Details of the Target
General Information of Target
| Target ID | LDTP01197 | |||||
|---|---|---|---|---|---|---|
| Target Name | Filamin-B (FLNB) | |||||
| Gene Name | FLNB | |||||
| Gene ID | 2317 | |||||
| Synonyms |
FLN1L; FLN3; TABP; TAP; Filamin-B; FLN-B; ABP-278; ABP-280 homolog; Actin-binding-like protein; Beta-filamin; Filamin homolog 1; Fh1; Filamin-3; Thyroid autoantigen; Truncated actin-binding protein; Truncated ABP
|
|||||
| 3D Structure | ||||||
| Sequence |
MPVTEKDLAEDAPWKKIQQNTFTRWCNEHLKCVNKRIGNLQTDLSDGLRLIALLEVLSQK
RMYRKYHQRPTFRQMQLENVSVALEFLDRESIKLVSIDSKAIVDGNLKLILGLVWTLILH YSISMPVWEDEGDDDAKKQTPKQRLLGWIQNKIPYLPITNFNQNWQDGKALGALVDSCAP GLCPDWESWDPQKPVDNAREAMQQADDWLGVPQVITPEEIIHPDVDEHSVMTYLSQFPKA KLKPGAPLKPKLNPKKARAYGRGIEPTGNMVKQPAKFTVDTISAGQGDVMVFVEDPEGNK EEAQVTPDSDKNKTYSVEYLPKVTGLHKVTVLFAGQHISKSPFEVSVDKAQGDASKVTAK GPGLEAVGNIANKPTYFDIYTAGAGVGDIGVEVEDPQGKNTVELLVEDKGNQVYRCVYKP MQPGPHVVKIFFAGDTIPKSPFVVQVGEACNPNACRASGRGLQPKGVRIRETTDFKVDTK AAGSGELGVTMKGPKGLEELVKQKDFLDGVYAFEYYPSTPGRYSIAITWGGHHIPKSPFE VQVGPEAGMQKVRAWGPGLHGGIVGRSADFVVESIGSEVGSLGFAIEGPSQAKIEYNDQN DGSCDVKYWPKEPGEYAVHIMCDDEDIKDSPYMAFIHPATGGYNPDLVRAYGPGLEKSGC IVNNLAEFTVDPKDAGKAPLKIFAQDGEGQRIDIQMKNRMDGTYACSYTPVKAIKHTIAV VWGGVNIPHSPYRVNIGQGSHPQKVKVFGPGVERSGLKANEPTHFTVDCTEAGEGDVSVG IKCDARVLSEDEEDVDFDIIHNANDTFTVKYVPPAAGRYTIKVLFASQEIPASPFRVKVD PSHDASKVKAEGPGLSKAGVENGKPTHFTVYTKGAGKAPLNVQFNSPLPGDAVKDLDIID NYDYSHTVKYTPTQQGNMQVLVTYGGDPIPKSPFTVGVAAPLDLSKIKLNGLENRVEVGK DQEFTVDTRGAGGQGKLDVTILSPSRKVVPCLVTPVTGRENSTAKFIPREEGLYAVDVTY DGHPVPGSPYTVEASLPPDPSKVKAHGPGLEGGLVGKPAEFTIDTKGAGTGGLGLTVEGP CEAKIECSDNGDGTCSVSYLPTKPGEYFVNILFEEVHIPGSPFKADIEMPFDPSKVVASG PGLEHGKVGEAGLLSVDCSEAGPGALGLEAVSDSGTKAEVSIQNNKDGTYAVTYVPLTAG MYTLTMKYGGELVPHFPARVKVEPAVDTSRIKVFGPGIEGKDVFREATTDFTVDSRPLTQ VGGDHIKAHIANPSGASTECFVTDNADGTYQVEYTPFEKGLHVVEVTYDDVPIPNSPFKV AVTEGCQPSRVQAQGPGLKEAFTNKPNVFTVVTRGAGIGGLGITVEGPSESKINCRDNKD GSCSAEYIPFAPGDYDVNITYGGAHIPGSPFRVPVKDVVDPSKVKIAGPGLGSGVRARVL QSFTVDSSKAGLAPLEVRVLGPRGLVEPVNVVDNGDGTHTVTYTPSQEGPYMVSVKYADE EIPRSPFKVKVLPTYDASKVTASGPGLSSYGVPASLPVDFAIDARDAGEGLLAVQITDQE GKPKRAIVHDNKDGTYAVTYIPDKTGRYMIGVTYGGDDIPLSPYRIRATQTGDASKCLAT GPGIASTVKTGEEVGFVVDAKTAGKGKVTCTVLTPDGTEAEADVIENEDGTYDIFYTAAK PGTYVIYVRFGGVDIPNSPFTVMATDGEVTAVEEAPVNACPPGFRPWVTEEAYVPVSDMN GLGFKPFDLVIPFAVRKGEITGEVHMPSGKTATPEIVDNKDGTVTVRYAPTEVGLHEMHI KYMGSHIPESPLQFYVNYPNSGSVSAYGPGLVYGVANKTATFTIVTEDAGEGGLDLAIEG PSKAEISCIDNKDGTCTVTYLPTLPGDYSILVKYNDKHIPGSPFTAKITDDSRRCSQVKL GSAADFLLDISETDLSSLTASIKAPSGRDEPCLLKRLPNNHIGISFIPREVGEHLVSIKK NGNHVANSPVSIMVVQSEIGDARRAKVYGRGLSEGRTFEMSDFIVDTRDAGYGGISLAVE GPSKVDIQTEDLEDGTCKVSYFPTVPGVYIVSTKFADEHVPGSPFTVKISGEGRVKESIT RTSRAPSVATVGSICDLNLKIPEINSSDMSAHVTSPSGRVTEAEIVPMGKNSHCVRFVPQ EMGVHTVSVKYRGQHVTGSPFQFTVGPLGEGGAHKVRAGGPGLERGEAGVPAEFSIWTRE AGAGGLSIAVEGPSKAEITFDDHKNGSCGVSYIAQEPGNYEVSIKFNDEHIPESPYLVPV IAPSDDARRLTVMSLQESGLKVNQPASFAIRLNGAKGKIDAKVHSPSGAVEECHVSELEP DKYAVRFIPHENGVHTIDVKFNGSHVVGSPFKVRVGEPGQAGNPALVSAYGTGLEGGTTG IQSEFFINTTRAGPGTLSVTIEGPSKVKMDCQETPEGYKVMYTPMAPGNYLISVKYGGPN HIVGSPFKAKVTGQRLVSPGSANETSSILVESVTRSSTETCYSAIPKASSDASKVTSKGA GLSKAFVGQKSSFLVDCSKAGSNMLLIGVHGPTTPCEEVSMKHVGNQQYNVTYVVKERGD YVLAVKWGEEHIPGSPFHVTVP |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Filamin family
|
|||||
| Subcellular location |
Cytoplasm, cytoskeleton, stress fiber; Cytoplasm, cell cortex; Cytoplasm, cytoskeleton
|
|||||
| Function |
Connects cell membrane constituents to the actin cytoskeleton. May promote orthogonal branching of actin filaments and links actin filaments to membrane glycoproteins. Anchors various transmembrane proteins to the actin cytoskeleton. Interaction with FLNA may allow neuroblast migration from the ventricular zone into the cortical plate. Various interactions and localizations of isoforms affect myotube morphology and myogenesis. Isoform 6 accelerates muscle differentiation in vitro.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| COLO792 | SNV: p.K1745R | DBIA Probe Info | |||
| CORL88 | SNV: p.D624G | DBIA Probe Info | |||
| DANG | SNV: p.D1738E | DBIA Probe Info | |||
| HCT15 | SNV: p.N2259K | DBIA Probe Info | |||
| HT1376 | SNV: p.P557L | DBIA Probe Info | |||
| IM95 | SNV: p.Y515H | DBIA Probe Info | |||
| IPC298 | SNV: p.S1529F | DBIA Probe Info | |||
| JURKAT | SNV: p.V103M; p.M270T | Compound 10 Probe Info | |||
| LNCaP clone FGC | SNV: p.Q2255Ter | DBIA Probe Info | |||
| MDST8 | SNV: p.D1706N | DBIA Probe Info | |||
| MFE319 | SNV: p.A705T; p.G959E | DBIA Probe Info | |||
| MOLT4 | SNV: p.V279L | IA-alkyne Probe Info | |||
| NALM6 | SNV: p.P1752L | DBIA Probe Info | |||
| ONS76 | SNV: p.P247A | DBIA Probe Info | |||
| PFSK1 | SNV: p.Y2260Ter | DBIA Probe Info | |||
| RL952 | Deletion: p.A971QfsTer9 SNV: p.R818Ter |
DBIA Probe Info | |||
| SNGM | SNV: p.R553C; p.S1041N | DBIA Probe Info | |||
| SW403 | SNV: p.K2006E | DBIA Probe Info | |||
| TE1 | SNV: p.Q503E | DBIA Probe Info | |||
| YSCCC | SNV: p.D474H | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
10.72 | LDD0402 | [1] | |
|
P2 Probe Info |
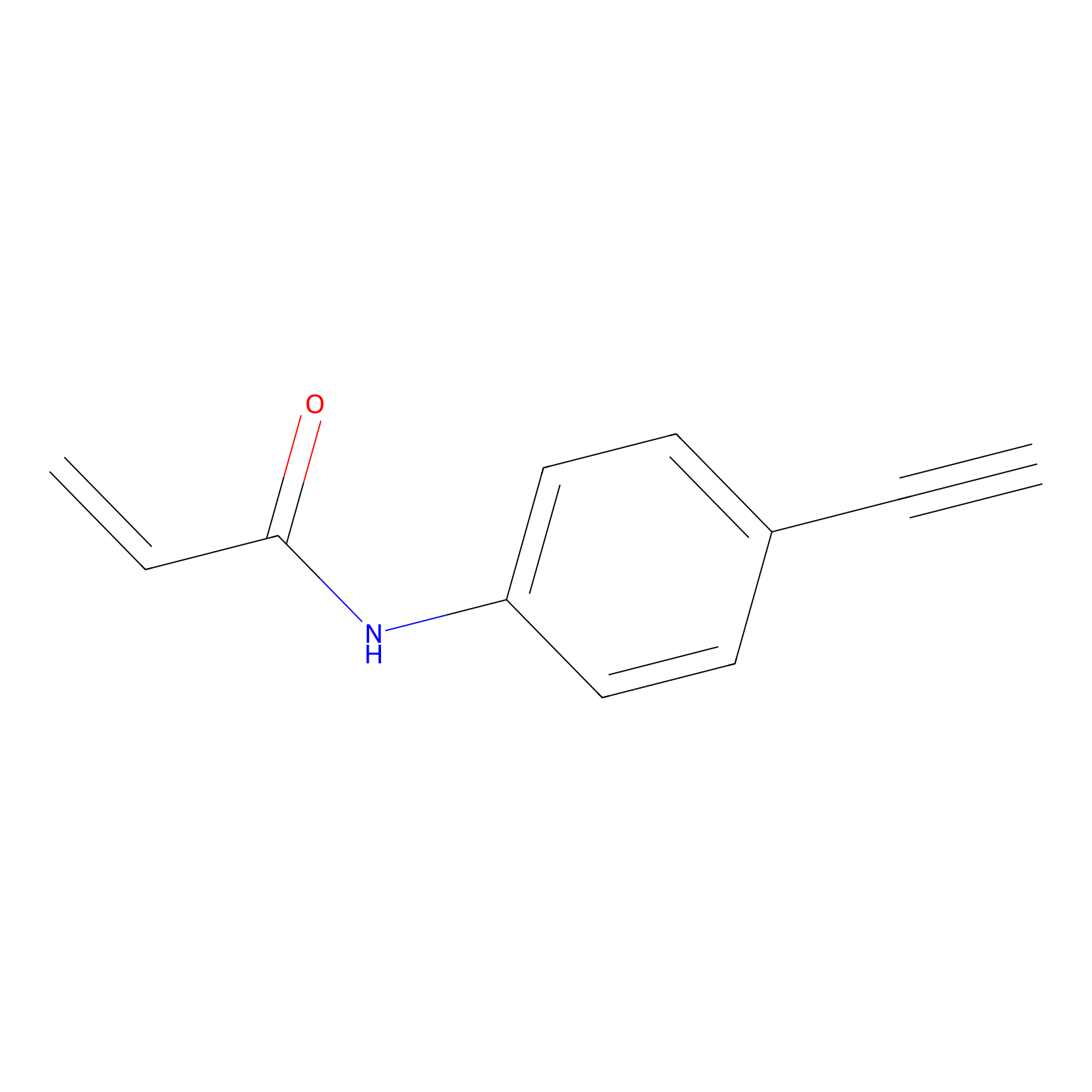 |
2.22 | LDD0449 | [2] | |
|
P3 Probe Info |
 |
4.31 | LDD0450 | [2] | |
|
A-EBA Probe Info |
 |
4.21 | LDD0215 | [3] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [4] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [4] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [4] | |
|
W1 Probe Info |
 |
10.74 | LDD0235 | [5] | |
|
TH211 Probe Info |
 |
Y596(20.00); Y2502(16.34); Y2069(10.53) | LDD0257 | [6] | |
|
TH214 Probe Info |
 |
Y596(20.00) | LDD0258 | [6] | |
|
TH216 Probe Info |
 |
Y2502(20.00); Y596(20.00) | LDD0259 | [6] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [7] | |
|
ONAyne Probe Info |
 |
K1005(3.69); K1057(1.02); K1221(2.88); K1241(4.24) | LDD0274 | [8] | |
|
OPA-S-S-alkyne Probe Info |
 |
K1955(1.23); K1241(1.42); K960(1.54); K2470(1.78) | LDD3494 | [9] | |
|
Probe 1 Probe Info |
 |
Y66(11.19); Y511(9.64); Y596(5.47); Y904(9.56) | LDD3495 | [10] | |
|
JZ128-DTB Probe Info |
 |
C2248(0.00); C178(0.00) | LDD0462 | [11] | |
|
DBIA Probe Info |
 |
C2556(9.72) | LDD0209 | [12] | |
|
HHS-475 Probe Info |
 |
Y902(0.71); Y596(0.85); Y1530(0.98); Y904(1.96) | LDD0264 | [13] | |
|
HHS-465 Probe Info |
 |
Y1530(10.00); Y596(10.00); Y904(9.98) | LDD2237 | [14] | |
|
5E-2FA Probe Info |
 |
H2243(0.00); H2550(0.00); H867(0.00); H1046(0.00) | LDD2235 | [15] | |
|
ATP probe Probe Info |
 |
K681(0.00); K2088(0.00); K2170(0.00); K409(0.00) | LDD0199 | [16] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C1868(0.00); C2556(0.00); C1280(0.00); C2115(0.00) | LDD0038 | [17] | |
|
IA-alkyne Probe Info |
 |
C450(0.00); C1617(0.00); C706(0.00); C2537(0.00) | LDD0032 | [18] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [19] | |
|
Lodoacetamide azide Probe Info |
 |
C1868(0.00); C2556(0.00); C1280(0.00); C769(0.00) | LDD0037 | [17] | |
|
ATP probe Probe Info |
 |
K2470(0.00); K16(0.00); K2507(0.00); K60(0.00) | LDD0035 | [20] | |
|
BTD Probe Info |
 |
C2333(0.00); C2115(0.00) | LDD0004 | [21] | |
|
NAIA_4 Probe Info |
 |
C706(0.00); C1326(0.00); C1375(0.00); C2115(0.00) | LDD2226 | [22] | |
|
TFBX Probe Info |
 |
C2501(0.00); C450(0.00); C660(0.00); C2537(0.00) | LDD0027 | [23] | |
|
WYneC Probe Info |
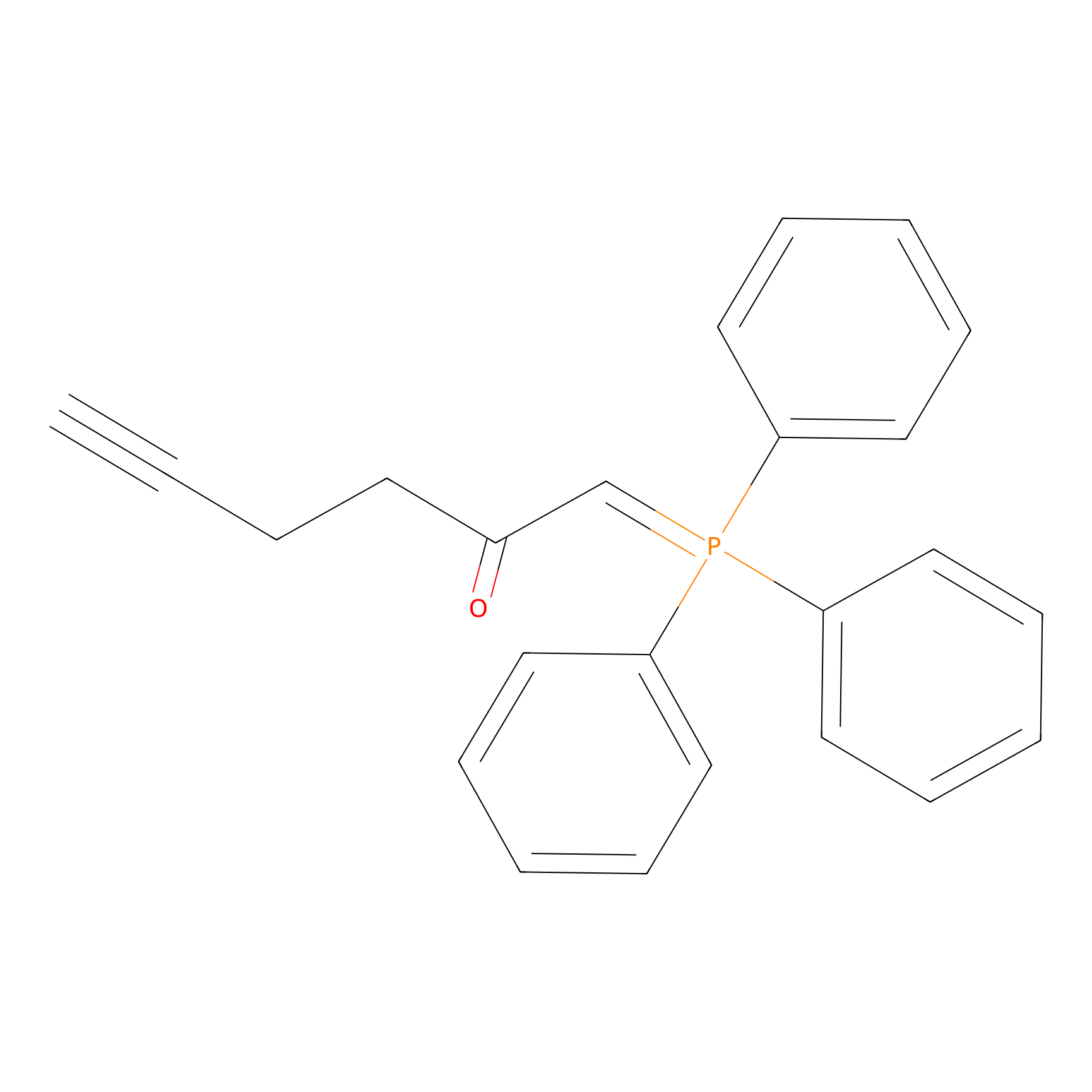 |
C2333(0.00); C1617(0.00); C2431(0.00) | LDD0014 | [21] | |
|
WYneN Probe Info |
 |
C2333(0.00); C706(0.00); C2431(0.00); C660(0.00) | LDD0021 | [21] | |
|
WYneO Probe Info |
 |
C1326(0.00); C2501(0.00); C604(0.00); C2154(0.00) | LDD0022 | [21] | |
|
DA-P3 Probe Info |
 |
N.A. | LDD0178 | [24] | |
|
Compound 10 Probe Info |
 |
C1617(0.00); C2115(0.00); C2501(0.00); C660(0.00) | LDD2216 | [25] | |
|
Compound 11 Probe Info |
 |
C2501(0.00); C660(0.00) | LDD2213 | [25] | |
|
ENE Probe Info |
 |
C1081(0.00); C1868(0.00); C455(0.00); C660(0.00) | LDD0006 | [21] | |
|
IPM Probe Info |
 |
C2501(0.00); C2154(0.00); C1326(0.00) | LDD0005 | [21] | |
|
NHS Probe Info |
 |
K1221(0.00); K838(0.00); K681(0.00); K1510(0.00) | LDD0010 | [21] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [26] | |
|
PPMS Probe Info |
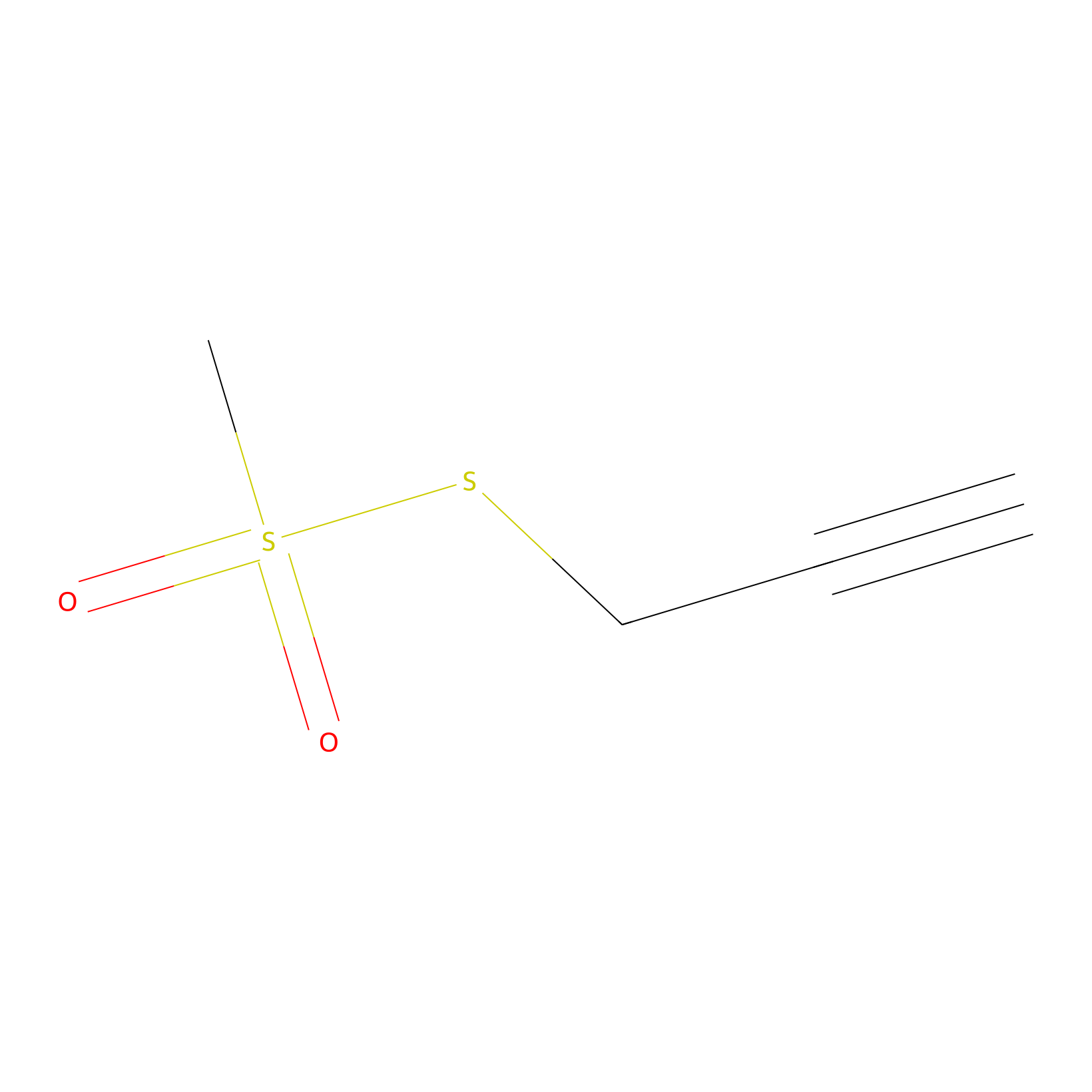 |
C2501(0.00); C1081(0.00) | LDD0008 | [21] | |
|
SF Probe Info |
 |
Y904(0.00); K6(0.00); Y902(0.00) | LDD0028 | [27] | |
|
STPyne Probe Info |
 |
K681(0.00); K838(0.00); K2428(0.00) | LDD0009 | [21] | |
|
VSF Probe Info |
 |
C1617(0.00); C450(0.00); C660(0.00); C455(0.00) | LDD0007 | [21] | |
|
YN-1 Probe Info |
 |
D505(0.00); E498(0.00) | LDD0447 | [7] | |
|
Phosphinate-6 Probe Info |
 |
C1915(0.00); C1326(0.00); C1617(0.00); C2501(0.00) | LDD0018 | [28] | |
|
1c-yne Probe Info |
 |
K2576(0.00); K1232(0.00); K2244(0.00); K877(0.00) | LDD0228 | [29] | |
|
1d-yne Probe Info |
 |
K504(0.00); K822(0.00) | LDD0357 | [29] | |
|
Acrolein Probe Info |
 |
C1280(0.00); H2132(0.00); C2115(0.00); C660(0.00) | LDD0217 | [30] | |
|
Cinnamaldehyde Probe Info |
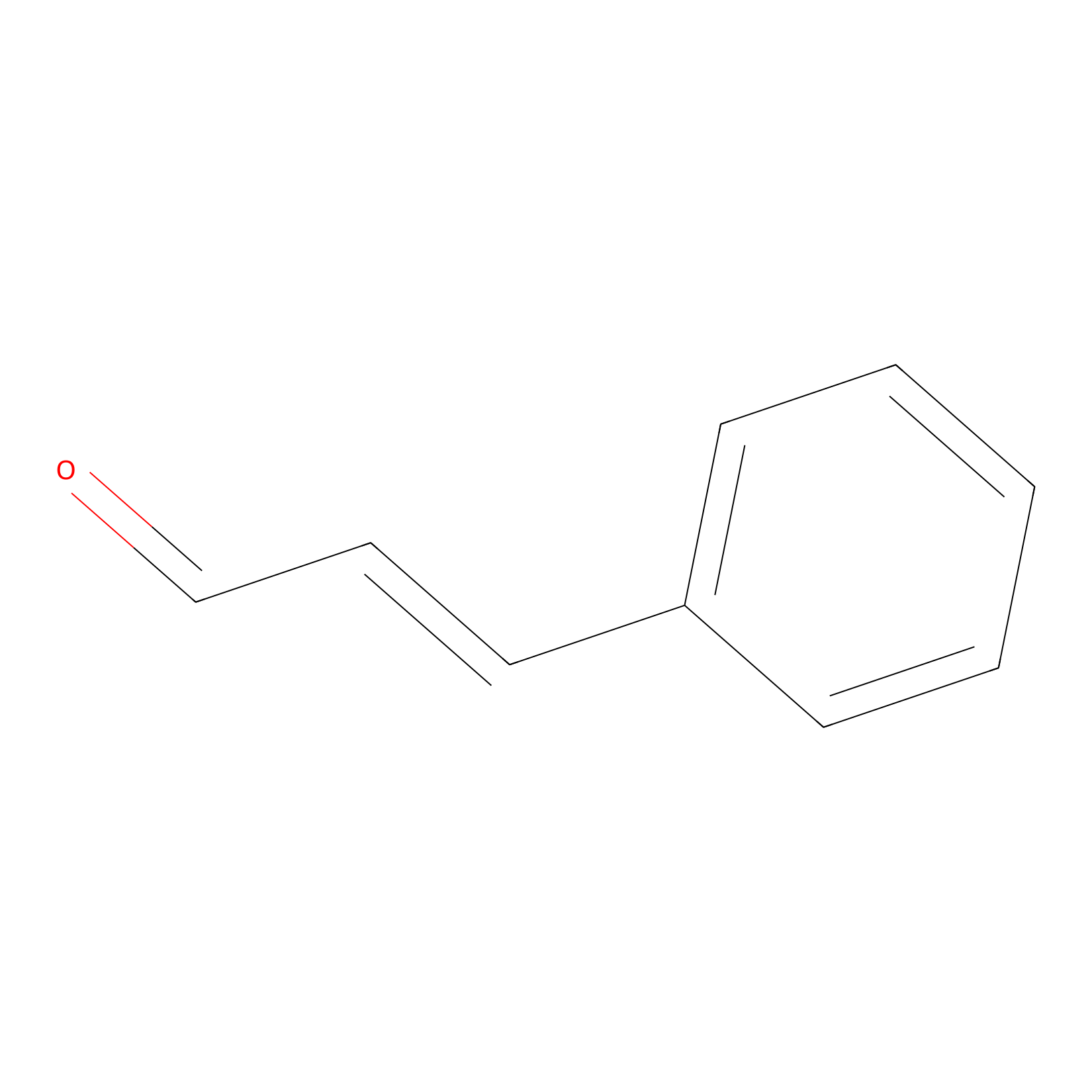 |
C1326(0.00); C991(0.00); C2537(0.00) | LDD0220 | [30] | |
|
Crotonaldehyde Probe Info |
 |
H560(0.00); C1326(0.00); C604(0.00); C991(0.00) | LDD0219 | [30] | |
|
Methacrolein Probe Info |
 |
C2556(0.00); C450(0.00); C660(0.00); C2057(0.00) | LDD0218 | [30] | |
|
NAIA_5 Probe Info |
 |
C2556(0.00); C1081(0.00); C2115(0.00); C1617(0.00) | LDD2223 | [22] | |
|
HHS-482 Probe Info |
 |
Y1530(1.38); Y155(1.04); Y1818(0.79); Y2343(1.31) | LDD2239 | [14] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-2 Probe Info |
 |
6.50 | LDD0138 | [31] | |
|
Staurosporine capture compound Probe Info |
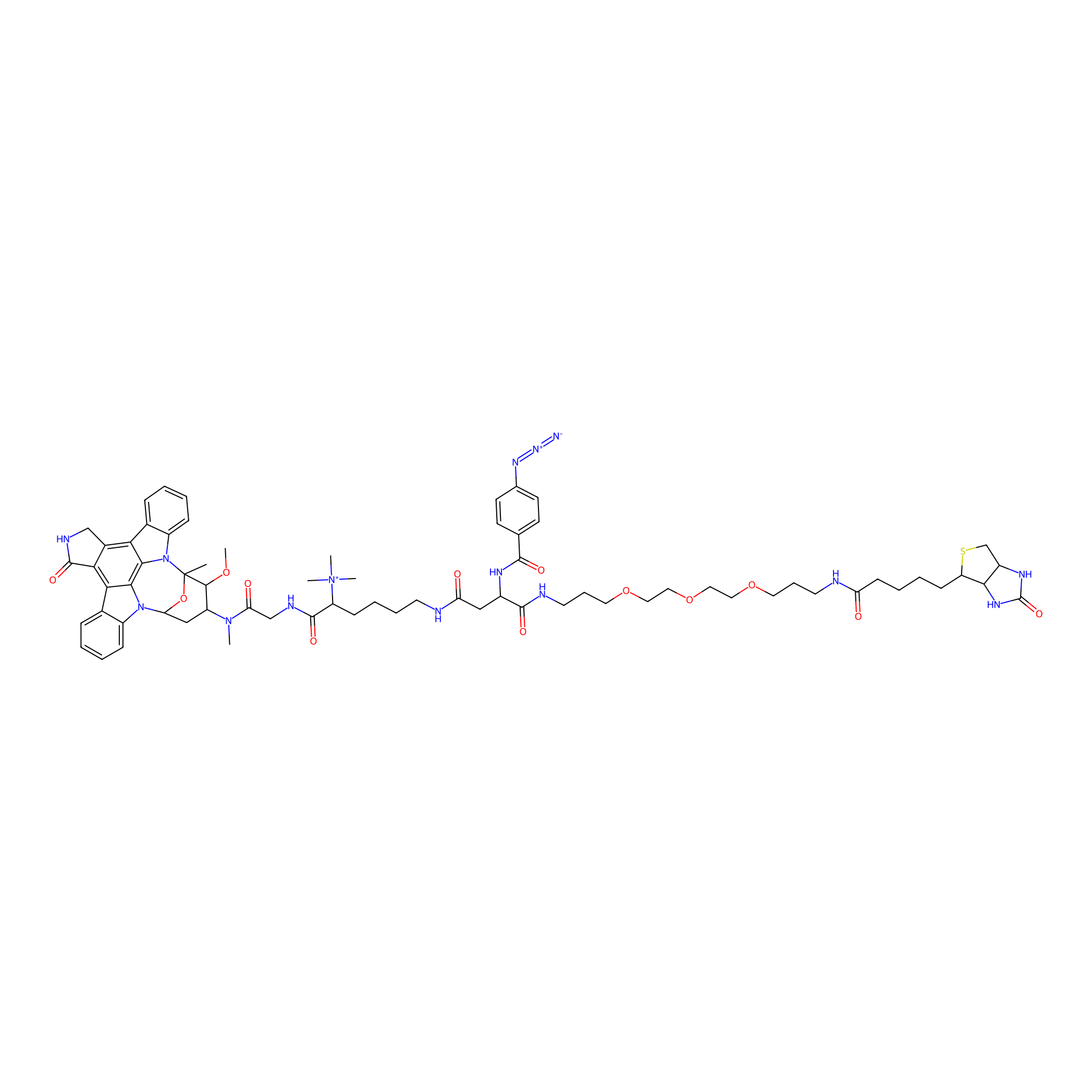 |
2.00 | LDD0083 | [32] | |
|
Photocelecoxib Probe Info |
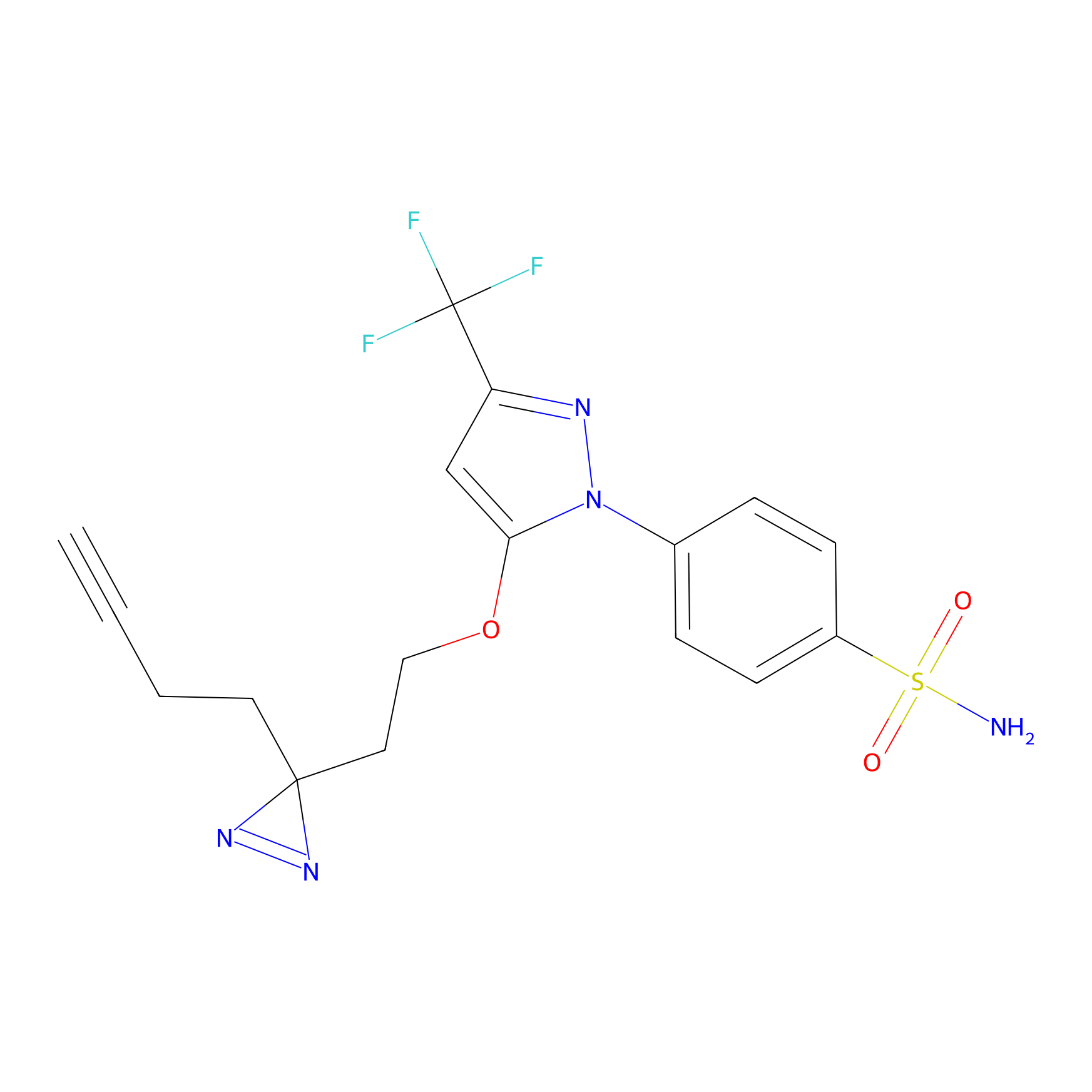 |
N.A. | LDD0153 | [33] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [34] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0069 | [35] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C1868(1.06); C2501(0.84); C450(0.55); C2431(0.45) | LDD2142 | [36] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C1868(0.76); C450(0.59); C2431(0.89); C2333(0.70) | LDD2112 | [36] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C1868(1.13); C2501(0.68); C450(0.67); C2556(0.59) | LDD2095 | [36] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C2501(0.83); C2431(0.61); C2333(0.86); C2115(0.84) | LDD2130 | [36] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C1868(1.07); C2501(0.92); C450(0.69); C2556(1.50) | LDD2117 | [36] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C1868(0.87); C2501(0.99); C450(1.15); C2431(1.48) | LDD2152 | [36] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C1868(0.98); C2501(1.08); C450(0.96); C2333(2.12) | LDD2103 | [36] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C2501(0.59); C450(0.66); C416(0.52); C2537(0.48) | LDD2132 | [36] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C1868(1.04); C2501(0.71); C2333(0.89); C1952(0.49) | LDD2131 | [36] |
| LDCM0025 | 4SU-RNA | HEK-293T | C26(3.09) | LDD0371 | [37] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C1617(6.66) | LDD0372 | [37] |
| LDCM0214 | AC1 | HEK-293T | C1326(0.86); C2501(1.07); C706(1.07); C1952(0.85) | LDD1507 | [38] |
| LDCM0215 | AC10 | HEK-293T | C1326(0.96); C2501(1.01); C706(0.95); C1952(0.84) | LDD1508 | [38] |
| LDCM0226 | AC11 | HEK-293T | C1326(0.90); C2501(1.07); C706(1.01); C1952(0.94) | LDD1509 | [38] |
| LDCM0237 | AC12 | HEK-293T | C1326(0.98); C2501(1.03); C706(0.92); C1952(0.92) | LDD1510 | [38] |
| LDCM0259 | AC14 | HEK-293T | C1326(0.92); C2501(0.98); C706(0.95); C1952(0.95) | LDD1512 | [38] |
| LDCM0270 | AC15 | HEK-293T | C1326(0.89); C2501(0.96); C706(1.13); C1952(0.96) | LDD1513 | [38] |
| LDCM0276 | AC17 | HEK-293T | C1326(0.92); C2501(1.08); C706(1.09); C1952(0.90) | LDD1515 | [38] |
| LDCM0277 | AC18 | HEK-293T | C1326(1.02); C2501(1.02); C706(1.00); C1952(0.90) | LDD1516 | [38] |
| LDCM0278 | AC19 | HEK-293T | C1326(0.78); C2501(1.03); C706(1.05); C1952(0.92) | LDD1517 | [38] |
| LDCM0279 | AC2 | HEK-293T | C1326(1.05); C2501(0.97); C706(1.04); C1952(0.97) | LDD1518 | [38] |
| LDCM0280 | AC20 | HEK-293T | C1326(1.04); C2501(1.01); C706(1.00); C1952(0.92) | LDD1519 | [38] |
| LDCM0281 | AC21 | HEK-293T | C1326(0.97); C2501(1.07); C706(0.97); C1952(0.92) | LDD1520 | [38] |
| LDCM0282 | AC22 | HEK-293T | C1326(0.99); C2501(0.97); C706(1.01); C1952(0.87) | LDD1521 | [38] |
| LDCM0283 | AC23 | HEK-293T | C1326(0.95); C2501(0.97); C706(1.04); C1952(0.94) | LDD1522 | [38] |
| LDCM0284 | AC24 | HEK-293T | C1326(0.95); C2501(0.97); C706(0.98); C1952(0.98) | LDD1523 | [38] |
| LDCM0285 | AC25 | HEK-293T | C1326(0.94); C2501(1.06); C706(1.05); C1952(0.85) | LDD1524 | [38] |
| LDCM0286 | AC26 | HEK-293T | C1326(1.02); C2501(1.00); C706(0.96); C1952(0.87) | LDD1525 | [38] |
| LDCM0287 | AC27 | HEK-293T | C1326(0.95); C2501(1.01); C706(0.98); C1952(0.95) | LDD1526 | [38] |
| LDCM0288 | AC28 | HEK-293T | C1326(0.95); C2501(0.96); C706(1.00); C1952(0.83) | LDD1527 | [38] |
| LDCM0289 | AC29 | HEK-293T | C1326(1.01); C2501(1.11); C706(0.98); C1952(1.00) | LDD1528 | [38] |
| LDCM0290 | AC3 | HEK-293T | C1326(0.83); C2501(1.03); C706(1.05); C1952(0.96) | LDD1529 | [38] |
| LDCM0291 | AC30 | HEK-293T | C1326(1.02); C2501(0.99); C706(0.97); C1952(0.99) | LDD1530 | [38] |
| LDCM0292 | AC31 | HEK-293T | C1326(0.94); C2501(0.96); C706(1.06); C1952(0.95) | LDD1531 | [38] |
| LDCM0293 | AC32 | HEK-293T | C1326(0.95); C2501(0.98); C706(0.96); C1952(0.93) | LDD1532 | [38] |
| LDCM0294 | AC33 | HEK-293T | C1326(0.90); C2501(0.96); C706(0.99); C1952(0.82) | LDD1533 | [38] |
| LDCM0295 | AC34 | HEK-293T | C1326(0.99); C2501(0.97); C706(1.02); C1952(0.81) | LDD1534 | [38] |
| LDCM0296 | AC35 | HEK-293T | C1326(0.87); C2501(1.06); C706(0.97); C1952(0.93) | LDD1535 | [38] |
| LDCM0297 | AC36 | HEK-293T | C1326(0.92); C2501(0.96); C706(0.97); C1952(0.89) | LDD1536 | [38] |
| LDCM0298 | AC37 | HEK-293T | C1326(1.01); C2501(0.94); C706(1.05); C1952(1.01) | LDD1537 | [38] |
| LDCM0299 | AC38 | HEK-293T | C1326(0.93); C2501(0.99); C706(0.93); C1952(0.91) | LDD1538 | [38] |
| LDCM0300 | AC39 | HEK-293T | C1326(0.98); C2501(0.95); C706(1.14); C1952(0.96) | LDD1539 | [38] |
| LDCM0301 | AC4 | HEK-293T | C1326(0.93); C2501(1.04); C706(0.91); C1952(0.95) | LDD1540 | [38] |
| LDCM0302 | AC40 | HEK-293T | C1326(1.01); C2501(0.96); C706(0.93); C1952(0.93) | LDD1541 | [38] |
| LDCM0303 | AC41 | HEK-293T | C1326(0.86); C2501(0.98); C706(1.03); C1952(0.91) | LDD1542 | [38] |
| LDCM0304 | AC42 | HEK-293T | C1326(1.00); C2501(1.00); C706(1.02); C1952(0.85) | LDD1543 | [38] |
| LDCM0305 | AC43 | HEK-293T | C1326(0.98); C2501(1.07); C706(1.01); C1952(0.88) | LDD1544 | [38] |
| LDCM0306 | AC44 | HEK-293T | C1326(0.98); C2501(0.96); C706(0.92); C1952(0.89) | LDD1545 | [38] |
| LDCM0307 | AC45 | HEK-293T | C1326(1.14); C2501(1.04); C706(0.99); C1952(0.94) | LDD1546 | [38] |
| LDCM0308 | AC46 | HEK-293T | C1326(1.05); C2501(0.93); C706(1.02); C1952(0.92) | LDD1547 | [38] |
| LDCM0309 | AC47 | HEK-293T | C1326(0.99); C2501(0.88); C706(1.05); C1952(0.96) | LDD1548 | [38] |
| LDCM0310 | AC48 | HEK-293T | C1326(0.97); C2501(1.03); C706(0.97); C1952(0.94) | LDD1549 | [38] |
| LDCM0311 | AC49 | HEK-293T | C1326(0.92); C2501(0.99); C706(1.05); C1952(0.83) | LDD1550 | [38] |
| LDCM0312 | AC5 | HEK-293T | C1326(1.13); C2501(1.11); C706(0.95); C1952(1.01) | LDD1551 | [38] |
| LDCM0313 | AC50 | HEK-293T | C1326(0.97); C2501(0.98); C706(0.98); C1952(0.85) | LDD1552 | [38] |
| LDCM0314 | AC51 | HEK-293T | C1326(0.89); C2501(1.02); C706(0.99); C1952(0.89) | LDD1553 | [38] |
| LDCM0315 | AC52 | HEK-293T | C1326(1.01); C2501(0.98); C706(0.98); C1952(0.85) | LDD1554 | [38] |
| LDCM0316 | AC53 | HEK-293T | C1326(1.11); C2501(0.96); C706(0.97); C1952(0.91) | LDD1555 | [38] |
| LDCM0317 | AC54 | HEK-293T | C1326(0.99); C2501(0.94); C706(1.01); C1952(0.91) | LDD1556 | [38] |
| LDCM0318 | AC55 | HEK-293T | C1326(0.91); C2501(0.91); C706(1.10); C1952(0.99) | LDD1557 | [38] |
| LDCM0319 | AC56 | HEK-293T | C1326(0.93); C2501(0.97); C706(0.91); C1952(0.91) | LDD1558 | [38] |
| LDCM0320 | AC57 | HEK-293T | C1326(0.82); C2501(1.00); C706(1.09); C1952(0.78) | LDD1559 | [38] |
| LDCM0321 | AC58 | HEK-293T | C1326(0.93); C2501(0.98); C706(0.98); C1952(0.78) | LDD1560 | [38] |
| LDCM0322 | AC59 | HEK-293T | C1326(0.92); C2501(1.03); C706(0.99); C1952(0.90) | LDD1561 | [38] |
| LDCM0323 | AC6 | HEK-293T | C1326(0.97); C2501(1.03); C706(0.98); C1952(0.86) | LDD1562 | [38] |
| LDCM0324 | AC60 | HEK-293T | C1326(0.89); C2501(0.96); C706(0.96); C1952(0.75) | LDD1563 | [38] |
| LDCM0325 | AC61 | HEK-293T | C1326(0.99); C2501(1.04); C706(1.03); C1952(0.79) | LDD1564 | [38] |
| LDCM0326 | AC62 | HEK-293T | C1326(0.97); C2501(0.95); C706(0.96); C1952(0.74) | LDD1565 | [38] |
| LDCM0327 | AC63 | HEK-293T | C1326(0.94); C2501(0.96); C706(0.98); C1952(0.81) | LDD1566 | [38] |
| LDCM0328 | AC64 | HEK-293T | C1326(1.08); C2501(0.96); C706(0.96); C1952(0.88) | LDD1567 | [38] |
| LDCM0334 | AC7 | HEK-293T | C1326(0.96); C2501(1.01); C706(1.13); C1952(1.01) | LDD1568 | [38] |
| LDCM0345 | AC8 | HEK-293T | C1326(0.98); C2501(1.02); C706(0.96); C1952(1.09) | LDD1569 | [38] |
| LDCM0545 | Acetamide | MDA-MB-231 | C416(0.91); C2333(0.69); C1952(0.79); C2115(0.87) | LDD2138 | [36] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C1868(0.65); C2501(0.69); C450(0.70); C2431(0.58) | LDD2113 | [36] |
| LDCM0248 | AKOS034007472 | HEK-293T | C1326(1.09); C2501(1.02); C706(1.01); C1952(0.95) | LDD1511 | [38] |
| LDCM0356 | AKOS034007680 | HEK-293T | C1326(0.90); C2501(1.04); C706(1.02); C1952(0.91) | LDD1570 | [38] |
| LDCM0275 | AKOS034007705 | HEK-293T | C1326(0.97); C2501(0.97); C706(0.93); C1952(0.95) | LDD1514 | [38] |
| LDCM0156 | Aniline | NCI-H1299 | C706(0.00); C1326(0.00) | LDD0404 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C1868(1.31); C450(1.24); C416(0.91); C2537(1.08) | LDD2091 | [36] |
| LDCM0630 | CCW28-3 | 231MFP | C1095(1.22); C1087(1.03) | LDD2214 | [39] |
| LDCM0108 | Chloroacetamide | HeLa | C1280(0.00); C2556(0.00); H2132(0.00); C2115(0.00) | LDD0222 | [30] |
| LDCM0632 | CL-Sc | Hep-G2 | C2115(2.20); C660(1.53); C2115(1.45); C2556(1.35) | LDD2227 | [22] |
| LDCM0367 | CL1 | HEK-293T | C1326(1.02); C2501(1.00); C706(0.93); C1952(0.96) | LDD1571 | [38] |
| LDCM0368 | CL10 | HEK-293T | C1326(0.81); C2501(0.90); C706(1.05); C1952(0.97) | LDD1572 | [38] |
| LDCM0369 | CL100 | HEK-293T | C1326(1.00); C2501(1.03); C706(0.94); C1952(0.87) | LDD1573 | [38] |
| LDCM0370 | CL101 | HEK-293T | C1326(1.01); C2501(1.00); C706(1.01); C1952(1.00) | LDD1574 | [38] |
| LDCM0371 | CL102 | HEK-293T | C1326(1.09); C2501(0.94); C706(1.02); C1952(0.84) | LDD1575 | [38] |
| LDCM0372 | CL103 | HEK-293T | C1326(0.99); C2501(0.95); C706(1.01); C1952(0.99) | LDD1576 | [38] |
| LDCM0373 | CL104 | HEK-293T | C1326(0.94); C2501(0.98); C706(0.95); C1952(1.00) | LDD1577 | [38] |
| LDCM0374 | CL105 | HEK-293T | C1326(1.04); C2501(0.97); C706(1.00); C1952(0.92) | LDD1578 | [38] |
| LDCM0375 | CL106 | HEK-293T | C1326(0.94); C2501(0.99); C706(0.91); C1952(0.91) | LDD1579 | [38] |
| LDCM0376 | CL107 | HEK-293T | C1326(0.97); C2501(0.89); C706(0.99); C1952(0.95) | LDD1580 | [38] |
| LDCM0377 | CL108 | HEK-293T | C1326(0.88); C2501(1.00); C706(0.95); C1952(0.88) | LDD1581 | [38] |
| LDCM0378 | CL109 | HEK-293T | C1326(1.06); C2501(1.03); C706(1.04); C1952(0.92) | LDD1582 | [38] |
| LDCM0379 | CL11 | HEK-293T | C1326(0.90); C2501(0.94); C706(1.05); C1952(1.28) | LDD1583 | [38] |
| LDCM0380 | CL110 | HEK-293T | C1326(1.03); C2501(0.92); C706(1.07); C1952(0.84) | LDD1584 | [38] |
| LDCM0381 | CL111 | HEK-293T | C1326(1.12); C2501(0.95); C706(1.03); C1952(1.03) | LDD1585 | [38] |
| LDCM0382 | CL112 | HEK-293T | C1326(0.96); C2501(1.08); C706(1.02); C1952(0.93) | LDD1586 | [38] |
| LDCM0383 | CL113 | HEK-293T | C1326(0.98); C2501(1.04); C706(0.97); C1952(0.94) | LDD1587 | [38] |
| LDCM0384 | CL114 | HEK-293T | C1326(1.09); C2501(0.88); C706(1.01); C1952(0.92) | LDD1588 | [38] |
| LDCM0385 | CL115 | HEK-293T | C1326(0.97); C2501(0.98); C706(0.99); C1952(0.98) | LDD1589 | [38] |
| LDCM0386 | CL116 | HEK-293T | C1326(0.97); C2501(1.00); C706(0.91); C1952(0.96) | LDD1590 | [38] |
| LDCM0387 | CL117 | HEK-293T | C1326(0.96); C2501(1.02); C706(0.92); C1952(1.00) | LDD1591 | [38] |
| LDCM0388 | CL118 | HEK-293T | C1326(1.02); C2501(0.90); C706(0.99); C1952(0.87) | LDD1592 | [38] |
| LDCM0389 | CL119 | HEK-293T | C1326(1.02); C2501(0.94); C706(1.05); C1952(1.02) | LDD1593 | [38] |
| LDCM0390 | CL12 | HEK-293T | C1326(0.85); C2501(0.99); C706(1.02); C1952(1.23) | LDD1594 | [38] |
| LDCM0391 | CL120 | HEK-293T | C1326(0.92); C2501(1.01); C706(1.01); C1952(0.93) | LDD1595 | [38] |
| LDCM0392 | CL121 | HEK-293T | C1326(1.01); C2501(0.99); C706(1.04); C1952(0.94) | LDD1596 | [38] |
| LDCM0393 | CL122 | HEK-293T | C1326(1.02); C2501(0.85); C706(0.97); C1952(0.90) | LDD1597 | [38] |
| LDCM0394 | CL123 | HEK-293T | C1326(0.87); C2501(0.86); C706(0.99); C1952(0.84) | LDD1598 | [38] |
| LDCM0395 | CL124 | HEK-293T | C1326(0.94); C2501(1.03); C706(1.08); C1952(0.89) | LDD1599 | [38] |
| LDCM0396 | CL125 | HEK-293T | C1326(1.06); C2501(0.92); C706(1.06); C1952(0.86) | LDD1600 | [38] |
| LDCM0397 | CL126 | HEK-293T | C1326(1.10); C2501(0.90); C706(0.99); C1952(0.82) | LDD1601 | [38] |
| LDCM0398 | CL127 | HEK-293T | C1326(1.12); C2501(0.92); C706(1.04); C1952(0.86) | LDD1602 | [38] |
| LDCM0399 | CL128 | HEK-293T | C1326(0.91); C2501(0.96); C706(0.96); C1952(0.82) | LDD1603 | [38] |
| LDCM0400 | CL13 | HEK-293T | C1326(1.01); C2501(1.03); C706(1.03); C1952(0.99) | LDD1604 | [38] |
| LDCM0401 | CL14 | HEK-293T | C1326(1.00); C2501(1.01); C706(1.03); C1952(1.04) | LDD1605 | [38] |
| LDCM0402 | CL15 | HEK-293T | C1326(0.87); C2501(0.89); C706(1.02); C1952(1.00) | LDD1606 | [38] |
| LDCM0403 | CL16 | HEK-293T | C1326(0.97); C2501(1.06); C706(1.00); C1952(0.95) | LDD1607 | [38] |
| LDCM0404 | CL17 | HEK-293T | C1326(0.85); C2501(1.09); C706(1.03); C1952(0.93) | LDD1608 | [38] |
| LDCM0405 | CL18 | HEK-293T | C1326(0.93); C2501(1.04); C706(0.99); C1952(0.86) | LDD1609 | [38] |
| LDCM0406 | CL19 | HEK-293T | C1326(0.90); C2501(1.12); C706(1.06); C1952(1.17) | LDD1610 | [38] |
| LDCM0407 | CL2 | HEK-293T | C1326(0.98); C2501(1.10); C706(0.99); C1952(1.02) | LDD1611 | [38] |
| LDCM0408 | CL20 | HEK-293T | C1326(0.96); C2501(1.09); C706(0.97); C1952(1.15) | LDD1612 | [38] |
| LDCM0409 | CL21 | HEK-293T | C1326(1.10); C2501(1.09); C706(1.00); C1952(0.96) | LDD1613 | [38] |
| LDCM0410 | CL22 | HEK-293T | C1326(0.90); C2501(1.07); C706(1.11); C1952(1.20) | LDD1614 | [38] |
| LDCM0411 | CL23 | HEK-293T | C1326(1.02); C2501(1.00); C706(1.38); C1952(1.23) | LDD1615 | [38] |
| LDCM0412 | CL24 | HEK-293T | C1326(0.94); C2501(1.05); C706(1.03); C1952(1.20) | LDD1616 | [38] |
| LDCM0413 | CL25 | HEK-293T | C1326(1.09); C2501(0.97); C706(0.93); C1952(0.94) | LDD1617 | [38] |
| LDCM0414 | CL26 | HEK-293T | C1326(1.02); C2501(0.92); C706(0.97); C1952(1.01) | LDD1618 | [38] |
| LDCM0415 | CL27 | HEK-293T | C1326(1.04); C2501(0.96); C706(1.00); C1952(1.08) | LDD1619 | [38] |
| LDCM0416 | CL28 | HEK-293T | C1326(0.95); C2501(0.96); C706(1.08); C1952(1.08) | LDD1620 | [38] |
| LDCM0417 | CL29 | HEK-293T | C1326(0.99); C2501(1.00); C706(1.05); C1952(1.01) | LDD1621 | [38] |
| LDCM0418 | CL3 | HEK-293T | C1326(0.94); C2501(1.06); C706(1.09); C1952(1.00) | LDD1622 | [38] |
| LDCM0419 | CL30 | HEK-293T | C1326(1.04); C2501(1.01); C706(0.94); C1952(0.91) | LDD1623 | [38] |
| LDCM0420 | CL31 | HEK-293T | C1326(0.88); C2501(1.09); C706(0.99); C1952(1.02) | LDD1624 | [38] |
| LDCM0421 | CL32 | HEK-293T | C1326(1.05); C2501(1.11); C706(0.96); C1952(1.03) | LDD1625 | [38] |
| LDCM0422 | CL33 | HEK-293T | C1326(0.84); C2501(0.92); C706(1.00); C1952(0.86) | LDD1626 | [38] |
| LDCM0423 | CL34 | HEK-293T | C1326(0.85); C2501(1.01); C706(1.00); C1952(1.16) | LDD1627 | [38] |
| LDCM0424 | CL35 | HEK-293T | C1326(0.96); C2501(1.11); C706(1.30); C1952(1.17) | LDD1628 | [38] |
| LDCM0425 | CL36 | HEK-293T | C1326(1.00); C2501(0.98); C706(0.92); C1952(1.07) | LDD1629 | [38] |
| LDCM0426 | CL37 | HEK-293T | C1326(1.04); C2501(0.98); C706(0.91); C1952(0.94) | LDD1630 | [38] |
| LDCM0428 | CL39 | HEK-293T | C1326(0.91); C2501(0.87); C706(1.04); C1952(0.96) | LDD1632 | [38] |
| LDCM0429 | CL4 | HEK-293T | C1326(0.83); C2501(1.01); C706(0.96); C1952(1.06) | LDD1633 | [38] |
| LDCM0430 | CL40 | HEK-293T | C1326(1.03); C2501(0.95); C706(1.09); C1952(1.10) | LDD1634 | [38] |
| LDCM0431 | CL41 | HEK-293T | C1326(0.96); C2501(1.00); C706(1.11); C1952(0.87) | LDD1635 | [38] |
| LDCM0432 | CL42 | HEK-293T | C1326(1.04); C2501(1.02); C706(0.99); C1952(0.92) | LDD1636 | [38] |
| LDCM0433 | CL43 | HEK-293T | C1326(0.93); C2501(0.99); C706(1.01); C1952(1.02) | LDD1637 | [38] |
| LDCM0434 | CL44 | HEK-293T | C1326(1.06); C2501(1.08); C706(0.90); C1952(0.98) | LDD1638 | [38] |
| LDCM0435 | CL45 | HEK-293T | C1326(0.96); C2501(1.10); C706(0.94); C1952(1.03) | LDD1639 | [38] |
| LDCM0436 | CL46 | HEK-293T | C1326(0.89); C2501(1.08); C706(1.20); C1952(1.12) | LDD1640 | [38] |
| LDCM0437 | CL47 | HEK-293T | C1326(0.91); C2501(0.98); C706(1.07); C1952(1.13) | LDD1641 | [38] |
| LDCM0438 | CL48 | HEK-293T | C1326(1.02); C2501(1.00); C706(0.93); C1952(1.06) | LDD1642 | [38] |
| LDCM0439 | CL49 | HEK-293T | C1326(1.01); C2501(0.88); C706(1.07); C1952(0.94) | LDD1643 | [38] |
| LDCM0440 | CL5 | HEK-293T | C1326(0.90); C2501(1.12); C706(1.09); C1952(1.11) | LDD1644 | [38] |
| LDCM0441 | CL50 | HEK-293T | C1326(1.06); C2501(0.96); C706(0.95); C1952(0.91) | LDD1645 | [38] |
| LDCM0443 | CL52 | HEK-293T | C1326(0.98); C2501(0.96); C706(1.00); C1952(0.99) | LDD1646 | [38] |
| LDCM0444 | CL53 | HEK-293T | C1326(0.94); C2501(0.97); C706(0.99); C1952(0.85) | LDD1647 | [38] |
| LDCM0445 | CL54 | HEK-293T | C1326(0.86); C2501(0.91); C706(0.84); C1952(0.81) | LDD1648 | [38] |
| LDCM0446 | CL55 | HEK-293T | C1326(0.93); C2501(1.06); C706(0.99); C1952(1.09) | LDD1649 | [38] |
| LDCM0447 | CL56 | HEK-293T | C1326(1.00); C2501(1.02); C706(0.87); C1952(1.03) | LDD1650 | [38] |
| LDCM0448 | CL57 | HEK-293T | C1326(0.92); C2501(0.98); C706(1.02); C1952(1.00) | LDD1651 | [38] |
| LDCM0449 | CL58 | HEK-293T | C1326(0.90); C2501(0.91); C706(0.96); C1952(1.06) | LDD1652 | [38] |
| LDCM0450 | CL59 | HEK-293T | C1326(0.85); C2501(0.98); C706(1.02); C1952(1.15) | LDD1653 | [38] |
| LDCM0451 | CL6 | HEK-293T | C1326(0.88); C2501(0.96); C706(0.99); C1952(0.86) | LDD1654 | [38] |
| LDCM0452 | CL60 | HEK-293T | C1326(0.91); C2501(0.94); C706(0.99); C1952(1.12) | LDD1655 | [38] |
| LDCM0453 | CL61 | HEK-293T | C1326(1.01); C2501(0.92); C706(1.05); C1952(1.01) | LDD1656 | [38] |
| LDCM0454 | CL62 | HEK-293T | C1326(1.01); C2501(0.92); C706(0.98); C1952(0.95) | LDD1657 | [38] |
| LDCM0455 | CL63 | HEK-293T | C1326(0.97); C2501(0.94); C706(0.98); C1952(1.00) | LDD1658 | [38] |
| LDCM0456 | CL64 | HEK-293T | C1326(0.79); C2501(0.98); C706(1.20); C1952(0.85) | LDD1659 | [38] |
| LDCM0457 | CL65 | HEK-293T | C1326(0.94); C2501(1.02); C706(1.02); C1952(0.90) | LDD1660 | [38] |
| LDCM0458 | CL66 | HEK-293T | C1326(1.05); C2501(0.99); C706(0.95); C1952(0.83) | LDD1661 | [38] |
| LDCM0459 | CL67 | HEK-293T | C1326(0.98); C2501(1.09); C706(1.05); C1952(1.02) | LDD1662 | [38] |
| LDCM0460 | CL68 | HEK-293T | C1326(1.02); C2501(0.93); C706(0.90); C1952(1.02) | LDD1663 | [38] |
| LDCM0461 | CL69 | HEK-293T | C1326(1.09); C2501(1.01); C706(1.01); C1952(1.00) | LDD1664 | [38] |
| LDCM0462 | CL7 | HEK-293T | C1326(0.85); C2501(1.11); C706(1.03); C1952(1.12) | LDD1665 | [38] |
| LDCM0463 | CL70 | HEK-293T | C1326(0.88); C2501(1.01); C706(0.94); C1952(0.98) | LDD1666 | [38] |
| LDCM0464 | CL71 | HEK-293T | C1326(0.97); C2501(0.96); C706(1.30); C1952(1.07) | LDD1667 | [38] |
| LDCM0465 | CL72 | HEK-293T | C1326(0.96); C2501(1.04); C706(0.98); C1952(1.01) | LDD1668 | [38] |
| LDCM0466 | CL73 | HEK-293T | C1326(1.05); C2501(0.86); C706(1.03); C1952(0.95) | LDD1669 | [38] |
| LDCM0467 | CL74 | HEK-293T | C1326(1.02); C2501(0.89); C706(1.03); C1952(0.90) | LDD1670 | [38] |
| LDCM0469 | CL76 | HEK-293T | C1326(0.99); C2501(1.03); C706(1.04); C1952(0.97) | LDD1672 | [38] |
| LDCM0470 | CL77 | HEK-293T | C1326(0.85); C2501(0.96); C706(0.96); C1952(0.86) | LDD1673 | [38] |
| LDCM0471 | CL78 | HEK-293T | C1326(1.04); C2501(1.04); C706(0.94); C1952(0.93) | LDD1674 | [38] |
| LDCM0472 | CL79 | HEK-293T | C1326(0.92); C2501(1.11); C706(0.98); C1952(1.10) | LDD1675 | [38] |
| LDCM0473 | CL8 | HEK-293T | C1326(0.88); C2501(0.83); C706(0.98); C1952(0.63) | LDD1676 | [38] |
| LDCM0474 | CL80 | HEK-293T | C1326(0.99); C2501(0.95); C706(0.93); C1952(1.05) | LDD1677 | [38] |
| LDCM0475 | CL81 | HEK-293T | C1326(1.01); C2501(0.98); C706(1.04); C1952(1.08) | LDD1678 | [38] |
| LDCM0476 | CL82 | HEK-293T | C1326(0.87); C2501(0.98); C706(1.04); C1952(1.02) | LDD1679 | [38] |
| LDCM0477 | CL83 | HEK-293T | C1326(1.06); C2501(0.96); C706(1.09); C1952(1.03) | LDD1680 | [38] |
| LDCM0478 | CL84 | HEK-293T | C1326(0.90); C2501(0.99); C706(0.94); C1952(0.98) | LDD1681 | [38] |
| LDCM0479 | CL85 | HEK-293T | C1326(1.10); C2501(0.91); C706(0.89); C1952(0.98) | LDD1682 | [38] |
| LDCM0480 | CL86 | HEK-293T | C1326(1.07); C2501(0.91); C706(0.98); C1952(0.89) | LDD1683 | [38] |
| LDCM0481 | CL87 | HEK-293T | C1326(1.00); C2501(0.84); C706(0.94); C1952(0.90) | LDD1684 | [38] |
| LDCM0482 | CL88 | HEK-293T | C1326(0.90); C2501(1.01); C706(0.97); C1952(0.90) | LDD1685 | [38] |
| LDCM0483 | CL89 | HEK-293T | C1326(0.97); C2501(0.94); C706(0.99); C1952(0.86) | LDD1686 | [38] |
| LDCM0484 | CL9 | HEK-293T | C1326(1.09); C2501(1.04); C706(1.05); C1952(1.06) | LDD1687 | [38] |
| LDCM0485 | CL90 | HEK-293T | C1326(0.68); C2501(0.78); C706(0.94); C1952(0.62) | LDD1688 | [38] |
| LDCM0486 | CL91 | HEK-293T | C1326(1.01); C2501(1.04); C706(1.03); C1952(0.96) | LDD1689 | [38] |
| LDCM0487 | CL92 | HEK-293T | C1326(1.01); C2501(0.95); C706(0.95); C1952(0.85) | LDD1690 | [38] |
| LDCM0488 | CL93 | HEK-293T | C1326(1.03); C2501(0.94); C706(1.03); C1952(0.88) | LDD1691 | [38] |
| LDCM0489 | CL94 | HEK-293T | C1326(0.93); C2501(1.00); C706(0.96); C1952(0.95) | LDD1692 | [38] |
| LDCM0490 | CL95 | HEK-293T | C1326(0.77); C2501(0.89); C706(1.21); C1952(0.85) | LDD1693 | [38] |
| LDCM0491 | CL96 | HEK-293T | C1326(0.94); C2501(0.83); C706(0.92); C1952(0.91) | LDD1694 | [38] |
| LDCM0492 | CL97 | HEK-293T | C1326(1.03); C2501(0.92); C706(1.03); C1952(0.95) | LDD1695 | [38] |
| LDCM0493 | CL98 | HEK-293T | C1326(1.01); C2501(0.92); C706(0.92); C1952(0.99) | LDD1696 | [38] |
| LDCM0494 | CL99 | HEK-293T | C1326(0.85); C2501(1.03); C706(1.17); C1952(0.94) | LDD1697 | [38] |
| LDCM0634 | CY-0357 | Hep-G2 | C1868(1.53); C2501(0.91) | LDD2228 | [22] |
| LDCM0495 | E2913 | HEK-293T | C1326(0.98); C2501(0.97); C706(0.96); C1952(1.03) | LDD1698 | [38] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C706(1.12); C2501(1.12); C2556(0.99); C2115(0.86) | LDD1702 | [36] |
| LDCM0625 | F8 | Ramos | C2501(1.48); C2556(1.14) | LDD2187 | [40] |
| LDCM0572 | Fragment10 | Ramos | C2501(1.32); C2556(0.78) | LDD2189 | [40] |
| LDCM0573 | Fragment11 | Ramos | C2501(1.67); C2556(2.29) | LDD2190 | [40] |
| LDCM0574 | Fragment12 | Ramos | C2501(0.69); C2556(0.78) | LDD2191 | [40] |
| LDCM0575 | Fragment13 | Ramos | C2501(1.03); C2556(1.13) | LDD2192 | [40] |
| LDCM0576 | Fragment14 | Ramos | C2501(0.98); C2556(1.71) | LDD2193 | [40] |
| LDCM0579 | Fragment20 | Ramos | C2501(0.65); C2556(0.72) | LDD2194 | [40] |
| LDCM0580 | Fragment21 | Ramos | C2501(1.00); C2556(2.82) | LDD2195 | [40] |
| LDCM0582 | Fragment23 | Ramos | C2501(1.03) | LDD2196 | [40] |
| LDCM0578 | Fragment27 | Ramos | C2501(0.96); C2556(1.00) | LDD2197 | [40] |
| LDCM0586 | Fragment28 | Ramos | C2501(0.85); C2556(1.51) | LDD2198 | [40] |
| LDCM0588 | Fragment30 | Ramos | C2501(1.09); C2556(0.83) | LDD2199 | [40] |
| LDCM0589 | Fragment31 | Ramos | C2501(0.91) | LDD2200 | [40] |
| LDCM0590 | Fragment32 | Ramos | C2501(1.37); C2556(0.85) | LDD2201 | [40] |
| LDCM0468 | Fragment33 | HEK-293T | C1326(0.98); C2501(0.87); C706(1.04); C1952(0.98) | LDD1671 | [38] |
| LDCM0596 | Fragment38 | Ramos | C2501(0.75) | LDD2203 | [40] |
| LDCM0566 | Fragment4 | Ramos | C2501(0.86); C2556(0.90) | LDD2184 | [40] |
| LDCM0427 | Fragment51 | HEK-293T | C1326(1.10); C2501(1.05); C706(0.97); C1952(0.92) | LDD1631 | [38] |
| LDCM0610 | Fragment52 | Ramos | C2501(1.25); C2556(0.86) | LDD2204 | [40] |
| LDCM0614 | Fragment56 | Ramos | C2501(1.22) | LDD2205 | [40] |
| LDCM0569 | Fragment7 | Ramos | C2501(0.86); C2556(0.50) | LDD2186 | [40] |
| LDCM0571 | Fragment9 | Ramos | C2501(1.01); C2556(0.54) | LDD2188 | [40] |
| LDCM0116 | HHS-0101 | DM93 | Y902(0.71); Y596(0.85); Y1530(0.98); Y904(1.96) | LDD0264 | [13] |
| LDCM0117 | HHS-0201 | DM93 | Y596(0.71); Y1530(1.16); Y904(1.71) | LDD0265 | [13] |
| LDCM0118 | HHS-0301 | DM93 | Y596(0.77); Y904(1.18); Y1530(1.48) | LDD0266 | [13] |
| LDCM0119 | HHS-0401 | DM93 | Y596(0.81); Y904(1.24); Y1530(1.65); Y902(2.98) | LDD0267 | [13] |
| LDCM0120 | HHS-0701 | DM93 | Y904(1.02); Y596(1.47); Y902(1.84); Y1530(1.91) | LDD0268 | [13] |
| LDCM0107 | IAA | HeLa | C1280(0.00); H2132(0.00); C2115(0.00); H906(0.00) | LDD0221 | [30] |
| LDCM0179 | JZ128 | PC-3 | C2248(0.00); C178(0.00) | LDD0462 | [11] |
| LDCM0022 | KB02 | HEK-293T | C1326(1.02); C604(0.93); C2501(1.04); C1868(1.10) | LDD1492 | [38] |
| LDCM0023 | KB03 | Jurkat | C2556(9.72) | LDD0209 | [12] |
| LDCM0024 | KB05 | G361 | C1648(2.34); C991(1.42); C2568(1.66); C2532(1.56) | LDD3311 | [41] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C1868(0.91); C2501(1.00); C450(1.08); C2333(0.99) | LDD2102 | [36] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C2501(0.63); C416(1.11); C2333(1.14); C1952(0.78) | LDD2121 | [36] |
| LDCM0109 | NEM | HeLa | H2132(0.00); C1280(0.00); H906(0.00); H1046(0.00) | LDD0223 | [30] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C1868(0.74); C2501(0.64); C450(1.50); C416(1.03) | LDD2089 | [36] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C2501(1.18); C450(1.00); C1952(1.57); C2115(1.29) | LDD2090 | [36] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C1868(0.91); C450(1.12); C2333(1.53); C1952(1.16) | LDD2092 | [36] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C1868(0.86); C2501(1.03); C450(0.88); C2537(1.20) | LDD2093 | [36] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C1868(1.37); C2333(1.79); C2115(2.01); C2154(3.54) | LDD2094 | [36] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C450(0.60); C2333(0.12); C2154(0.23); C991(0.30) | LDD2096 | [36] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C1868(0.77); C2501(0.77); C450(1.08); C2431(0.88) | LDD2097 | [36] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C1868(0.78); C2501(0.86); C450(0.87); C2333(1.29) | LDD2098 | [36] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C1868(1.27); C2501(0.82); C416(1.27); C2537(1.53) | LDD2099 | [36] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C2501(1.07); C450(0.65); C2431(0.36); C2333(0.46) | LDD2100 | [36] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C1868(0.69); C2501(0.78); C450(0.98); C2537(2.17) | LDD2101 | [36] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C1868(0.70); C2501(1.10); C2333(0.29); C1952(0.30) | LDD2104 | [36] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C1868(1.18); C2501(1.07); C450(1.94); C2431(1.08) | LDD2105 | [36] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C1868(0.64); C2501(0.93); C450(0.33); C416(0.26) | LDD2106 | [36] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C2501(0.81); C450(0.93); C416(0.86); C2537(1.17) | LDD2107 | [36] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C1868(0.63); C2501(0.65); C2333(0.61); C1952(0.32) | LDD2108 | [36] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C2501(0.71); C450(0.55); C416(0.95); C2431(0.63) | LDD2109 | [36] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C1868(1.09); C450(0.35); C2333(0.39); C1952(0.28) | LDD2110 | [36] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C1868(1.24); C2501(0.86); C450(0.87); C2431(1.17) | LDD2111 | [36] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C1868(1.06); C2501(0.88); C450(0.48); C2431(0.35) | LDD2114 | [36] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C2501(0.48); C450(0.34); C2333(0.40); C2115(0.36) | LDD2115 | [36] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C1868(1.43); C450(0.88); C416(1.06); C2431(0.31) | LDD2116 | [36] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C2501(0.74); C2537(0.61); C2154(0.30); C1326(0.37) | LDD2118 | [36] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C2501(2.52); C2537(1.80); C2431(2.20); C2333(1.95) | LDD2119 | [36] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C2501(0.98); C450(0.70); C416(0.84); C2333(0.61) | LDD2120 | [36] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C1868(1.23); C2501(0.62); C450(0.61); C2537(0.32) | LDD2122 | [36] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C1868(0.88); C2501(0.79); C450(0.68); C2556(1.52) | LDD2123 | [36] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C1868(1.44); C2501(0.76); C450(0.88); C2537(0.39) | LDD2124 | [36] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C2501(0.76); C450(0.62); C2537(0.90); C2431(0.93) | LDD2125 | [36] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C1868(1.44); C2501(0.68); C450(0.65); C2431(1.56) | LDD2126 | [36] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C1868(0.98); C2501(0.83); C450(0.60); C2556(1.48) | LDD2127 | [36] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C1868(0.95); C2501(0.81); C450(0.80); C2333(1.18) | LDD2128 | [36] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C450(0.79); C2537(1.42); C2431(1.42); C2333(1.88) | LDD2129 | [36] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C1868(0.49); C2501(0.67); C450(0.76); C2115(0.44) | LDD2133 | [36] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C1868(0.29); C2501(0.48); C450(0.70); C2537(0.58) | LDD2134 | [36] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C2501(0.80); C450(0.74); C2537(1.57); C2431(1.53) | LDD2135 | [36] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C1868(1.00); C450(1.28); C2556(1.12); C2537(1.36) | LDD2136 | [36] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C1868(0.98); C2501(0.79); C450(0.58); C2556(1.09) | LDD2137 | [36] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C2333(3.40); C2537(3.36); C1952(2.26); C1617(2.17) | LDD1700 | [36] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C1868(1.03); C2501(0.67); C450(0.98); C2537(1.10) | LDD2140 | [36] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C1868(0.74); C2501(0.60); C2431(0.81); C2333(0.70) | LDD2141 | [36] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C2501(1.12); C2537(0.74); C2333(0.94); C1952(0.57) | LDD2143 | [36] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C2501(3.26); C2537(1.91); C2431(2.61); C2333(1.84) | LDD2144 | [36] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C1868(1.81); C2501(3.70); C450(0.45); C2333(0.06) | LDD2145 | [36] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C1868(0.83); C450(0.74); C2556(1.02); C2333(1.24) | LDD2146 | [36] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C2501(2.60); C2333(1.35); C2154(2.04); C1326(3.45) | LDD2147 | [36] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C2501(0.47); C450(0.47); C2537(0.60); C2431(0.35) | LDD2148 | [36] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C1868(1.46); C2501(0.62); C450(0.79); C2154(0.21) | LDD2149 | [36] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C1868(0.83); C2501(0.41); C2556(0.98); C416(0.96) | LDD2150 | [36] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C1868(1.44); C2154(0.39) | LDD2151 | [36] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C1868(1.27); C2501(1.06); C450(2.86); C1952(1.38) | LDD2153 | [36] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C2501(0.85) | LDD2206 | [42] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C2501(0.65) | LDD2207 | [42] |
| LDCM0019 | Staurosporine | Hep-G2 | 2.00 | LDD0083 | [32] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Filamin-A (FLNA) | Filamin family | P21333 | |||
| Filamin-B (FLNB) | Filamin family | O75369 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Growth factor receptor-bound protein 2 (GRB2) | GRB2/sem-5/DRK family | P62993 | |||
| SH2/SH3 adapter protein NCK1 (NCK1) | . | P16333 | |||
| Ubiquitin-like protein ISG15 (ISG15) | . | P05161 | |||
References
