Details of the Target
General Information of Target
| Target ID | LDTP00267 | |||||
|---|---|---|---|---|---|---|
| Target Name | DNA-directed RNA polymerase, mitochondrial (POLRMT) | |||||
| Gene Name | POLRMT | |||||
| Gene ID | 5442 | |||||
| Synonyms |
DNA-directed RNA polymerase, mitochondrial; MtRPOL; EC 2.7.7.6 |
|||||
| 3D Structure | ||||||
| Sequence |
MSALCWGRGAAGLKRALRPCGRPGLPGKEGTAGGVCGPRRSSSASPQEQDQDRRKDWGHV
ELLEVLQARVRQLQAESVSEVVVNRVDVARLPECGSGDGSLQPPRKVQMGAKDATPVPCG RWAKILEKDKRTQQMRMQRLKAKLQMPFQSGEFKALTRRLQVEPRLLSKQMAGCLEDCTR QAPESPWEEQLARLLQEAPGKLSLDVEQAPSGQHSQAQLSGQQQRLLAFFKCCLLTDQLP LAHHLLVVHHGQRQKRKLLTLDMYNAVMLGWARQGAFKELVYVLFMVKDAGLTPDLLSYA AALQCMGRQDQDAGTIERCLEQMSQEGLKLQALFTAVLLSEEDRATVLKAVHKVKPTFSL PPQLPPPVNTSKLLRDVYAKDGRVSYPKLHLPLKTLQCLFEKQLHMELASRVCVVSVEKP TLPSKEVKHARKTLKTLRDQWEKALCRALRETKNRLEREVYEGRFSLYPFLCLLDEREVV RMLLQVLQALPAQGESFTTLARELSARTFSRHVVQRQRVSGQVQALQNHYRKYLCLLASD AEVPEPCLPRQYWEELGAPEALREQPWPLPVQMELGKLLAEMLVQATQMPCSLDKPHRSS RLVPVLYHVYSFRNVQQIGILKPHPAYVQLLEKAAEPTLTFEAVDVPMLCPPLPWTSPHS GAFLLSPTKLMRTVEGATQHQELLETCPPTALHGALDALTQLGNCAWRVNGRVLDLVLQL FQAKGCPQLGVPAPPSEAPQPPEAHLPHSAAPARKAELRRELAHCQKVAREMHSLRAEAL YRLSLAQHLRDRVFWLPHNMDFRGRTYPCPPHFNHLGSDVARALLEFAQGRPLGPHGLDW LKIHLVNLTGLKKREPLRKRLAFAEEVMDDILDSADQPLTGRKWWMGAEEPWQTLACCME VANAVRASDPAAYVSHLPVHQDGSCNGLQHYAALGRDSVGAASVNLEPSDVPQDVYSGVA AQVEVFRRQDAQRGMRVAQVLEGFITRKVVKQTVMTVVYGVTRYGGRLQIEKRLRELSDF PQEFVWEASHYLVRQVFKSLQEMFSGTRAIQHWLTESARLISHMGSVVEWVTPLGVPVIQ PYRLDSKVKQIGGGIQSITYTHNGDISRKPNTRKQKNGFPPNFIHSLDSSHMMLTALHCY RKGLTFVSVHDCYWTHAADVSVMNQVCREQFVRLHSEPILQDLSRFLVKRFCSEPQKILE ASQLKETLQAVPKPGAFDLEQVKRSTYFFS |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Phage and mitochondrial RNA polymerase family
|
|||||
| Subcellular location |
Mitochondrion
|
|||||
| Function |
DNA-dependent RNA polymerase catalyzes the transcription of mitochondrial DNA into RNA using the four ribonucleoside triphosphates as substrates. Component of the mitochondrial transcription initiation complex, composed at least of TFB2M, TFAM and POLRMT that is required for basal transcription of mitochondrial DNA. In this complex, TFAM recruits POLRMT to a specific promoter whereas TFB2M induces structural changes in POLRMT to enable promoter opening and trapping of the DNA non-template strand. Has DNA primase activity. Catalyzes the synthesis of short RNA primers that are necessary for the initiation of lagging-strand DNA synthesis from the origin of light-strand DNA replication (OriL).
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
9.92 | LDD0402 | [1] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
STPyne Probe Info |
 |
K14(6.67) | LDD0277 | [3] | |
|
BTD Probe Info |
 |
C1192(1.01); C413(1.00) | LDD2093 | [4] | |
|
Sulforaphane-probe2 Probe Info |
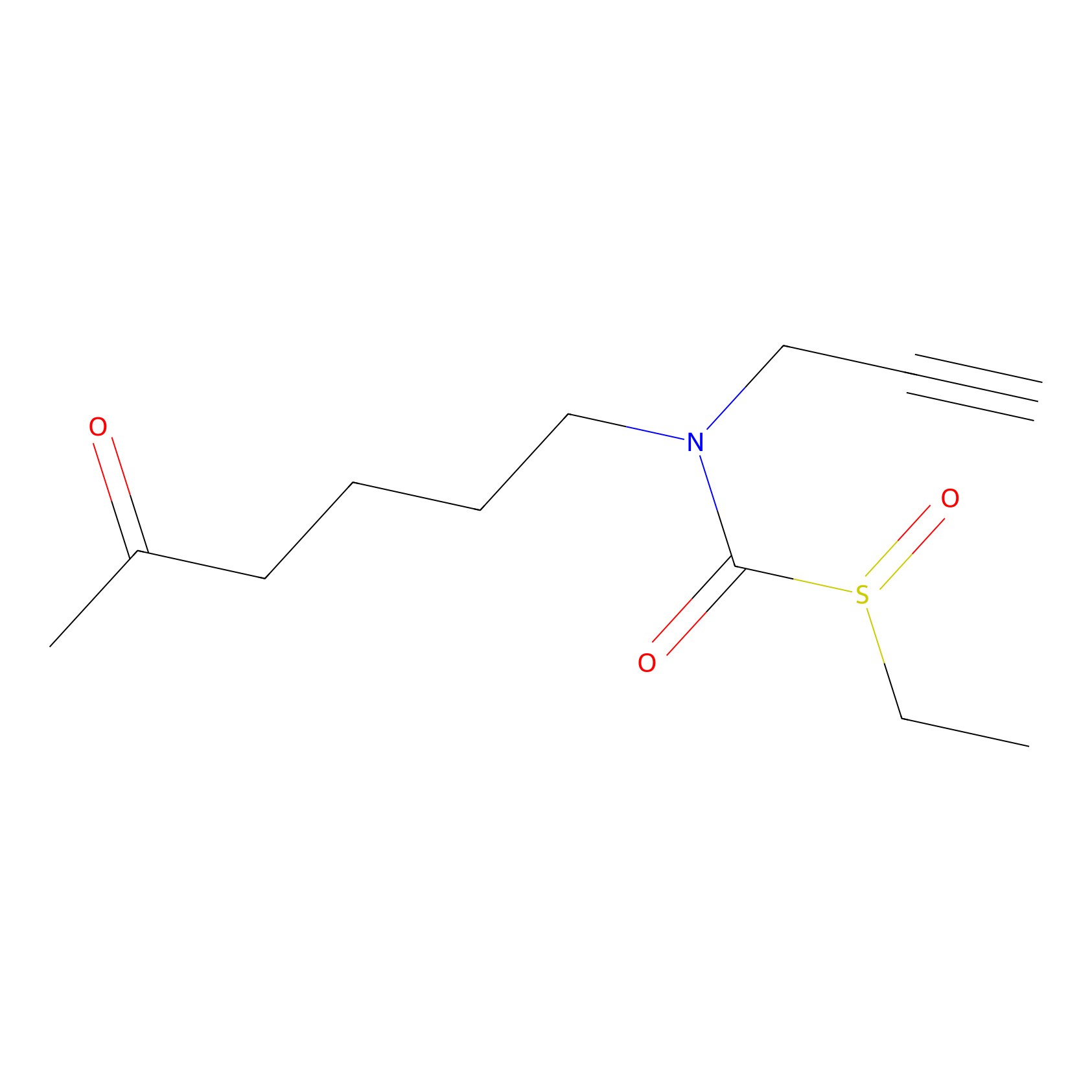 |
2.60 | LDD0160 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C705(5.26); C413(2.17) | LDD0168 | [6] | |
|
DBIA Probe Info |
 |
C174(1.03); C178(1.03); C398(1.01); C119(0.95) | LDD0078 | [7] | |
|
W1 Probe Info |
 |
C119(1.00) | LDD0239 | [8] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [9] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C472(0.00); C305(0.00); C413(0.00) | LDD0038 | [10] | |
|
IA-alkyne Probe Info |
 |
C591(0.00); C398(0.00); C413(0.00) | LDD0036 | [10] | |
|
Lodoacetamide azide Probe Info |
 |
C319(0.00); C305(0.00); C472(0.00); C765(0.00) | LDD0037 | [10] | |
|
TFBX Probe Info |
 |
C319(0.00); C174(0.00) | LDD0027 | [11] | |
|
WYneN Probe Info |
 |
C413(0.00); C94(0.00); C119(0.00) | LDD0021 | [12] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [13] | |
|
Compound 10 Probe Info |
 |
C119(0.00); C398(0.00) | LDD2216 | [14] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [12] | |
|
IPM Probe Info |
 |
C174(0.00); C726(0.00); C413(0.00); C119(0.00) | LDD0005 | [12] | |
|
VSF Probe Info |
 |
C726(0.00); C413(0.00) | LDD0007 | [12] | |
|
Phosphinate-6 Probe Info |
 |
C119(0.00); C94(0.00) | LDD0018 | [15] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [13] | |
|
Acrolein Probe Info |
 |
C94(0.00); C809(0.00); C119(0.00) | LDD0217 | [16] | |
|
Crotonaldehyde Probe Info |
 |
C119(0.00); C94(0.00) | LDD0219 | [16] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [16] | |
|
AOyne Probe Info |
 |
4.40 | LDD0443 | [17] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [18] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C004 Probe Info |
 |
5.82 | LDD1714 | [19] | |
|
C158 Probe Info |
 |
9.13 | LDD1838 | [19] | |
|
C231 Probe Info |
 |
12.73 | LDD1904 | [19] | |
|
C232 Probe Info |
 |
37.27 | LDD1905 | [19] | |
|
C338 Probe Info |
 |
10.63 | LDD2001 | [19] | |
|
C348 Probe Info |
 |
10.48 | LDD2009 | [19] | |
|
C366 Probe Info |
 |
5.46 | LDD2027 | [19] | |
|
FFF probe13 Probe Info |
 |
17.71 | LDD0475 | [20] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [20] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [20] | |
|
FFF probe3 Probe Info |
 |
11.86 | LDD0465 | [20] | |
|
FFF probe6 Probe Info |
 |
9.38 | LDD0467 | [20] | |
|
JN0003 Probe Info |
 |
7.60 | LDD0469 | [20] | |
|
OEA-DA Probe Info |
 |
8.54 | LDD0046 | [21] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C765(0.83) | LDD2117 | [4] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C765(1.11) | LDD2152 | [4] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C413(0.66) | LDD2132 | [4] |
| LDCM0025 | 4SU-RNA | HEK-293T | C705(5.26); C413(2.17) | LDD0168 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C705(2.20); C687(2.01) | LDD0169 | [6] |
| LDCM0214 | AC1 | HCT 116 | C174(1.14); C726(0.94); C535(0.76); C547(0.76) | LDD0531 | [7] |
| LDCM0215 | AC10 | HCT 116 | C174(1.16); C726(1.04); C535(0.72); C547(0.72) | LDD0532 | [7] |
| LDCM0216 | AC100 | HCT 116 | C174(1.09); C726(0.88); C119(1.09); C1192(1.17) | LDD0533 | [7] |
| LDCM0217 | AC101 | HCT 116 | C174(1.02); C726(1.03); C119(1.15); C1192(0.95) | LDD0534 | [7] |
| LDCM0218 | AC102 | HCT 116 | C174(0.91); C726(1.06); C119(1.13); C1192(0.93) | LDD0535 | [7] |
| LDCM0219 | AC103 | HCT 116 | C174(1.20); C726(1.09); C119(1.37); C1192(0.91) | LDD0536 | [7] |
| LDCM0220 | AC104 | HCT 116 | C174(1.08); C726(1.09); C119(1.10); C1192(1.01) | LDD0537 | [7] |
| LDCM0221 | AC105 | HCT 116 | C174(1.07); C726(0.92); C119(1.39); C1192(1.00) | LDD0538 | [7] |
| LDCM0222 | AC106 | HCT 116 | C174(1.00); C726(0.95); C119(1.23); C1192(1.05) | LDD0539 | [7] |
| LDCM0223 | AC107 | HCT 116 | C174(0.72); C726(0.98); C119(1.27); C1192(1.02) | LDD0540 | [7] |
| LDCM0224 | AC108 | HCT 116 | C174(1.61); C726(0.88); C119(1.08); C1192(1.12) | LDD0541 | [7] |
| LDCM0225 | AC109 | HCT 116 | C174(1.95); C726(0.83); C119(1.13); C1192(1.17) | LDD0542 | [7] |
| LDCM0226 | AC11 | HCT 116 | C174(1.19); C726(1.24); C535(0.79); C547(0.79) | LDD0543 | [7] |
| LDCM0227 | AC110 | HCT 116 | C174(1.35); C726(0.96); C119(1.08); C1192(0.96) | LDD0544 | [7] |
| LDCM0228 | AC111 | HCT 116 | C174(1.32); C726(1.19); C119(1.18); C1192(0.87) | LDD0545 | [7] |
| LDCM0229 | AC112 | HCT 116 | C174(0.97); C726(1.09); C119(1.19); C1192(0.87) | LDD0546 | [7] |
| LDCM0230 | AC113 | HCT 116 | C174(1.23); C726(1.24); C119(0.95); C1192(1.04) | LDD0547 | [7] |
| LDCM0231 | AC114 | HCT 116 | C174(1.01); C726(1.74); C119(0.88); C1192(1.13) | LDD0548 | [7] |
| LDCM0232 | AC115 | HCT 116 | C174(0.93); C726(1.63); C119(0.85); C1192(1.03) | LDD0549 | [7] |
| LDCM0233 | AC116 | HCT 116 | C174(1.05); C726(1.46); C119(0.99); C1192(1.06) | LDD0550 | [7] |
| LDCM0234 | AC117 | HCT 116 | C174(1.02); C726(1.00); C119(0.76); C1192(0.89) | LDD0551 | [7] |
| LDCM0235 | AC118 | HCT 116 | C174(1.00); C726(1.27); C119(0.75); C1192(0.96) | LDD0552 | [7] |
| LDCM0236 | AC119 | HCT 116 | C174(1.32); C726(1.18); C119(1.10); C1192(0.90) | LDD0553 | [7] |
| LDCM0237 | AC12 | HCT 116 | C174(1.02); C726(0.98); C535(1.00); C547(1.00) | LDD0554 | [7] |
| LDCM0238 | AC120 | HCT 116 | C174(1.37); C726(1.39); C119(0.89); C1192(1.07) | LDD0555 | [7] |
| LDCM0239 | AC121 | HCT 116 | C174(1.29); C726(1.18); C119(0.84); C1192(0.93) | LDD0556 | [7] |
| LDCM0240 | AC122 | HCT 116 | C174(1.02); C726(1.14); C119(0.99); C1192(0.96) | LDD0557 | [7] |
| LDCM0241 | AC123 | HCT 116 | C174(1.03); C726(1.15); C119(0.91); C1192(0.88) | LDD0558 | [7] |
| LDCM0242 | AC124 | HCT 116 | C174(1.05); C726(1.27); C119(0.73); C1192(0.87) | LDD0559 | [7] |
| LDCM0243 | AC125 | HCT 116 | C174(1.00); C726(1.13); C119(0.89); C1192(0.99) | LDD0560 | [7] |
| LDCM0244 | AC126 | HCT 116 | C174(0.96); C726(1.50); C119(0.82); C1192(1.01) | LDD0561 | [7] |
| LDCM0245 | AC127 | HCT 116 | C174(0.94); C726(1.56); C119(0.88); C1192(0.97) | LDD0562 | [7] |
| LDCM0246 | AC128 | HCT 116 | C174(1.36); C726(1.00); C535(0.64); C547(0.64) | LDD0563 | [7] |
| LDCM0247 | AC129 | HCT 116 | C174(0.93); C726(0.77); C535(1.19); C547(1.19) | LDD0564 | [7] |
| LDCM0249 | AC130 | HCT 116 | C174(1.05); C726(1.07); C535(0.69); C547(0.69) | LDD0566 | [7] |
| LDCM0250 | AC131 | HCT 116 | C174(0.86); C726(0.90); C535(1.34); C547(1.34) | LDD0567 | [7] |
| LDCM0251 | AC132 | HCT 116 | C174(0.95); C726(1.05); C535(1.41); C547(1.41) | LDD0568 | [7] |
| LDCM0252 | AC133 | HCT 116 | C174(1.03); C726(0.85); C535(0.90); C547(0.90) | LDD0569 | [7] |
| LDCM0253 | AC134 | HCT 116 | C174(1.23); C726(1.00); C535(0.70); C547(0.70) | LDD0570 | [7] |
| LDCM0254 | AC135 | HCT 116 | C174(1.16); C726(1.07); C535(0.73); C547(0.73) | LDD0571 | [7] |
| LDCM0255 | AC136 | HCT 116 | C174(1.11); C726(1.10); C535(0.69); C547(0.69) | LDD0572 | [7] |
| LDCM0256 | AC137 | HCT 116 | C174(1.04); C726(0.86); C535(1.16); C547(1.16) | LDD0573 | [7] |
| LDCM0257 | AC138 | HCT 116 | C174(1.08); C726(0.92); C535(0.74); C547(0.74) | LDD0574 | [7] |
| LDCM0258 | AC139 | HCT 116 | C174(1.16); C726(1.12); C535(1.00); C547(1.00) | LDD0575 | [7] |
| LDCM0259 | AC14 | HCT 116 | C174(1.01); C726(0.92); C535(0.90); C547(0.90) | LDD0576 | [7] |
| LDCM0260 | AC140 | HCT 116 | C174(1.29); C726(1.00); C535(0.67); C547(0.67) | LDD0577 | [7] |
| LDCM0261 | AC141 | HCT 116 | C174(1.19); C726(1.11); C535(1.02); C547(1.02) | LDD0578 | [7] |
| LDCM0262 | AC142 | HCT 116 | C174(1.05); C726(0.79); C535(1.55); C547(1.55) | LDD0579 | [7] |
| LDCM0263 | AC143 | HCT 116 | C174(1.30); C726(1.33); C535(1.14); C547(1.14) | LDD0580 | [7] |
| LDCM0264 | AC144 | HCT 116 | C535(0.38); C547(0.38); C413(0.81); C398(0.99) | LDD0581 | [7] |
| LDCM0265 | AC145 | HCT 116 | C535(0.82); C547(0.82); C413(0.87); C178(0.94) | LDD0582 | [7] |
| LDCM0266 | AC146 | HCT 116 | C535(0.46); C547(0.46); C413(0.76); C398(0.89) | LDD0583 | [7] |
| LDCM0267 | AC147 | HCT 116 | C535(0.37); C547(0.37); C413(0.87); C398(0.90) | LDD0584 | [7] |
| LDCM0268 | AC148 | HCT 116 | C535(0.42); C547(0.42); C413(0.71); C1192(0.79) | LDD0585 | [7] |
| LDCM0269 | AC149 | HCT 116 | C535(0.39); C547(0.39); C178(0.88); C413(0.97) | LDD0586 | [7] |
| LDCM0270 | AC15 | HCT 116 | C765(0.59); C232(0.60); C233(0.60); C535(0.73) | LDD0587 | [7] |
| LDCM0271 | AC150 | HCT 116 | C535(0.73); C547(0.73); C413(0.85); C1192(0.89) | LDD0588 | [7] |
| LDCM0272 | AC151 | HCT 116 | C178(0.82); C413(0.82); C765(0.93); C1192(0.97) | LDD0589 | [7] |
| LDCM0273 | AC152 | HCT 116 | C535(0.34); C547(0.34); C178(0.85); C413(0.90) | LDD0590 | [7] |
| LDCM0274 | AC153 | HCT 116 | C535(0.43); C547(0.43); C178(0.89); C1192(0.90) | LDD0591 | [7] |
| LDCM0621 | AC154 | HCT 116 | C174(1.53); C726(1.68); C535(0.50); C547(0.50) | LDD2158 | [7] |
| LDCM0622 | AC155 | HCT 116 | C174(1.27); C726(1.63); C535(0.43); C547(0.43) | LDD2159 | [7] |
| LDCM0623 | AC156 | HCT 116 | C174(1.04); C726(1.29); C535(1.40); C547(1.40) | LDD2160 | [7] |
| LDCM0624 | AC157 | HCT 116 | C174(1.04); C726(1.24); C535(0.87); C547(0.87) | LDD2161 | [7] |
| LDCM0276 | AC17 | HCT 116 | C305(0.84); C119(0.91); C765(0.92); C94(0.94) | LDD0593 | [7] |
| LDCM0277 | AC18 | HCT 116 | C305(0.54); C535(0.67); C547(0.67); C119(0.84) | LDD0594 | [7] |
| LDCM0278 | AC19 | HCT 116 | C305(0.59); C119(0.86); C765(0.88); C178(0.92) | LDD0595 | [7] |
| LDCM0279 | AC2 | HCT 116 | C687(0.60); C705(0.60); C319(0.87); C809(0.87) | LDD0596 | [7] |
| LDCM0280 | AC20 | HCT 116 | C809(0.86); C305(0.86); C119(0.86); C319(0.91) | LDD0597 | [7] |
| LDCM0281 | AC21 | HCT 116 | C305(0.85); C765(0.85); C319(0.91); C413(0.95) | LDD0598 | [7] |
| LDCM0282 | AC22 | HCT 116 | C319(0.86); C765(0.89); C119(0.93); C94(0.94) | LDD0599 | [7] |
| LDCM0283 | AC23 | HCT 116 | C94(0.88); C765(0.88); C119(0.93); C535(0.94) | LDD0600 | [7] |
| LDCM0284 | AC24 | HCT 116 | C765(0.80); C94(0.89); C119(0.91); C305(0.92) | LDD0601 | [7] |
| LDCM0285 | AC25 | HCT 116 | C305(0.56); C535(0.77); C547(0.77); C1192(0.81) | LDD0602 | [7] |
| LDCM0286 | AC26 | HCT 116 | C305(0.37); C535(0.71); C547(0.71); C687(0.71) | LDD0603 | [7] |
| LDCM0287 | AC27 | HCT 116 | C305(0.46); C535(0.59); C547(0.59); C1192(0.59) | LDD0604 | [7] |
| LDCM0288 | AC28 | HCT 116 | C305(0.45); C535(0.61); C547(0.61); C1192(0.67) | LDD0605 | [7] |
| LDCM0289 | AC29 | HCT 116 | C305(0.57); C687(0.64); C705(0.64); C1192(0.70) | LDD0606 | [7] |
| LDCM0290 | AC3 | HCT 116 | C687(0.81); C705(0.81); C178(0.84); C809(0.85) | LDD0607 | [7] |
| LDCM0291 | AC30 | HCT 116 | C305(0.44); C687(0.52); C705(0.52); C1192(0.63) | LDD0608 | [7] |
| LDCM0292 | AC31 | HCT 116 | C305(0.52); C535(0.61); C547(0.61); C1192(0.62) | LDD0609 | [7] |
| LDCM0293 | AC32 | HCT 116 | C305(0.29); C535(0.40); C547(0.40); C687(0.40) | LDD0610 | [7] |
| LDCM0294 | AC33 | HCT 116 | C305(0.40); C535(0.43); C547(0.43); C687(0.43) | LDD0611 | [7] |
| LDCM0295 | AC34 | HCT 116 | C535(0.37); C547(0.37); C305(0.39); C687(0.40) | LDD0612 | [7] |
| LDCM0296 | AC35 | HCT 116 | C232(0.78); C233(0.78); C765(0.79); C174(0.83) | LDD0613 | [7] |
| LDCM0297 | AC36 | HCT 116 | C94(0.88); C174(0.90); C319(0.94); C178(0.97) | LDD0614 | [7] |
| LDCM0298 | AC37 | HCT 116 | C94(0.89); C765(0.91); C413(0.91); C178(0.92) | LDD0615 | [7] |
| LDCM0299 | AC38 | HCT 116 | C765(0.90); C94(0.93); C398(0.94); C178(0.95) | LDD0616 | [7] |
| LDCM0300 | AC39 | HCT 116 | C765(0.76); C232(0.81); C233(0.81); C413(0.94) | LDD0617 | [7] |
| LDCM0301 | AC4 | HCT 116 | C687(0.63); C705(0.63); C809(0.73); C94(0.85) | LDD0618 | [7] |
| LDCM0302 | AC40 | HCT 116 | C319(0.76); C765(0.79); C232(0.86); C233(0.86) | LDD0619 | [7] |
| LDCM0303 | AC41 | HCT 116 | C765(0.73); C94(0.93); C178(0.96); C726(1.05) | LDD0620 | [7] |
| LDCM0304 | AC42 | HCT 116 | C765(0.72); C94(0.94); C535(0.95); C547(0.95) | LDD0621 | [7] |
| LDCM0305 | AC43 | HCT 116 | C765(0.71); C232(0.91); C233(0.91); C319(0.93) | LDD0622 | [7] |
| LDCM0306 | AC44 | HCT 116 | C765(0.67); C232(0.83); C233(0.83); C94(0.86) | LDD0623 | [7] |
| LDCM0307 | AC45 | HCT 116 | C765(0.75); C687(0.78); C705(0.78); C319(0.80) | LDD0624 | [7] |
| LDCM0308 | AC46 | HCT 116 | C472(0.88); C319(0.93); C1192(0.95); C535(0.95) | LDD0625 | [7] |
| LDCM0309 | AC47 | HCT 116 | C687(0.54); C705(0.54); C535(0.76); C547(0.76) | LDD0626 | [7] |
| LDCM0310 | AC48 | HCT 116 | C687(0.56); C705(0.56); C472(0.83); C1192(0.87) | LDD0627 | [7] |
| LDCM0311 | AC49 | HCT 116 | C687(0.53); C705(0.53); C472(0.57); C535(0.77) | LDD0628 | [7] |
| LDCM0312 | AC5 | HCT 116 | C687(0.59); C705(0.59); C535(0.78); C547(0.78) | LDD0629 | [7] |
| LDCM0313 | AC50 | HCT 116 | C687(0.54); C705(0.54); C472(0.57); C535(0.68) | LDD0630 | [7] |
| LDCM0314 | AC51 | HCT 116 | C1192(0.88); C765(0.90); C472(0.95); C398(0.97) | LDD0631 | [7] |
| LDCM0315 | AC52 | HCT 116 | C687(0.61); C705(0.61); C535(0.74); C547(0.74) | LDD0632 | [7] |
| LDCM0316 | AC53 | HCT 116 | C687(0.51); C705(0.51); C535(0.67); C547(0.67) | LDD0633 | [7] |
| LDCM0317 | AC54 | HCT 116 | C687(0.54); C705(0.54); C472(0.66); C535(0.67) | LDD0634 | [7] |
| LDCM0318 | AC55 | HCT 116 | C687(0.53); C705(0.53); C472(0.61); C535(0.78) | LDD0635 | [7] |
| LDCM0319 | AC56 | HCT 116 | C687(0.63); C705(0.63); C472(0.71); C319(0.78) | LDD0636 | [7] |
| LDCM0320 | AC57 | HCT 116 | C535(0.45); C547(0.45); C472(0.55); C178(0.61) | LDD0637 | [7] |
| LDCM0321 | AC58 | HCT 116 | C535(0.49); C547(0.49); C472(0.53); C178(0.63) | LDD0638 | [7] |
| LDCM0322 | AC59 | HCT 116 | C535(0.43); C547(0.43); C472(0.51); C178(0.63) | LDD0639 | [7] |
| LDCM0323 | AC6 | HCT 116 | C535(0.71); C547(0.71); C232(0.73); C233(0.73) | LDD0640 | [7] |
| LDCM0324 | AC60 | HCT 116 | C535(0.58); C547(0.58); C472(0.64); C178(0.69) | LDD0641 | [7] |
| LDCM0325 | AC61 | HCT 116 | C535(0.61); C547(0.61); C174(0.65); C178(0.67) | LDD0642 | [7] |
| LDCM0326 | AC62 | HCT 116 | C535(0.45); C547(0.45); C472(0.55); C178(0.62) | LDD0643 | [7] |
| LDCM0327 | AC63 | HCT 116 | C535(0.57); C547(0.57); C472(0.59); C178(0.79) | LDD0644 | [7] |
| LDCM0328 | AC64 | HCT 116 | C535(0.41); C547(0.41); C472(0.52); C178(0.69) | LDD0645 | [7] |
| LDCM0329 | AC65 | HCT 116 | C535(0.44); C547(0.44); C178(0.59); C174(0.65) | LDD0646 | [7] |
| LDCM0330 | AC66 | HCT 116 | C535(0.54); C547(0.54); C178(0.59); C174(0.64) | LDD0647 | [7] |
| LDCM0331 | AC67 | HCT 116 | C472(0.53); C535(0.54); C547(0.54); C178(0.59) | LDD0648 | [7] |
| LDCM0332 | AC68 | HCT 116 | C1192(0.73); C413(0.89); C765(0.91); C535(0.92) | LDD0649 | [7] |
| LDCM0333 | AC69 | HCT 116 | C535(0.60); C547(0.60); C1192(0.72); C413(0.81) | LDD0650 | [7] |
| LDCM0334 | AC7 | HCT 116 | C232(0.68); C233(0.68); C535(0.80); C547(0.80) | LDD0651 | [7] |
| LDCM0335 | AC70 | HCT 116 | C535(0.53); C547(0.53); C1192(0.64); C232(0.74) | LDD0652 | [7] |
| LDCM0336 | AC71 | HCT 116 | C413(0.85); C1192(0.88); C178(0.94); C398(0.95) | LDD0653 | [7] |
| LDCM0337 | AC72 | HCT 116 | C1192(0.66); C535(0.71); C547(0.71); C232(0.89) | LDD0654 | [7] |
| LDCM0338 | AC73 | HCT 116 | C1192(0.83); C535(0.85); C547(0.85); C232(0.87) | LDD0655 | [7] |
| LDCM0339 | AC74 | HCT 116 | C1192(0.84); C413(0.90); C178(0.90); C232(0.92) | LDD0656 | [7] |
| LDCM0340 | AC75 | HCT 116 | C1192(0.86); C232(0.96); C233(0.96); C178(1.00) | LDD0657 | [7] |
| LDCM0341 | AC76 | HCT 116 | C535(0.69); C547(0.69); C1192(0.70); C232(0.80) | LDD0658 | [7] |
| LDCM0342 | AC77 | HCT 116 | C535(0.61); C547(0.61); C1192(0.67); C232(0.85) | LDD0659 | [7] |
| LDCM0343 | AC78 | HCT 116 | C232(0.93); C233(0.93); C1192(0.93); C535(0.98) | LDD0660 | [7] |
| LDCM0344 | AC79 | HCT 116 | C1192(0.73); C535(0.84); C547(0.84); C232(0.91) | LDD0661 | [7] |
| LDCM0345 | AC8 | HCT 116 | C472(0.62); C232(0.62); C233(0.62); C687(0.69) | LDD0662 | [7] |
| LDCM0346 | AC80 | HCT 116 | C1192(0.71); C535(0.72); C547(0.72); C178(0.82) | LDD0663 | [7] |
| LDCM0347 | AC81 | HCT 116 | C232(0.77); C233(0.77); C119(0.91); C765(0.94) | LDD0664 | [7] |
| LDCM0348 | AC82 | HCT 116 | C1192(0.91); C232(0.93); C233(0.93); C413(1.02) | LDD0665 | [7] |
| LDCM0349 | AC83 | HCT 116 | C535(0.40); C547(0.40); C472(0.46); C1192(0.62) | LDD0666 | [7] |
| LDCM0350 | AC84 | HCT 116 | C535(0.35); C547(0.35); C472(0.47); C1192(0.64) | LDD0667 | [7] |
| LDCM0351 | AC85 | HCT 116 | C535(0.59); C547(0.59); C809(0.71); C472(0.71) | LDD0668 | [7] |
| LDCM0352 | AC86 | HCT 116 | C535(0.48); C547(0.48); C1192(0.77); C809(0.79) | LDD0669 | [7] |
| LDCM0353 | AC87 | HCT 116 | C472(0.80); C1192(0.88); C398(0.98); C809(0.99) | LDD0670 | [7] |
| LDCM0354 | AC88 | HCT 116 | C535(0.72); C547(0.72); C472(0.75); C1192(0.82) | LDD0671 | [7] |
| LDCM0355 | AC89 | HCT 116 | C535(0.38); C547(0.38); C1192(0.61); C472(0.67) | LDD0672 | [7] |
| LDCM0357 | AC90 | HCT 116 | C765(0.92); C398(1.04); C726(1.04); C809(1.05) | LDD0674 | [7] |
| LDCM0358 | AC91 | HCT 116 | C535(0.31); C547(0.31); C472(0.39); C1192(0.61) | LDD0675 | [7] |
| LDCM0359 | AC92 | HCT 116 | C535(0.31); C547(0.31); C472(0.38); C1192(0.66) | LDD0676 | [7] |
| LDCM0360 | AC93 | HCT 116 | C535(0.63); C547(0.63); C1192(0.74); C809(0.82) | LDD0677 | [7] |
| LDCM0361 | AC94 | HCT 116 | C472(0.84); C809(0.85); C1192(0.87); C765(0.90) | LDD0678 | [7] |
| LDCM0362 | AC95 | HCT 116 | C472(0.64); C1192(0.84); C809(0.87); C535(0.88) | LDD0679 | [7] |
| LDCM0363 | AC96 | HCT 116 | C535(0.46); C547(0.46); C472(0.60); C1192(0.71) | LDD0680 | [7] |
| LDCM0364 | AC97 | HCT 116 | C535(0.27); C547(0.27); C472(0.49); C1192(0.55) | LDD0681 | [7] |
| LDCM0365 | AC98 | HCT 116 | C472(0.30); C178(0.75); C319(0.89); C809(0.90) | LDD0682 | [7] |
| LDCM0366 | AC99 | HCT 116 | C472(0.65); C232(0.84); C233(0.84); C765(0.86) | LDD0683 | [7] |
| LDCM0248 | AKOS034007472 | HCT 116 | C174(1.10); C726(1.02); C535(1.01); C547(1.01) | LDD0565 | [7] |
| LDCM0356 | AKOS034007680 | HCT 116 | C765(0.66); C232(0.74); C233(0.74); C535(0.78) | LDD0673 | [7] |
| LDCM0275 | AKOS034007705 | HCT 116 | C535(0.41); C547(0.41); C687(0.48); C705(0.48) | LDD0592 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | N.A. | LDD0405 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C174(1.03); C178(1.03); C398(1.01); C119(0.95) | LDD0078 | [7] |
| LDCM0102 | BDHI 8 | Jurkat | C726(6.81) | LDD0204 | [22] |
| LDCM0632 | CL-Sc | Hep-G2 | C591(20.00) | LDD2227 | [18] |
| LDCM0367 | CL1 | HCT 116 | C765(0.86); C319(0.89); C809(0.92); C413(0.92) | LDD0684 | [7] |
| LDCM0368 | CL10 | HCT 116 | C687(0.46); C705(0.46); C535(0.66); C547(0.66) | LDD0685 | [7] |
| LDCM0369 | CL100 | HCT 116 | C687(0.58); C705(0.58); C178(0.79); C809(0.88) | LDD0686 | [7] |
| LDCM0370 | CL101 | HCT 116 | C535(0.63); C547(0.63); C232(0.65); C233(0.65) | LDD0687 | [7] |
| LDCM0371 | CL102 | HCT 116 | C413(0.82); C232(0.86); C233(0.86); C535(0.87) | LDD0688 | [7] |
| LDCM0372 | CL103 | HCT 116 | C398(0.76); C472(0.78); C413(0.85); C765(0.85) | LDD0689 | [7] |
| LDCM0373 | CL104 | HCT 116 | C765(0.75); C398(0.77); C413(0.82); C535(0.89) | LDD0690 | [7] |
| LDCM0374 | CL105 | HCT 116 | C305(0.68); C535(0.72); C547(0.72); C94(0.90) | LDD0691 | [7] |
| LDCM0375 | CL106 | HCT 116 | C305(0.49); C535(0.54); C547(0.54); C178(0.83) | LDD0692 | [7] |
| LDCM0376 | CL107 | HCT 116 | C305(0.49); C535(0.49); C547(0.49); C178(0.73) | LDD0693 | [7] |
| LDCM0377 | CL108 | HCT 116 | C535(0.45); C547(0.45); C305(0.55); C178(0.72) | LDD0694 | [7] |
| LDCM0378 | CL109 | HCT 116 | C305(0.80); C319(0.86); C535(0.88); C547(0.88) | LDD0695 | [7] |
| LDCM0379 | CL11 | HCT 116 | C687(0.47); C705(0.47); C535(0.67); C547(0.67) | LDD0696 | [7] |
| LDCM0380 | CL110 | HCT 116 | C305(0.60); C535(0.79); C547(0.79); C319(0.92) | LDD0697 | [7] |
| LDCM0381 | CL111 | HCT 116 | C305(0.69); C809(0.90); C178(0.93); C319(0.94) | LDD0698 | [7] |
| LDCM0382 | CL112 | HCT 116 | C305(0.46); C1192(0.71); C535(0.80); C547(0.80) | LDD0699 | [7] |
| LDCM0383 | CL113 | HCT 116 | C305(0.42); C535(0.51); C547(0.51); C1192(0.52) | LDD0700 | [7] |
| LDCM0384 | CL114 | HCT 116 | C305(0.57); C687(0.63); C705(0.63); C319(0.74) | LDD0701 | [7] |
| LDCM0385 | CL115 | HCT 116 | C305(0.49); C687(0.56); C705(0.56); C1192(0.69) | LDD0702 | [7] |
| LDCM0386 | CL116 | HCT 116 | C305(0.38); C535(0.64); C547(0.64); C1192(0.69) | LDD0703 | [7] |
| LDCM0387 | CL117 | HCT 116 | C765(0.83); C232(0.89); C233(0.89); C319(0.90) | LDD0704 | [7] |
| LDCM0388 | CL118 | HCT 116 | C765(0.69); C178(0.87); C174(0.91); C535(0.92) | LDD0705 | [7] |
| LDCM0389 | CL119 | HCT 116 | C765(0.64); C319(0.71); C232(0.78); C233(0.78) | LDD0706 | [7] |
| LDCM0390 | CL12 | HCT 116 | C687(0.41); C705(0.41); C535(0.67); C547(0.67) | LDD0707 | [7] |
| LDCM0391 | CL120 | HCT 116 | C765(0.77); C94(0.89); C319(0.90); C178(0.96) | LDD0708 | [7] |
| LDCM0392 | CL121 | HCT 116 | C472(0.77); C1192(0.77); C726(0.83); C765(0.84) | LDD0709 | [7] |
| LDCM0393 | CL122 | HCT 116 | C687(0.76); C705(0.76); C1192(0.79); C535(0.81) | LDD0710 | [7] |
| LDCM0394 | CL123 | HCT 116 | C687(0.41); C705(0.41); C535(0.64); C547(0.64) | LDD0711 | [7] |
| LDCM0395 | CL124 | HCT 116 | C687(0.48); C705(0.48); C472(0.61); C535(0.71) | LDD0712 | [7] |
| LDCM0396 | CL125 | HCT 116 | C472(0.84); C178(0.85); C119(0.89); C232(0.95) | LDD0713 | [7] |
| LDCM0397 | CL126 | HCT 116 | C472(0.70); C535(0.79); C547(0.79); C1192(0.82) | LDD0714 | [7] |
| LDCM0398 | CL127 | HCT 116 | C472(0.74); C178(0.85); C1192(0.92); C119(0.92) | LDD0715 | [7] |
| LDCM0399 | CL128 | HCT 116 | C535(0.39); C547(0.39); C472(0.57); C1192(0.72) | LDD0716 | [7] |
| LDCM0400 | CL13 | HCT 116 | C687(0.62); C705(0.62); C535(0.74); C547(0.74) | LDD0717 | [7] |
| LDCM0401 | CL14 | HCT 116 | C687(0.82); C705(0.82); C1192(0.85); C319(0.90) | LDD0718 | [7] |
| LDCM0402 | CL15 | HCT 116 | C94(0.82); C687(0.83); C705(0.83); C413(0.93) | LDD0719 | [7] |
| LDCM0403 | CL16 | HCT 116 | C687(0.64); C705(0.64); C535(0.70); C547(0.70) | LDD0720 | [7] |
| LDCM0404 | CL17 | HCT 116 | C535(0.72); C547(0.72); C809(0.74); C1192(0.94) | LDD0721 | [7] |
| LDCM0405 | CL18 | HCT 116 | C687(0.46); C705(0.46); C535(0.48); C547(0.48) | LDD0722 | [7] |
| LDCM0406 | CL19 | HCT 116 | C687(0.62); C705(0.62); C535(0.62); C547(0.62) | LDD0723 | [7] |
| LDCM0407 | CL2 | HCT 116 | C765(0.93); C687(0.97); C705(0.97); C319(0.97) | LDD0724 | [7] |
| LDCM0408 | CL20 | HCT 116 | C535(0.39); C547(0.39); C687(0.58); C705(0.58) | LDD0725 | [7] |
| LDCM0409 | CL21 | HCT 116 | C535(0.47); C547(0.47); C687(0.62); C705(0.62) | LDD0726 | [7] |
| LDCM0410 | CL22 | HCT 116 | C687(0.43); C705(0.43); C535(0.64); C547(0.64) | LDD0727 | [7] |
| LDCM0411 | CL23 | HCT 116 | C535(0.51); C547(0.51); C687(0.61); C705(0.61) | LDD0728 | [7] |
| LDCM0412 | CL24 | HCT 116 | C687(0.33); C705(0.33); C535(0.45); C547(0.45) | LDD0729 | [7] |
| LDCM0413 | CL25 | HCT 116 | C174(2.89); C726(0.99); C535(0.46); C547(0.46) | LDD0730 | [7] |
| LDCM0414 | CL26 | HCT 116 | C174(1.20); C726(1.08); C535(0.53); C547(0.53) | LDD0731 | [7] |
| LDCM0415 | CL27 | HCT 116 | C174(1.21); C726(0.86); C535(0.58); C547(0.58) | LDD0732 | [7] |
| LDCM0416 | CL28 | HCT 116 | C174(1.27); C726(1.00); C535(0.55); C547(0.55) | LDD0733 | [7] |
| LDCM0417 | CL29 | HCT 116 | C174(1.15); C726(1.24); C535(0.48); C547(0.48) | LDD0734 | [7] |
| LDCM0418 | CL3 | HCT 116 | C174(1.17); C726(1.10); C535(1.13); C547(1.13) | LDD0735 | [7] |
| LDCM0419 | CL30 | HCT 116 | C174(1.01); C726(0.97); C535(0.76); C547(0.76) | LDD0736 | [7] |
| LDCM0420 | CL31 | HCT 116 | C174(1.34); C726(1.00); C119(1.05); C1192(0.90) | LDD0737 | [7] |
| LDCM0421 | CL32 | HCT 116 | C174(1.41); C726(1.28); C119(1.07); C1192(0.77) | LDD0738 | [7] |
| LDCM0422 | CL33 | HCT 116 | C174(1.94); C726(1.94); C119(2.89); C1192(0.85) | LDD0739 | [7] |
| LDCM0423 | CL34 | HCT 116 | C174(2.41); C726(1.19); C119(1.09); C1192(0.75) | LDD0740 | [7] |
| LDCM0424 | CL35 | HCT 116 | C174(2.35); C726(1.29); C119(1.08); C1192(0.60) | LDD0741 | [7] |
| LDCM0425 | CL36 | HCT 116 | C174(2.13); C726(1.81); C119(0.99); C1192(0.61) | LDD0742 | [7] |
| LDCM0426 | CL37 | HCT 116 | C174(2.50); C726(1.19); C119(1.03); C1192(0.73) | LDD0743 | [7] |
| LDCM0428 | CL39 | HCT 116 | C174(3.41); C726(1.03); C119(1.65); C1192(0.74) | LDD0745 | [7] |
| LDCM0429 | CL4 | HCT 116 | C174(1.21); C726(1.06); C535(0.94); C547(0.94) | LDD0746 | [7] |
| LDCM0430 | CL40 | HCT 116 | C174(1.90); C726(2.56); C119(1.18); C1192(0.65) | LDD0747 | [7] |
| LDCM0431 | CL41 | HCT 116 | C174(1.80); C726(2.05); C119(0.99); C1192(0.71) | LDD0748 | [7] |
| LDCM0432 | CL42 | HCT 116 | C174(6.17); C726(1.16); C119(1.19); C1192(0.80) | LDD0749 | [7] |
| LDCM0433 | CL43 | HCT 116 | C174(2.67); C726(1.20); C119(1.03); C1192(0.84) | LDD0750 | [7] |
| LDCM0434 | CL44 | HCT 116 | C174(2.42); C726(1.04); C119(1.19); C1192(0.71) | LDD0751 | [7] |
| LDCM0435 | CL45 | HCT 116 | C174(2.62); C726(1.98); C119(1.26); C1192(0.72) | LDD0752 | [7] |
| LDCM0436 | CL46 | HCT 116 | C174(1.30); C535(1.70); C547(1.70); C119(1.55) | LDD0753 | [7] |
| LDCM0437 | CL47 | HCT 116 | C174(1.21); C535(1.48); C547(1.48); C119(1.35) | LDD0754 | [7] |
| LDCM0438 | CL48 | HCT 116 | C174(1.29); C535(1.83); C547(1.83); C119(1.05) | LDD0755 | [7] |
| LDCM0439 | CL49 | HCT 116 | C174(1.24); C535(0.90); C547(0.90); C119(0.88) | LDD0756 | [7] |
| LDCM0440 | CL5 | HCT 116 | C174(1.06); C726(0.82); C535(0.84); C547(0.84) | LDD0757 | [7] |
| LDCM0441 | CL50 | HCT 116 | C174(1.11); C535(1.22); C547(1.22); C119(1.11) | LDD0758 | [7] |
| LDCM0442 | CL51 | HCT 116 | C174(1.36); C535(1.27); C547(1.27); C119(0.93) | LDD0759 | [7] |
| LDCM0443 | CL52 | HCT 116 | C174(1.26); C535(1.29); C547(1.29); C119(1.22) | LDD0760 | [7] |
| LDCM0444 | CL53 | HCT 116 | C174(1.79); C535(1.81); C547(1.81); C119(0.99) | LDD0761 | [7] |
| LDCM0445 | CL54 | HCT 116 | C174(1.27); C535(1.15); C547(1.15); C119(1.16) | LDD0762 | [7] |
| LDCM0446 | CL55 | HCT 116 | C174(1.06); C535(1.31); C547(1.31); C119(0.99) | LDD0763 | [7] |
| LDCM0447 | CL56 | HCT 116 | C174(1.13); C535(1.20); C547(1.20); C119(1.03) | LDD0764 | [7] |
| LDCM0448 | CL57 | HCT 116 | C174(1.35); C535(1.54); C547(1.54); C119(0.91) | LDD0765 | [7] |
| LDCM0449 | CL58 | HCT 116 | C174(1.32); C535(1.44); C547(1.44); C119(1.25) | LDD0766 | [7] |
| LDCM0450 | CL59 | HCT 116 | C174(1.23); C535(1.30); C547(1.30); C119(1.37) | LDD0767 | [7] |
| LDCM0451 | CL6 | HCT 116 | C174(1.31); C726(1.84); C535(0.88); C547(0.88) | LDD0768 | [7] |
| LDCM0452 | CL60 | HCT 116 | C174(1.35); C535(1.21); C547(1.21); C119(1.05) | LDD0769 | [7] |
| LDCM0453 | CL61 | HCT 116 | C174(0.96); C535(0.58); C547(0.58); C119(1.39) | LDD0770 | [7] |
| LDCM0454 | CL62 | HCT 116 | C174(0.93); C535(0.60); C547(0.60); C119(1.30) | LDD0771 | [7] |
| LDCM0455 | CL63 | HCT 116 | C174(1.05); C535(0.45); C547(0.45); C119(1.27) | LDD0772 | [7] |
| LDCM0456 | CL64 | HCT 116 | C174(0.98); C535(0.29); C547(0.29); C119(1.20) | LDD0773 | [7] |
| LDCM0457 | CL65 | HCT 116 | C174(0.92); C535(0.43); C547(0.43); C119(1.05) | LDD0774 | [7] |
| LDCM0458 | CL66 | HCT 116 | C174(0.83); C535(0.34); C547(0.34); C119(1.09) | LDD0775 | [7] |
| LDCM0459 | CL67 | HCT 116 | C174(0.91); C535(0.35); C547(0.35); C119(1.07) | LDD0776 | [7] |
| LDCM0460 | CL68 | HCT 116 | C174(0.92); C535(0.42); C547(0.42); C119(1.09) | LDD0777 | [7] |
| LDCM0461 | CL69 | HCT 116 | C174(1.06); C535(0.45); C547(0.45); C119(1.65) | LDD0778 | [7] |
| LDCM0462 | CL7 | HCT 116 | C174(1.15); C726(0.80); C535(0.63); C547(0.63) | LDD0779 | [7] |
| LDCM0463 | CL70 | HCT 116 | C174(0.90); C535(0.51); C547(0.51); C119(1.16) | LDD0780 | [7] |
| LDCM0464 | CL71 | HCT 116 | C174(1.08); C535(0.51); C547(0.51); C119(1.38) | LDD0781 | [7] |
| LDCM0465 | CL72 | HCT 116 | C174(1.25); C535(0.72); C547(0.72); C119(1.37) | LDD0782 | [7] |
| LDCM0466 | CL73 | HCT 116 | C174(0.94); C535(0.43); C547(0.43); C119(1.20) | LDD0783 | [7] |
| LDCM0467 | CL74 | HCT 116 | C174(1.00); C535(0.45); C547(0.45); C119(1.32) | LDD0784 | [7] |
| LDCM0469 | CL76 | HCT 116 | C174(0.99); C535(0.97); C547(0.97); C119(1.13) | LDD0786 | [7] |
| LDCM0470 | CL77 | HCT 116 | C174(1.13); C535(1.08); C547(1.08); C119(1.15) | LDD0787 | [7] |
| LDCM0471 | CL78 | HCT 116 | C174(1.09); C535(0.78); C547(0.78); C119(1.05) | LDD0788 | [7] |
| LDCM0472 | CL79 | HCT 116 | C174(0.95); C535(0.39); C547(0.39); C119(1.04) | LDD0789 | [7] |
| LDCM0473 | CL8 | HCT 116 | C174(2.58); C726(14.48); C535(0.64); C547(0.64) | LDD0790 | [7] |
| LDCM0474 | CL80 | HCT 116 | C174(0.97); C535(0.84); C547(0.84); C119(1.13) | LDD0791 | [7] |
| LDCM0475 | CL81 | HCT 116 | C174(1.01); C535(0.59); C547(0.59); C119(1.01) | LDD0792 | [7] |
| LDCM0476 | CL82 | HCT 116 | C174(0.82); C535(0.47); C547(0.47); C119(1.04) | LDD0793 | [7] |
| LDCM0477 | CL83 | HCT 116 | C174(1.49); C535(0.48); C547(0.48); C119(1.17) | LDD0794 | [7] |
| LDCM0478 | CL84 | HCT 116 | C174(1.00); C535(0.76); C547(0.76); C119(1.20) | LDD0795 | [7] |
| LDCM0479 | CL85 | HCT 116 | C174(1.01); C535(0.63); C547(0.63); C119(1.06) | LDD0796 | [7] |
| LDCM0480 | CL86 | HCT 116 | C174(1.01); C535(1.26); C547(1.26); C119(1.02) | LDD0797 | [7] |
| LDCM0481 | CL87 | HCT 116 | C174(1.07); C535(0.68); C547(0.68); C119(1.04) | LDD0798 | [7] |
| LDCM0482 | CL88 | HCT 116 | C174(0.98); C535(0.54); C547(0.54); C119(0.93) | LDD0799 | [7] |
| LDCM0483 | CL89 | HCT 116 | C174(0.98); C535(0.80); C547(0.80); C119(1.15) | LDD0800 | [7] |
| LDCM0484 | CL9 | HCT 116 | C174(1.50); C726(1.43); C535(0.67); C547(0.67) | LDD0801 | [7] |
| LDCM0485 | CL90 | HCT 116 | C174(1.35); C535(1.39); C547(1.39); C119(1.65) | LDD0802 | [7] |
| LDCM0486 | CL91 | HCT 116 | C174(1.16); C726(1.43); C535(0.96); C547(0.96) | LDD0803 | [7] |
| LDCM0487 | CL92 | HCT 116 | C174(1.25); C726(1.20); C535(1.07); C547(1.07) | LDD0804 | [7] |
| LDCM0488 | CL93 | HCT 116 | C174(1.21); C726(1.01); C535(1.02); C547(1.02) | LDD0805 | [7] |
| LDCM0489 | CL94 | HCT 116 | C174(1.80); C726(1.32); C535(0.81); C547(0.81) | LDD0806 | [7] |
| LDCM0490 | CL95 | HCT 116 | C174(1.62); C726(3.54); C535(0.95); C547(0.95) | LDD0807 | [7] |
| LDCM0491 | CL96 | HCT 116 | C174(1.32); C726(1.07); C535(1.03); C547(1.03) | LDD0808 | [7] |
| LDCM0492 | CL97 | HCT 116 | C174(1.40); C726(1.30); C535(1.00); C547(1.00) | LDD0809 | [7] |
| LDCM0493 | CL98 | HCT 116 | C174(1.34); C726(1.15); C535(0.84); C547(0.84) | LDD0810 | [7] |
| LDCM0494 | CL99 | HCT 116 | C174(1.11); C726(1.15); C535(0.92); C547(0.92) | LDD0811 | [7] |
| LDCM0198 | Dimethyl Fumarate(DMF) | T cell | C726(10.43); C398(7.26) | LDD0513 | [23] |
| LDCM0495 | E2913 | HEK-293T | C765(0.97); C94(0.97); C726(1.20); C413(0.92) | LDD1698 | [24] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C413(2.92); C119(1.76) | LDD1702 | [4] |
| LDCM0625 | F8 | Ramos | C94(1.54); C765(1.44); C591(0.63) | LDD2187 | [25] |
| LDCM0572 | Fragment10 | Ramos | C94(1.59) | LDD2189 | [25] |
| LDCM0573 | Fragment11 | Ramos | C94(3.95); C765(7.41); C591(18.25) | LDD2190 | [25] |
| LDCM0574 | Fragment12 | Ramos | C94(1.65); C398(1.07) | LDD2191 | [25] |
| LDCM0575 | Fragment13 | Ramos | C94(1.54); C726(1.19) | LDD2192 | [25] |
| LDCM0576 | Fragment14 | Ramos | C94(2.17); C765(0.59); C591(1.03) | LDD2193 | [25] |
| LDCM0579 | Fragment20 | Ramos | C94(1.40); C398(1.31) | LDD2194 | [25] |
| LDCM0580 | Fragment21 | Ramos | C94(1.80); C398(1.08); C726(0.74) | LDD2195 | [25] |
| LDCM0582 | Fragment23 | Ramos | C94(1.46) | LDD2196 | [25] |
| LDCM0578 | Fragment27 | Ramos | C94(1.22); C398(1.48); C726(0.84) | LDD2197 | [25] |
| LDCM0586 | Fragment28 | Ramos | C94(0.93); C765(0.39); C398(0.97); C591(0.83) | LDD2198 | [25] |
| LDCM0588 | Fragment30 | Ramos | C94(1.68); C398(1.78); C726(1.05) | LDD2199 | [25] |
| LDCM0589 | Fragment31 | Ramos | C94(2.28); C398(1.53); C726(3.13) | LDD2200 | [25] |
| LDCM0590 | Fragment32 | Ramos | C94(2.51) | LDD2201 | [25] |
| LDCM0468 | Fragment33 | HCT 116 | C174(1.03); C535(0.35); C547(0.35); C119(1.21) | LDD0785 | [7] |
| LDCM0596 | Fragment38 | Ramos | C94(1.36); C398(0.80); C726(1.87) | LDD2203 | [25] |
| LDCM0566 | Fragment4 | Ramos | C94(0.70); C765(0.68); C591(0.93); C726(1.13) | LDD2184 | [25] |
| LDCM0427 | Fragment51 | HCT 116 | C174(3.66); C726(7.52); C119(1.10); C1192(0.69) | LDD0744 | [7] |
| LDCM0610 | Fragment52 | Ramos | C94(1.98) | LDD2204 | [25] |
| LDCM0614 | Fragment56 | Ramos | C94(1.28); C726(1.84) | LDD2205 | [25] |
| LDCM0569 | Fragment7 | Ramos | C94(0.63); C765(0.60); C591(0.78) | LDD2186 | [25] |
| LDCM0571 | Fragment9 | Ramos | C94(1.47) | LDD2188 | [25] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [16] |
| LDCM0022 | KB02 | HCT 116 | C174(6.54); C119(6.32) | LDD0080 | [7] |
| LDCM0023 | KB03 | HCT 116 | C174(20.00); C119(2.46) | LDD0081 | [7] |
| LDCM0024 | KB05 | HCT 116 | C174(4.85); C119(2.65) | LDD0082 | [7] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [16] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C1192(1.01); C413(1.00) | LDD2093 | [4] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C413(0.89) | LDD2098 | [4] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C413(0.94); C119(1.33) | LDD2107 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C413(0.75) | LDD2109 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C765(1.01); C1192(0.99) | LDD2111 | [4] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C765(2.37); C413(1.20) | LDD2119 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C398(0.87) | LDD2123 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C413(0.75) | LDD2125 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C413(0.86); C398(1.06) | LDD2127 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C765(1.13); C1192(1.05) | LDD2129 | [4] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C765(0.97) | LDD2135 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C413(1.31) | LDD2136 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C413(0.85) | LDD2137 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C413(0.90); C119(1.39) | LDD2140 | [4] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C765(1.00); C413(1.25) | LDD2144 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C413(0.94) | LDD2146 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C413(0.51) | LDD2150 | [4] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C119(0.31) | LDD2207 | [26] |
| LDCM0131 | RA190 | MM1.R | C119(1.85) | LDD0304 | [27] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 2.60 | LDD0160 | [5] |
| LDCM0021 | THZ1 | HCT 116 | C174(1.05); C398(1.10); C119(1.00) | LDD2173 | [7] |
| LDCM0112 | W16 | Hep-G2 | C119(1.00) | LDD0239 | [8] |
The Interaction Atlas With This Target
References
