Details of the Target
General Information of Target
| Target ID | LDTP14058 | |||||
|---|---|---|---|---|---|---|
| Target Name | Rap guanine nucleotide exchange factor 2 (RAPGEF2) | |||||
| Gene Name | RAPGEF2 | |||||
| Gene ID | 9693 | |||||
| Synonyms |
KIAA0313; NRAPGEP; PDZGEF1; Rap guanine nucleotide exchange factor 2; Cyclic nucleotide ras GEF; CNrasGEF; Neural RAP guanine nucleotide exchange protein; nRap GEP; PDZ domain-containing guanine nucleotide exchange factor 1; PDZ-GEF1; RA-GEF-1; Ras/Rap1-associating GEF-1
|
|||||
| 3D Structure | ||||||
| Sequence |
MVALSLKICVRHCNVVKTMQFEPSTAVYDACRVIRERVPEAQTGQASDYGLFLSDEDPRK
GIWLEAGRTLDYYMLRNGDILEYKKKQRPQKIRMLDGSVKTVMVDDSKTVGELLVTICSR IGITNYEEYSLIQETIEEKKEEGTGTLKKDRTLLRDERKMEKLKAKLHTDDDLNWLDHSR TFREQGVDENETLLLRRKFFYSDQNVDSRDPVQLNLLYVQARDDILNGSHPVSFEKACEF GGFQAQIQFGPHVEHKHKPGFLDLKEFLPKEYIKQRGAEKRIFQEHKNCGEMSEIEAKVK YVKLARSLRTYGVSFFLVKEKMKGKNKLVPRLLGITKDSVMRVDEKTKEVLQEWPLTTVK RWAASPKSFTLDFGEYQESYYSVQTTEGEQISQLIAGYIDIILKKKQSKDRFGLEGDEES TMLEESVSPKKSTILQQQFNRTGKAEHGSVALPAVMRSGSSGPETFNVGSMPSPQQQVMV GQMHRGHMPPLTSAQQALMGTINTSMHAVQQAQDDLSELDSLPPLGQDMASRVWVQNKVD ESKHEIHSQVDAITAGTASVVNLTAGDPADTDYTAVGCAITTISSNLTEMSKGVKLLAAL MDDEVGSGEDLLRAARTLAGAVSDLLKAVQPTSGEPRQTVLTAAGSIGQASGDLLRQIGE NETDERFQDVLMSLAKAVANAAAMLVLKAKNVAQVAEDTVLQNRVIAAATQCALSTSQLV ACAKVVSPTISSPVCQEQLIEAGKLVDRSVENCVRACQAATTDSELLKQVSAAASVVSQA LHDLLQHVRQFASRGEPIGRYDQATDTIMCVTESIFSSMGDAGEMVRQARVLAQATSDLV NAMRSDAEAEIDMENSKKLLAAAKLLADSTARMVEAAKGAAANPENEDQQQRLREAAEGL RVATNAAAQNAIKKKIVNRLEVAAKQAAAAATQTIAASQNAAVSNKNPAAQQQLVQSCKA VADHIPQLVQGVRGSQAQAEDLSAQLALIISSQNFLQPGSKMVSSAKAAVPTVSDQAAAM QLSQCAKNLATSLAELRTASQKAHEACGPMEIDSALNTVQTLKNELQDAKMAAVESQLKP LPGETLEKCAQDLGSTSKAVGSSMAQLLTCAAQGNEHYTGVAARETAQALKTLAQAARGV AASTTDPAAAHAMLDSARDVMEGSAMLIQEAKQALIAPGDAERQQRLAQVAKAVSHSLNN CVNCLPGQKDVDVALKSIGESSKKLLVDSLPPSTKPFQEAQSELNQAAADLNQSAGEVVH ATRGQSGELAAASGKFSDDFDEFLDAGIEMAGQAQTKEDQIQVIGNLKNISMASSKLLLA AKSLSVDPGAPNAKNLLAAAARAVTESINQLITLCTQQAPGQKECDNALRELETVKGMLD NPNEPVSDLSYFDCIESVMENSKVLGESMAGISQNAKTGDLPAFGECVGIASKALCGLTE AAAQAAYLVGISDPNSQAGHQGLVDPIQFARANQAIQMACQNLVDPGSSPSQVLSAATIV AKHTSALCNACRIASSKTANPVAKRHFVQSAKEVANSTANLVKTIKALDGDFSEDNRNKC RIATAPLIEAVENLTAFASNPEFVSIPAQISSEGSQAQEPILVSAKTMLESSSYLIRTAR SLAINPKDPPTWSVLAGHSHTVSDSIKSLITSIRDKAPGQRECDYSIDGINRCIRDIEQA SLAAVSQSLATRDDISVEALQEQLTSVVQEIGHLIDPIATAARGEAAQLGHKVTQLASYF EPLILAAVGVASKILDHQQQMTVLDQTKTLAESALQMLYAAKEGGGNPKAQHTHDAITEA AQLMKEAVDDIMVTLNEAASEVGLVGGMVDAIAEAMSKLDEGTPPEPKGTFVDYQTTVVK YSKAIAVTAQEMMTKSVTNPEELGGLASQMTSDYGHLAFQGQMAAATAEPEEIGFQIRTR VQDLGHGCIFLVQKAGALQVCPTDSYTKRELIECARAVTEKVSLVLSALQAGNKGTQACI TAATAVSGIIADLDTTIMFATAGTLNAENSETFADHRENILKTAKALVEDTKLLVSGAAS TPDKLAQAAQSSAATITQLAEVVKLGAASLGSDDPETQVVLINAIKDVAKALSDLISATK GAASKPVDDPSMYQLKGAAKVMVTNVTSLLKTVKAVEDEATRGTRALEATIECIKQELTV FQSKDVPEKTSSPEESIRMTKGITMATAKAVAAGNSCRQEDVIATANLSRKAVSDMLTAC KQASFHPDVSDEVRTRALRFGTECTLGYLDLLEHVLVILQKPTPEFKQQLAAFSKRVAGA VTELIQAAEAMKGTEWVDPEDPTVIAETELLGAAASIEAAAKKLEQLKPRAKPKQADETL DFEEQILEAAKSIAAATSALVKSASAAQRELVAQGKVGSIPANAADDGQWSQGLISAARM VAAATSSLCEAANASVQGHASEEKLISSAKQVAASTAQLLVACKVKADQDSEAMRRLQAA GNAVKRASDNLVRAAQKAAFGKADDDDVVVKTKFVGGIAQIIAAQEEMLKKERELEEARK KLAQIRQQQYKFLPTELREDEG |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
RAPGEF2 family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Functions as a guanine nucleotide exchange factor (GEF), which activates Rap and Ras family of small GTPases by exchanging bound GDP for free GTP in a cAMP-dependent manner. Serves as a link between cell surface receptors and Rap/Ras GTPases in intracellular signaling cascades. Acts also as an effector for Rap1 by direct association with Rap1-GTP thereby leading to the amplification of Rap1-mediated signaling. Shows weak activity on HRAS. It is controversial whether RAPGEF2 binds cAMP and cGMP or not. Its binding to ligand-activated beta-1 adrenergic receptor ADRB1 leads to the Ras activation through the G(s)-alpha signaling pathway. Involved in the cAMP-induced Ras and Erk1/2 signaling pathway that leads to sustained inhibition of long term melanogenesis by reducing dendrite extension and melanin synthesis. Provides also inhibitory signals for cell proliferation of melanoma cells and promotes their apoptosis in a cAMP-independent nanner. Regulates cAMP-induced neuritogenesis by mediating the Rap1/B-Raf/ERK signaling through a pathway that is independent on both PKA and RAPGEF3/RAPGEF4. Involved in neuron migration and in the formation of the major forebrain fiber connections forming the corpus callosum, the anterior commissure and the hippocampal commissure during brain development. Involved in neuronal growth factor (NGF)-induced sustained activation of Rap1 at late endosomes and in brain-derived neurotrophic factor (BDNF)-induced axon outgrowth of hippocampal neurons. Plays a role in the regulation of embryonic blood vessel formation and in the establishment of basal junction integrity and endothelial barrier function. May be involved in the regulation of the vascular endothelial growth factor receptor KDR and cadherin CDH5 expression at allantois endothelial cell-cell junctions.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C194(3.77) | LDD3318 | [1] | |
|
AHL-Pu-1 Probe Info |
 |
C1252(6.41) | LDD0170 | [2] | |
|
IA-alkyne Probe Info |
 |
C377(0.00); C798(0.00) | LDD0166 | [3] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [4] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [5] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C010 Probe Info |
 |
6.59 | LDD1719 | [6] | |
|
C083 Probe Info |
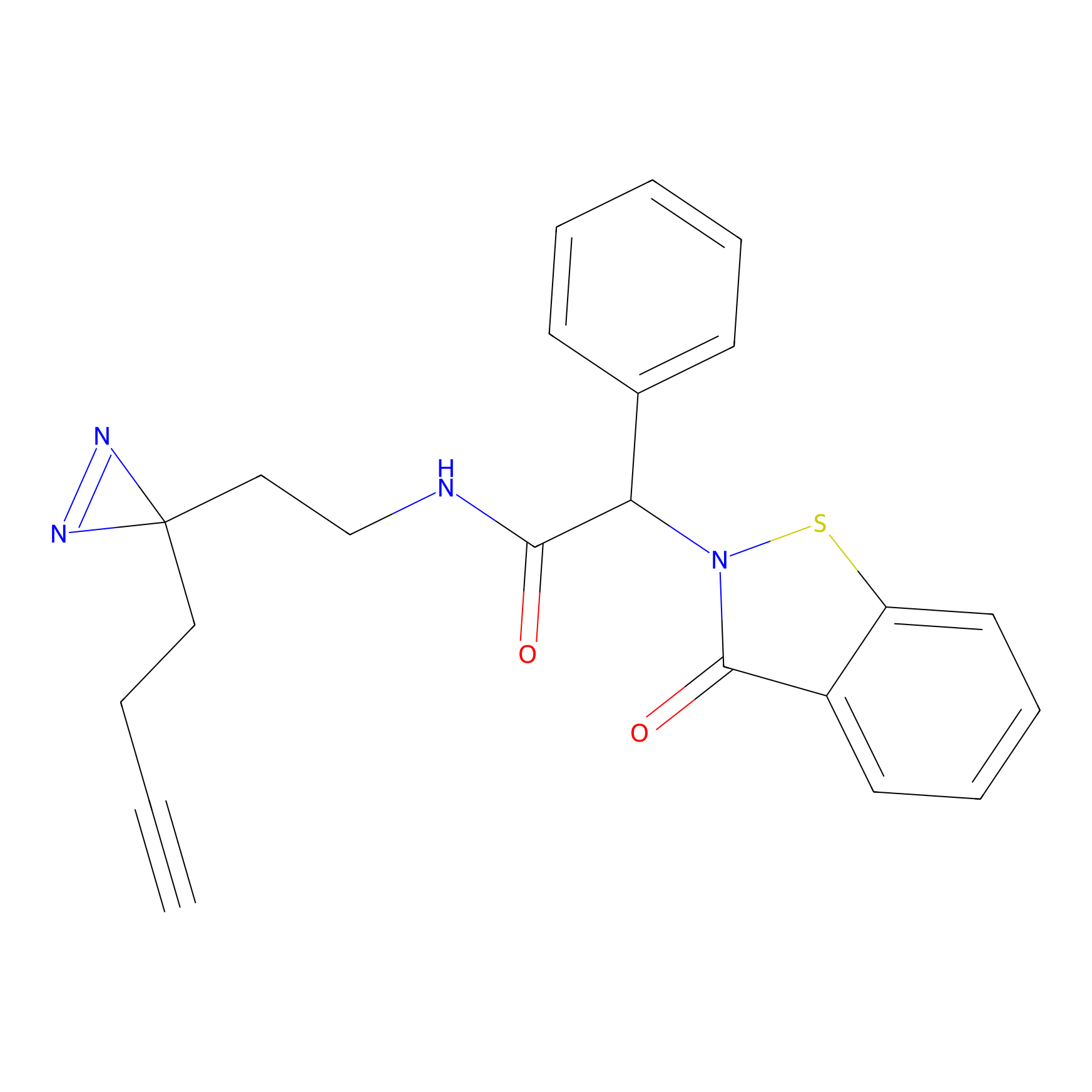 |
6.41 | LDD1775 | [6] | |
|
C086 Probe Info |
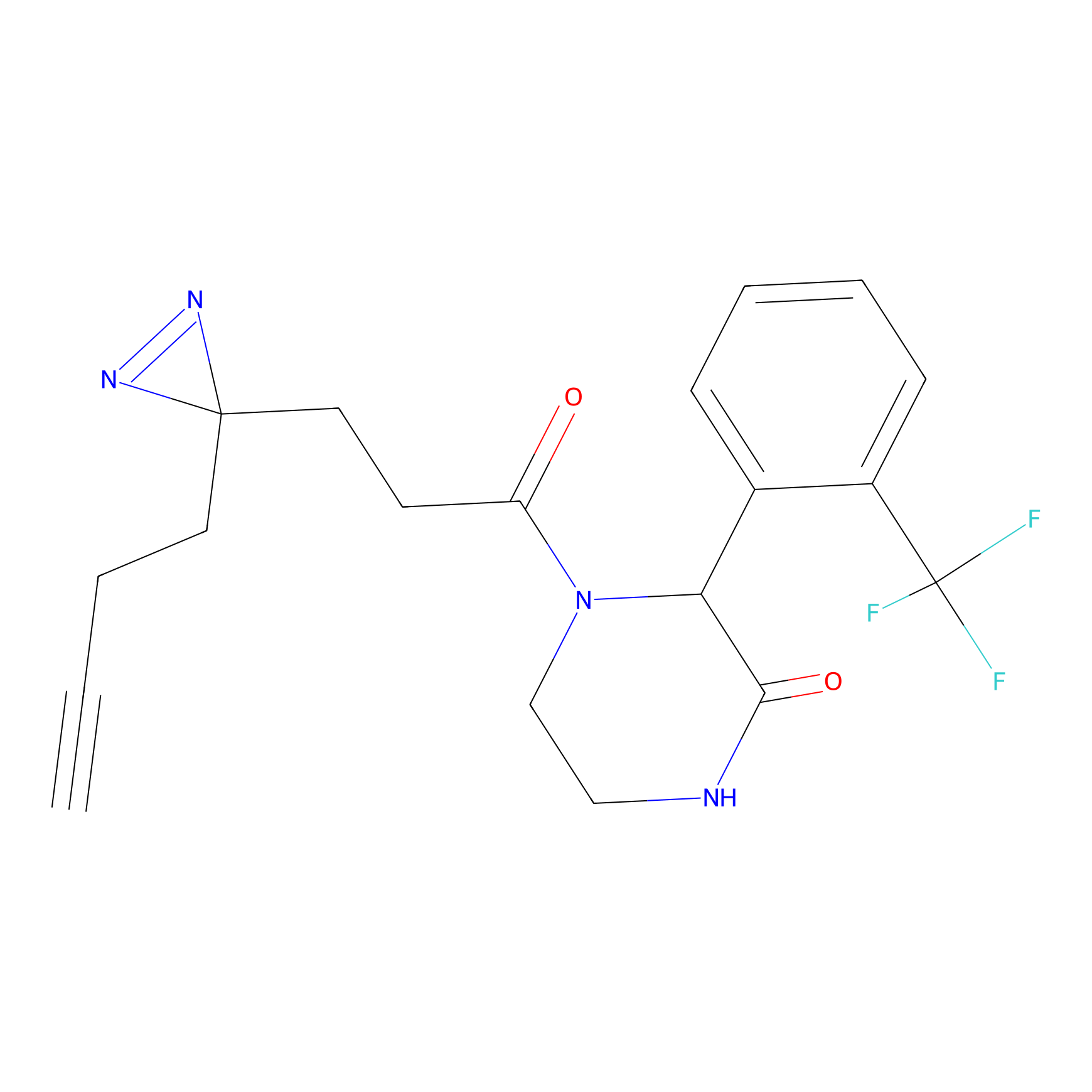 |
6.02 | LDD1778 | [6] | |
|
C095 Probe Info |
 |
4.99 | LDD1786 | [6] | |
|
C145 Probe Info |
 |
8.94 | LDD1827 | [6] | |
|
C173 Probe Info |
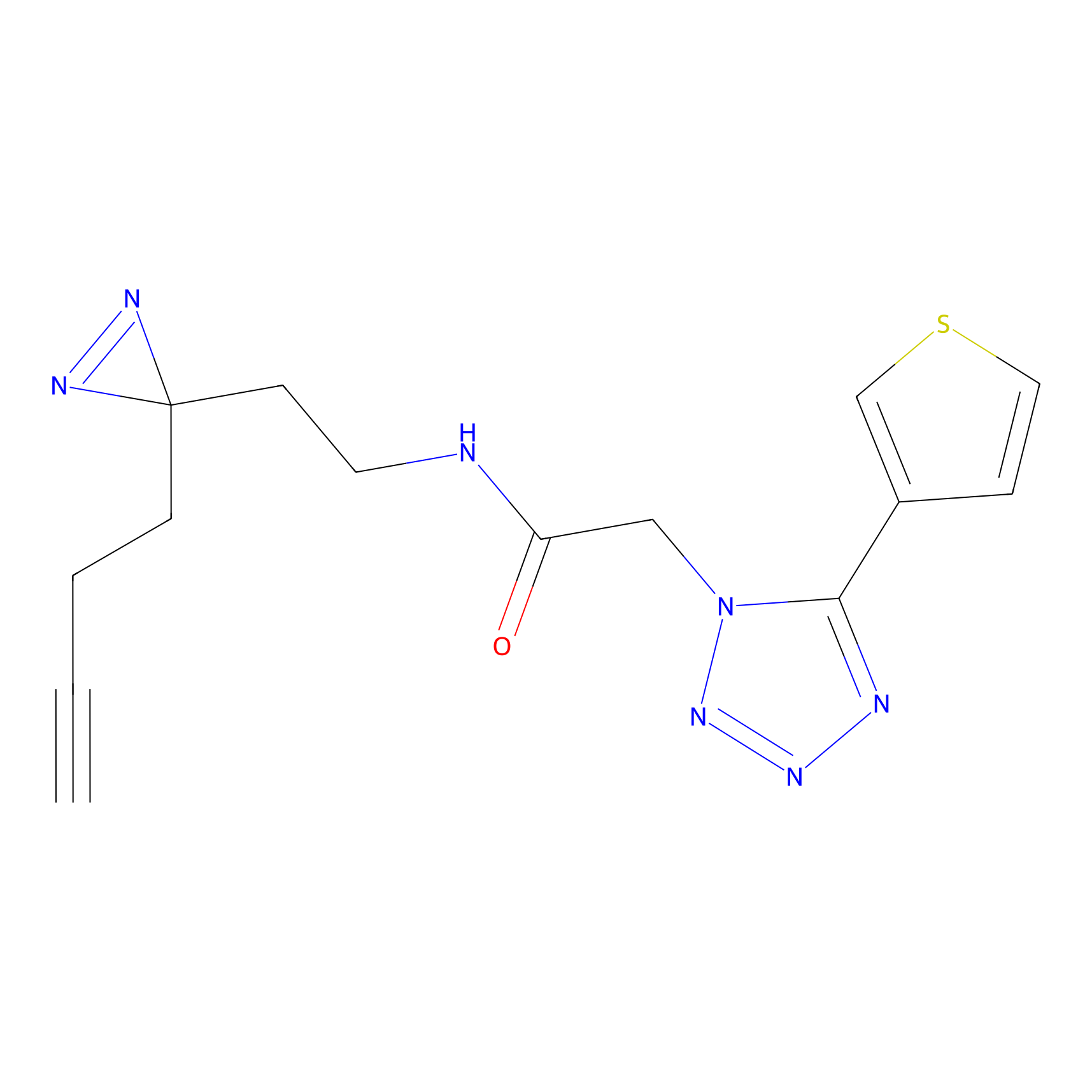 |
9.65 | LDD1853 | [6] | |
|
C185 Probe Info |
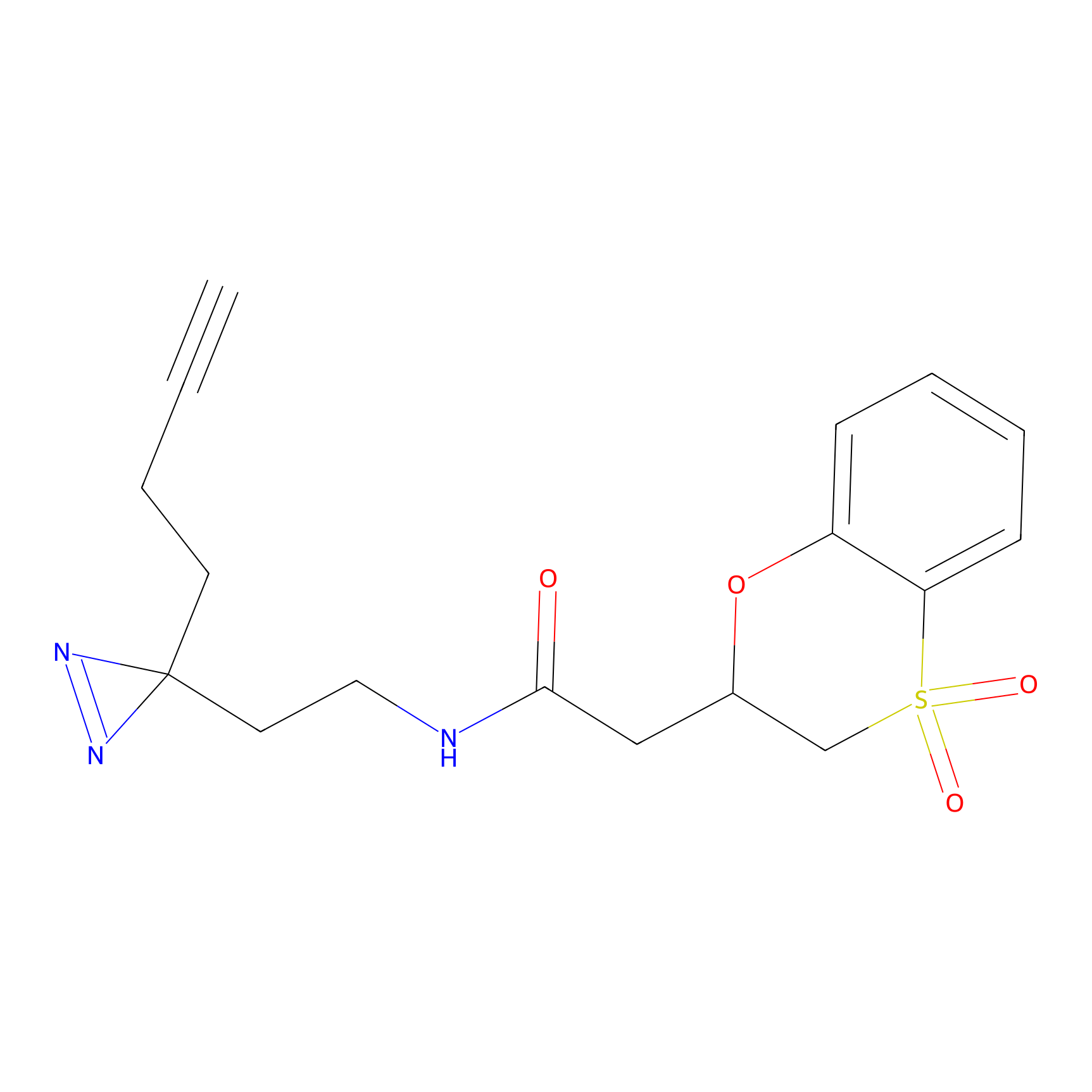 |
4.92 | LDD1863 | [6] | |
|
C193 Probe Info |
 |
7.31 | LDD1869 | [6] | |
|
C214 Probe Info |
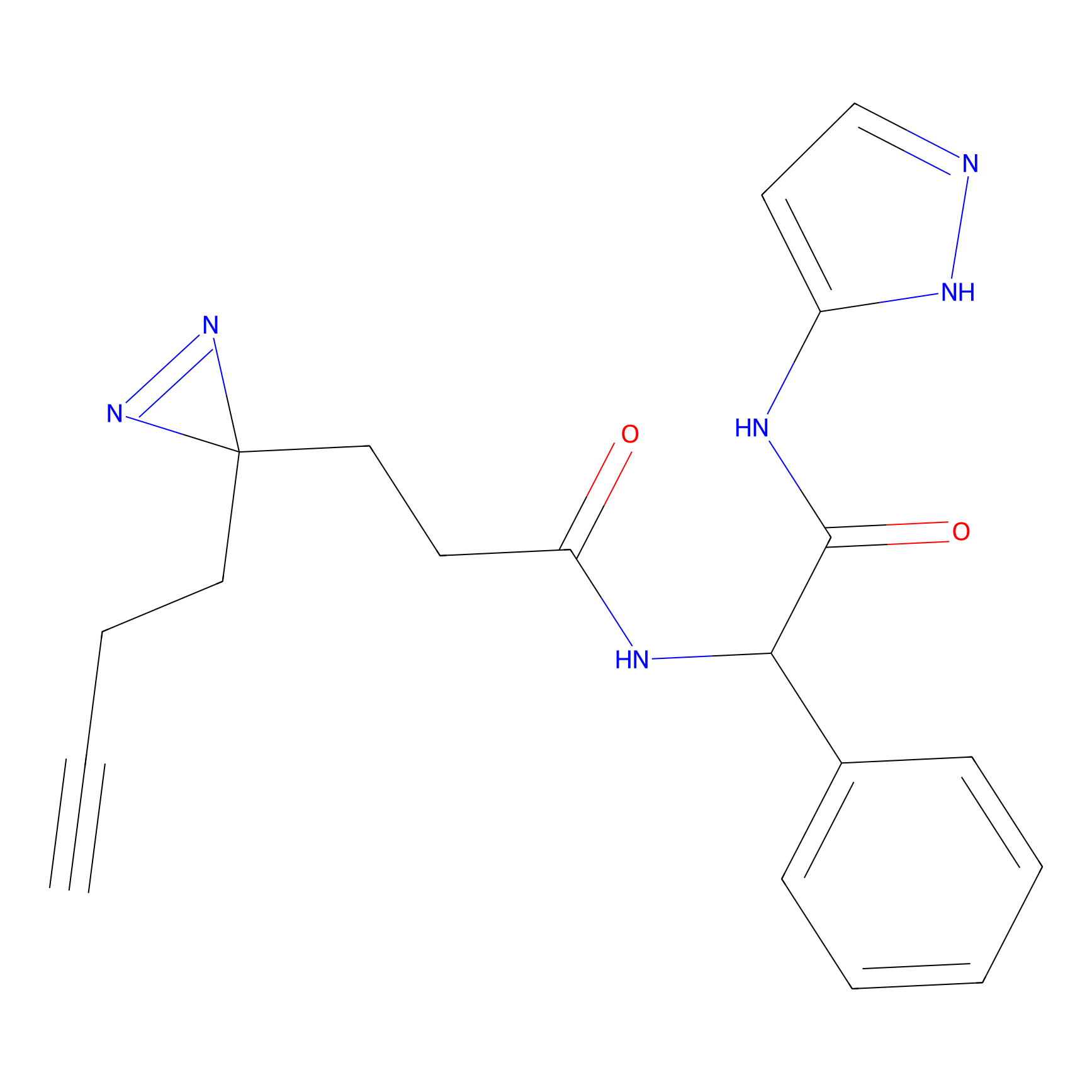 |
5.39 | LDD1888 | [6] | |
|
C252 Probe Info |
 |
9.65 | LDD1925 | [6] | |
|
C255 Probe Info |
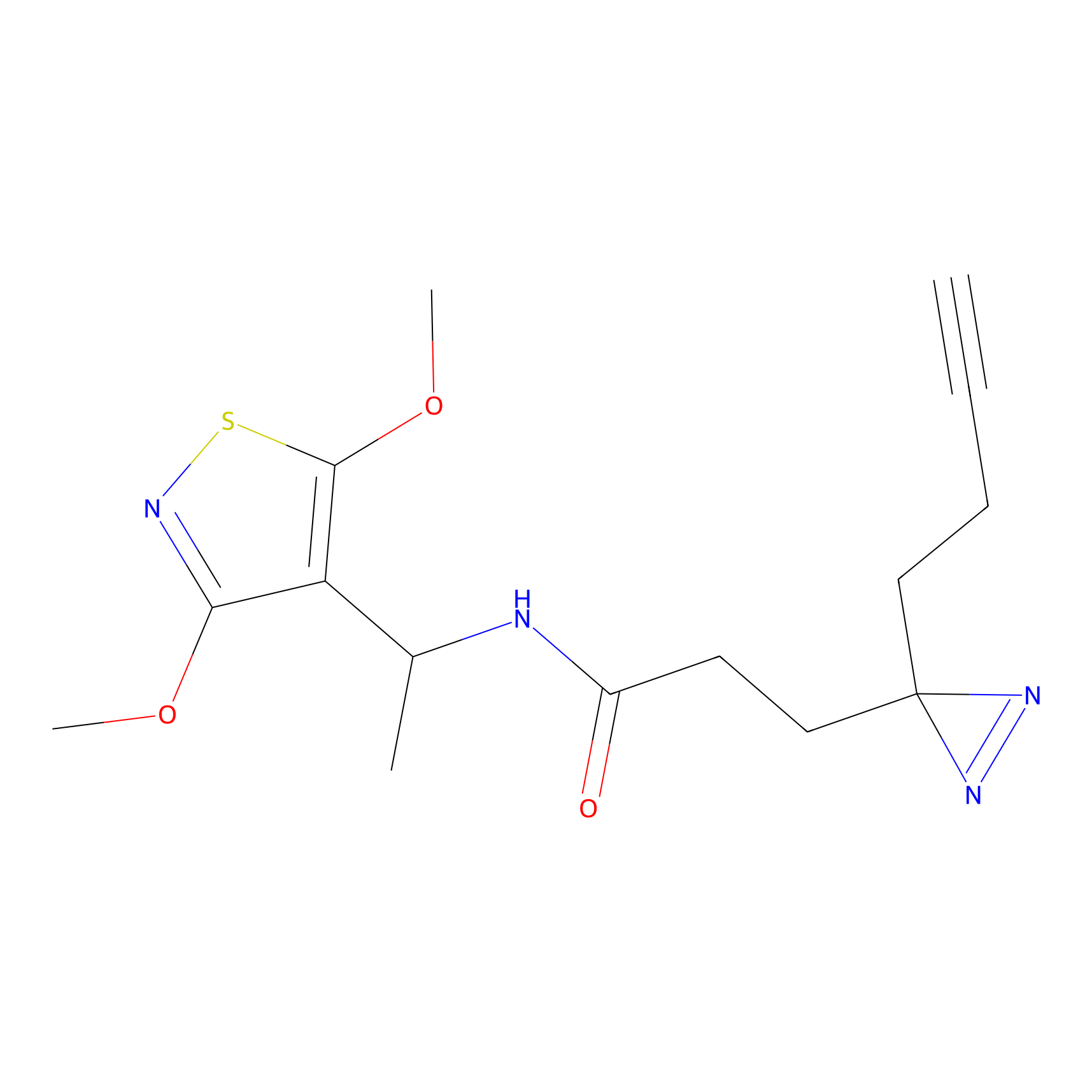 |
7.62 | LDD1928 | [6] | |
|
C272 Probe Info |
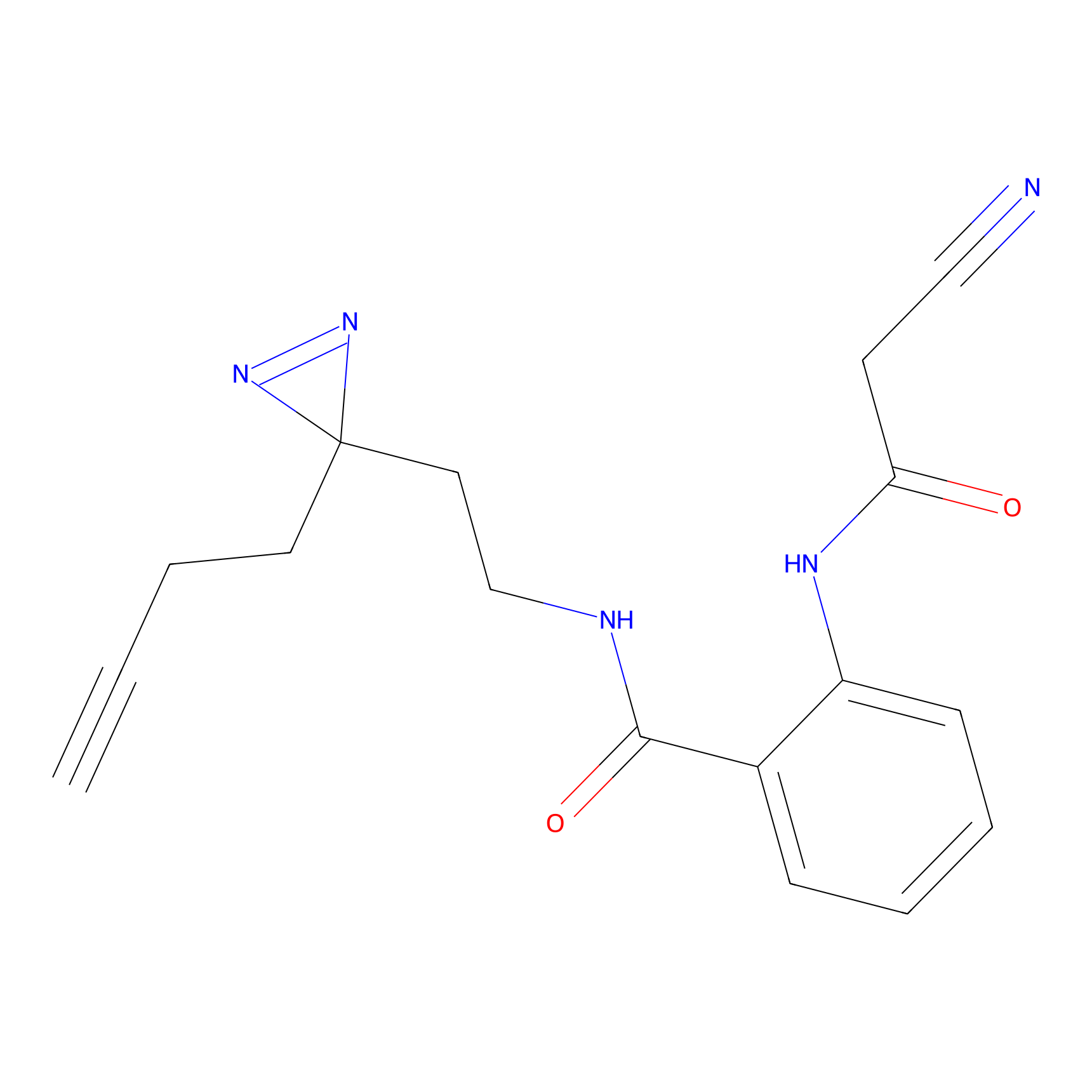 |
4.96 | LDD1942 | [6] | |
|
C288 Probe Info |
 |
6.68 | LDD1958 | [6] | |
|
C297 Probe Info |
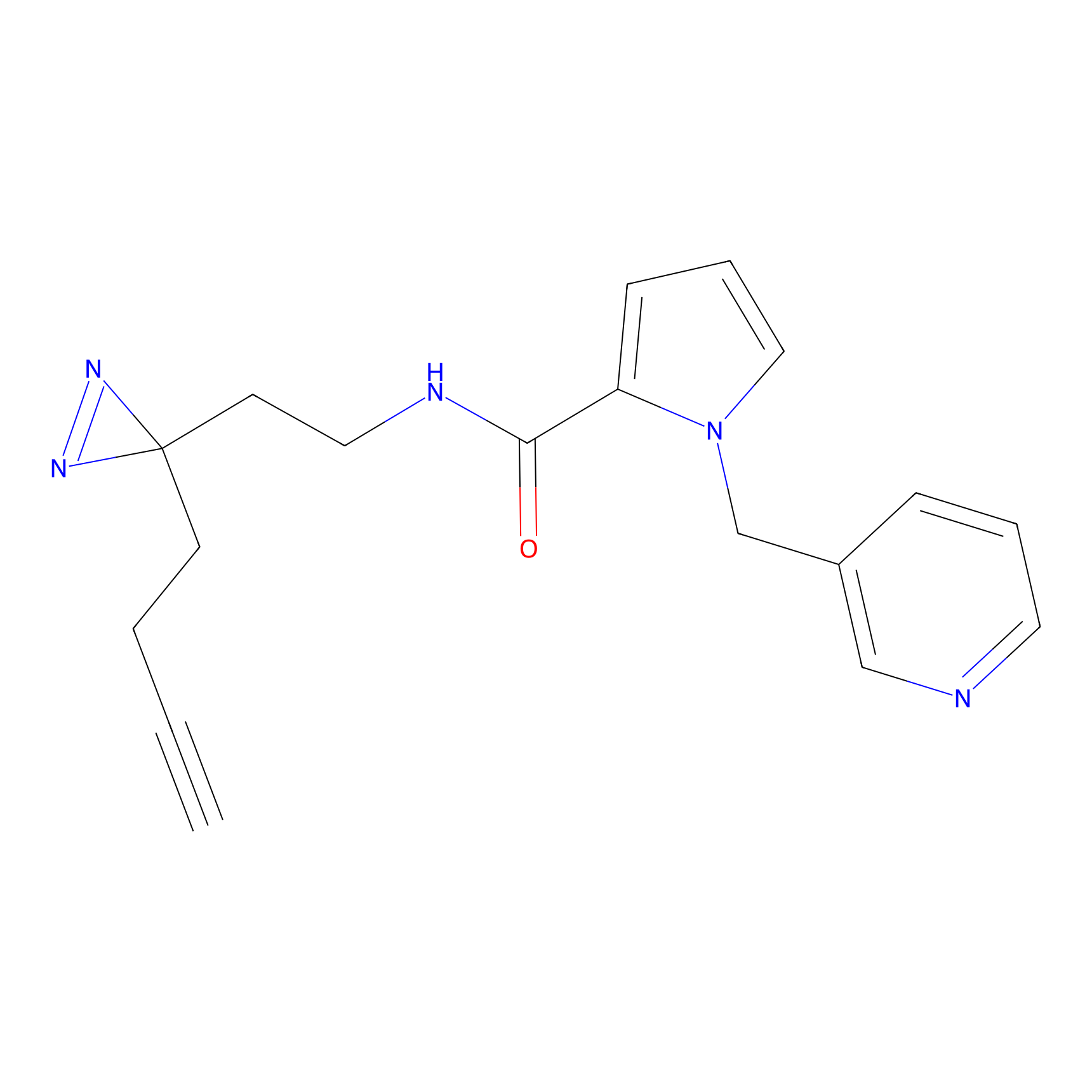 |
7.94 | LDD1967 | [6] | |
|
C347 Probe Info |
 |
6.63 | LDD2008 | [6] | |
|
C356 Probe Info |
 |
8.00 | LDD2017 | [6] | |
|
C366 Probe Info |
 |
5.54 | LDD2027 | [6] | |
|
C380 Probe Info |
 |
5.24 | LDD2039 | [6] | |
|
C394 Probe Info |
 |
5.86 | LDD2053 | [6] | |
|
C419 Probe Info |
 |
9.99 | LDD2074 | [6] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | DM93 | C1252(6.41) | LDD0170 | [2] |
| LDCM0214 | AC1 | HEK-293T | C377(0.86) | LDD1507 | [7] |
| LDCM0215 | AC10 | HEK-293T | C377(0.94) | LDD1508 | [7] |
| LDCM0226 | AC11 | HEK-293T | C377(0.94) | LDD1509 | [7] |
| LDCM0237 | AC12 | HEK-293T | C377(0.96); C194(1.22) | LDD1510 | [7] |
| LDCM0259 | AC14 | HEK-293T | C377(1.05); C194(1.00) | LDD1512 | [7] |
| LDCM0270 | AC15 | HEK-293T | C377(0.91) | LDD1513 | [7] |
| LDCM0276 | AC17 | HEK-293T | C377(0.98) | LDD1515 | [7] |
| LDCM0277 | AC18 | HEK-293T | C377(0.97) | LDD1516 | [7] |
| LDCM0278 | AC19 | HEK-293T | C377(0.92) | LDD1517 | [7] |
| LDCM0279 | AC2 | HEK-293T | C377(0.90) | LDD1518 | [7] |
| LDCM0280 | AC20 | HEK-293T | C377(0.94); C194(1.09) | LDD1519 | [7] |
| LDCM0281 | AC21 | HEK-293T | C377(0.98); C194(1.12) | LDD1520 | [7] |
| LDCM0282 | AC22 | HEK-293T | C377(1.02); C194(0.82) | LDD1521 | [7] |
| LDCM0283 | AC23 | HEK-293T | C377(1.04) | LDD1522 | [7] |
| LDCM0284 | AC24 | HEK-293T | C377(1.01); C194(1.01) | LDD1523 | [7] |
| LDCM0285 | AC25 | HEK-293T | C377(0.93) | LDD1524 | [7] |
| LDCM0286 | AC26 | HEK-293T | C377(0.96) | LDD1525 | [7] |
| LDCM0287 | AC27 | HEK-293T | C377(1.06) | LDD1526 | [7] |
| LDCM0288 | AC28 | HEK-293T | C377(1.05); C194(0.97) | LDD1527 | [7] |
| LDCM0289 | AC29 | HEK-293T | C377(1.00); C194(1.19) | LDD1528 | [7] |
| LDCM0290 | AC3 | HEK-293T | C377(0.85) | LDD1529 | [7] |
| LDCM0291 | AC30 | HEK-293T | C377(1.07); C194(0.93) | LDD1530 | [7] |
| LDCM0292 | AC31 | HEK-293T | C377(1.03) | LDD1531 | [7] |
| LDCM0293 | AC32 | HEK-293T | C377(0.99); C194(0.93) | LDD1532 | [7] |
| LDCM0294 | AC33 | HEK-293T | C377(0.89) | LDD1533 | [7] |
| LDCM0295 | AC34 | HEK-293T | C377(0.93) | LDD1534 | [7] |
| LDCM0296 | AC35 | HEK-293T | C377(0.90) | LDD1535 | [7] |
| LDCM0297 | AC36 | HEK-293T | C377(0.95); C194(0.98) | LDD1536 | [7] |
| LDCM0298 | AC37 | HEK-293T | C377(1.00); C194(1.27) | LDD1537 | [7] |
| LDCM0299 | AC38 | HEK-293T | C377(1.09); C194(1.00) | LDD1538 | [7] |
| LDCM0300 | AC39 | HEK-293T | C377(0.93) | LDD1539 | [7] |
| LDCM0301 | AC4 | HEK-293T | C377(0.85); C194(1.03) | LDD1540 | [7] |
| LDCM0302 | AC40 | HEK-293T | C377(0.85); C194(0.93) | LDD1541 | [7] |
| LDCM0303 | AC41 | HEK-293T | C377(0.89) | LDD1542 | [7] |
| LDCM0304 | AC42 | HEK-293T | C377(0.93) | LDD1543 | [7] |
| LDCM0305 | AC43 | HEK-293T | C377(0.95) | LDD1544 | [7] |
| LDCM0306 | AC44 | HEK-293T | C377(0.91); C194(1.08) | LDD1545 | [7] |
| LDCM0307 | AC45 | HEK-293T | C377(1.03); C194(1.38) | LDD1546 | [7] |
| LDCM0308 | AC46 | HEK-293T | C377(0.99); C194(0.94) | LDD1547 | [7] |
| LDCM0309 | AC47 | HEK-293T | C377(1.03) | LDD1548 | [7] |
| LDCM0310 | AC48 | HEK-293T | C377(0.88); C194(1.00) | LDD1549 | [7] |
| LDCM0311 | AC49 | HEK-293T | C377(0.99) | LDD1550 | [7] |
| LDCM0312 | AC5 | HEK-293T | C377(0.97); C194(1.09) | LDD1551 | [7] |
| LDCM0313 | AC50 | HEK-293T | C377(0.98) | LDD1552 | [7] |
| LDCM0314 | AC51 | HEK-293T | C377(0.91) | LDD1553 | [7] |
| LDCM0315 | AC52 | HEK-293T | C377(0.95); C194(0.97) | LDD1554 | [7] |
| LDCM0316 | AC53 | HEK-293T | C377(1.00); C194(1.22) | LDD1555 | [7] |
| LDCM0317 | AC54 | HEK-293T | C377(0.94); C194(0.91) | LDD1556 | [7] |
| LDCM0318 | AC55 | HEK-293T | C377(0.95) | LDD1557 | [7] |
| LDCM0319 | AC56 | HEK-293T | C377(0.97); C194(0.97) | LDD1558 | [7] |
| LDCM0320 | AC57 | HEK-293T | C377(0.98) | LDD1559 | [7] |
| LDCM0321 | AC58 | HEK-293T | C377(0.97) | LDD1560 | [7] |
| LDCM0322 | AC59 | HEK-293T | C377(0.93) | LDD1561 | [7] |
| LDCM0323 | AC6 | HEK-293T | C377(0.95); C194(1.15) | LDD1562 | [7] |
| LDCM0324 | AC60 | HEK-293T | C377(1.01); C194(1.16) | LDD1563 | [7] |
| LDCM0325 | AC61 | HEK-293T | C377(0.99); C194(1.25) | LDD1564 | [7] |
| LDCM0326 | AC62 | HEK-293T | C377(0.94); C194(0.99) | LDD1565 | [7] |
| LDCM0327 | AC63 | HEK-293T | C377(0.94) | LDD1566 | [7] |
| LDCM0328 | AC64 | HEK-293T | C377(0.87); C194(0.99) | LDD1567 | [7] |
| LDCM0334 | AC7 | HEK-293T | C377(0.89) | LDD1568 | [7] |
| LDCM0345 | AC8 | HEK-293T | C377(0.86); C194(0.97) | LDD1569 | [7] |
| LDCM0248 | AKOS034007472 | HEK-293T | C377(0.96); C194(1.18) | LDD1511 | [7] |
| LDCM0356 | AKOS034007680 | HEK-293T | C377(0.93) | LDD1570 | [7] |
| LDCM0275 | AKOS034007705 | HEK-293T | C377(0.94); C194(0.98) | LDD1514 | [7] |
| LDCM0367 | CL1 | HEK-293T | C377(1.11) | LDD1571 | [7] |
| LDCM0368 | CL10 | HEK-293T | C377(1.73); C194(0.96) | LDD1572 | [7] |
| LDCM0369 | CL100 | HEK-293T | C377(0.90) | LDD1573 | [7] |
| LDCM0370 | CL101 | HEK-293T | C377(1.00) | LDD1574 | [7] |
| LDCM0371 | CL102 | HEK-293T | C377(0.97) | LDD1575 | [7] |
| LDCM0372 | CL103 | HEK-293T | C377(0.96); C194(0.89) | LDD1576 | [7] |
| LDCM0373 | CL104 | HEK-293T | C377(0.98) | LDD1577 | [7] |
| LDCM0374 | CL105 | HEK-293T | C377(1.06) | LDD1578 | [7] |
| LDCM0375 | CL106 | HEK-293T | C377(1.11) | LDD1579 | [7] |
| LDCM0376 | CL107 | HEK-293T | C377(0.98); C194(0.99) | LDD1580 | [7] |
| LDCM0377 | CL108 | HEK-293T | C377(0.98) | LDD1581 | [7] |
| LDCM0378 | CL109 | HEK-293T | C377(1.09) | LDD1582 | [7] |
| LDCM0379 | CL11 | HEK-293T | C377(0.98) | LDD1583 | [7] |
| LDCM0380 | CL110 | HEK-293T | C377(1.06) | LDD1584 | [7] |
| LDCM0381 | CL111 | HEK-293T | C377(0.93); C194(0.87) | LDD1585 | [7] |
| LDCM0382 | CL112 | HEK-293T | C377(1.10) | LDD1586 | [7] |
| LDCM0383 | CL113 | HEK-293T | C377(0.95) | LDD1587 | [7] |
| LDCM0384 | CL114 | HEK-293T | C377(1.03) | LDD1588 | [7] |
| LDCM0385 | CL115 | HEK-293T | C377(0.96); C194(0.73) | LDD1589 | [7] |
| LDCM0386 | CL116 | HEK-293T | C377(0.95) | LDD1590 | [7] |
| LDCM0387 | CL117 | HEK-293T | C377(1.02) | LDD1591 | [7] |
| LDCM0388 | CL118 | HEK-293T | C377(0.98) | LDD1592 | [7] |
| LDCM0389 | CL119 | HEK-293T | C377(0.90); C194(0.83) | LDD1593 | [7] |
| LDCM0390 | CL12 | HEK-293T | C377(1.13); C194(1.02) | LDD1594 | [7] |
| LDCM0391 | CL120 | HEK-293T | C377(0.98) | LDD1595 | [7] |
| LDCM0392 | CL121 | HEK-293T | C377(0.98) | LDD1596 | [7] |
| LDCM0393 | CL122 | HEK-293T | C377(0.99) | LDD1597 | [7] |
| LDCM0394 | CL123 | HEK-293T | C377(1.09); C194(0.86) | LDD1598 | [7] |
| LDCM0395 | CL124 | HEK-293T | C377(0.97) | LDD1599 | [7] |
| LDCM0396 | CL125 | HEK-293T | C377(1.03) | LDD1600 | [7] |
| LDCM0397 | CL126 | HEK-293T | C377(0.93) | LDD1601 | [7] |
| LDCM0398 | CL127 | HEK-293T | C377(0.95); C194(1.06) | LDD1602 | [7] |
| LDCM0399 | CL128 | HEK-293T | C377(0.99) | LDD1603 | [7] |
| LDCM0400 | CL13 | HEK-293T | C377(0.96) | LDD1604 | [7] |
| LDCM0401 | CL14 | HEK-293T | C377(0.95) | LDD1605 | [7] |
| LDCM0402 | CL15 | HEK-293T | C377(0.85); C194(0.86) | LDD1606 | [7] |
| LDCM0403 | CL16 | HEK-293T | C377(0.94) | LDD1607 | [7] |
| LDCM0404 | CL17 | HEK-293T | C377(0.98) | LDD1608 | [7] |
| LDCM0405 | CL18 | HEK-293T | C377(0.87) | LDD1609 | [7] |
| LDCM0406 | CL19 | HEK-293T | C377(0.90) | LDD1610 | [7] |
| LDCM0407 | CL2 | HEK-293T | C377(0.93) | LDD1611 | [7] |
| LDCM0408 | CL20 | HEK-293T | C377(0.98); C194(1.30) | LDD1612 | [7] |
| LDCM0409 | CL21 | HEK-293T | C377(1.08); C194(0.54) | LDD1613 | [7] |
| LDCM0410 | CL22 | HEK-293T | C377(1.45); C194(0.99) | LDD1614 | [7] |
| LDCM0411 | CL23 | HEK-293T | C377(0.97) | LDD1615 | [7] |
| LDCM0412 | CL24 | HEK-293T | C377(1.21); C194(1.04) | LDD1616 | [7] |
| LDCM0413 | CL25 | HEK-293T | C377(1.03) | LDD1617 | [7] |
| LDCM0414 | CL26 | HEK-293T | C377(0.98) | LDD1618 | [7] |
| LDCM0415 | CL27 | HEK-293T | C377(0.88); C194(0.97) | LDD1619 | [7] |
| LDCM0416 | CL28 | HEK-293T | C377(0.94) | LDD1620 | [7] |
| LDCM0417 | CL29 | HEK-293T | C377(1.02) | LDD1621 | [7] |
| LDCM0418 | CL3 | HEK-293T | C377(0.92); C194(1.06) | LDD1622 | [7] |
| LDCM0419 | CL30 | HEK-293T | C377(0.94) | LDD1623 | [7] |
| LDCM0420 | CL31 | HEK-293T | C377(0.96) | LDD1624 | [7] |
| LDCM0421 | CL32 | HEK-293T | C377(0.99); C194(1.03) | LDD1625 | [7] |
| LDCM0422 | CL33 | HEK-293T | C377(1.10); C194(1.24) | LDD1626 | [7] |
| LDCM0423 | CL34 | HEK-293T | C377(1.47); C194(0.93) | LDD1627 | [7] |
| LDCM0424 | CL35 | HEK-293T | C377(1.12) | LDD1628 | [7] |
| LDCM0425 | CL36 | HEK-293T | C377(1.24); C194(0.96) | LDD1629 | [7] |
| LDCM0426 | CL37 | HEK-293T | C377(1.13) | LDD1630 | [7] |
| LDCM0428 | CL39 | HEK-293T | C377(1.05); C194(1.08) | LDD1632 | [7] |
| LDCM0429 | CL4 | HEK-293T | C377(1.00) | LDD1633 | [7] |
| LDCM0430 | CL40 | HEK-293T | C377(0.97) | LDD1634 | [7] |
| LDCM0431 | CL41 | HEK-293T | C377(0.97) | LDD1635 | [7] |
| LDCM0432 | CL42 | HEK-293T | C377(1.05) | LDD1636 | [7] |
| LDCM0433 | CL43 | HEK-293T | C377(1.00) | LDD1637 | [7] |
| LDCM0434 | CL44 | HEK-293T | C377(1.05); C194(1.02) | LDD1638 | [7] |
| LDCM0435 | CL45 | HEK-293T | C377(1.14); C194(0.92) | LDD1639 | [7] |
| LDCM0436 | CL46 | HEK-293T | C377(1.38); C194(1.18) | LDD1640 | [7] |
| LDCM0437 | CL47 | HEK-293T | C377(1.09) | LDD1641 | [7] |
| LDCM0438 | CL48 | HEK-293T | C377(1.16); C194(0.87) | LDD1642 | [7] |
| LDCM0439 | CL49 | HEK-293T | C377(1.00) | LDD1643 | [7] |
| LDCM0440 | CL5 | HEK-293T | C377(0.91) | LDD1644 | [7] |
| LDCM0441 | CL50 | HEK-293T | C377(1.01) | LDD1645 | [7] |
| LDCM0443 | CL52 | HEK-293T | C377(0.92) | LDD1646 | [7] |
| LDCM0444 | CL53 | HEK-293T | C377(0.97) | LDD1647 | [7] |
| LDCM0445 | CL54 | HEK-293T | C377(0.92) | LDD1648 | [7] |
| LDCM0446 | CL55 | HEK-293T | C377(0.96) | LDD1649 | [7] |
| LDCM0447 | CL56 | HEK-293T | C377(1.01); C194(1.28) | LDD1650 | [7] |
| LDCM0448 | CL57 | HEK-293T | C377(0.99); C194(0.92) | LDD1651 | [7] |
| LDCM0449 | CL58 | HEK-293T | C377(1.11); C194(1.20) | LDD1652 | [7] |
| LDCM0450 | CL59 | HEK-293T | C377(0.97) | LDD1653 | [7] |
| LDCM0451 | CL6 | HEK-293T | C377(1.03) | LDD1654 | [7] |
| LDCM0452 | CL60 | HEK-293T | C377(1.08); C194(0.98) | LDD1655 | [7] |
| LDCM0453 | CL61 | HEK-293T | C377(0.96) | LDD1656 | [7] |
| LDCM0454 | CL62 | HEK-293T | C377(0.98) | LDD1657 | [7] |
| LDCM0455 | CL63 | HEK-293T | C377(0.99); C194(0.99) | LDD1658 | [7] |
| LDCM0456 | CL64 | HEK-293T | C377(1.03) | LDD1659 | [7] |
| LDCM0457 | CL65 | HEK-293T | C377(0.99) | LDD1660 | [7] |
| LDCM0458 | CL66 | HEK-293T | C377(0.94) | LDD1661 | [7] |
| LDCM0459 | CL67 | HEK-293T | C377(0.86) | LDD1662 | [7] |
| LDCM0460 | CL68 | HEK-293T | C377(1.04); C194(1.22) | LDD1663 | [7] |
| LDCM0461 | CL69 | HEK-293T | C377(1.07); C194(1.03) | LDD1664 | [7] |
| LDCM0462 | CL7 | HEK-293T | C377(0.89) | LDD1665 | [7] |
| LDCM0463 | CL70 | HEK-293T | C377(1.46); C194(1.02) | LDD1666 | [7] |
| LDCM0464 | CL71 | HEK-293T | C377(1.05) | LDD1667 | [7] |
| LDCM0465 | CL72 | HEK-293T | C377(1.06); C194(1.07) | LDD1668 | [7] |
| LDCM0466 | CL73 | HEK-293T | C377(1.19) | LDD1669 | [7] |
| LDCM0467 | CL74 | HEK-293T | C377(1.06) | LDD1670 | [7] |
| LDCM0469 | CL76 | HEK-293T | C377(0.91) | LDD1672 | [7] |
| LDCM0470 | CL77 | HEK-293T | C377(1.03) | LDD1673 | [7] |
| LDCM0471 | CL78 | HEK-293T | C377(0.84) | LDD1674 | [7] |
| LDCM0472 | CL79 | HEK-293T | C377(0.99) | LDD1675 | [7] |
| LDCM0473 | CL8 | HEK-293T | C377(1.13); C194(1.03) | LDD1676 | [7] |
| LDCM0474 | CL80 | HEK-293T | C377(1.00); C194(0.87) | LDD1677 | [7] |
| LDCM0475 | CL81 | HEK-293T | C377(1.04); C194(1.17) | LDD1678 | [7] |
| LDCM0476 | CL82 | HEK-293T | C377(1.47); C194(0.93) | LDD1679 | [7] |
| LDCM0477 | CL83 | HEK-293T | C377(1.08) | LDD1680 | [7] |
| LDCM0478 | CL84 | HEK-293T | C377(1.08); C194(1.00) | LDD1681 | [7] |
| LDCM0479 | CL85 | HEK-293T | C377(0.96) | LDD1682 | [7] |
| LDCM0480 | CL86 | HEK-293T | C377(1.12) | LDD1683 | [7] |
| LDCM0481 | CL87 | HEK-293T | C377(0.92); C194(0.81) | LDD1684 | [7] |
| LDCM0482 | CL88 | HEK-293T | C377(0.91) | LDD1685 | [7] |
| LDCM0483 | CL89 | HEK-293T | C377(0.94) | LDD1686 | [7] |
| LDCM0484 | CL9 | HEK-293T | C377(1.00); C194(1.11) | LDD1687 | [7] |
| LDCM0485 | CL90 | HEK-293T | C377(1.06) | LDD1688 | [7] |
| LDCM0486 | CL91 | HEK-293T | C377(1.02) | LDD1689 | [7] |
| LDCM0487 | CL92 | HEK-293T | C377(1.03); C194(0.98) | LDD1690 | [7] |
| LDCM0488 | CL93 | HEK-293T | C377(1.06); C194(1.32) | LDD1691 | [7] |
| LDCM0489 | CL94 | HEK-293T | C377(1.21); C194(0.93) | LDD1692 | [7] |
| LDCM0490 | CL95 | HEK-293T | C377(1.12) | LDD1693 | [7] |
| LDCM0491 | CL96 | HEK-293T | C377(1.20); C194(0.82) | LDD1694 | [7] |
| LDCM0492 | CL97 | HEK-293T | C377(1.16) | LDD1695 | [7] |
| LDCM0493 | CL98 | HEK-293T | C377(1.03) | LDD1696 | [7] |
| LDCM0494 | CL99 | HEK-293T | C377(0.96); C194(0.84) | LDD1697 | [7] |
| LDCM0495 | E2913 | HEK-293T | C377(1.07); C194(0.93) | LDD1698 | [7] |
| LDCM0468 | Fragment33 | HEK-293T | C377(0.91); C194(0.92) | LDD1671 | [7] |
| LDCM0427 | Fragment51 | HEK-293T | C377(1.01) | LDD1631 | [7] |
| LDCM0022 | KB02 | Calu-1 | C194(1.41) | LDD2292 | [1] |
| LDCM0023 | KB03 | Calu-1 | C194(1.47) | LDD2709 | [1] |
| LDCM0024 | KB05 | MELHO | C194(3.77) | LDD3318 | [1] |
References
