Details of the Target
General Information of Target
| Target ID | LDTP04309 | |||||
|---|---|---|---|---|---|---|
| Target Name | Nuclear pore complex protein Nup153 (NUP153) | |||||
| Gene Name | NUP153 | |||||
| Gene ID | 9972 | |||||
| Synonyms |
Nuclear pore complex protein Nup153; 153 kDa nucleoporin; Nucleoporin Nup153 |
|||||
| 3D Structure | ||||||
| Sequence |
MASGAGGVGGGGGGKIRTRRCHQGPIKPYQQGRQQHQGILSRVTESVKNIVPGWLQRYFN
KNEDVCSCSTDTSEVPRWPENKEDHLVYADEESSNITDGRITPEPAVSNTEEPSTTSTAS NYPDVLTRPSLHRSHLNFSMLESPALHCQPSTSSAFPIGSSGFSLVKEIKDSTSQHDDDN ISTTSGFSSRASDKDITVSKNTSLPPLWSPEAERSHSLSQHTATSSKKPAFNLSAFGTLS PSLGNSSILKTSQLGDSPFYPGKTTYGGAAAAVRQSKLRNTPYQAPVRRQMKAKQLSAQS YGVTSSTARRILQSLEKMSSPLADAKRIPSIVSSPLNSPLDRSGIDITDFQAKREKVDSQ YPPVQRLMTPKPVSIATNRSVYFKPSLTPSGEFRKTNQRIDNKCSTGYEKNMTPGQNREQ RESGFSYPNFSLPAANGLSSGVGGGGGKMRRERTRFVASKPLEEEEMEVPVLPKISLPIT SSSLPTFNFSSPEITTSSPSPINSSQALTNKVQMTSPSSTGSPMFKFSSPIVKSTEANVL PPSSIGFTFSVPVAKTAELSGSSSTLEPIISSSAHHVTTVNSTNCKKTPPEDCEGPFRPA EILKEGSVLDILKSPGFASPKIDSVAAQPTATSPVVYTRPAISSFSSSGIGFGESLKAGS SWQCDTCLLQNKVTDNKCIACQAAKLSPRDTAKQTGIETPNKSGKTTLSASGTGFGDKFK PVIGTWDCDTCLVQNKPEAIKCVACETPKPGTCVKRALTLTVVSESAETMTASSSSCTVT TGTLGFGDKFKRPIGSWECSVCCVSNNAEDNKCVSCMSEKPGSSVPASSSSTVPVSLPSG GSLGLEKFKKPEGSWDCELCLVQNKADSTKCLACESAKPGTKSGFKGFDTSSSSSNSAAS SSFKFGVSSSSSGPSQTLTSTGNFKFGDQGGFKIGVSSDSGSINPMSEGFKFSKPIGDFK FGVSSESKPEEVKKDSKNDNFKFGLSSGLSNPVSLTPFQFGVSNLGQEEKKEELPKSSSA GFSFGTGVINSTPAPANTIVTSENKSSFNLGTIETKSASVAPFTCKTSEAKKEEMPATKG GFSFGNVEPASLPSASVFVLGRTEEKQQEPVTSTSLVFGKKADNEEPKCQPVFSFGNSEQ TKDENSSKSTFSFSMTKPSEKESEQPAKATFAFGAQTSTTADQGAAKPVFSFLNNSSSSS STPATSAGGGIFGSSTSSSNPPVATFVFGQSSNPVSSSAFGNTAESSTSQSLLFSQDSKL ATTSSTGTAVTPFVFGPGASSNNTTTSGFGFGATTTSSSAGSSFVFGTGPSAPSASPAFG ANQTPTFGQSQGASQPNPPGFGSISSSTALFPTGSQPAPPTFGTVSSSSQPPVFGQQPSQ SAFGSGTTPNSSSAFQFGSSTTNFNFTNNSPSGVFTFGANSSTPAASAQPSGSGGFPFNQ SPAAFTVGSNGKNVFSSSGTSFSGRKIKTAVRRRK |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
NUP153 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Component of the nuclear pore complex (NPC), a complex required for the trafficking across the nuclear envelope. Functions as a scaffolding element in the nuclear phase of the NPC essential for normal nucleocytoplasmic transport of proteins and mRNAs. Involved in the quality control and retention of unspliced mRNAs in the nucleus; in association with TPR, regulates the nuclear export of unspliced mRNA species bearing constitutive transport element (CTE) in a NXF1- and KHDRBS1-independent manner. Mediates TPR anchoring to the nuclear membrane at NPC. The repeat-containing domain may be involved in anchoring other components of the NPC to the pore membrane. Possible DNA-binding subunit of the nuclear pore complex (NPC).; (Microbial infection) Interacts with HIV-1 caspid protein P24 and thereby promotes the integration of the virus in the nucleus of non-dividing cells (in vitro).; (Microbial infection) Binds HIV-2 protein vpx and thereby promotes the nuclear translocation of the lentiviral genome (in vitro).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AGS | SNV: p.E409G | DBIA Probe Info | |||
| EFO21 | SNV: p.N1322S | DBIA Probe Info | |||
| HCT15 | SNV: p.G1184C | DBIA Probe Info | |||
| NCIH1155 | SNV: p.V1454M | DBIA Probe Info | |||
| SUPT1 | SNV: p.R455C | DBIA Probe Info | |||
| TE1 | SNV: p.S1237N | DBIA Probe Info | |||
| TE10 | SNV: p.L165V | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
14.88 | LDD0402 | [1] | |
|
DA-P2 Probe Info |
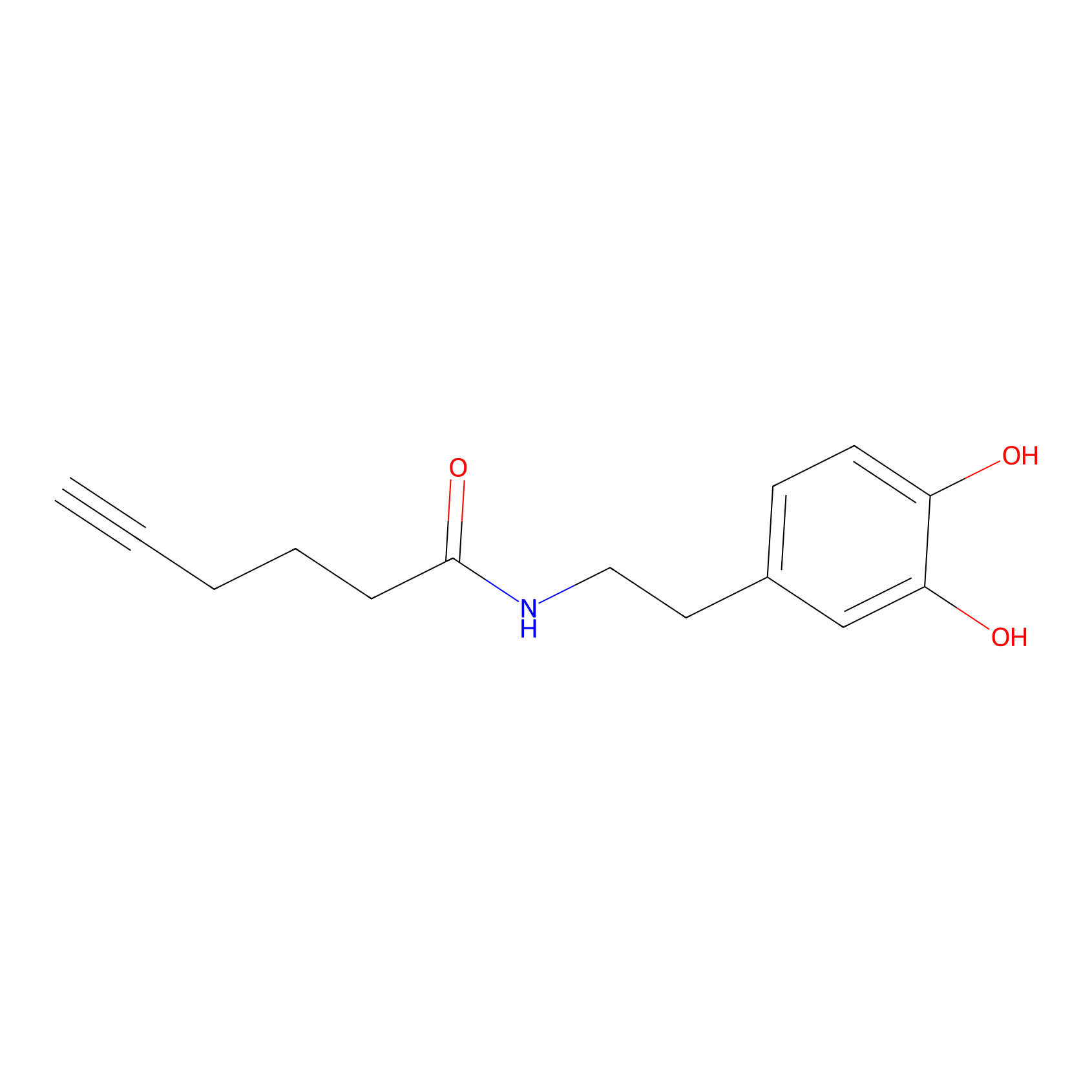 |
11.15 | LDD0348 | [2] | |
|
HRP Probe Info |
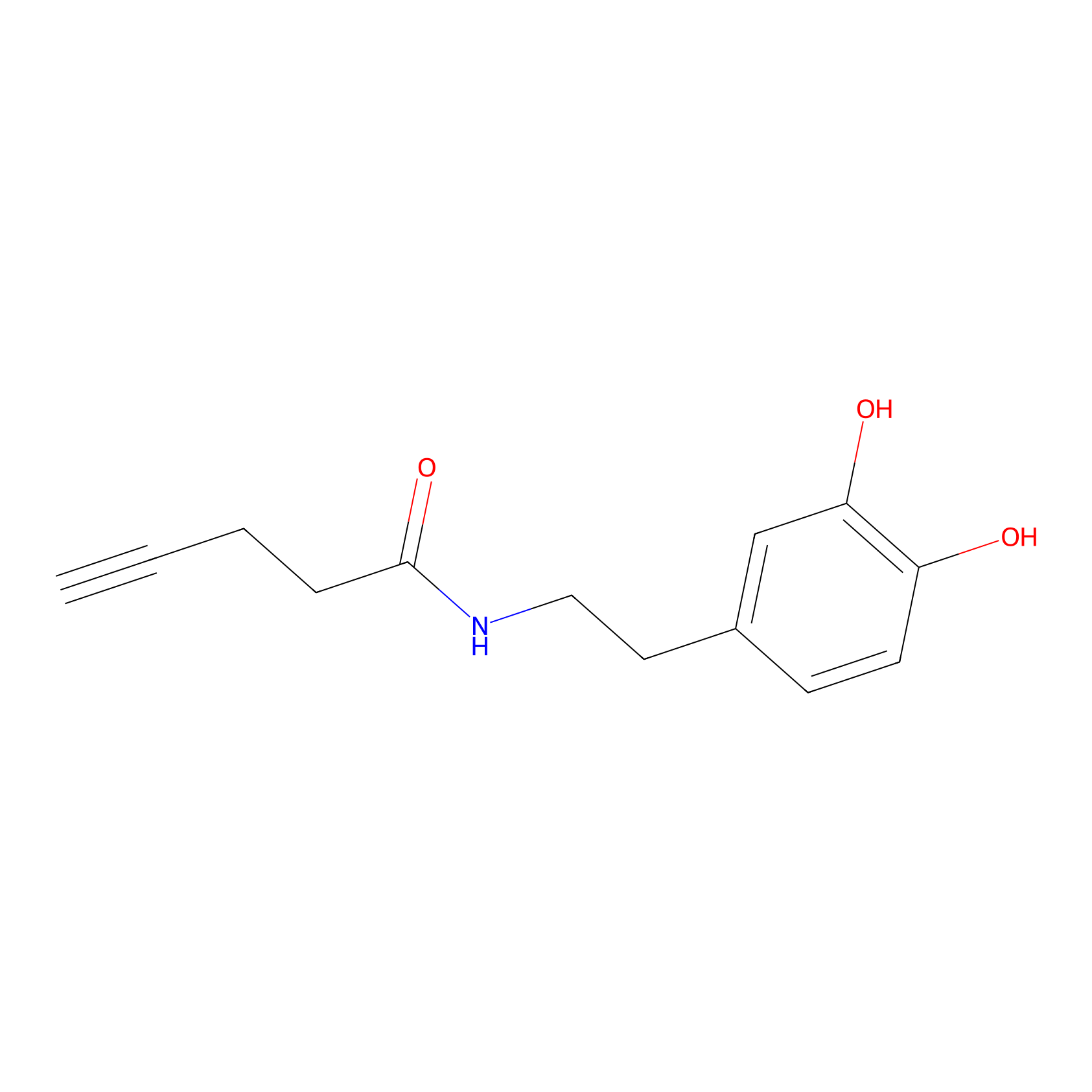 |
9.83 | LDD0347 | [2] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [3] | |
|
TH211 Probe Info |
 |
Y29(7.39) | LDD0260 | [4] | |
|
TH216 Probe Info |
 |
Y382(19.00); Y29(16.02) | LDD0259 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
STPyne Probe Info |
 |
K227(10.00); K27(3.58); K317(10.00); K326(0.51) | LDD0277 | [6] | |
|
P12 Probe Info |
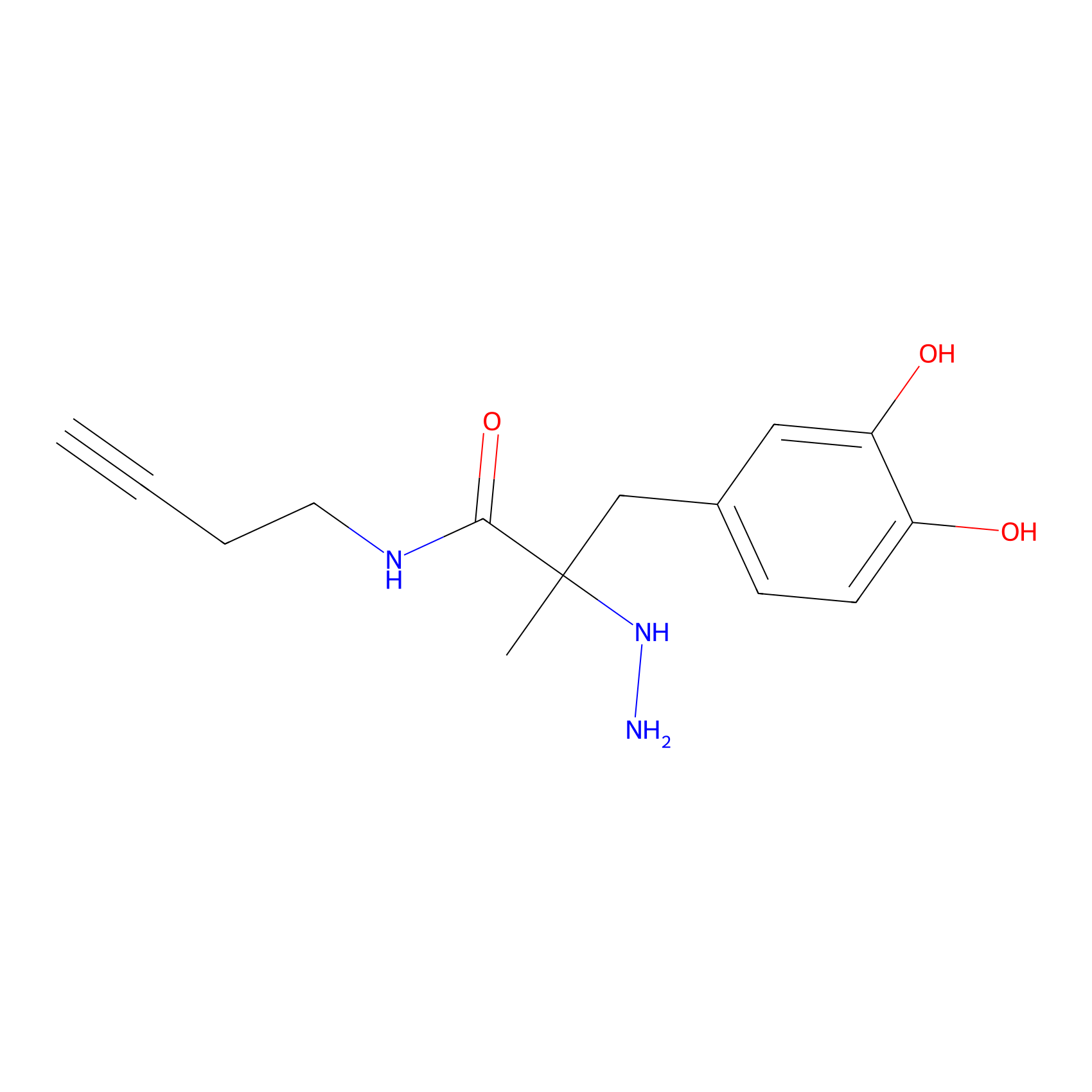 |
16.80 | LDD0202 | [7] | |
|
AHL-Pu-1 Probe Info |
 |
C68(2.31); C66(2.06); C1065(2.65) | LDD0168 | [8] | |
|
Acrolein Probe Info |
 |
H221(0.00); H216(0.00) | LDD0221 | [9] | |
|
DBIA Probe Info |
 |
C68(2.85); C66(3.04); C1129(1.61); C667(1.49) | LDD0080 | [10] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [11] | |
|
ATP probe Probe Info |
 |
K849(0.00); K850(0.00); K755(0.00) | LDD0199 | [12] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C1129(0.00); C1065(0.00) | LDD0038 | [13] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [14] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [15] | |
|
Lodoacetamide azide Probe Info |
 |
C1129(0.00); C1065(0.00); C404(0.00) | LDD0037 | [13] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [16] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [17] | |
|
NAIA_4 Probe Info |
 |
C585(0.00); C593(0.00); C1065(0.00); C1129(0.00) | LDD2226 | [18] | |
|
TFBX Probe Info |
 |
C21(0.00); C66(0.00); C68(0.00) | LDD0027 | [17] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [16] | |
|
DA-P3 Probe Info |
 |
C68(0.00); C21(0.00); C1065(0.00) | LDD0178 | [2] | |
|
1d-yne Probe Info |
 |
K61(0.00); K294(0.00) | LDD0356 | [19] | |
|
Compound 10 Probe Info |
 |
C1065(0.00); C1129(0.00); C148(0.00); C21(0.00) | LDD2216 | [20] | |
|
Compound 11 Probe Info |
 |
C1065(0.00); C1129(0.00); C148(0.00); C21(0.00) | LDD2213 | [20] | |
|
ENE Probe Info |
 |
C1129(0.00); C68(0.00) | LDD0006 | [16] | |
|
IPM Probe Info |
 |
C1129(0.00); C678(0.00); C1065(0.00); C404(0.00) | LDD0005 | [16] | |
|
NHS Probe Info |
 |
K384(0.00); K353(0.00) | LDD0010 | [16] | |
|
NPM Probe Info |
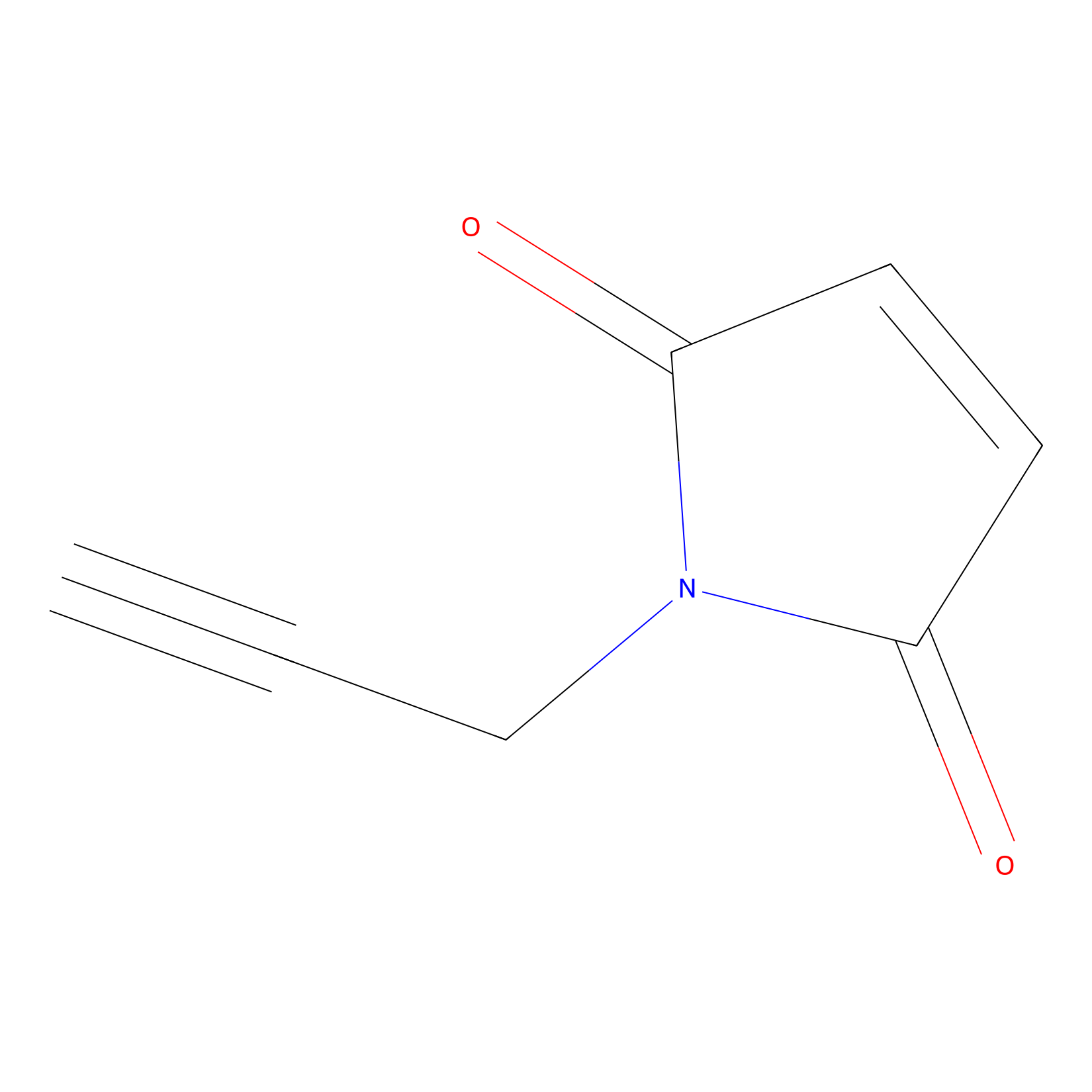 |
C404(0.00); C1129(0.00); C21(0.00) | LDD0016 | [16] | |
|
SF Probe Info |
 |
Y260(0.00); Y382(0.00) | LDD0028 | [21] | |
|
VSF Probe Info |
 |
C1129(0.00); C66(0.00); C585(0.00); C68(0.00) | LDD0007 | [16] | |
|
Phosphinate-6 Probe Info |
 |
C874(0.00); C753(0.00); C404(0.00); C148(0.00) | LDD0018 | [22] | |
|
1c-yne Probe Info |
 |
K693(0.00); K263(0.00); K613(0.00); K228(0.00) | LDD0228 | [19] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [9] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [9] | |
|
W1 Probe Info |
 |
C66(0.00); C68(0.00) | LDD0236 | [23] | |
|
AOyne Probe Info |
 |
5.90 | LDD0443 | [24] | |
|
NAIA_5 Probe Info |
 |
C21(0.00); C404(0.00) | LDD2223 | [18] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [25] | |
|
FFF probe2 Probe Info |
 |
18.23 | LDD0463 | [25] | |
|
FFF probe9 Probe Info |
 |
5.22 | LDD0470 | [25] | |
|
OEA-DA Probe Info |
 |
4.25 | LDD0046 | [26] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C404(0.60) | LDD2142 | [27] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C404(0.93); C678(0.61); C1129(0.86); C753(1.14) | LDD2112 | [27] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C1065(0.85) | LDD2130 | [27] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C1065(1.02); C404(0.83) | LDD2117 | [27] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C1065(0.79) | LDD2152 | [27] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C1065(0.99); C404(1.08) | LDD2103 | [27] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C1065(0.54); C404(0.59) | LDD2132 | [27] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C1065(0.78) | LDD2131 | [27] |
| LDCM0025 | 4SU-RNA | HEK-293T | C68(2.31); C66(2.06); C1065(2.65) | LDD0168 | [8] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C68(3.34); C21(3.03); C593(2.14); C585(20.00) | LDD0169 | [8] |
| LDCM0214 | AC1 | HCT 116 | C1065(0.89); C1129(0.86); C404(0.80); C593(0.78) | LDD0531 | [10] |
| LDCM0215 | AC10 | HCT 116 | C1065(1.19); C1129(1.29); C21(1.00); C404(1.03) | LDD0532 | [10] |
| LDCM0216 | AC100 | HCT 116 | C1065(1.03); C1129(1.17); C21(1.12); C404(1.12) | LDD0533 | [10] |
| LDCM0217 | AC101 | HCT 116 | C1065(1.06); C1129(1.28); C21(1.02); C404(0.95) | LDD0534 | [10] |
| LDCM0218 | AC102 | HCT 116 | C1065(1.02); C1129(0.98); C21(1.01); C404(0.90) | LDD0535 | [10] |
| LDCM0219 | AC103 | HCT 116 | C1065(1.17); C1129(1.42); C21(1.20); C404(0.87) | LDD0536 | [10] |
| LDCM0220 | AC104 | HCT 116 | C1065(1.44); C1129(1.24); C21(0.80); C404(0.75) | LDD0537 | [10] |
| LDCM0221 | AC105 | HCT 116 | C1065(1.15); C1129(1.58); C21(1.24); C404(1.00) | LDD0538 | [10] |
| LDCM0222 | AC106 | HCT 116 | C1065(1.14); C1129(1.67); C21(1.28); C404(0.84) | LDD0539 | [10] |
| LDCM0223 | AC107 | HCT 116 | C1065(1.14); C1129(1.35); C21(1.15); C404(0.57) | LDD0540 | [10] |
| LDCM0224 | AC108 | HCT 116 | C1065(0.92); C1129(1.19); C21(1.22); C404(0.59) | LDD0541 | [10] |
| LDCM0225 | AC109 | HCT 116 | C1065(1.05); C1129(0.92); C21(0.89); C404(0.40) | LDD0542 | [10] |
| LDCM0226 | AC11 | HCT 116 | C1065(1.17); C1129(1.43); C21(0.94); C404(1.08) | LDD0543 | [10] |
| LDCM0227 | AC110 | HCT 116 | C1065(1.09); C1129(1.12); C21(0.97); C404(0.53) | LDD0544 | [10] |
| LDCM0228 | AC111 | HCT 116 | C1065(1.31); C1129(1.32); C21(1.09); C404(0.61) | LDD0545 | [10] |
| LDCM0229 | AC112 | HCT 116 | C1065(1.32); C1129(1.26); C21(1.05); C404(0.54) | LDD0546 | [10] |
| LDCM0230 | AC113 | HCT 116 | C1065(0.99); C1129(0.98); C21(0.85); C404(0.76) | LDD0547 | [10] |
| LDCM0231 | AC114 | HCT 116 | C1065(1.00); C1129(1.36); C21(0.92); C404(0.89) | LDD0548 | [10] |
| LDCM0232 | AC115 | HCT 116 | C1065(1.28); C1129(1.51); C21(1.01); C404(0.91) | LDD0549 | [10] |
| LDCM0233 | AC116 | HCT 116 | C1065(1.95); C1129(1.46); C21(0.86); C404(0.76) | LDD0550 | [10] |
| LDCM0234 | AC117 | HCT 116 | C1065(0.98); C1129(1.34); C21(0.86); C404(0.75) | LDD0551 | [10] |
| LDCM0235 | AC118 | HCT 116 | C1065(1.02); C1129(1.26); C21(0.84); C404(0.81) | LDD0552 | [10] |
| LDCM0236 | AC119 | HCT 116 | C1065(1.31); C1129(1.27); C21(1.01); C404(0.70) | LDD0553 | [10] |
| LDCM0237 | AC12 | HCT 116 | C1065(1.08); C1129(1.05); C21(0.92); C404(0.97) | LDD0554 | [10] |
| LDCM0238 | AC120 | HCT 116 | C1065(1.04); C1129(1.04); C21(1.05); C404(0.97) | LDD0555 | [10] |
| LDCM0239 | AC121 | HCT 116 | C1065(0.96); C1129(1.19); C21(1.08); C404(0.57) | LDD0556 | [10] |
| LDCM0240 | AC122 | HCT 116 | C1065(1.11); C1129(1.28); C21(0.74); C404(0.63) | LDD0557 | [10] |
| LDCM0241 | AC123 | HCT 116 | C1065(1.05); C1129(1.36); C21(0.86); C404(0.55) | LDD0558 | [10] |
| LDCM0242 | AC124 | HCT 116 | C1065(1.08); C1129(1.21); C21(0.94); C404(0.55) | LDD0559 | [10] |
| LDCM0243 | AC125 | HCT 116 | C1065(0.96); C1129(1.30); C21(0.82); C404(0.68) | LDD0560 | [10] |
| LDCM0244 | AC126 | HCT 116 | C1065(1.14); C1129(1.42); C21(0.91); C404(0.58) | LDD0561 | [10] |
| LDCM0245 | AC127 | HCT 116 | C1065(1.22); C1129(1.59); C21(0.83); C404(0.59) | LDD0562 | [10] |
| LDCM0246 | AC128 | HCT 116 | C1065(0.49); C1129(1.17); C404(0.33); C593(0.69) | LDD0563 | [10] |
| LDCM0247 | AC129 | HCT 116 | C1065(0.46); C1129(0.88); C404(0.40); C593(0.85) | LDD0564 | [10] |
| LDCM0249 | AC130 | HCT 116 | C1065(0.59); C1129(1.15); C404(0.50); C593(0.84) | LDD0566 | [10] |
| LDCM0250 | AC131 | HCT 116 | C1065(0.55); C1129(0.99); C404(0.30); C593(0.98) | LDD0567 | [10] |
| LDCM0251 | AC132 | HCT 116 | C1065(0.63); C1129(1.00); C404(0.35); C593(1.07) | LDD0568 | [10] |
| LDCM0252 | AC133 | HCT 116 | C1065(0.59); C1129(1.05); C404(0.32); C593(0.92) | LDD0569 | [10] |
| LDCM0253 | AC134 | HCT 116 | C1065(0.66); C1129(1.32); C404(0.34); C593(1.01) | LDD0570 | [10] |
| LDCM0254 | AC135 | HCT 116 | C1065(0.59); C1129(1.27); C404(0.36); C593(1.02) | LDD0571 | [10] |
| LDCM0255 | AC136 | HCT 116 | C1065(0.70); C1129(1.05); C404(0.30); C593(1.10) | LDD0572 | [10] |
| LDCM0256 | AC137 | HCT 116 | C1065(0.65); C1129(0.93); C404(0.23); C593(0.81) | LDD0573 | [10] |
| LDCM0257 | AC138 | HCT 116 | C1065(0.98); C1129(1.10); C404(0.25); C593(0.94) | LDD0574 | [10] |
| LDCM0258 | AC139 | HCT 116 | C1065(0.65); C1129(1.19); C404(0.24); C593(1.02) | LDD0575 | [10] |
| LDCM0259 | AC14 | HCT 116 | C1065(1.15); C1129(1.25); C21(0.89); C404(1.07) | LDD0576 | [10] |
| LDCM0260 | AC140 | HCT 116 | C1065(0.85); C1129(1.26); C404(0.23); C593(1.19) | LDD0577 | [10] |
| LDCM0261 | AC141 | HCT 116 | C1065(0.82); C1129(1.33); C404(0.25); C593(1.16) | LDD0578 | [10] |
| LDCM0262 | AC142 | HCT 116 | C1065(0.63); C1129(1.14); C404(0.20); C593(1.03) | LDD0579 | [10] |
| LDCM0263 | AC143 | HCT 116 | C404(0.62); C799(0.88); C802(0.88); C803(0.88) | LDD0580 | [10] |
| LDCM0264 | AC144 | HCT 116 | C404(0.44); C664(0.82); C1129(0.98); C871(1.00) | LDD0581 | [10] |
| LDCM0265 | AC145 | HCT 116 | C404(0.62); C664(0.81); C799(0.91); C1129(0.92) | LDD0582 | [10] |
| LDCM0266 | AC146 | HCT 116 | C404(0.54); C1129(0.92); C802(0.93); C664(0.98) | LDD0583 | [10] |
| LDCM0267 | AC147 | HCT 116 | C404(0.55); C1129(0.93); C871(0.94); C874(0.94) | LDD0584 | [10] |
| LDCM0268 | AC148 | HCT 116 | C404(0.60); C1129(0.83); C664(0.94); C802(1.03) | LDD0585 | [10] |
| LDCM0269 | AC149 | HCT 116 | C404(0.63); C664(0.87); C802(0.94); C1129(1.01) | LDD0586 | [10] |
| LDCM0270 | AC15 | HCT 116 | C667(0.80); C871(0.81); C874(0.81); C664(0.81) | LDD0587 | [10] |
| LDCM0271 | AC150 | HCT 116 | C802(0.78); C799(0.80); C404(0.82); C803(0.84) | LDD0588 | [10] |
| LDCM0272 | AC151 | HCT 116 | C404(0.62); C871(0.82); C874(0.82); C799(0.88) | LDD0589 | [10] |
| LDCM0273 | AC152 | HCT 116 | C404(0.51); C1129(0.88); C871(1.01); C874(1.01) | LDD0590 | [10] |
| LDCM0274 | AC153 | HCT 116 | C404(0.51); C664(0.78); C1129(0.90); C802(0.98) | LDD0591 | [10] |
| LDCM0621 | AC154 | HCT 116 | C1065(1.20); C1129(0.91); C21(1.27); C404(0.58) | LDD2158 | [10] |
| LDCM0622 | AC155 | HCT 116 | C1065(1.13); C1129(0.92); C21(1.50); C404(0.71) | LDD2159 | [10] |
| LDCM0623 | AC156 | HCT 116 | C1065(0.98); C1129(0.97); C21(1.30); C404(0.45) | LDD2160 | [10] |
| LDCM0624 | AC157 | HCT 116 | C1065(1.04); C1129(0.87); C21(1.46); C404(0.61) | LDD2161 | [10] |
| LDCM0276 | AC17 | HCT 116 | C404(0.55); C21(0.79); C1065(0.83); C871(0.83) | LDD0593 | [10] |
| LDCM0277 | AC18 | HCT 116 | C404(0.57); C66(0.78); C1065(0.83); C871(0.85) | LDD0594 | [10] |
| LDCM0278 | AC19 | HCT 116 | C404(0.50); C21(0.68); C1065(0.81); C871(0.83) | LDD0595 | [10] |
| LDCM0279 | AC2 | HCT 116 | C678(0.81); C681(0.81); C404(0.82); C1065(0.84) | LDD0596 | [10] |
| LDCM0280 | AC20 | HCT 116 | C404(0.61); C871(0.81); C874(0.81); C21(0.86) | LDD0597 | [10] |
| LDCM0281 | AC21 | HCT 116 | C404(0.55); C871(0.61); C874(0.61); C21(0.72) | LDD0598 | [10] |
| LDCM0282 | AC22 | HCT 116 | C404(0.53); C871(0.72); C874(0.72); C21(0.90) | LDD0599 | [10] |
| LDCM0283 | AC23 | HCT 116 | C404(0.41); C871(0.68); C874(0.68); C664(0.79) | LDD0600 | [10] |
| LDCM0284 | AC24 | HCT 116 | C404(0.63); C871(0.71); C874(0.71); C21(0.78) | LDD0601 | [10] |
| LDCM0285 | AC25 | HCT 116 | C593(0.89); C799(0.91); C857(0.97); C860(0.97) | LDD0602 | [10] |
| LDCM0286 | AC26 | HCT 116 | C857(1.01); C860(1.01); C664(1.09); C799(1.12) | LDD0603 | [10] |
| LDCM0287 | AC27 | HCT 116 | C799(1.08); C664(1.13); C857(1.16); C860(1.16) | LDD0604 | [10] |
| LDCM0288 | AC28 | HCT 116 | C664(1.07); C799(1.07); C1129(1.08); C667(1.09) | LDD0605 | [10] |
| LDCM0289 | AC29 | HCT 116 | C799(1.08); C802(1.08); C803(1.08); C593(1.12) | LDD0606 | [10] |
| LDCM0290 | AC3 | HCT 116 | C404(0.75); C1065(0.76); C678(0.79); C681(0.79) | LDD0607 | [10] |
| LDCM0291 | AC30 | HCT 116 | C803(1.07); C802(1.10); C799(1.12); C667(1.12) | LDD0608 | [10] |
| LDCM0292 | AC31 | HCT 116 | C799(0.90); C667(1.07); C664(1.12); C857(1.18) | LDD0609 | [10] |
| LDCM0293 | AC32 | HCT 116 | C799(1.00); C667(1.16); C664(1.20); C802(1.22) | LDD0610 | [10] |
| LDCM0294 | AC33 | HCT 116 | C667(0.88); C664(1.05); C803(1.10); C802(1.11) | LDD0611 | [10] |
| LDCM0295 | AC34 | HCT 116 | C803(0.72); C802(0.90); C667(0.97); C799(1.08) | LDD0612 | [10] |
| LDCM0296 | AC35 | HCT 116 | C593(0.68); C68(0.77); C66(0.78); C1129(0.80) | LDD0613 | [10] |
| LDCM0297 | AC36 | HCT 116 | C593(0.69); C68(0.90); C1129(0.92); C66(0.93) | LDD0614 | [10] |
| LDCM0298 | AC37 | HCT 116 | C593(0.75); C1129(0.88); C681(0.97); C678(1.01) | LDD0615 | [10] |
| LDCM0299 | AC38 | HCT 116 | C1129(0.91); C593(0.96); C664(0.98); C66(1.00) | LDD0616 | [10] |
| LDCM0300 | AC39 | HCT 116 | C1129(0.87); C799(0.93); C802(0.93); C593(0.94) | LDD0617 | [10] |
| LDCM0301 | AC4 | HCT 116 | C404(0.67); C593(0.77); C1129(0.86); C1065(0.89) | LDD0618 | [10] |
| LDCM0302 | AC40 | HCT 116 | C1129(0.93); C799(1.06); C802(1.06); C664(1.09) | LDD0619 | [10] |
| LDCM0303 | AC41 | HCT 116 | C593(0.70); C1129(0.93); C68(0.98); C66(1.00) | LDD0620 | [10] |
| LDCM0304 | AC42 | HCT 116 | C1129(0.90); C68(0.98); C678(1.00); C664(1.00) | LDD0621 | [10] |
| LDCM0305 | AC43 | HCT 116 | C593(0.93); C1129(1.00); C681(1.03); C678(1.05) | LDD0622 | [10] |
| LDCM0306 | AC44 | HCT 116 | C593(0.84); C681(0.92); C678(0.96); C1129(0.99) | LDD0623 | [10] |
| LDCM0307 | AC45 | HCT 116 | C1129(0.91); C664(1.06); C667(1.07); C799(1.12) | LDD0624 | [10] |
| LDCM0308 | AC46 | HCT 116 | C799(0.85); C802(0.85); C678(0.86); C681(0.86) | LDD0625 | [10] |
| LDCM0309 | AC47 | HCT 116 | C799(0.76); C802(0.76); C21(1.00); C66(1.03) | LDD0626 | [10] |
| LDCM0310 | AC48 | HCT 116 | C799(0.71); C802(0.71); C678(1.02); C681(1.02) | LDD0627 | [10] |
| LDCM0311 | AC49 | HCT 116 | C799(0.59); C802(0.59); C678(1.14); C681(1.14) | LDD0628 | [10] |
| LDCM0312 | AC5 | HCT 116 | C404(0.60); C593(0.73); C1065(0.83); C1129(0.85) | LDD0629 | [10] |
| LDCM0313 | AC50 | HCT 116 | C799(0.76); C802(0.76); C21(1.10); C678(1.12) | LDD0630 | [10] |
| LDCM0314 | AC51 | HCT 116 | C799(0.67); C802(0.67); C678(0.94); C681(0.94) | LDD0631 | [10] |
| LDCM0315 | AC52 | HCT 116 | C799(0.67); C802(0.67); C678(0.94); C681(0.94) | LDD0632 | [10] |
| LDCM0316 | AC53 | HCT 116 | C799(0.72); C802(0.72); C664(0.88); C678(0.93) | LDD0633 | [10] |
| LDCM0317 | AC54 | HCT 116 | C799(0.77); C802(0.77); C664(0.92); C21(0.97) | LDD0634 | [10] |
| LDCM0318 | AC55 | HCT 116 | C799(0.80); C802(0.80); C664(1.02); C21(1.04) | LDD0635 | [10] |
| LDCM0319 | AC56 | HCT 116 | C799(0.81); C802(0.81); C664(0.99); C21(1.02) | LDD0636 | [10] |
| LDCM0320 | AC57 | HCT 116 | C871(0.77); C874(0.77); C21(0.80); C1065(0.92) | LDD0637 | [10] |
| LDCM0321 | AC58 | HCT 116 | C871(0.94); C874(0.94); C21(0.95); C799(1.14) | LDD0638 | [10] |
| LDCM0322 | AC59 | HCT 116 | C871(0.78); C874(0.78); C21(0.92); C1065(1.07) | LDD0639 | [10] |
| LDCM0323 | AC6 | HCT 116 | C871(0.85); C874(0.85); C667(0.92); C664(0.93) | LDD0640 | [10] |
| LDCM0324 | AC60 | HCT 116 | C871(0.83); C874(0.83); C21(0.84); C1129(0.98) | LDD0641 | [10] |
| LDCM0325 | AC61 | HCT 116 | C21(0.86); C871(0.91); C874(0.91); C1065(1.01) | LDD0642 | [10] |
| LDCM0326 | AC62 | HCT 116 | C21(0.85); C871(0.95); C874(0.95); C1065(1.00) | LDD0643 | [10] |
| LDCM0327 | AC63 | HCT 116 | C871(0.88); C874(0.88); C21(0.89); C799(1.11) | LDD0644 | [10] |
| LDCM0328 | AC64 | HCT 116 | C21(0.78); C871(0.88); C874(0.88); C1065(0.98) | LDD0645 | [10] |
| LDCM0329 | AC65 | HCT 116 | C21(0.87); C68(0.94); C66(0.99); C871(1.03) | LDD0646 | [10] |
| LDCM0330 | AC66 | HCT 116 | C68(0.87); C66(0.92); C21(0.93); C871(0.97) | LDD0647 | [10] |
| LDCM0331 | AC67 | HCT 116 | C21(0.92); C871(1.00); C874(1.00); C68(1.05) | LDD0648 | [10] |
| LDCM0332 | AC68 | HCT 116 | C593(0.71); C803(0.76); C799(0.88); C802(0.88) | LDD0649 | [10] |
| LDCM0333 | AC69 | HCT 116 | C803(0.77); C799(0.81); C802(0.81); C593(0.87) | LDD0650 | [10] |
| LDCM0334 | AC7 | HCT 116 | C667(0.93); C857(0.94); C860(0.94); C871(0.94) | LDD0651 | [10] |
| LDCM0335 | AC70 | HCT 116 | C803(0.62); C799(0.76); C802(0.76); C667(0.83) | LDD0652 | [10] |
| LDCM0336 | AC71 | HCT 116 | C799(0.79); C802(0.79); C678(0.82); C681(0.82) | LDD0653 | [10] |
| LDCM0337 | AC72 | HCT 116 | C803(0.56); C799(0.71); C802(0.71); C593(0.82) | LDD0654 | [10] |
| LDCM0338 | AC73 | HCT 116 | C803(0.60); C799(0.72); C802(0.72); C593(0.95) | LDD0655 | [10] |
| LDCM0339 | AC74 | HCT 116 | C803(0.57); C593(0.74); C799(0.75); C802(0.75) | LDD0656 | [10] |
| LDCM0340 | AC75 | HCT 116 | C803(0.71); C799(0.81); C802(0.81); C593(1.01) | LDD0657 | [10] |
| LDCM0341 | AC76 | HCT 116 | C803(0.63); C799(0.73); C802(0.73); C593(0.84) | LDD0658 | [10] |
| LDCM0342 | AC77 | HCT 116 | C799(0.80); C802(0.80); C803(0.82); C593(0.92) | LDD0659 | [10] |
| LDCM0343 | AC78 | HCT 116 | C803(0.63); C799(0.69); C802(0.69); C857(0.96) | LDD0660 | [10] |
| LDCM0344 | AC79 | HCT 116 | C593(0.82); C803(0.84); C799(0.88); C802(0.88) | LDD0661 | [10] |
| LDCM0345 | AC8 | HCT 116 | C667(0.83); C664(0.86); C871(0.90); C874(0.90) | LDD0662 | [10] |
| LDCM0346 | AC80 | HCT 116 | C803(0.81); C799(0.86); C802(0.86); C593(0.93) | LDD0663 | [10] |
| LDCM0347 | AC81 | HCT 116 | C593(0.74); C803(0.87); C799(0.88); C802(0.88) | LDD0664 | [10] |
| LDCM0348 | AC82 | HCT 116 | C803(0.71); C799(0.78); C802(0.78); C593(0.86) | LDD0665 | [10] |
| LDCM0349 | AC83 | HCT 116 | C1065(0.95); C871(1.10); C874(1.10); C21(1.13) | LDD0666 | [10] |
| LDCM0350 | AC84 | HCT 116 | C21(1.01); C1065(1.04); C857(1.06); C860(1.06) | LDD0667 | [10] |
| LDCM0351 | AC85 | HCT 116 | C404(0.86); C1065(0.96); C857(1.02); C860(1.02) | LDD0668 | [10] |
| LDCM0352 | AC86 | HCT 116 | C1065(0.80); C404(0.96); C21(1.06); C871(1.08) | LDD0669 | [10] |
| LDCM0353 | AC87 | HCT 116 | C404(0.69); C1065(0.82); C871(0.96); C874(0.96) | LDD0670 | [10] |
| LDCM0354 | AC88 | HCT 116 | C404(0.81); C1065(0.84); C21(1.00); C871(1.02) | LDD0671 | [10] |
| LDCM0355 | AC89 | HCT 116 | C404(0.81); C1065(0.85); C21(1.04); C871(1.17) | LDD0672 | [10] |
| LDCM0357 | AC90 | HCT 116 | C1065(0.73); C667(1.04); C1129(1.04); C593(1.09) | LDD0674 | [10] |
| LDCM0358 | AC91 | HCT 116 | C404(0.57); C1065(0.79); C21(1.21); C871(1.27) | LDD0675 | [10] |
| LDCM0359 | AC92 | HCT 116 | C404(0.49); C1065(0.82); C21(0.92); C871(1.20) | LDD0676 | [10] |
| LDCM0360 | AC93 | HCT 116 | C404(0.49); C21(0.82); C1065(0.82); C678(1.11) | LDD0677 | [10] |
| LDCM0361 | AC94 | HCT 116 | C404(0.27); C1065(0.79); C21(0.87); C68(1.05) | LDD0678 | [10] |
| LDCM0362 | AC95 | HCT 116 | C404(0.53); C1065(0.78); C857(0.97); C860(0.97) | LDD0679 | [10] |
| LDCM0363 | AC96 | HCT 116 | C404(0.28); C1065(0.77); C21(1.02); C857(1.12) | LDD0680 | [10] |
| LDCM0364 | AC97 | HCT 116 | C404(0.56); C1065(1.00); C21(1.04); C678(1.26) | LDD0681 | [10] |
| LDCM0365 | AC98 | HCT 116 | C404(1.19); C667(1.20); C678(1.21); C681(1.21) | LDD0682 | [10] |
| LDCM0366 | AC99 | HCT 116 | C66(0.80); C68(0.82); C678(0.85); C681(0.85) | LDD0683 | [10] |
| LDCM0545 | Acetamide | MDA-MB-231 | C1065(0.52); C404(0.30) | LDD2138 | [27] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C1065(1.00) | LDD2113 | [27] |
| LDCM0248 | AKOS034007472 | HCT 116 | C1065(0.95); C1129(1.02); C21(0.88); C404(1.24) | LDD0565 | [10] |
| LDCM0356 | AKOS034007680 | HCT 116 | C667(0.82); C664(0.86); C871(0.91); C874(0.91) | LDD0673 | [10] |
| LDCM0275 | AKOS034007705 | HCT 116 | C871(0.83); C874(0.83); C667(0.86); C664(0.91) | LDD0592 | [10] |
| LDCM0156 | Aniline | NCI-H1299 | N.A. | LDD0404 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C799(2.16); C802(2.16); C803(2.16); C404(0.89) | LDD2171 | [10] |
| LDCM0102 | BDHI 8 | Jurkat | C857(15.04); C860(15.04) | LDD0204 | [28] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C1065(0.50); C404(0.36) | LDD2091 | [27] |
| LDCM0088 | C45 | HEK-293T | 16.80 | LDD0202 | [7] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [9] |
| LDCM0632 | CL-Sc | Hep-G2 | C21(0.76); C1129(0.53); C21(0.51) | LDD2227 | [18] |
| LDCM0367 | CL1 | HCT 116 | C857(0.82); C860(0.82); C799(0.84); C802(0.84) | LDD0684 | [10] |
| LDCM0368 | CL10 | HCT 116 | C404(0.91); C21(1.05); C664(1.06); C799(1.08) | LDD0685 | [10] |
| LDCM0369 | CL100 | HCT 116 | C404(0.81); C593(0.88); C1129(0.88); C1065(0.97) | LDD0686 | [10] |
| LDCM0370 | CL101 | HCT 116 | C667(0.88); C664(0.89); C21(0.99); C404(1.09) | LDD0687 | [10] |
| LDCM0371 | CL102 | HCT 116 | C667(0.69); C664(0.71); C871(0.88); C874(0.88) | LDD0688 | [10] |
| LDCM0372 | CL103 | HCT 116 | C667(0.88); C404(0.91); C664(0.91); C871(0.92) | LDD0689 | [10] |
| LDCM0373 | CL104 | HCT 116 | C667(0.80); C871(0.83); C874(0.83); C21(0.84) | LDD0690 | [10] |
| LDCM0374 | CL105 | HCT 116 | C404(0.55); C871(0.74); C874(0.74); C21(0.80) | LDD0691 | [10] |
| LDCM0375 | CL106 | HCT 116 | C404(0.42); C21(0.74); C1129(0.83); C871(0.83) | LDD0692 | [10] |
| LDCM0376 | CL107 | HCT 116 | C404(0.48); C21(0.72); C1065(0.91); C1129(1.01) | LDD0693 | [10] |
| LDCM0377 | CL108 | HCT 116 | C404(0.44); C21(0.72); C1065(0.75); C871(0.85) | LDD0694 | [10] |
| LDCM0378 | CL109 | HCT 116 | C404(0.60); C21(0.71); C871(0.87); C874(0.87) | LDD0695 | [10] |
| LDCM0379 | CL11 | HCT 116 | C871(0.76); C874(0.76); C404(0.86); C667(0.89) | LDD0696 | [10] |
| LDCM0380 | CL110 | HCT 116 | C404(0.35); C21(0.62); C1065(0.72); C871(0.87) | LDD0697 | [10] |
| LDCM0381 | CL111 | HCT 116 | C404(0.39); C21(0.52); C871(0.83); C874(0.83) | LDD0698 | [10] |
| LDCM0382 | CL112 | HCT 116 | C799(0.84); C593(0.89); C857(0.93); C860(0.93) | LDD0699 | [10] |
| LDCM0383 | CL113 | HCT 116 | C799(0.95); C802(0.98); C803(1.00); C664(1.07) | LDD0700 | [10] |
| LDCM0384 | CL114 | HCT 116 | C803(0.73); C802(0.82); C857(0.88); C860(0.88) | LDD0701 | [10] |
| LDCM0385 | CL115 | HCT 116 | C803(0.84); C802(0.88); C799(0.93); C664(0.99) | LDD0702 | [10] |
| LDCM0386 | CL116 | HCT 116 | C799(0.94); C802(0.98); C593(1.01); C803(1.03) | LDD0703 | [10] |
| LDCM0387 | CL117 | HCT 116 | C1129(1.02); C799(1.12); C802(1.12); C664(1.13) | LDD0704 | [10] |
| LDCM0388 | CL118 | HCT 116 | C593(0.85); C1129(0.91); C21(1.02); C664(1.02) | LDD0705 | [10] |
| LDCM0389 | CL119 | HCT 116 | C799(0.97); C802(0.97); C21(0.98); C1129(0.99) | LDD0706 | [10] |
| LDCM0390 | CL12 | HCT 116 | C404(0.87); C667(1.04); C21(1.07); C593(1.10) | LDD0707 | [10] |
| LDCM0391 | CL120 | HCT 116 | C593(0.87); C681(0.99); C678(1.05); C1129(1.08) | LDD0708 | [10] |
| LDCM0392 | CL121 | HCT 116 | C799(0.76); C802(0.76); C664(0.91); C21(0.99) | LDD0709 | [10] |
| LDCM0393 | CL122 | HCT 116 | C799(0.74); C802(0.74); C678(0.99); C681(0.99) | LDD0710 | [10] |
| LDCM0394 | CL123 | HCT 116 | C799(0.62); C802(0.62); C664(0.82); C678(0.90) | LDD0711 | [10] |
| LDCM0395 | CL124 | HCT 116 | C799(0.76); C802(0.76); C664(0.88); C678(1.06) | LDD0712 | [10] |
| LDCM0396 | CL125 | HCT 116 | C1129(0.90); C871(0.96); C874(0.96); C857(0.99) | LDD0713 | [10] |
| LDCM0397 | CL126 | HCT 116 | C871(0.76); C874(0.76); C857(0.86); C860(0.86) | LDD0714 | [10] |
| LDCM0398 | CL127 | HCT 116 | C21(0.79); C871(0.96); C874(0.96); C857(0.96) | LDD0715 | [10] |
| LDCM0399 | CL128 | HCT 116 | C21(0.78); C871(0.82); C874(0.82); C1065(0.98) | LDD0716 | [10] |
| LDCM0400 | CL13 | HCT 116 | C404(0.85); C667(0.88); C871(0.91); C874(0.91) | LDD0717 | [10] |
| LDCM0401 | CL14 | HCT 116 | C667(0.85); C678(0.89); C681(0.89); C871(0.91) | LDD0718 | [10] |
| LDCM0402 | CL15 | HCT 116 | C667(0.68); C404(0.78); C593(0.87); C871(0.92) | LDD0719 | [10] |
| LDCM0403 | CL16 | HCT 116 | C21(0.95); C664(1.00); C1065(1.01); C667(1.04) | LDD0720 | [10] |
| LDCM0404 | CL17 | HCT 116 | C664(0.86); C404(0.88); C667(0.88); C1065(0.96) | LDD0721 | [10] |
| LDCM0405 | CL18 | HCT 116 | C593(0.95); C1065(0.95); C664(1.02); C21(1.13) | LDD0722 | [10] |
| LDCM0406 | CL19 | HCT 116 | C664(0.94); C593(0.98); C1065(1.00); C21(1.03) | LDD0723 | [10] |
| LDCM0407 | CL2 | HCT 116 | C857(0.89); C860(0.89); C799(0.89); C802(0.89) | LDD0724 | [10] |
| LDCM0408 | CL20 | HCT 116 | C1065(0.99); C664(1.05); C21(1.08); C593(1.10) | LDD0725 | [10] |
| LDCM0409 | CL21 | HCT 116 | C664(0.76); C593(0.92); C1065(0.94); C667(1.08) | LDD0726 | [10] |
| LDCM0410 | CL22 | HCT 116 | C593(0.89); C21(0.90); C664(0.90); C1065(0.97) | LDD0727 | [10] |
| LDCM0411 | CL23 | HCT 116 | C593(0.91); C21(0.93); C667(1.00); C1065(1.01) | LDD0728 | [10] |
| LDCM0412 | CL24 | HCT 116 | C593(0.87); C1065(0.89); C21(0.89); C664(1.06) | LDD0729 | [10] |
| LDCM0413 | CL25 | HCT 116 | C1065(0.93); C1129(1.36); C21(0.92); C404(1.78) | LDD0730 | [10] |
| LDCM0414 | CL26 | HCT 116 | C1065(1.21); C1129(1.18); C21(0.99); C404(1.46) | LDD0731 | [10] |
| LDCM0415 | CL27 | HCT 116 | C1065(0.88); C1129(1.31); C21(0.93); C404(1.21) | LDD0732 | [10] |
| LDCM0416 | CL28 | HCT 116 | C1065(0.90); C1129(1.31); C21(1.06); C404(1.85) | LDD0733 | [10] |
| LDCM0417 | CL29 | HCT 116 | C1065(1.00); C1129(1.31); C21(1.07); C404(1.43) | LDD0734 | [10] |
| LDCM0418 | CL3 | HCT 116 | C1065(0.94); C1129(0.96); C21(1.11); C404(0.89) | LDD0735 | [10] |
| LDCM0419 | CL30 | HCT 116 | C1065(0.85); C1129(1.15); C21(1.09); C404(1.14) | LDD0736 | [10] |
| LDCM0420 | CL31 | HCT 116 | C1065(0.92); C1129(1.17); C21(0.96); C593(0.94) | LDD0737 | [10] |
| LDCM0421 | CL32 | HCT 116 | C1065(1.09); C1129(1.41); C21(1.04); C404(0.82) | LDD0738 | [10] |
| LDCM0422 | CL33 | HCT 116 | C1065(1.07); C1129(1.19); C21(1.11); C404(0.87) | LDD0739 | [10] |
| LDCM0423 | CL34 | HCT 116 | C1065(1.04); C1129(1.35); C21(1.37); C404(0.81) | LDD0740 | [10] |
| LDCM0424 | CL35 | HCT 116 | C1065(1.18); C1129(1.56); C21(0.91); C404(0.61) | LDD0741 | [10] |
| LDCM0425 | CL36 | HCT 116 | C1065(1.29); C1129(1.47); C21(0.96); C404(0.56) | LDD0742 | [10] |
| LDCM0426 | CL37 | HCT 116 | C1065(1.26); C1129(1.45); C21(0.98); C404(0.54) | LDD0743 | [10] |
| LDCM0428 | CL39 | HCT 116 | C1065(1.08); C1129(1.53); C21(1.00); C404(0.52) | LDD0745 | [10] |
| LDCM0429 | CL4 | HCT 116 | C1065(0.96); C1129(1.07); C21(1.11); C404(1.09) | LDD0746 | [10] |
| LDCM0430 | CL40 | HCT 116 | C1065(1.09); C1129(1.15); C21(0.91); C404(0.48) | LDD0747 | [10] |
| LDCM0431 | CL41 | HCT 116 | C1065(1.05); C1129(1.20); C21(1.01); C404(0.53) | LDD0748 | [10] |
| LDCM0432 | CL42 | HCT 116 | C1065(1.13); C1129(1.70); C21(0.70); C404(0.40) | LDD0749 | [10] |
| LDCM0433 | CL43 | HCT 116 | C1065(1.07); C1129(1.39); C21(0.94); C404(0.44) | LDD0750 | [10] |
| LDCM0434 | CL44 | HCT 116 | C1065(1.06); C1129(1.38); C21(1.06); C404(0.46) | LDD0751 | [10] |
| LDCM0435 | CL45 | HCT 116 | C1065(0.96); C1129(1.53); C21(0.89); C404(0.47) | LDD0752 | [10] |
| LDCM0436 | CL46 | HCT 116 | C1065(0.97); C1129(0.92); C21(1.27); C404(0.83) | LDD0753 | [10] |
| LDCM0437 | CL47 | HCT 116 | C1065(1.02); C1129(0.90); C21(1.12); C404(0.68) | LDD0754 | [10] |
| LDCM0438 | CL48 | HCT 116 | C1065(1.05); C1129(0.91); C21(1.18); C404(0.94) | LDD0755 | [10] |
| LDCM0439 | CL49 | HCT 116 | C1065(0.81); C1129(0.95); C21(1.10); C404(0.65) | LDD0756 | [10] |
| LDCM0440 | CL5 | HCT 116 | C1065(0.97); C1129(1.06); C21(1.17); C404(0.96) | LDD0757 | [10] |
| LDCM0441 | CL50 | HCT 116 | C1065(1.01); C1129(0.96); C21(1.13); C404(0.93) | LDD0758 | [10] |
| LDCM0442 | CL51 | HCT 116 | C1065(0.97); C1129(0.99); C21(1.27); C404(0.73) | LDD0759 | [10] |
| LDCM0443 | CL52 | HCT 116 | C1065(1.13); C1129(1.04); C21(1.31); C404(0.88) | LDD0760 | [10] |
| LDCM0444 | CL53 | HCT 116 | C1065(1.40); C1129(0.97); C21(1.12); C404(0.78) | LDD0761 | [10] |
| LDCM0445 | CL54 | HCT 116 | C1065(1.06); C1129(0.92); C21(1.02); C404(0.86) | LDD0762 | [10] |
| LDCM0446 | CL55 | HCT 116 | C1065(1.04); C1129(0.92); C21(0.98); C404(0.91) | LDD0763 | [10] |
| LDCM0447 | CL56 | HCT 116 | C1065(1.06); C1129(0.96); C21(0.98); C404(0.78) | LDD0764 | [10] |
| LDCM0448 | CL57 | HCT 116 | C1065(0.96); C1129(0.95); C21(0.99); C404(0.68) | LDD0765 | [10] |
| LDCM0449 | CL58 | HCT 116 | C1065(1.01); C1129(0.90); C21(1.02); C404(1.04) | LDD0766 | [10] |
| LDCM0450 | CL59 | HCT 116 | C1065(0.92); C1129(1.04); C21(0.84); C404(0.78) | LDD0767 | [10] |
| LDCM0451 | CL6 | HCT 116 | C1065(1.07); C1129(1.11); C21(1.25); C404(1.16) | LDD0768 | [10] |
| LDCM0452 | CL60 | HCT 116 | C1065(0.96); C1129(0.99); C21(1.13); C404(0.91) | LDD0769 | [10] |
| LDCM0453 | CL61 | HCT 116 | C1065(1.03); C1129(0.93); C21(1.11); C66(1.10) | LDD0770 | [10] |
| LDCM0454 | CL62 | HCT 116 | C1065(1.08); C1129(1.04); C21(1.20); C66(1.17) | LDD0771 | [10] |
| LDCM0455 | CL63 | HCT 116 | C1065(1.20); C1129(0.99); C21(1.06); C66(1.29) | LDD0772 | [10] |
| LDCM0456 | CL64 | HCT 116 | C1065(1.24); C1129(0.97); C21(0.95); C66(1.29) | LDD0773 | [10] |
| LDCM0457 | CL65 | HCT 116 | C1065(1.00); C1129(0.99); C21(0.99); C66(1.02) | LDD0774 | [10] |
| LDCM0458 | CL66 | HCT 116 | C1065(1.40); C1129(1.00); C21(0.97); C66(1.50) | LDD0775 | [10] |
| LDCM0459 | CL67 | HCT 116 | C1065(1.47); C1129(0.96); C21(1.05); C66(1.15) | LDD0776 | [10] |
| LDCM0460 | CL68 | HCT 116 | C1065(1.40); C1129(0.89); C21(0.89); C66(1.31) | LDD0777 | [10] |
| LDCM0461 | CL69 | HCT 116 | C1065(1.08); C1129(1.04); C21(1.15); C66(1.25) | LDD0778 | [10] |
| LDCM0462 | CL7 | HCT 116 | C1065(1.12); C1129(1.22); C21(1.02); C404(0.95) | LDD0779 | [10] |
| LDCM0463 | CL70 | HCT 116 | C1065(1.21); C1129(1.02); C21(0.92); C66(1.27) | LDD0780 | [10] |
| LDCM0464 | CL71 | HCT 116 | C1065(1.34); C1129(1.04); C21(0.83); C66(1.15) | LDD0781 | [10] |
| LDCM0465 | CL72 | HCT 116 | C1065(1.23); C1129(1.08); C21(0.87); C66(1.07) | LDD0782 | [10] |
| LDCM0466 | CL73 | HCT 116 | C1065(1.19); C1129(0.95); C21(0.76); C66(1.28) | LDD0783 | [10] |
| LDCM0467 | CL74 | HCT 116 | C1065(1.31); C1129(1.03); C21(0.98); C66(1.38) | LDD0784 | [10] |
| LDCM0469 | CL76 | HCT 116 | C1065(1.14); C1129(0.90); C21(1.05); C404(0.86) | LDD0786 | [10] |
| LDCM0470 | CL77 | HCT 116 | C1065(1.30); C1129(1.04); C21(0.87); C404(0.87) | LDD0787 | [10] |
| LDCM0471 | CL78 | HCT 116 | C1065(1.08); C1129(1.07); C21(1.01); C404(1.00) | LDD0788 | [10] |
| LDCM0472 | CL79 | HCT 116 | C1065(1.18); C1129(0.85); C21(1.26); C404(0.72) | LDD0789 | [10] |
| LDCM0473 | CL8 | HCT 116 | C1065(1.70); C1129(1.51); C21(1.47); C404(1.17) | LDD0790 | [10] |
| LDCM0474 | CL80 | HCT 116 | C1065(1.06); C1129(0.93); C21(1.11); C404(0.83) | LDD0791 | [10] |
| LDCM0475 | CL81 | HCT 116 | C1065(1.03); C1129(0.90); C21(1.04); C404(0.75) | LDD0792 | [10] |
| LDCM0476 | CL82 | HCT 116 | C1065(0.82); C1129(0.81); C21(1.38); C404(0.65) | LDD0793 | [10] |
| LDCM0477 | CL83 | HCT 116 | C1065(1.61); C1129(0.86); C21(1.23); C404(0.78) | LDD0794 | [10] |
| LDCM0478 | CL84 | HCT 116 | C1065(1.43); C1129(1.06); C21(1.54); C404(1.04) | LDD0795 | [10] |
| LDCM0479 | CL85 | HCT 116 | C1065(1.49); C1129(0.96); C21(0.90); C404(0.94) | LDD0796 | [10] |
| LDCM0480 | CL86 | HCT 116 | C1065(1.37); C1129(0.99); C21(0.82); C404(0.93) | LDD0797 | [10] |
| LDCM0481 | CL87 | HCT 116 | C1065(1.79); C1129(0.83); C21(1.05); C404(0.80) | LDD0798 | [10] |
| LDCM0482 | CL88 | HCT 116 | C1065(1.54); C1129(0.95); C21(1.18); C404(0.82) | LDD0799 | [10] |
| LDCM0483 | CL89 | HCT 116 | C1065(1.23); C1129(0.96); C21(1.80); C404(0.74) | LDD0800 | [10] |
| LDCM0484 | CL9 | HCT 116 | C1065(1.17); C1129(1.12); C21(1.13); C404(1.12) | LDD0801 | [10] |
| LDCM0485 | CL90 | HCT 116 | C1065(1.35); C1129(0.90); C21(0.78); C404(0.77) | LDD0802 | [10] |
| LDCM0486 | CL91 | HCT 116 | C1065(1.02); C1129(1.03); C404(1.10); C593(1.03) | LDD0803 | [10] |
| LDCM0487 | CL92 | HCT 116 | C1065(0.96); C1129(0.87); C404(1.19); C593(0.97) | LDD0804 | [10] |
| LDCM0488 | CL93 | HCT 116 | C1065(0.92); C1129(0.89); C404(1.03); C593(0.83) | LDD0805 | [10] |
| LDCM0489 | CL94 | HCT 116 | C1065(1.01); C1129(0.87); C404(1.08); C593(0.95) | LDD0806 | [10] |
| LDCM0490 | CL95 | HCT 116 | C1065(1.12); C1129(0.92); C404(0.99); C593(0.93) | LDD0807 | [10] |
| LDCM0491 | CL96 | HCT 116 | C1065(0.90); C1129(0.94); C404(0.89); C593(0.89) | LDD0808 | [10] |
| LDCM0492 | CL97 | HCT 116 | C1065(0.87); C1129(1.08); C404(1.20); C593(1.02) | LDD0809 | [10] |
| LDCM0493 | CL98 | HCT 116 | C1065(0.91); C1129(1.10); C404(1.14); C593(1.05) | LDD0810 | [10] |
| LDCM0494 | CL99 | HCT 116 | C1065(0.96); C1129(0.94); C404(0.88); C593(0.87) | LDD0811 | [10] |
| LDCM0634 | CY-0357 | Hep-G2 | C21(1.14); C1065(0.75) | LDD2228 | [18] |
| LDCM0027 | Dopamine | HEK-293T | 4.94 | LDD0179 | [2] |
| LDCM0495 | E2913 | HEK-293T | C21(1.01); C1065(1.25); C585(1.04); C1129(1.04) | LDD1698 | [29] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C404(2.25); C681(1.26); C678(1.26); C585(1.15) | LDD1702 | [27] |
| LDCM0625 | F8 | Ramos | C404(1.95); C1065(1.43); C21(2.13); C148(0.36) | LDD2187 | [30] |
| LDCM0572 | Fragment10 | Ramos | C404(1.37); C21(1.05) | LDD2189 | [30] |
| LDCM0573 | Fragment11 | Ramos | C404(1.94); C1065(0.15); C21(0.30); C148(1.54) | LDD2190 | [30] |
| LDCM0574 | Fragment12 | Ramos | C404(0.90); C21(0.61) | LDD2191 | [30] |
| LDCM0575 | Fragment13 | Ramos | C404(0.93); C21(0.54) | LDD2192 | [30] |
| LDCM0576 | Fragment14 | Ramos | C404(1.01); C1065(1.07); C21(0.89); C148(0.95) | LDD2193 | [30] |
| LDCM0579 | Fragment20 | Ramos | C404(1.16); C21(1.07) | LDD2194 | [30] |
| LDCM0580 | Fragment21 | Ramos | C404(0.97); C21(0.50) | LDD2195 | [30] |
| LDCM0582 | Fragment23 | Ramos | C404(1.19); C21(0.95) | LDD2196 | [30] |
| LDCM0578 | Fragment27 | Ramos | C404(1.10); C21(0.91) | LDD2197 | [30] |
| LDCM0586 | Fragment28 | Ramos | C404(0.72); C21(0.74); C148(1.27) | LDD2198 | [30] |
| LDCM0588 | Fragment30 | Ramos | C404(2.24); C21(0.60) | LDD2199 | [30] |
| LDCM0589 | Fragment31 | Ramos | C404(1.50); C21(0.42) | LDD2200 | [30] |
| LDCM0590 | Fragment32 | Ramos | C404(1.73); C21(1.06) | LDD2201 | [30] |
| LDCM0468 | Fragment33 | HCT 116 | C1065(1.19); C1129(0.93); C21(0.81); C66(1.34) | LDD0785 | [10] |
| LDCM0596 | Fragment38 | Ramos | C404(1.34); C21(1.01) | LDD2203 | [30] |
| LDCM0566 | Fragment4 | Ramos | C1065(0.64); C21(1.08); C148(0.65) | LDD2184 | [30] |
| LDCM0427 | Fragment51 | HCT 116 | C1065(2.10); C1129(1.72); C21(1.13); C404(0.59) | LDD0744 | [10] |
| LDCM0610 | Fragment52 | Ramos | C404(2.09); C21(0.76) | LDD2204 | [30] |
| LDCM0614 | Fragment56 | Ramos | C404(1.44); C21(0.50) | LDD2205 | [30] |
| LDCM0569 | Fragment7 | Ramos | C404(1.05); C1065(0.94); C21(1.17); C148(1.62) | LDD2186 | [30] |
| LDCM0571 | Fragment9 | Ramos | C404(1.17); C21(0.60) | LDD2188 | [30] |
| LDCM0107 | IAA | HeLa | H221(0.00); H216(0.00) | LDD0221 | [9] |
| LDCM0022 | KB02 | HCT 116 | C68(2.85); C66(3.04); C1129(1.61); C667(1.49) | LDD0080 | [10] |
| LDCM0023 | KB03 | HCT 116 | C68(2.04); C66(2.02); C1129(1.19); C667(1.29) | LDD0081 | [10] |
| LDCM0024 | KB05 | HCT 116 | C68(4.04); C66(3.23); C1129(1.84); C667(1.66) | LDD0082 | [10] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C1065(0.89); C404(1.17); C1129(0.85); C753(0.98) | LDD2102 | [27] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C1065(0.56); C404(0.39) | LDD2121 | [27] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [9] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C1065(0.57); C404(0.35) | LDD2089 | [27] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C1065(1.27); C404(1.44) | LDD2090 | [27] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C404(1.01); C1129(1.01) | LDD2092 | [27] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C1065(0.99) | LDD2093 | [27] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C404(1.42); C1129(1.01); C753(1.02) | LDD2094 | [27] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C1065(0.50); C404(0.51) | LDD2096 | [27] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C1065(0.88) | LDD2097 | [27] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C404(1.22) | LDD2098 | [27] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C1065(0.98); C404(0.56); C753(0.63) | LDD2099 | [27] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C404(0.78); C678(0.35); C1129(1.43) | LDD2100 | [27] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C404(1.10) | LDD2101 | [27] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C404(0.72); C1129(1.04) | LDD2104 | [27] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C404(2.26); C678(1.89); C1129(1.23) | LDD2105 | [27] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C404(0.42); C1129(0.60) | LDD2106 | [27] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C1065(0.85); C404(0.88) | LDD2107 | [27] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C404(0.67); C1129(0.86) | LDD2108 | [27] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C1065(1.09); C404(0.92) | LDD2109 | [27] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C404(1.49) | LDD2110 | [27] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C1065(0.80) | LDD2111 | [27] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C404(0.48); C678(0.63); C1129(0.71); C753(0.89) | LDD2114 | [27] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C1065(0.66); C404(0.69) | LDD2116 | [27] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C1065(0.54); C404(0.67) | LDD2118 | [27] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C1065(0.64) | LDD2119 | [27] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C1065(0.95); C404(0.88); C753(0.90) | LDD2120 | [27] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C404(0.97) | LDD2122 | [27] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C404(0.47) | LDD2123 | [27] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C1065(0.64); C404(0.68) | LDD2124 | [27] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C1065(0.82); C404(0.87); C753(0.50) | LDD2125 | [27] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C1065(0.72); C404(0.69) | LDD2126 | [27] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C1065(1.06); C404(0.77) | LDD2127 | [27] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C1065(0.99); C404(0.87) | LDD2128 | [27] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C1065(1.02); C404(0.48) | LDD2129 | [27] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C1065(0.84) | LDD2133 | [27] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C1065(0.55) | LDD2134 | [27] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C1065(0.94); C404(0.56) | LDD2135 | [27] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C1065(1.22); C404(0.99); C753(0.87) | LDD2136 | [27] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C1065(0.94); C404(0.87) | LDD2137 | [27] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C1065(0.97); C404(0.90) | LDD1700 | [27] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C1065(0.92); C404(0.57) | LDD2140 | [27] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C1065(0.24); C404(0.61); C1129(1.05) | LDD2141 | [27] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C1065(1.03); C404(0.94) | LDD2143 | [27] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C404(2.07) | LDD2145 | [27] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C1065(0.94); C404(0.56) | LDD2146 | [27] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C1065(2.59); C404(2.26); C753(3.14) | LDD2147 | [27] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C1065(0.54); C404(0.78) | LDD2149 | [27] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C1065(0.46); C404(0.28) | LDD2150 | [27] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C1065(1.04) | LDD2151 | [27] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C1065(1.37) | LDD2153 | [27] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C21(0.46) | LDD2206 | [31] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C21(0.73) | LDD2207 | [31] |
| LDCM0029 | Quercetin | HEK-293T | 6.48 | LDD0181 | [2] |
| LDCM0131 | RA190 | MM1.R | C66(20.00); C68(20.00) | LDD0304 | [32] |
| LDCM0021 | THZ1 | HCT 116 | C799(2.16); C802(2.16); C803(2.16); C404(0.89) | LDD2173 | [10] |
| LDCM0112 | W16 | Hep-G2 | C1065(0.95) | LDD0239 | [23] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Ubiquitin carboxyl-terminal hydrolase 7 (USP7) | Peptidase C19 family | Q93009 | |||
| Ephrin type-B receptor 2 (EPHB2) | Tyr protein kinase family | P29323 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Importin subunit beta-1 (KPNB1) | Importin beta family | Q14974 | |||
| Nuclear pore complex protein Nup153 (NUP153) | NUP153 family | P49790 | |||
References
