Details of the Target
General Information of Target
| Target ID | LDTP05378 | |||||
|---|---|---|---|---|---|---|
| Target Name | Large ribosomal subunit protein eL6 (RPL6) | |||||
| Gene Name | RPL6 | |||||
| Gene ID | 6128 | |||||
| Synonyms |
TXREB1; Large ribosomal subunit protein eL6; 60S ribosomal protein L6; Neoplasm-related protein C140; Tax-responsive enhancer element-binding protein 107; TaxREB107 |
|||||
| 3D Structure | ||||||
| Sequence |
MAGEKVEKPDTKEKKPEAKKVDAGGKVKKGNLKAKKPKKGKPHCSRNPVLVRGIGRYSRS
AMYSRKAMYKRKYSAAKSKVEKKKKEKVLATVTKPVGGDKNGGTRVVKLRKMPRYYPTED VPRKLLSHGKKPFSQHVRKLRASITPGTILIILTGRHRGKRVVFLKQLASGLLLVTGPLV LNRVPLRRTHQKFVIATSTKIDISNVKIPKHLTDAYFKKKKLRKPRHQEGEIFDTEKEKY EITEQRKIDQKAVDSQILPKIKAIPQLQGYLRSVFALTNGIYPHKLVF |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Eukaryotic ribosomal protein eL6 family
|
|||||
| Subcellular location |
Cytoplasm, cytosol
|
|||||
| Function |
Component of the large ribosomal subunit. The ribosome is a large ribonucleoprotein complex responsible for the synthesis of proteins in the cell.; (Microbial infection) Specifically binds to domain C of the Tax-responsive enhancer element in the long terminal repeat of HTLV-I.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
9.24 | LDD0402 | [1] | |
|
P3 Probe Info |
 |
1.69 | LDD0450 | [2] | |
|
A-EBA Probe Info |
 |
4.00 | LDD0215 | [3] | |
|
TH211 Probe Info |
 |
Y282(20.00); Y216(7.79) | LDD0260 | [4] | |
|
C-Sul Probe Info |
 |
4.02 | LDD0066 | [5] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [6] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [6] | |
|
BTD Probe Info |
 |
C44(6.71) | LDD1699 | [7] | |
|
ONAyne Probe Info |
 |
K20(0.00); K94(0.00); K207(0.00); K237(0.00) | LDD0273 | [8] | |
|
Probe 1 Probe Info |
 |
Y115(28.47); Y116(14.16); Y270(12.98) | LDD3495 | [9] | |
|
AZ-9 Probe Info |
 |
10.00 | LDD2154 | [10] | |
|
HHS-482 Probe Info |
 |
Y115(20.00) | LDD0285 | [11] | |
|
DBIA Probe Info |
 |
C44(1.64) | LDD0547 | [12] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [13] | |
|
ATP probe Probe Info |
 |
K87(0.00); K207(0.00); K237(0.00); K210(0.00) | LDD0199 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [14] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [14] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [15] | |
|
IPM Probe Info |
 |
N.A. | LDD0147 | [16] | |
|
NHS Probe Info |
 |
K260(0.00); K87(0.00); K5(0.00); K251(0.00) | LDD0010 | [15] | |
|
OSF Probe Info |
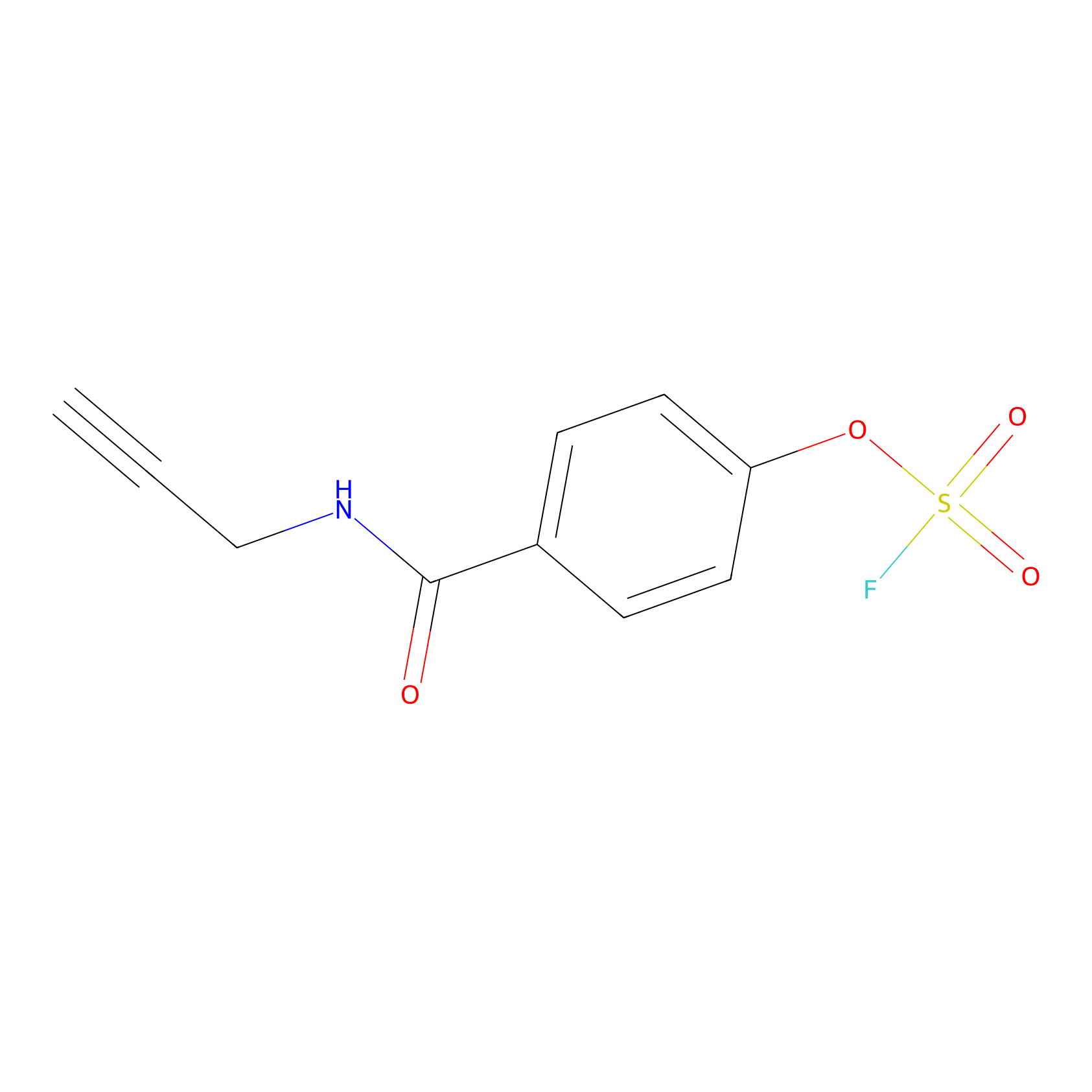 |
Y115(0.00); H128(0.00) | LDD0029 | [17] | |
|
SF Probe Info |
 |
Y240(0.00); K5(0.00); K260(0.00); Y282(0.00) | LDD0028 | [17] | |
|
STPyne Probe Info |
 |
K207(0.00); K251(0.00) | LDD0009 | [15] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [16] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [18] | |
|
1c-yne Probe Info |
 |
K237(0.00); K192(0.00); K251(0.00) | LDD0228 | [19] | |
|
Acrolein Probe Info |
 |
H227(0.00); H211(0.00); H128(0.00) | LDD0217 | [20] | |
|
Crotonaldehyde Probe Info |
 |
H227(0.00); H211(0.00); H136(0.00) | LDD0219 | [20] | |
|
Methacrolein Probe Info |
 |
C44(0.00); H136(0.00) | LDD0218 | [20] | |
|
AOyne Probe Info |
 |
5.50 | LDD0443 | [21] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [22] | |
|
HHS-475 Probe Info |
 |
Y115(0.93) | LDD2238 | [23] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-2 Probe Info |
 |
10.46 | LDD0138 | [24] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [25] | |
|
Dasatinib-CA-3PAP Probe Info |
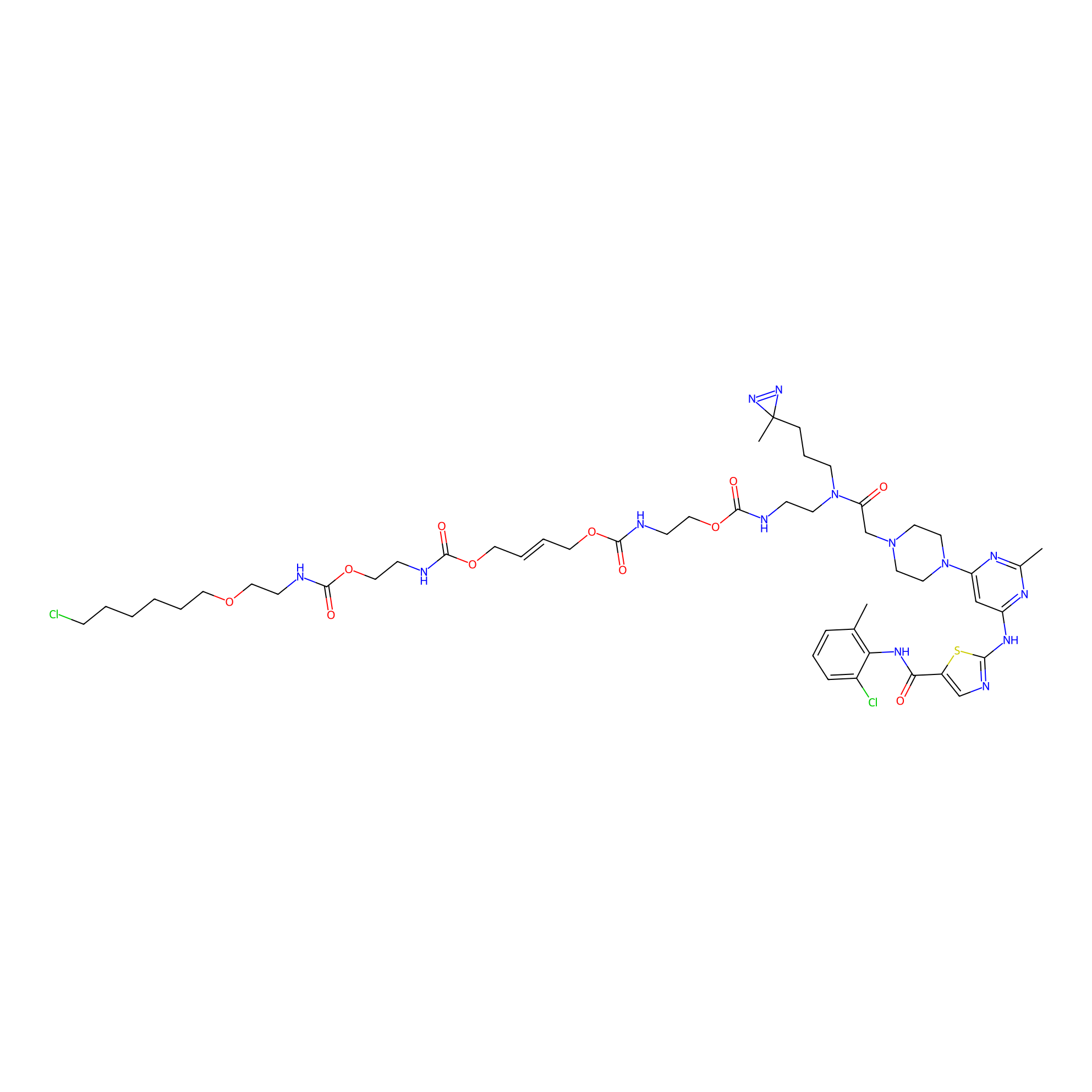 |
5.00 | LDD0365 | [26] | |
|
Diazir Probe Info |
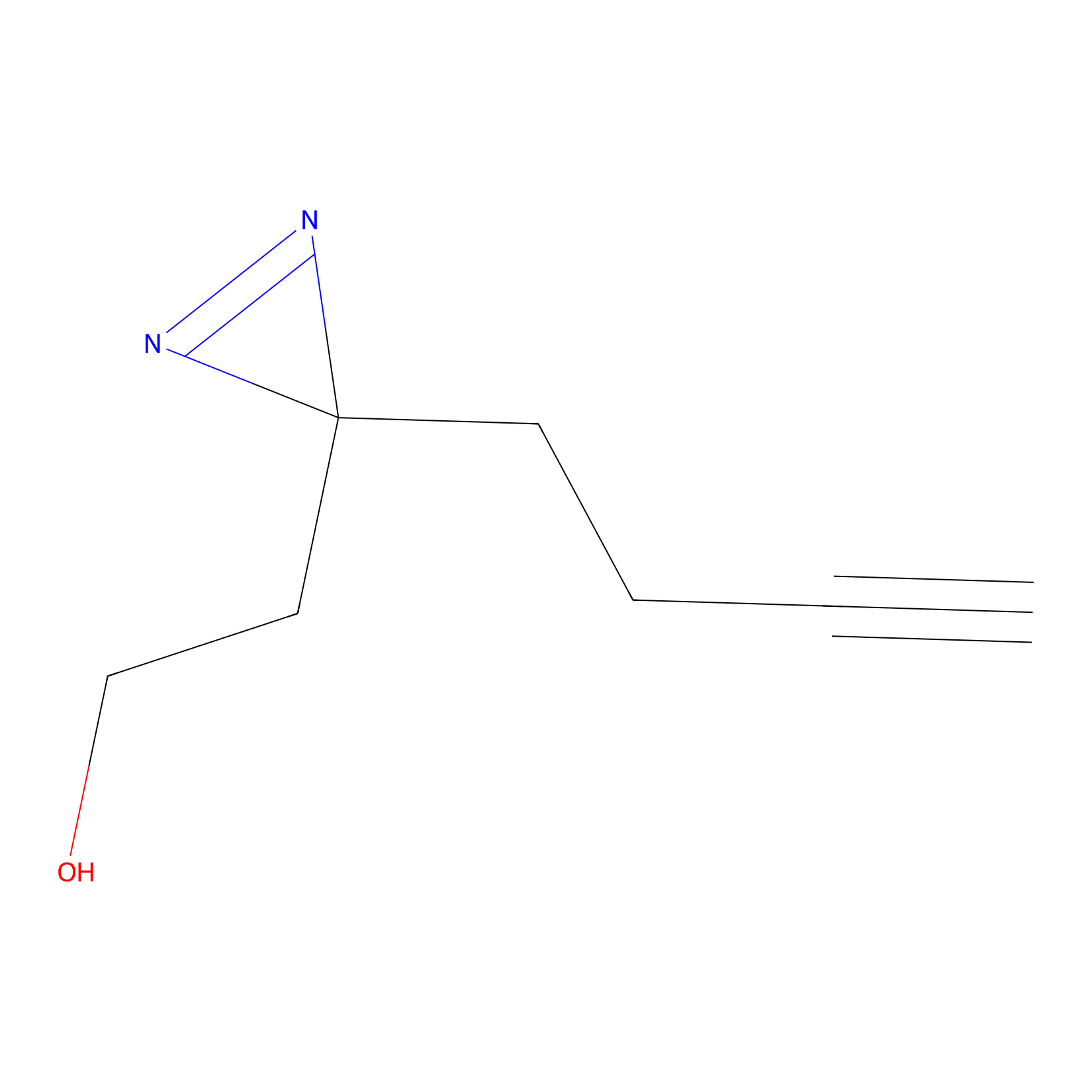 |
N.A. | LDD0011 | [15] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [27] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0069 | [28] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C44(0.84) | LDD2142 | [7] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C44(1.05) | LDD2112 | [7] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C44(0.67) | LDD2095 | [7] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C44(0.81) | LDD2130 | [7] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C44(1.09) | LDD2117 | [7] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C44(1.40) | LDD2152 | [7] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C44(0.55) | LDD2103 | [7] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C44(1.78) | LDD2132 | [7] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C44(0.53) | LDD2131 | [7] |
| LDCM0214 | AC1 | HEK-293T | C44(1.20) | LDD1507 | [29] |
| LDCM0215 | AC10 | HEK-293T | C44(1.08) | LDD1508 | [29] |
| LDCM0226 | AC11 | HEK-293T | C44(1.23) | LDD1509 | [29] |
| LDCM0230 | AC113 | HCT 116 | C44(1.64) | LDD0547 | [12] |
| LDCM0231 | AC114 | HCT 116 | C44(1.05) | LDD0548 | [12] |
| LDCM0232 | AC115 | HCT 116 | C44(1.09) | LDD0549 | [12] |
| LDCM0233 | AC116 | HCT 116 | C44(1.01) | LDD0550 | [12] |
| LDCM0234 | AC117 | HCT 116 | C44(1.02) | LDD0551 | [12] |
| LDCM0235 | AC118 | HCT 116 | C44(1.12) | LDD0552 | [12] |
| LDCM0236 | AC119 | HCT 116 | C44(0.95) | LDD0553 | [12] |
| LDCM0237 | AC12 | HEK-293T | C44(0.97) | LDD1510 | [29] |
| LDCM0238 | AC120 | HCT 116 | C44(0.97) | LDD0555 | [12] |
| LDCM0239 | AC121 | HCT 116 | C44(0.84) | LDD0556 | [12] |
| LDCM0240 | AC122 | HCT 116 | C44(0.97) | LDD0557 | [12] |
| LDCM0241 | AC123 | HCT 116 | C44(1.04) | LDD0558 | [12] |
| LDCM0242 | AC124 | HCT 116 | C44(0.94) | LDD0559 | [12] |
| LDCM0243 | AC125 | HCT 116 | C44(0.91) | LDD0560 | [12] |
| LDCM0244 | AC126 | HCT 116 | C44(1.01) | LDD0561 | [12] |
| LDCM0245 | AC127 | HCT 116 | C44(0.99) | LDD0562 | [12] |
| LDCM0259 | AC14 | HEK-293T | C44(0.99) | LDD1512 | [29] |
| LDCM0263 | AC143 | HEK-293T | C44(1.08) | LDD0861 | [12] |
| LDCM0264 | AC144 | HEK-293T | C44(1.06) | LDD0862 | [12] |
| LDCM0265 | AC145 | HEK-293T | C44(1.66) | LDD0863 | [12] |
| LDCM0266 | AC146 | HEK-293T | C44(1.71) | LDD0864 | [12] |
| LDCM0267 | AC147 | HEK-293T | C44(1.01) | LDD0865 | [12] |
| LDCM0268 | AC148 | HEK-293T | C44(1.92) | LDD0866 | [12] |
| LDCM0269 | AC149 | HEK-293T | C44(0.94) | LDD0867 | [12] |
| LDCM0270 | AC15 | HEK-293T | C44(1.02) | LDD1513 | [29] |
| LDCM0271 | AC150 | HEK-293T | C44(1.23) | LDD0869 | [12] |
| LDCM0272 | AC151 | HEK-293T | C44(0.95) | LDD0870 | [12] |
| LDCM0273 | AC152 | HEK-293T | C44(0.44) | LDD0871 | [12] |
| LDCM0274 | AC153 | HEK-293T | C44(0.55) | LDD0872 | [12] |
| LDCM0621 | AC154 | HEK-293T | C44(0.59) | LDD2162 | [12] |
| LDCM0622 | AC155 | HEK-293T | C44(0.84) | LDD2163 | [12] |
| LDCM0623 | AC156 | HEK-293T | C44(0.64) | LDD2164 | [12] |
| LDCM0624 | AC157 | HEK-293T | C44(0.48) | LDD2165 | [12] |
| LDCM0276 | AC17 | HCT 116 | C44(0.78) | LDD0593 | [12] |
| LDCM0277 | AC18 | HCT 116 | C44(0.83) | LDD0594 | [12] |
| LDCM0278 | AC19 | HCT 116 | C44(0.70) | LDD0595 | [12] |
| LDCM0279 | AC2 | HEK-293T | C44(1.04) | LDD1518 | [29] |
| LDCM0280 | AC20 | HCT 116 | C44(0.81) | LDD0597 | [12] |
| LDCM0281 | AC21 | HCT 116 | C44(0.70) | LDD0598 | [12] |
| LDCM0282 | AC22 | HCT 116 | C44(0.69) | LDD0599 | [12] |
| LDCM0283 | AC23 | HCT 116 | C44(0.73) | LDD0600 | [12] |
| LDCM0284 | AC24 | HCT 116 | C44(0.76) | LDD0601 | [12] |
| LDCM0285 | AC25 | HEK-293T | C44(0.90) | LDD0883 | [12] |
| LDCM0286 | AC26 | HEK-293T | C44(0.96) | LDD0884 | [12] |
| LDCM0287 | AC27 | HEK-293T | C44(1.16) | LDD0885 | [12] |
| LDCM0288 | AC28 | HEK-293T | C44(1.27) | LDD0886 | [12] |
| LDCM0289 | AC29 | HEK-293T | C44(1.10) | LDD0887 | [12] |
| LDCM0290 | AC3 | HEK-293T | C44(1.23) | LDD1529 | [29] |
| LDCM0291 | AC30 | HEK-293T | C44(1.03) | LDD0889 | [12] |
| LDCM0292 | AC31 | HEK-293T | C44(0.92) | LDD0890 | [12] |
| LDCM0293 | AC32 | HEK-293T | C44(0.97) | LDD0891 | [12] |
| LDCM0294 | AC33 | HEK-293T | C44(1.21) | LDD0892 | [12] |
| LDCM0295 | AC34 | HEK-293T | C44(1.00) | LDD0893 | [12] |
| LDCM0296 | AC35 | HCT 116 | C44(0.95) | LDD0613 | [12] |
| LDCM0297 | AC36 | HCT 116 | C44(1.03) | LDD0614 | [12] |
| LDCM0298 | AC37 | HCT 116 | C44(0.82) | LDD0615 | [12] |
| LDCM0299 | AC38 | HCT 116 | C44(1.29) | LDD0616 | [12] |
| LDCM0300 | AC39 | HCT 116 | C44(1.13) | LDD0617 | [12] |
| LDCM0301 | AC4 | HEK-293T | C44(1.00) | LDD1540 | [29] |
| LDCM0302 | AC40 | HCT 116 | C44(1.10) | LDD0619 | [12] |
| LDCM0303 | AC41 | HCT 116 | C44(0.73) | LDD0620 | [12] |
| LDCM0304 | AC42 | HCT 116 | C44(0.86) | LDD0621 | [12] |
| LDCM0305 | AC43 | HCT 116 | C44(0.43) | LDD0622 | [12] |
| LDCM0306 | AC44 | HCT 116 | C44(0.72) | LDD0623 | [12] |
| LDCM0307 | AC45 | HCT 116 | C44(0.67) | LDD0624 | [12] |
| LDCM0308 | AC46 | HEK-293T | C44(0.99) | LDD1547 | [29] |
| LDCM0309 | AC47 | HEK-293T | C44(0.99) | LDD1548 | [29] |
| LDCM0310 | AC48 | HEK-293T | C44(0.88) | LDD1549 | [29] |
| LDCM0311 | AC49 | HEK-293T | C44(1.09) | LDD1550 | [29] |
| LDCM0312 | AC5 | HEK-293T | C44(1.03) | LDD1551 | [29] |
| LDCM0313 | AC50 | HEK-293T | C44(1.09) | LDD1552 | [29] |
| LDCM0314 | AC51 | HEK-293T | C44(1.09) | LDD1553 | [29] |
| LDCM0315 | AC52 | HEK-293T | C44(0.97) | LDD1554 | [29] |
| LDCM0316 | AC53 | HEK-293T | C44(0.97) | LDD1555 | [29] |
| LDCM0317 | AC54 | HEK-293T | C44(0.99) | LDD1556 | [29] |
| LDCM0318 | AC55 | HEK-293T | C44(1.02) | LDD1557 | [29] |
| LDCM0319 | AC56 | HEK-293T | C44(0.99) | LDD1558 | [29] |
| LDCM0320 | AC57 | HEK-293T | C44(1.07) | LDD0918 | [12] |
| LDCM0321 | AC58 | HEK-293T | C44(1.13) | LDD0919 | [12] |
| LDCM0322 | AC59 | HEK-293T | C44(1.32) | LDD0920 | [12] |
| LDCM0323 | AC6 | HEK-293T | C44(1.07) | LDD1562 | [29] |
| LDCM0324 | AC60 | HEK-293T | C44(5.59) | LDD0922 | [12] |
| LDCM0325 | AC61 | HEK-293T | C44(1.57) | LDD0923 | [12] |
| LDCM0326 | AC62 | HEK-293T | C44(1.14) | LDD0924 | [12] |
| LDCM0327 | AC63 | HEK-293T | C44(1.39) | LDD0925 | [12] |
| LDCM0328 | AC64 | HEK-293T | C44(1.17) | LDD0926 | [12] |
| LDCM0329 | AC65 | HEK-293T | C44(1.33) | LDD0927 | [12] |
| LDCM0330 | AC66 | HEK-293T | C44(1.02) | LDD0928 | [12] |
| LDCM0331 | AC67 | HEK-293T | C44(1.34) | LDD0929 | [12] |
| LDCM0332 | AC68 | HEK-293T | C44(1.01) | LDD0930 | [12] |
| LDCM0333 | AC69 | HEK-293T | C44(1.14) | LDD0931 | [12] |
| LDCM0334 | AC7 | HEK-293T | C44(1.00) | LDD1568 | [29] |
| LDCM0335 | AC70 | HEK-293T | C44(1.18) | LDD0933 | [12] |
| LDCM0336 | AC71 | HEK-293T | C44(1.11) | LDD0934 | [12] |
| LDCM0337 | AC72 | HEK-293T | C44(1.25) | LDD0935 | [12] |
| LDCM0338 | AC73 | HEK-293T | C44(1.24) | LDD0936 | [12] |
| LDCM0339 | AC74 | HEK-293T | C44(1.18) | LDD0937 | [12] |
| LDCM0340 | AC75 | HEK-293T | C44(1.02) | LDD0938 | [12] |
| LDCM0341 | AC76 | HEK-293T | C44(0.87) | LDD0939 | [12] |
| LDCM0342 | AC77 | HEK-293T | C44(1.04) | LDD0940 | [12] |
| LDCM0343 | AC78 | HEK-293T | C44(1.10) | LDD0941 | [12] |
| LDCM0344 | AC79 | HEK-293T | C44(1.07) | LDD0942 | [12] |
| LDCM0345 | AC8 | HEK-293T | C44(1.06) | LDD1569 | [29] |
| LDCM0346 | AC80 | HEK-293T | C44(1.20) | LDD0944 | [12] |
| LDCM0347 | AC81 | HEK-293T | C44(1.04) | LDD0945 | [12] |
| LDCM0348 | AC82 | HEK-293T | C44(1.42) | LDD0946 | [12] |
| LDCM0349 | AC83 | HEK-293T | C44(0.79) | LDD0947 | [12] |
| LDCM0350 | AC84 | HEK-293T | C44(1.22) | LDD0948 | [12] |
| LDCM0351 | AC85 | HEK-293T | C44(1.00) | LDD0949 | [12] |
| LDCM0352 | AC86 | HEK-293T | C44(1.31) | LDD0950 | [12] |
| LDCM0353 | AC87 | HEK-293T | C44(0.99) | LDD0951 | [12] |
| LDCM0354 | AC88 | HEK-293T | C44(1.25) | LDD0952 | [12] |
| LDCM0355 | AC89 | HEK-293T | C44(1.49) | LDD0953 | [12] |
| LDCM0357 | AC90 | HEK-293T | C44(1.27) | LDD0955 | [12] |
| LDCM0358 | AC91 | HEK-293T | C44(0.82) | LDD0956 | [12] |
| LDCM0359 | AC92 | HEK-293T | C44(0.85) | LDD0957 | [12] |
| LDCM0360 | AC93 | HEK-293T | C44(0.83) | LDD0958 | [12] |
| LDCM0361 | AC94 | HEK-293T | C44(0.99) | LDD0959 | [12] |
| LDCM0362 | AC95 | HEK-293T | C44(0.90) | LDD0960 | [12] |
| LDCM0363 | AC96 | HEK-293T | C44(0.85) | LDD0961 | [12] |
| LDCM0364 | AC97 | HEK-293T | C44(1.32) | LDD0962 | [12] |
| LDCM0545 | Acetamide | MDA-MB-231 | C44(0.34) | LDD2138 | [7] |
| LDCM0248 | AKOS034007472 | HEK-293T | C44(0.97) | LDD1511 | [29] |
| LDCM0356 | AKOS034007680 | HEK-293T | C44(1.12) | LDD1570 | [29] |
| LDCM0275 | AKOS034007705 | HEK-293T | C44(0.99) | LDD1514 | [29] |
| LDCM0156 | Aniline | NCI-H1299 | 11.43 | LDD0403 | [1] |
| LDCM0151 | AZ-11 | HeLa | 10.00 | LDD2154 | [10] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C44(0.40) | LDD2091 | [7] |
| LDCM0108 | Chloroacetamide | HeLa | H227(0.00); H136(0.00); H128(0.00); H211(0.00) | LDD0222 | [20] |
| LDCM0632 | CL-Sc | Hep-G2 | C44(2.72); C44(0.80) | LDD2227 | [22] |
| LDCM0367 | CL1 | HEK-293T | C44(0.85) | LDD1571 | [29] |
| LDCM0368 | CL10 | HEK-293T | C44(0.89) | LDD1572 | [29] |
| LDCM0369 | CL100 | HEK-293T | C44(1.16) | LDD1573 | [29] |
| LDCM0370 | CL101 | HEK-293T | C44(0.99) | LDD1574 | [29] |
| LDCM0371 | CL102 | HEK-293T | C44(0.97) | LDD1575 | [29] |
| LDCM0372 | CL103 | HEK-293T | C44(0.99) | LDD1576 | [29] |
| LDCM0373 | CL104 | HEK-293T | C44(1.04) | LDD1577 | [29] |
| LDCM0374 | CL105 | HCT 116 | C44(0.74) | LDD0691 | [12] |
| LDCM0375 | CL106 | HCT 116 | C44(0.60) | LDD0692 | [12] |
| LDCM0376 | CL107 | HCT 116 | C44(0.78) | LDD0693 | [12] |
| LDCM0377 | CL108 | HCT 116 | C44(0.76) | LDD0694 | [12] |
| LDCM0378 | CL109 | HCT 116 | C44(0.73) | LDD0695 | [12] |
| LDCM0379 | CL11 | HEK-293T | C44(0.79) | LDD1583 | [29] |
| LDCM0380 | CL110 | HCT 116 | C44(0.57) | LDD0697 | [12] |
| LDCM0381 | CL111 | HCT 116 | C44(0.64) | LDD0698 | [12] |
| LDCM0382 | CL112 | HEK-293T | C44(1.38) | LDD0980 | [12] |
| LDCM0383 | CL113 | HEK-293T | C44(0.92) | LDD0981 | [12] |
| LDCM0384 | CL114 | HEK-293T | C44(1.39) | LDD0982 | [12] |
| LDCM0385 | CL115 | HEK-293T | C44(1.21) | LDD0983 | [12] |
| LDCM0386 | CL116 | HEK-293T | C44(1.30) | LDD0984 | [12] |
| LDCM0387 | CL117 | HCT 116 | C44(1.00) | LDD0704 | [12] |
| LDCM0388 | CL118 | HCT 116 | C44(0.89) | LDD0705 | [12] |
| LDCM0389 | CL119 | HCT 116 | C44(0.77) | LDD0706 | [12] |
| LDCM0390 | CL12 | HEK-293T | C44(0.69) | LDD1594 | [29] |
| LDCM0391 | CL120 | HCT 116 | C44(0.68) | LDD0708 | [12] |
| LDCM0392 | CL121 | HEK-293T | C44(0.99) | LDD1596 | [29] |
| LDCM0393 | CL122 | HEK-293T | C44(0.96) | LDD1597 | [29] |
| LDCM0394 | CL123 | HEK-293T | C44(0.94) | LDD1598 | [29] |
| LDCM0395 | CL124 | HEK-293T | C44(1.03) | LDD1599 | [29] |
| LDCM0396 | CL125 | HEK-293T | C44(1.08) | LDD0994 | [12] |
| LDCM0397 | CL126 | HEK-293T | C44(1.26) | LDD0995 | [12] |
| LDCM0398 | CL127 | HEK-293T | C44(1.01) | LDD0996 | [12] |
| LDCM0399 | CL128 | HEK-293T | C44(0.90) | LDD0997 | [12] |
| LDCM0400 | CL13 | HEK-293T | C44(0.91) | LDD1604 | [29] |
| LDCM0401 | CL14 | HEK-293T | C44(0.98) | LDD1605 | [29] |
| LDCM0402 | CL15 | HEK-293T | C44(0.91) | LDD1606 | [29] |
| LDCM0403 | CL16 | HEK-293T | C44(1.02) | LDD1607 | [29] |
| LDCM0404 | CL17 | HEK-293T | C44(0.92) | LDD1608 | [29] |
| LDCM0405 | CL18 | HEK-293T | C44(0.86) | LDD1609 | [29] |
| LDCM0406 | CL19 | HEK-293T | C44(0.69) | LDD1610 | [29] |
| LDCM0407 | CL2 | HEK-293T | C44(0.94) | LDD1611 | [29] |
| LDCM0408 | CL20 | HEK-293T | C44(0.88) | LDD1612 | [29] |
| LDCM0409 | CL21 | HEK-293T | C44(0.89) | LDD1613 | [29] |
| LDCM0410 | CL22 | HEK-293T | C44(0.72) | LDD1614 | [29] |
| LDCM0411 | CL23 | HEK-293T | C44(0.77) | LDD1615 | [29] |
| LDCM0412 | CL24 | HEK-293T | C44(0.76) | LDD1616 | [29] |
| LDCM0413 | CL25 | HEK-293T | C44(0.89) | LDD1617 | [29] |
| LDCM0414 | CL26 | HEK-293T | C44(1.06) | LDD1618 | [29] |
| LDCM0415 | CL27 | HEK-293T | C44(1.08) | LDD1619 | [29] |
| LDCM0416 | CL28 | HEK-293T | C44(1.07) | LDD1620 | [29] |
| LDCM0417 | CL29 | HEK-293T | C44(1.06) | LDD1621 | [29] |
| LDCM0418 | CL3 | HEK-293T | C44(0.89) | LDD1622 | [29] |
| LDCM0419 | CL30 | HEK-293T | C44(1.02) | LDD1623 | [29] |
| LDCM0420 | CL31 | HEK-293T | C44(0.95) | LDD1624 | [29] |
| LDCM0421 | CL32 | HEK-293T | C44(0.93) | LDD1625 | [29] |
| LDCM0422 | CL33 | HEK-293T | C44(0.93) | LDD1626 | [29] |
| LDCM0423 | CL34 | HEK-293T | C44(0.74) | LDD1627 | [29] |
| LDCM0424 | CL35 | HEK-293T | C44(0.80) | LDD1628 | [29] |
| LDCM0425 | CL36 | HEK-293T | C44(0.87) | LDD1629 | [29] |
| LDCM0426 | CL37 | HEK-293T | C44(0.90) | LDD1630 | [29] |
| LDCM0428 | CL39 | HEK-293T | C44(1.12) | LDD1632 | [29] |
| LDCM0429 | CL4 | HEK-293T | C44(0.98) | LDD1633 | [29] |
| LDCM0430 | CL40 | HEK-293T | C44(1.05) | LDD1634 | [29] |
| LDCM0431 | CL41 | HEK-293T | C44(1.04) | LDD1635 | [29] |
| LDCM0432 | CL42 | HEK-293T | C44(0.92) | LDD1636 | [29] |
| LDCM0433 | CL43 | HEK-293T | C44(0.86) | LDD1637 | [29] |
| LDCM0434 | CL44 | HEK-293T | C44(0.96) | LDD1638 | [29] |
| LDCM0435 | CL45 | HEK-293T | C44(0.99) | LDD1639 | [29] |
| LDCM0436 | CL46 | HEK-293T | C44(0.78) | LDD1640 | [29] |
| LDCM0437 | CL47 | HEK-293T | C44(0.80) | LDD1641 | [29] |
| LDCM0438 | CL48 | HEK-293T | C44(0.89) | LDD1642 | [29] |
| LDCM0439 | CL49 | HEK-293T | C44(0.94) | LDD1643 | [29] |
| LDCM0440 | CL5 | HEK-293T | C44(0.96) | LDD1644 | [29] |
| LDCM0441 | CL50 | HEK-293T | C44(1.10) | LDD1645 | [29] |
| LDCM0443 | CL52 | HEK-293T | C44(1.12) | LDD1646 | [29] |
| LDCM0444 | CL53 | HEK-293T | C44(1.01) | LDD1647 | [29] |
| LDCM0445 | CL54 | HEK-293T | C44(0.87) | LDD1648 | [29] |
| LDCM0446 | CL55 | HEK-293T | C44(0.92) | LDD1649 | [29] |
| LDCM0447 | CL56 | HEK-293T | C44(1.07) | LDD1650 | [29] |
| LDCM0448 | CL57 | HEK-293T | C44(1.04) | LDD1651 | [29] |
| LDCM0449 | CL58 | HEK-293T | C44(0.76) | LDD1652 | [29] |
| LDCM0450 | CL59 | HEK-293T | C44(0.83) | LDD1653 | [29] |
| LDCM0451 | CL6 | HEK-293T | C44(0.82) | LDD1654 | [29] |
| LDCM0452 | CL60 | HEK-293T | C44(0.85) | LDD1655 | [29] |
| LDCM0453 | CL61 | HCT 116 | C44(0.96) | LDD0770 | [12] |
| LDCM0454 | CL62 | HCT 116 | C44(0.60) | LDD0771 | [12] |
| LDCM0455 | CL63 | HCT 116 | C44(0.95) | LDD0772 | [12] |
| LDCM0456 | CL64 | HCT 116 | C44(0.77) | LDD0773 | [12] |
| LDCM0457 | CL65 | HCT 116 | C44(1.07) | LDD0774 | [12] |
| LDCM0458 | CL66 | HCT 116 | C44(0.83) | LDD0775 | [12] |
| LDCM0459 | CL67 | HCT 116 | C44(1.03) | LDD0776 | [12] |
| LDCM0460 | CL68 | HCT 116 | C44(1.16) | LDD0777 | [12] |
| LDCM0461 | CL69 | HCT 116 | C44(0.92) | LDD0778 | [12] |
| LDCM0462 | CL7 | HEK-293T | C44(0.88) | LDD1665 | [29] |
| LDCM0463 | CL70 | HCT 116 | C44(0.83) | LDD0780 | [12] |
| LDCM0464 | CL71 | HCT 116 | C44(0.59) | LDD0781 | [12] |
| LDCM0465 | CL72 | HCT 116 | C44(0.88) | LDD0782 | [12] |
| LDCM0466 | CL73 | HCT 116 | C44(1.02) | LDD0783 | [12] |
| LDCM0467 | CL74 | HCT 116 | C44(0.85) | LDD0784 | [12] |
| LDCM0469 | CL76 | HEK-293T | C44(1.12) | LDD1672 | [29] |
| LDCM0470 | CL77 | HEK-293T | C44(1.12) | LDD1673 | [29] |
| LDCM0471 | CL78 | HEK-293T | C44(1.03) | LDD1674 | [29] |
| LDCM0472 | CL79 | HEK-293T | C44(0.96) | LDD1675 | [29] |
| LDCM0473 | CL8 | HEK-293T | C44(0.71) | LDD1676 | [29] |
| LDCM0474 | CL80 | HEK-293T | C44(1.01) | LDD1677 | [29] |
| LDCM0475 | CL81 | HEK-293T | C44(0.98) | LDD1678 | [29] |
| LDCM0476 | CL82 | HEK-293T | C44(0.86) | LDD1679 | [29] |
| LDCM0477 | CL83 | HEK-293T | C44(0.87) | LDD1680 | [29] |
| LDCM0478 | CL84 | HEK-293T | C44(0.80) | LDD1681 | [29] |
| LDCM0479 | CL85 | HEK-293T | C44(0.74) | LDD1682 | [29] |
| LDCM0480 | CL86 | HEK-293T | C44(0.76) | LDD1683 | [29] |
| LDCM0481 | CL87 | HEK-293T | C44(0.86) | LDD1684 | [29] |
| LDCM0482 | CL88 | HEK-293T | C44(0.97) | LDD1685 | [29] |
| LDCM0483 | CL89 | HEK-293T | C44(0.97) | LDD1686 | [29] |
| LDCM0484 | CL9 | HEK-293T | C44(0.90) | LDD1687 | [29] |
| LDCM0485 | CL90 | HEK-293T | C44(0.90) | LDD1688 | [29] |
| LDCM0486 | CL91 | HEK-293T | C44(0.93) | LDD1689 | [29] |
| LDCM0487 | CL92 | HEK-293T | C44(0.82) | LDD1690 | [29] |
| LDCM0488 | CL93 | HEK-293T | C44(0.86) | LDD1691 | [29] |
| LDCM0489 | CL94 | HEK-293T | C44(0.80) | LDD1692 | [29] |
| LDCM0490 | CL95 | HEK-293T | C44(0.80) | LDD1693 | [29] |
| LDCM0491 | CL96 | HEK-293T | C44(0.61) | LDD1694 | [29] |
| LDCM0492 | CL97 | HEK-293T | C44(0.97) | LDD1695 | [29] |
| LDCM0493 | CL98 | HEK-293T | C44(1.11) | LDD1696 | [29] |
| LDCM0494 | CL99 | HEK-293T | C44(1.06) | LDD1697 | [29] |
| LDCM0097 | Dasatinib | K562 | 5.00 | LDD0365 | [26] |
| LDCM0495 | E2913 | HEK-293T | C44(1.12) | LDD1698 | [29] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C44(1.14) | LDD1702 | [7] |
| LDCM0625 | F8 | Ramos | C44(1.47) | LDD2187 | [30] |
| LDCM0572 | Fragment10 | Ramos | C44(0.23) | LDD2189 | [30] |
| LDCM0573 | Fragment11 | Ramos | C44(0.05) | LDD2190 | [30] |
| LDCM0574 | Fragment12 | Ramos | C44(0.33) | LDD2191 | [30] |
| LDCM0575 | Fragment13 | Ramos | C44(0.18) | LDD2192 | [30] |
| LDCM0576 | Fragment14 | Ramos | C44(0.17) | LDD2193 | [30] |
| LDCM0579 | Fragment20 | Ramos | C44(0.46) | LDD2194 | [30] |
| LDCM0580 | Fragment21 | Ramos | C44(0.35) | LDD2195 | [30] |
| LDCM0582 | Fragment23 | Ramos | C44(0.27) | LDD2196 | [30] |
| LDCM0578 | Fragment27 | Ramos | C44(0.59) | LDD2197 | [30] |
| LDCM0586 | Fragment28 | Ramos | C44(0.63) | LDD2198 | [30] |
| LDCM0588 | Fragment30 | Ramos | C44(0.34) | LDD2199 | [30] |
| LDCM0589 | Fragment31 | Ramos | C44(0.48) | LDD2200 | [30] |
| LDCM0590 | Fragment32 | Ramos | C44(0.39) | LDD2201 | [30] |
| LDCM0468 | Fragment33 | HCT 116 | C44(0.60) | LDD0785 | [12] |
| LDCM0596 | Fragment38 | Ramos | C44(0.41) | LDD2203 | [30] |
| LDCM0566 | Fragment4 | Ramos | C44(0.40) | LDD2184 | [30] |
| LDCM0427 | Fragment51 | HEK-293T | C44(1.23) | LDD1631 | [29] |
| LDCM0610 | Fragment52 | Ramos | C44(0.74) | LDD2204 | [30] |
| LDCM0614 | Fragment56 | Ramos | C44(0.16) | LDD2205 | [30] |
| LDCM0569 | Fragment7 | Ramos | C44(0.34) | LDD2186 | [30] |
| LDCM0571 | Fragment9 | Ramos | C44(0.13) | LDD2188 | [30] |
| LDCM0107 | IAA | HeLa | H227(0.00); H136(0.00); H211(0.00); H128(0.00) | LDD0221 | [20] |
| LDCM0123 | JWB131 | DM93 | Y115(20.00) | LDD0285 | [11] |
| LDCM0124 | JWB142 | DM93 | Y282(0.35) | LDD0286 | [11] |
| LDCM0125 | JWB146 | DM93 | Y282(0.90) | LDD0287 | [11] |
| LDCM0126 | JWB150 | DM93 | Y115(6.43); Y282(4.62) | LDD0288 | [11] |
| LDCM0127 | JWB152 | DM93 | Y115(3.12); Y282(1.33) | LDD0289 | [11] |
| LDCM0128 | JWB198 | DM93 | Y282(0.55) | LDD0290 | [11] |
| LDCM0129 | JWB202 | DM93 | Y282(0.36) | LDD0291 | [11] |
| LDCM0130 | JWB211 | DM93 | Y282(0.57) | LDD0292 | [11] |
| LDCM0022 | KB02 | HEK-293T | C44(0.99) | LDD1492 | [29] |
| LDCM0023 | KB03 | HEK-293T | C44(0.96) | LDD1497 | [29] |
| LDCM0024 | KB05 | SK-MEL-28 | C44(2.16) | LDD3427 | [31] |
| LDCM0109 | NEM | HeLa | H227(0.00); H136(0.00); H211(0.00); H284(0.00) | LDD0223 | [20] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C44(0.49) | LDD2089 | [7] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C44(1.05) | LDD2090 | [7] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C44(0.57) | LDD2092 | [7] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C44(1.10) | LDD2094 | [7] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C44(0.62) | LDD2096 | [7] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C44(1.85) | LDD2098 | [7] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C44(0.99) | LDD2100 | [7] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C44(1.15) | LDD2101 | [7] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C44(0.90) | LDD2104 | [7] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C44(1.23) | LDD2105 | [7] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C44(0.74) | LDD2106 | [7] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C44(1.05) | LDD2108 | [7] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C44(1.23) | LDD2109 | [7] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C44(1.07) | LDD2110 | [7] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C44(0.65) | LDD2116 | [7] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C44(0.66) | LDD2118 | [7] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C44(0.64) | LDD2120 | [7] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C44(0.83) | LDD2122 | [7] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C44(1.01) | LDD2123 | [7] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C44(0.77) | LDD2124 | [7] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C44(0.96) | LDD2125 | [7] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C44(0.98) | LDD2126 | [7] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C44(1.11) | LDD2127 | [7] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C44(0.60) | LDD2128 | [7] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C44(0.55) | LDD2129 | [7] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C44(0.27) | LDD2133 | [7] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C44(1.04) | LDD2134 | [7] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C44(0.57) | LDD2135 | [7] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C44(1.38) | LDD2136 | [7] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C44(1.02) | LDD2137 | [7] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C44(1.38) | LDD1700 | [7] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C44(1.39) | LDD2140 | [7] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C44(0.83) | LDD2143 | [7] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C44(2.04) | LDD2144 | [7] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C44(11.57) | LDD2145 | [7] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C44(1.02) | LDD2146 | [7] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C44(2.30) | LDD2147 | [7] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C44(0.42) | LDD2148 | [7] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C44(0.89) | LDD2149 | [7] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C44(0.35) | LDD2150 | [7] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C44(0.67) | LDD2151 | [7] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C44(1.98) | LDD2153 | [7] |
The Interaction Atlas With This Target
References
