922 | [37] |
| LDCM0325 | AC61 | HEK-293T | C300(1.43) | LDD0923 | [37] |
| LDCM0326 | AC62 | HEK-293T | C300(1.08) | LDD0924 | [37] |
| LDCM0327 | AC63 | HEK-293T | C300(1.32) | LDD0925 | [37] |
| LDCM0328 | AC64 | HEK-293T | C300(1.02) | LDD0926 | [37] |
| LDCM0329 | AC65 | HEK-293T | C300(1.39) | LDD0927 | [37] |
| LDCM0330 | AC66 | HEK-293T | C300(1.14) | LDD0928 | [37] |
| LDCM0331 | AC67 | HEK-293T | C300(1.34) | LDD0929 | [37] |
| LDCM0332 | AC68 | HEK-293T | C300(1.19) | LDD0930 | [37] |
| LDCM0333 | AC69 | HEK-293T | C300(1.10) | LDD0931 | [37] |
| LDCM0334 | AC7 | HEK-293T | C300(1.19) | LDD0932 | [37] |
| LDCM0335 | AC70 | HEK-293T | C300(1.52) | LDD0933 | [37] |
| LDCM0336 | AC71 | HEK-293T | C300(1.14) | LDD0934 | [37] |
| LDCM0337 | AC72 | HEK-293T | C300(1.31) | LDD0935 | [37] |
| LDCM0338 | AC73 | HEK-293T | C300(1.24) | LDD0936 | [37] |
| LDCM0339 | AC74 | HEK-293T | C300(1.04) | LDD0937 | [37] |
| LDCM0340 | AC75 | HEK-293T | C300(1.25) | LDD0938 | [37] |
| LDCM0341 | AC76 | HEK-293T | C300(1.36) | LDD0939 | [37] |
| LDCM0342 | AC77 | HEK-293T | C300(1.46) | LDD0940 | [37] |
| LDCM0343 | AC78 | HEK-293T | C300(1.26) | LDD0941 | [37] |
| LDCM0344 | AC79 | HEK-293T | C300(1.26) | LDD0942 | [37] |
| LDCM0345 | AC8 | HEK-293T | C300(1.15) | LDD0943 | [37] |
| LDCM0346 | AC80 | HEK-293T | C300(1.36) | LDD0944 | [37] |
| LDCM0347 | AC81 | HEK-293T | C300(1.40) | LDD0945 | [37] |
| LDCM0348 | AC82 | HEK-293T | C300(1.63) | LDD0946 | [37] |
| LDCM0349 | AC83 | HEK-293T | C300(0.92) | LDD0947 | [37] |
| LDCM0350 | AC84 | HEK-293T | C300(1.13) | LDD0948 | [37] |
| LDCM0351 | AC85 | HEK-293T | C300(1.09) | LDD0949 | [37] |
| LDCM0352 | AC86 | HEK-293T | C300(1.04) | LDD0950 | [37] |
| LDCM0353 | AC87 | HEK-293T | C300(1.10) | LDD0951 | [37] |
| LDCM0354 | AC88 | HEK-293T | C300(1.21) | LDD0952 | [37] |
| LDCM0355 | AC89 | HEK-293T | C300(1.12) | LDD0953 | [37] |
| LDCM0357 | AC90 | HEK-293T | C300(1.13) | LDD0955 | [37] |
| LDCM0358 | AC91 | HEK-293T | C300(1.10) | LDD0956 | [37] |
| LDCM0359 | AC92 | HEK-293T | C300(0.95) | LDD0957 | [37] |
| LDCM0360 | AC93 | HEK-293T | C300(1.10) | LDD0958 | [37] |
| LDCM0361 | AC94 | HEK-293T | C300(1.43) | LDD0959 | [37] |
| LDCM0362 | AC95 | HEK-293T | C300(1.13) | LDD0960 | [37] |
| LDCM0363 | AC96 | HEK-293T | C300(1.10) | LDD0961 | [37] |
| LDCM0364 | AC97 | HEK-293T | C300(1.64) | LDD0962 | [37] |
| LDCM0365 | AC98 | HEK-293T | C300(0.81) | LDD0963 | [37] |
| LDCM0366 | AC99 | HEK-293T | C300(0.74) | LDD0964 | [37] |
| LDCM0545 | Acetamide | MDA-MB-231 | C165(0.43); C239(0.42) | LDD2138 | [36] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C165(0.80); C239(0.83) | LDD2113 | [36] |
| LDCM0248 | AKOS034007472 | HEK-293T | C300(1.15) | LDD0846 | [37] |
| LDCM0356 | AKOS034007680 | HEK-293T | C300(1.33) | LDD0954 | [37] |
| LDCM0275 | AKOS034007705 | HEK-293T | C300(1.15) | LDD0873 | [37] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C165(0.50) | LDD2091 | [36] |
| LDCM0108 | Chloroacetamide | HeLa | C165(0.00); C198(0.00); H192(0.00); H167(0.00) | LDD0222 | [26] |
| LDCM0632 | CL-Sc | Hep-G2 | C198(1.18); C165(0.83) | LDD2227 | [22] |
| LDCM0367 | CL1 | HEK-293T | C300(1.53) | LDD0965 | [37] |
| LDCM0368 | CL10 | HEK-293T | C300(2.02) | LDD0966 | [37] |
| LDCM0369 | CL100 | HEK-293T | C300(1.24) | LDD0967 | [37] |
| LDCM0370 | CL101 | HEK-293T | C300(1.09) | LDD0968 | [37] |
| LDCM0371 | CL102 | HEK-293T | C300(1.10) | LDD0969 | [37] |
| LDCM0372 | CL103 | HEK-293T | C300(1.17) | LDD0970 | [37] |
| LDCM0373 | CL104 | HEK-293T | C300(1.05) | LDD0971 | [37] |
| LDCM0374 | CL105 | HEK-293T | C300(1.25) | LDD0972 | [37] |
| LDCM0375 | CL106 | HEK-293T | C300(1.37) | LDD0973 | [37] |
| LDCM0376 | CL107 | HEK-293T | C300(1.81) | LDD0974 | [37] |
| LDCM0377 | CL108 | HEK-293T | C300(1.73) | LDD0975 | [37] |
| LDCM0378 | CL109 | HEK-293T | C300(1.37) | LDD0976 | [37] |
| LDCM0379 | CL11 | HEK-293T | C300(1.93) | LDD0977 | [37] |
| LDCM0380 | CL110 | HEK-293T | C300(1.28) | LDD0978 | [37] |
| LDCM0381 | CL111 | HEK-293T | C300(1.23) | LDD0979 | [37] |
| LDCM0382 | CL112 | HEK-293T | C300(1.06) | LDD0980 | [37] |
| LDCM0383 | CL113 | HEK-293T | C300(1.06) | LDD0981 | [37] |
| LDCM0384 | CL114 | HEK-293T | C300(1.29) | LDD0982 | [37] |
| LDCM0385 | CL115 | HEK-293T | C300(1.07) | LDD0983 | [37] |
| LDCM0386 | CL116 | HEK-293T | C300(1.11) | LDD0984 | [37] |
| LDCM0387 | CL117 | HEK-293T | C300(1.18) | LDD0985 | [37] |
| LDCM0388 | CL118 | HEK-293T | C300(1.56) | LDD0986 | [37] |
| LDCM0389 | CL119 | HEK-293T | C300(1.30) | LDD0987 | [37] |
| LDCM0390 | CL12 | HEK-293T | C300(1.30) | LDD0988 | [37] |
| LDCM0391 | CL120 | HEK-293T | C300(1.44) | LDD0989 | [37] |
| LDCM0392 | CL121 | HEK-293T | C300(1.15) | LDD0990 | [37] |
| LDCM0393 | CL122 | HEK-293T | C300(1.26) | LDD0991 | [37] |
| LDCM0394 | CL123 | HEK-293T | C300(1.64) | LDD0992 | [37] |
| LDCM0395 | CL124 | HEK-293T | C300(0.98) | LDD0993 | [37] |
| LDCM0396 | CL125 | HEK-293T | C300(1.12) | LDD0994 | [37] |
| LDCM0397 | CL126 | HEK-293T | C300(1.73) | LDD0995 | [37] |
| LDCM0398 | CL127 | HEK-293T | C300(0.99) | LDD0996 | [37] |
| LDCM0399 | CL128 | HEK-293T | C300(1.02) | LDD0997 | [37] |
| LDCM0400 | CL13 | HEK-293T | C300(1.35) | LDD0998 | [37] |
| LDCM0401 | CL14 | HEK-293T | C300(1.14) | LDD0999 | [37] |
| LDCM0402 | CL15 | HEK-293T | C300(1.92) | LDD1000 | [37] |
| LDCM0403 | CL16 | PaTu 8988t | C198(0.86); C165(0.52) | LDD1282 | [37] |
| LDCM0404 | CL17 | PaTu 8988t | C198(0.81); C165(1.02) | LDD1283 | [37] |
| LDCM0405 | CL18 | PaTu 8988t | C198(0.70); C165(0.88) | LDD1284 | [37] |
| LDCM0406 | CL19 | PaTu 8988t | C198(0.78); C165(0.67) | LDD1285 | [37] |
| LDCM0407 | CL2 | HEK-293T | C300(1.03) | LDD1005 | [37] |
| LDCM0408 | CL20 | PaTu 8988t | C198(0.74); C165(0.66) | LDD1287 | [37] |
| LDCM0409 | CL21 | PaTu 8988t | C198(0.53); C165(0.50) | LDD1288 | [37] |
| LDCM0410 | CL22 | PaTu 8988t | C198(0.63); C165(0.69) | LDD1289 | [37] |
| LDCM0411 | CL23 | PaTu 8988t | C198(0.73); C165(0.89) | LDD1290 | [37] |
| LDCM0412 | CL24 | PaTu 8988t | C198(0.81); C165(0.91) | LDD1291 | [37] |
| LDCM0413 | CL25 | PaTu 8988t | C198(0.63); C165(0.68) | LDD1292 | [37] |
| LDCM0414 | CL26 | PaTu 8988t | C198(0.68); C165(0.60) | LDD1293 | [37] |
| LDCM0415 | CL27 | PaTu 8988t | C198(0.74); C165(0.86) | LDD1294 | [37] |
| LDCM0416 | CL28 | PaTu 8988t | C198(0.67); C165(0.69) | LDD1295 | [37] |
| LDCM0417 | CL29 | PaTu 8988t | C198(0.68); C165(0.64) | LDD1296 | [37] |
| LDCM0418 | CL3 | HEK-293T | C300(0.96) | LDD1016 | [37] |
| LDCM0419 | CL30 | PaTu 8988t | C198(0.52); C165(0.48) | LDD1298 | [37] |
| LDCM0420 | CL31 | HEK-293T | C300(1.19) | LDD1018 | [37] |
| LDCM0421 | CL32 | HEK-293T | C300(1.75) | LDD1019 | [37] |
| LDCM0422 | CL33 | HEK-293T | C300(1.41) | LDD1020 | [37] |
| LDCM0423 | CL34 | HEK-293T | C300(1.28) | LDD1021 | [37] |
| LDCM0424 | CL35 | HEK-293T | C300(1.30) | LDD1022 | [37] |
| LDCM0425 | CL36 | HEK-293T | C300(1.58) | LDD1023 | [37] |
| LDCM0426 | CL37 | HEK-293T | C300(1.30) | LDD1024 | [37] |
| LDCM0428 | CL39 | HEK-293T | C300(1.85) | LDD1026 | [37] |
| LDCM0429 | CL4 | HEK-293T | C300(1.18) | LDD1027 | [37] |
| LDCM0430 | CL40 | HEK-293T | C300(1.49) | LDD1028 | [37] |
| LDCM0431 | CL41 | HEK-293T | C300(1.89) | LDD1029 | [37] |
| LDCM0432 | CL42 | HEK-293T | C300(1.76) | LDD1030 | [37] |
| LDCM0433 | CL43 | HEK-293T | C300(2.01) | LDD1031 | [37] |
| LDCM0434 | CL44 | HEK-293T | C300(1.56) | LDD1032 | [37] |
| LDCM0435 | CL45 | HEK-293T | C300(1.54) | LDD1033 | [37] |
| LDCM0436 | CL46 | HEK-293T | C300(1.11) | LDD1034 | [37] |
| LDCM0437 | CL47 | HEK-293T | C300(1.04) | LDD1035 | [37] |
| LDCM0438 | CL48 | HEK-293T | C300(0.91) | LDD1036 | [37] |
| LDCM0439 | CL49 | HEK-293T | C300(1.00) | LDD1037 | [37] |
| LDCM0440 | CL5 | HEK-293T | C300(1.38) | LDD1038 | [37] |
| LDCM0441 | CL50 | HEK-293T | C300(1.28) | LDD1039 | [37] |
| LDCM0442 | CL51 | HEK-293T | C300(1.13) | LDD1040 | [37] |
| LDCM0443 | CL52 | HEK-293T | C300(0.88) | LDD1041 | [37] |
| LDCM0444 | CL53 | HEK-293T | C300(0.87) | LDD1042 | [37] |
| LDCM0445 | CL54 | HEK-293T | C300(1.08) | LDD1043 | [37] |
| LDCM0446 | CL55 | HEK-293T | C300(1.11) | LDD1044 | [37] |
| LDCM0447 | CL56 | HEK-293T | C300(1.02) | LDD1045 | [37] |
| LDCM0448 | CL57 | HEK-293T | C300(0.93) | LDD1046 | [37] |
| LDCM0449 | CL58 | HEK-293T | C300(0.95) | LDD1047 | [37] |
| LDCM0450 | CL59 | HEK-293T | C300(1.15) | LDD1048 | [37] |
| LDCM0451 | CL6 | HEK-293T | C300(1.64) | LDD1049 | [37] |
| LDCM0452 | CL60 | HEK-293T | C300(1.36) | LDD1050 | [37] |
| LDCM0453 | CL61 | HEK-293T | C300(1.05) | LDD1051 | [37] |
| LDCM0454 | CL62 | HEK-293T | C300(1.19) | LDD1052 | [37] |
| LDCM0455 | CL63 | HEK-293T | C300(1.31) | LDD1053 | [37] |
| LDCM0456 | CL64 | HEK-293T | C300(0.93) | LDD1054 | [37] |
| LDCM0457 | CL65 | HEK-293T | C300(1.31) | LDD1055 | [37] |
| LDCM0458 | CL66 | HEK-293T | C300(1.17) | LDD1056 | [37] |
| LDCM0459 | CL67 | HEK-293T | C300(1.21) | LDD1057 | [37] |
| LDCM0460 | CL68 | HEK-293T | C300(1.49) | LDD1058 | [37] |
| LDCM0461 | CL69 | HEK-293T | C300(1.29) | LDD1059 | [37] |
| LDCM0462 | CL7 | HEK-293T | C300(1.29) | LDD1060 | [37] |
| LDCM0463 | CL70 | HEK-293T | C300(1.31) | LDD1061 | [37] |
| LDCM0464 | CL71 | HEK-293T | C300(1.30) | LDD1062 | [37] |
| LDCM0465 | CL72 | HEK-293T | C300(1.10) | LDD1063 | [37] |
| LDCM0466 | CL73 | HEK-293T | C300(1.25) | LDD1064 | [37] |
| LDCM0467 | CL74 | HEK-293T | C300(1.72) | LDD1065 | [37] |
| LDCM0469 | CL76 | PaTu 8988t | C198(0.74); C165(0.81) | LDD1348 | [37] |
| LDCM0470 | CL77 | PaTu 8988t | C198(0.86); C165(0.83) | LDD1349 | [37] |
| LDCM0471 | CL78 | PaTu 8988t | C198(1.05); C165(1.33) | LDD1350 | [37] |
| LDCM0472 | CL79 | PaTu 8988t | C198(1.24); C165(1.42) | LDD1351 | [37] |
| LDCM0473 | CL8 | HEK-293T | C300(1.70) | LDD1071 | [37] |
| LDCM0474 | CL80 | PaTu 8988t | C198(0.72); C165(0.91) | LDD1353 | [37] |
| LDCM0475 | CL81 | PaTu 8988t | C198(0.90); C165(1.09) | LDD1354 | [37] |
| LDCM0476 | CL82 | PaTu 8988t | C198(0.80); C165(0.78) | LDD1355 | [37] |
| LDCM0477 | CL83 | PaTu 8988t | C198(0.91); C165(1.15) | LDD1356 | [37] |
| LDCM0478 | CL84 | PaTu 8988t | C198(0.93); C165(1.17) | LDD1357 | [37] |
| LDCM0479 | CL85 | PaTu 8988t | C198(1.06); C165(1.23) | LDD1358 | [37] |
| LDCM0480 | CL86 | PaTu 8988t | C198(0.70); C165(0.87) | LDD1359 | [37] |
| LDCM0481 | CL87 | PaTu 8988t | C198(0.78); C165(0.92) | LDD1360 | [37] |
| LDCM0482 | CL88 | PaTu 8988t | C198(0.79); C165(1.01) | LDD1361 | [37] |
| LDCM0483 | CL89 | PaTu 8988t | C198(0.96); C165(1.18) | LDD1362 | [37] |
| LDCM0484 | CL9 | HEK-293T | C300(1.26) | LDD1082 | [37] |
| LDCM0485 | CL90 | PaTu 8988t | C198(0.99); C165(1.08) | LDD1364 | [37] |
| LDCM0486 | CL91 | HEK-293T | C300(1.18) | LDD1084 | [37] |
| LDCM0487 | CL92 | HEK-293T | C300(1.22) | LDD1085 | [37] |
| LDCM0488 | CL93 | HEK-293T | C300(1.11) | LDD1086 | [37] |
| LDCM0489 | CL94 | HEK-293T | C300(1.23) | LDD1087 | [37] |
| LDCM0490 | CL95 | HEK-293T | C300(1.45) | LDD1088 | [37] |
| LDCM0491 | CL96 | HEK-293T | C300(1.08) | LDD1089 | [37] |
| LDCM0492 | CL97 | HEK-293T | C300(1.07) | LDD1090 | [37] |
| LDCM0493 | CL98 | HEK-293T | C300(1.37) | LDD1091 | [37] |
| LDCM0494 | CL99 | HEK-293T | C300(1.26) | LDD1092 | [37] |
| LDCM0634 | CY-0357 | Hep-G2 | C198(2.08) | LDD2228 | [22] |
| LDCM0495 | E2913 | HEK-293T | C239(0.95); C165(1.00); C198(1.03); C63(1.07) | LDD1698 | [38] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C165(0.70); C239(0.67); C198(0.62) | LDD1702 | [36] |
| LDCM0625 | F8 | Ramos | C165(0.90); C239(0.99); C300(0.50) | LDD2187 | [39] |
| LDCM0572 | Fragment10 | Ramos | C165(0.63); C239(0.70); C300(0.74) | LDD2189 | [39] |
| LDCM0573 | Fragment11 | Ramos | C165(2.22); C239(0.01); C300(0.76); C63(0.38) | LDD2190 | [39] |
| LDCM0574 | Fragment12 | Ramos | C165(0.57); C239(0.42); C300(0.98) | LDD2191 | [39] |
| LDCM0575 | Fragment13 | Ramos | C165(1.15); C239(1.23) | LDD2192 | [39] |
| LDCM0576 | Fragment14 | Ramos | C165(0.42); C239(0.43); C300(1.38) | LDD2193 | [39] |
| LDCM0579 | Fragment20 | Ramos | C165(0.41); C239(0.36); C300(0.80) | LDD2194 | [39] |
| LDCM0580 | Fragment21 | Ramos | C165(0.85); C239(0.86); C300(1.74) | LDD2195 | [39] |
| LDCM0582 | Fragment23 | Ramos | C165(0.13); C239(0.17); C300(1.54); C63(4.60) | LDD2196 | [39] |
| LDCM0578 | Fragment27 | Ramos | C165(1.04); C239(1.03) | LDD2197 | [39] |
| LDCM0586 | Fragment28 | Ramos | C165(0.80); C239(0.48); C300(0.48) | LDD2198 | [39] |
| LDCM0588 | Fragment30 | Ramos | C165(1.42); C239(1.04); C300(0.97) | LDD2199 | [39] |
| LDCM0589 | Fragment31 | Ramos | C165(1.00); C239(0.96) | LDD2200 | [39] |
| LDCM0590 | Fragment32 | Ramos | C165(0.50); C239(0.40); C300(0.74) | LDD2201 | [39] |
| LDCM0468 | Fragment33 | HEK-293T | C300(1.69) | LDD1066 | [37] |
| LDCM0596 | Fragment38 | Ramos | C165(1.14); C239(1.06) | LDD2203 | [39] |
| LDCM0566 | Fragment4 | Ramos | C165(0.79); C239(0.66); C300(0.57) | LDD2184 | [39] |
| LDCM0427 | Fragment51 | HEK-293T | C300(1.47) | LDD1025 | [37] |
| LDCM0610 | Fragment52 | Ramos | C165(1.68); C239(1.22) | LDD2204 | [39] |
| LDCM0614 | Fragment56 | Ramos | C165(1.19); C239(1.09); C300(0.80) | LDD2205 | [39] |
| LDCM0569 | Fragment7 | Ramos | C165(0.55); C239(0.41); C300(0.35); C63(2.73) | LDD2186 | [39] |
| LDCM0571 | Fragment9 | Ramos | C165(0.42); C239(0.39); C300(0.85) | LDD2188 | [39] |
| LDCM0116 | HHS-0101 | DM93 | Y39(0.75); Y266(1.15); Y265(1.70) | LDD0264 | [12] |
| LDCM0117 | HHS-0201 | DM93 | Y39(0.69); Y266(0.87); Y265(1.38) | LDD0265 | [12] |
| LDCM0118 | HHS-0301 | DM93 | Y39(0.63); Y266(1.13); Y265(1.80) | LDD0266 | [12] |
| LDCM0119 | HHS-0401 | DM93 | Y39(0.77); Y266(1.46); Y265(1.71) | LDD0267 | [12] |
| LDCM0120 | HHS-0701 | DM93 | Y39(0.80); Y265(0.91); Y266(1.32) | LDD0268 | [12] |
| LDCM0107 | IAA | HeLa | H192(0.00); C198(0.00); C165(0.00); H167(0.00) | LDD0221 | [26] |
| LDCM0123 | JWB131 | DM93 | Y39(0.61) | LDD0285 | [11] |
| LDCM0125 | JWB146 | DM93 | Y39(0.72) | LDD0287 | [11] |
| LDCM0126 | JWB150 | DM93 | Y39(2.35) | LDD0288 | [11] |
| LDCM0127 | JWB152 | DM93 | Y39(1.23) | LDD0289 | [11] |
| LDCM0128 | JWB198 | DM93 | Y39(1.35) | LDD0290 | [11] |
| LDCM0129 | JWB202 | DM93 | Y39(0.67) | LDD0291 | [11] |
| LDCM0130 | JWB211 | DM93 | Y39(0.62) | LDD0292 | [11] |
| LDCM0022 | KB02 | HEK-293T | C239(1.00); C165(0.97); C198(1.00); C63(1.14) | LDD1492 | [38] |
| LDCM0023 | KB03 | HEK-293T | C239(0.95); C165(1.02); C198(0.95); C63(0.98) | LDD1497 | [38] |
| LDCM0024 | KB05 | G361 | C315(1.53); C180(2.01) | LDD3311 | [8] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C165(0.78); C239(0.88) | LDD2102 | [36] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C165(0.53); C239(0.45) | LDD2121 | [36] |
| LDCM0109 | NEM | HeLa | H373(0.00); H192(0.00); C198(0.00); H167(0.00) | LDD0223 | [26] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C165(0.55); C239(0.81) | LDD2089 | [36] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C165(1.32); C239(1.07) | LDD2090 | [36] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C165(0.73); C239(0.95) | LDD2092 | [36] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C165(0.96); C239(1.10) | LDD2093 | [36] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C165(1.15); C239(1.19) | LDD2094 | [36] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C165(0.96); C239(0.14) | LDD2096 | [36] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C165(0.66); C239(0.73) | LDD2097 | [36] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C165(0.79); C239(0.71) | LDD2098 | [36] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C165(1.01); C239(0.99) | LDD2099 | [36] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C165(0.68); C239(0.64) | LDD2100 | [36] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C165(0.69); C239(1.39) | LDD2101 | [36] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C165(0.84); C239(0.98) | LDD2104 | [36] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C165(1.37); C239(1.34) | LDD2105 | [36] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C165(0.61); C239(0.65) | LDD2106 | [36] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C165(0.97); C239(0.98) | LDD2107 | [36] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C165(0.87); C239(0.73) | LDD2108 | [36] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C165(0.67); C239(0.65) | LDD2109 | [36] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C165(0.88); C239(2.14) | LDD2110 | [36] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C165(0.99); C239(1.11) | LDD2111 | [36] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C165(0.62); C239(0.84) | LDD2114 | [36] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C165(0.44); C239(0.65); C198(0.94) | LDD2115 | [36] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C165(1.13); C239(0.24) | LDD2116 | [36] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C165(0.97); C239(0.21) | LDD2118 | [36] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C165(1.26); C239(2.83) | LDD2119 | [36] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C165(0.79); C239(0.96) | LDD2120 | [36] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C165(1.02); C239(0.19) | LDD2122 | [36] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C165(0.86); C239(0.97); C198(1.23) | LDD2123 | [36] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C165(1.02); C239(0.27) | LDD2124 | [36] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C165(0.84); C239(0.81) | LDD2125 | [36] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C165(0.81); C239(0.30) | LDD2126 | [36] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C165(0.97); C239(1.07); C198(1.21) | LDD2127 | [36] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C165(0.85); C239(1.05) | LDD2128 | [36] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C165(1.03); C239(1.18) | LDD2129 | [36] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C165(0.49); C198(0.55) | LDD2134 | [36] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C165(1.10); C239(1.48) | LDD2135 | [36] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C165(1.19); C239(1.33); C198(2.39) | LDD2136 | [36] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C165(0.93); C239(1.02); C198(2.33) | LDD2137 | [36] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C239(1.85); C165(1.48) | LDD1700 | [36] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C165(0.82); C239(0.77) | LDD2140 | [36] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C165(0.65); C239(0.76) | LDD2141 | [36] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C165(0.98); C239(1.01) | LDD2143 | [36] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C165(1.97); C239(2.78); C198(1.28) | LDD2144 | [36] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C165(4.81); C239(6.62) | LDD2145 | [36] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C165(0.85); C239(0.92) | LDD2146 | [36] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C165(2.54); C239(3.30) | LDD2147 | [36] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C165(0.49); C239(0.51) | LDD2148 | [36] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C165(1.37) | LDD2149 | [36] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C165(0.66); C239(0.39) | LDD2150 | [36] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C165(1.08); C239(0.20) | LDD2151 | [36] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C165(2.04); C239(2.00) | LDD2153 | [36] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C63(2.86) | LDD2206 | [40] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C63(0.84); C165(0.42) | LDD2207 | [40] |
| LDCM0019 | Staurosporine | Hep-G2 | N.A. | LDD0083 | [33] |
| LDCM0021 | THZ1 | HeLa S3 | C75(1.11) | LDD0460 | [9] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| THO complex subunit 4 (ALYREF) | ALYREF family | Q86V81 | |||
| UAP56-interacting factor (FYTTD1) | UIF family | Q96QD9 | |||
| Syntenin-1 (SDCBP) | . | O00560 | |||
Other
References
