Details of the Target
General Information of Target
| Target ID | LDTP00400 | |||||
|---|---|---|---|---|---|---|
| Target Name | Torsin-1B (TOR1B) | |||||
| Gene Name | TOR1B | |||||
| Gene ID | 27348 | |||||
| Synonyms |
DQ1; Torsin-1B; Torsin ATPase-1B; EC 3.6.4.-; Torsin family 1 member B |
|||||
| 3D Structure | ||||||
| Sequence |
MLRAGWLRGAAALALLLAARVVAAFEPITVGLAIGAASAITGYLSYNDIYCRFAECCREE
RPLNASALKLDLEEKLFGQHLATEVIFKALTGFRNNKNPKKPLTLSLHGWAGTGKNFVSQ IVAENLHPKGLKSNFVHLFVSTLHFPHEQKIKLYQDQLQKWIRGNVSACANSVFIFDEMD KLHPGIIDAIKPFLDYYEQVDGVSYRKAIFIFLSNAGGDLITKTALDFWRAGRKREDIQL KDLEPVLSVGVFNNKHSGLWHSGLIDKNLIDYFIPFLPLEYRHVKMCVRAEMRARGSAID EDIVTRVAEEMTFFPRDEKIYSDKGCKTVQSRLDFH |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
ClpA/ClpB family, Torsin subfamily
|
|||||
| Subcellular location |
Endoplasmic reticulum lumen
|
|||||
| Function |
May serve as a molecular chaperone assisting in the proper folding of secreted and/or membrane proteins. Plays a role in non-neural cells nuclear envelope and endoplasmic reticulum integrity. May have a redundant function with TOR1A in non-neural tissues.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
3.29 | LDD0318 | [1] | |
|
BTD Probe Info |
 |
C57(0.57) | LDD2141 | [2] | |
|
IPM Probe Info |
 |
C57(2.14) | LDD1702 | [2] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [3] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [4] | |
|
DBIA Probe Info |
 |
C56(4.38) | LDD2236 | [5] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [6] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [6] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C056 Probe Info |
 |
18.00 | LDD1753 | [8] | |
|
C091 Probe Info |
 |
12.64 | LDD1782 | [8] | |
|
C092 Probe Info |
 |
23.92 | LDD1783 | [8] | |
|
C094 Probe Info |
 |
24.59 | LDD1785 | [8] | |
|
C112 Probe Info |
 |
21.26 | LDD1799 | [8] | |
|
C210 Probe Info |
 |
59.30 | LDD1884 | [8] | |
|
C218 Probe Info |
 |
14.93 | LDD1892 | [8] | |
|
C220 Probe Info |
 |
12.38 | LDD1894 | [8] | |
|
C228 Probe Info |
 |
17.15 | LDD1901 | [8] | |
|
C231 Probe Info |
 |
12.04 | LDD1904 | [8] | |
|
C235 Probe Info |
 |
17.75 | LDD1908 | [8] | |
|
C278 Probe Info |
 |
72.50 | LDD1948 | [8] | |
|
C282 Probe Info |
 |
18.13 | LDD1952 | [8] | |
|
C289 Probe Info |
 |
41.93 | LDD1959 | [8] | |
|
C293 Probe Info |
 |
17.75 | LDD1963 | [8] | |
|
C310 Probe Info |
 |
7.89 | LDD1977 | [8] | |
|
C362 Probe Info |
 |
42.81 | LDD2023 | [8] | |
|
C388 Probe Info |
 |
95.01 | LDD2047 | [8] | |
|
C390 Probe Info |
 |
23.10 | LDD2049 | [8] | |
|
C403 Probe Info |
 |
19.29 | LDD2061 | [8] | |
|
C407 Probe Info |
 |
11.16 | LDD2064 | [8] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [9] | |
|
Dasatinib-CA-8PAP Probe Info |
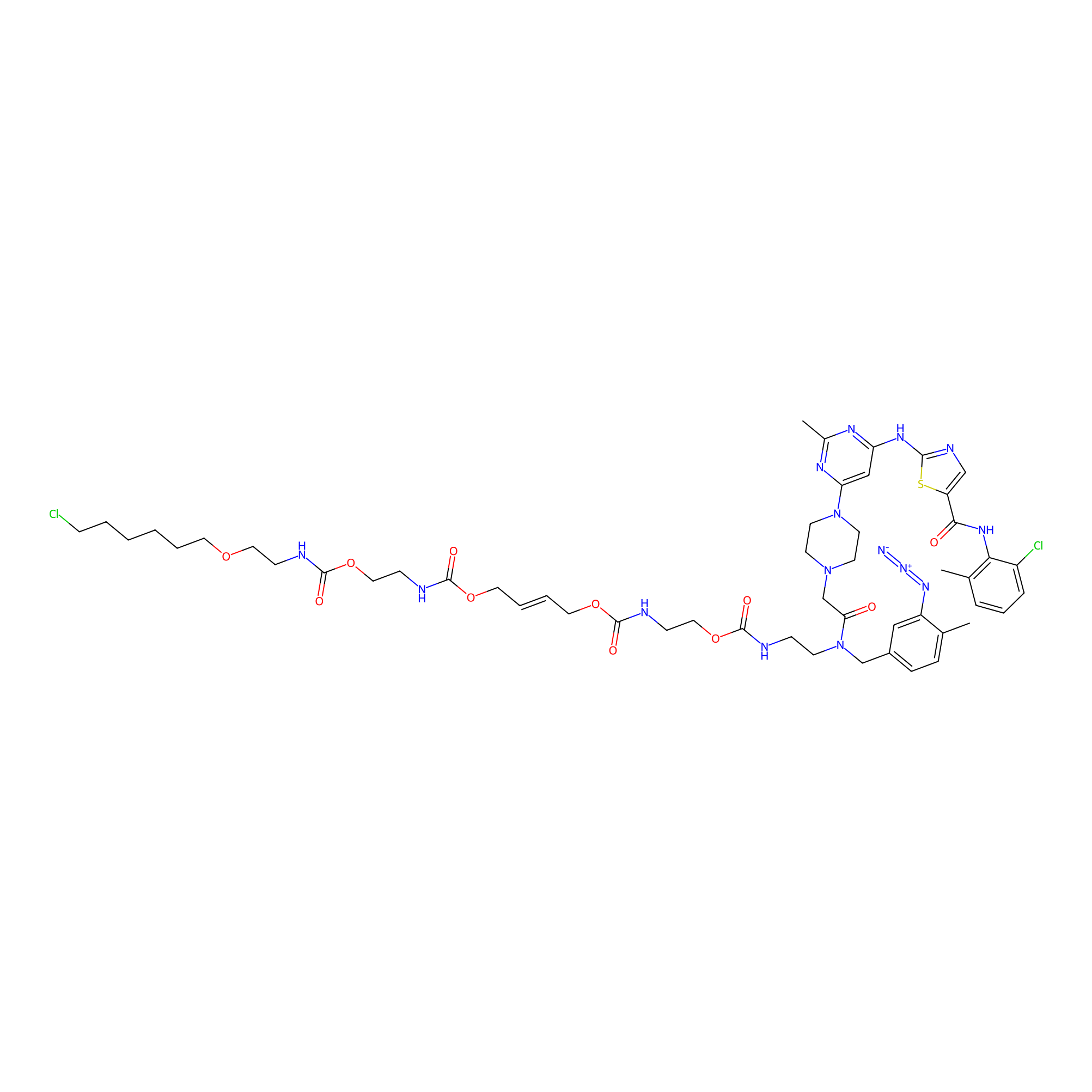 |
4.00 | LDD0364 | [10] | |
|
OEA-DA Probe Info |
 |
11.25 | LDD0046 | [11] | |
Competitor(s) Related to This Target
References
