Details of the Target
General Information of Target
| Target ID | LDTP10533 | |||||
|---|---|---|---|---|---|---|
| Target Name | Sorting nexin-27 (SNX27) | |||||
| Gene Name | SNX27 | |||||
| Gene ID | 81609 | |||||
| Synonyms |
KIAA0488; Sorting nexin-27 |
|||||
| 3D Structure | ||||||
| Sequence |
MHHGTGPQNVQHQLQRSRACPGSEGEEQPAHPNPPPSPAAPFAPSASPSAPQSPSYQIQQ
LMNRSPATGQNVNITLQSVGPVVGGNQQITLAPLPLPSPTSPGFQFSAQPRRFEHGSPSY IQVTSPLSQQVQTQSPTQPSPGPGQALQNVRAGAPGPGLGLCSSSPTGGFVDASVLVRQI SLSPSSGGHFVFQDGSGLTQIAQGAQVQLQHPGTPITVRERRPSQPHTQSGGTIHHLGPQ SPAAAGGAGLQPLASPSHITTANLPPQISSIIQGQLVQQQQVLQGPPLPRPLGFERTPGV LLPGAGGAAGFGMTSPPPPTSPSRTAVPPGLSSLPLTSVGNTGMKKVPKKLEEIPPASPE MAQMRKQCLDYHYQEMQALKEVFKEYLIELFFLQHFQGNMMDFLAFKKKHYAPLQAYLRQ NDLDIEEEEEEEEEEEEKSEVINDEVKVVTGKDGQTGTPVAIATQLPPKVSAAFSSQQQP FQQALAGSLVAGAGSTVETDLFKRQQAMPSTGMAEQSKRPRLEVGHQGVVFQHPGADAGV PLQQLMPTAQGGMPPTPQAAQLAGQRQSQQQYDPSTGPPVQNAASLHTPLPQLPGRLPPA GVPTAALSSALQFAQQPQVVEAQTQLQIPVKTQQPNVPIPAPPSSQLPIPPSQPAQLALH VPTPGKVQVQASQLSSLPQMVASTRLPVDPAPPCPRPLPTSSTSSLAPVSGSGPGPSPAR SSPVNRPSSATNKALSPVTSRTPGVVASAPTKPQSPAQNATSSQDSSQDTLTEQITLENQ VHQRIAELRKAGLWSQRRLPKLQEAPRPKSHWDYLLEEMQWMATDFAQERRWKVAAAKKL VRTVVRHHEEKQLREERGKKEEQSRLRRIAASTAREIECFWSNIEQVVEIKLRVELEEKR KKALNLQKVSRRGKELRPKGFDALQESSLDSGMSGRKRKASISLTDDEVDDEEETIEEEE ANEGVVDHQTELSNLAKEAELPLLDLMKLYEGAFLPSSQWPRPKPDGEDTSGEEDADDCP GDRESRKDLVLIDSLFIMDQFKAAERMNIGKPNAKDIADVTAVAEAILPKGSARVTTSVK FNAPSLLYGALRDYQKIGLDWLAKLYRKNLNGILADEAGLGKTVQIIAFFAHLACNEGNW GPHLVVVRSCNILKWELELKRWCPGLKILSYIGSHRELKAKRQEWAEPNSFHVCITSYTQ FFRGLTAFTRVRWKCLVIDEMQRVKGMTERHWEAVFTLQSQQRLLLIDSPLHNTFLELWT MVHFLVPGISRPYLSSPLRAPSEESQDYYHKVVIRLHRVTQPFILRRTKRDVEKQLTKKY EHVLKCRLSNRQKALYEDVILQPGTQEALKSGHFVNVLSILVRLQRICNHPGLVEPRHPG SSYVAGPLEYPSASLILKALERDFWKEADLSMFDLIGLENKITRHEAELLSKKKIPRKLM EEISTSAAPAARPAAAKLKASRLFQPVQYGQKPEGRTVAFPSTHPPRTAAPTTASAAPQG PLRGRPPIATFSANPEAKAAAAPFQTSQASASAPRHQPASASSTAASPAHPAKLRAQTTA QASTPGQPPPQPQAPSHAAGQSALPQRLVLPSQAQARLPSGEVVKIAQLASITGPQSRVA QPETPVTLQFQGSKFTLSHSQLRQLTAGQPLQLQGSVLQIVSAPGQPYLRAPGPVVMQTV SQAGAVHGALGSKPPAGGPSPAPLTPQVGVPGRVAVNALAVGEPGTASKPASPIGGPTQE EKTRLLKERLDQIYLVNERRCSQAPVYGRDLLRICALPSHGRVQWRGSLDGRRGKEAGPA HSYTSSSESPSELMLTLCRCGESLQDVIDRVAFVIPPVVAAPPSLRVPRPPPLYSHRMRI LRQGLREHAAPYFQQLRQTTAPRLLQFPELRLVQFDSGKLEALAILLQKLKSEGRRVLIL SQMILMLDILEMFLNFHYLTYVRIDENASSEQRQELMRSFNRDRRIFCAILSTHSRTTGI NLVEADTVVFYDNDLNPVMDAKAQEWCDRIGRCKDIHIYRLVSGNSIEEKLLKNGTKDLI REVAAQGNDYSMAFLTQRTIQELFEVYSPMDDAGFPVKAEEFVVLSQEPSVTETIAPKIA RPFIEALKSIEYLEEDAQKSAQEGVLGPHTDALSSDSENMPCDEEPSQLEELADFMEQLT PIEKYALNYLELFHTSIEQEKERNSEDAVMTAVRAWEFWNLKTLQEREARLRLEQEEAEL LTYTREDAYSMEYVYEDVDGQTEVMPLWTPPTPPQDDSDIYLDSVMCLMYEATPIPEAKL PPVYVRKERKRHKTDPSAAGRKKKQRHGEAVVPPRSLFDRATPGLLKIRREGKEQKKNIL LKQQVPFAKPLPTFAKPTAEPGQDNPEWLISEDWALLQAVKQLLELPLNLTIVSPAHTPN WDLVSDVVNSCSRIYRSSKQCRNRYENVIIPREEGKSKNNRPLRTSQIYAQDENATHTQL YTSHFDLMKMTAGKRSPPIKPLLGMNPFQKNPKHASVLAESGINYDKPLPPIQVASLRAE RIAKEKKALADQQKAQQPAVAQPPPPQPQPPPPPQQPPPPLPQPQAAGSQPPAGPPAVQP QPQPQPQTQPQPVQAPAKAQPAITTGGSAAVLAGTIKTSVTGTSMPTGAVSGNVIVNTIA GVPAATFQSINKRLASPVAPGALTTPGGSAPAQVVHTQPPPRAVGSPATATPDLVSMATT QGVRAVTSVTASAVVTTNLTPVQTPARSLVPQVSQATGVQLPGKTITPAHFQLLRQQQQQ QQQQQQQQQQQQQQQQQQQQQQQQTTTTSQVQVPQIQGQAQSPAQIKAVGKLTPEHLIKM QKQKLQMPPQPPPPQAQSAPPQPTAQVQVQTSQPPQQQSPQLTTVTAPRPGALLTGTTVA NLQVARLTRVPTSQLQAQGQMQTQAPQPAQVALAKPPVVSVPAAVVSSPGVTTLPMNVAG ISVAIGQPQKAAGQTVVAQPVHMQQLLKLKQQAVQQQKAIQPQAAQGPAAVQQKITAQQI TTPGAQQKVAYAAQPALKTQFLTTPISQAQKLAGAQQVQTQIQVAKLPQVVQQQTPVASI QQVASASQQASPQTVALTQATAAGQQVQMIPAVTATAQVVQQKLIQQQVVTTASAPLQTP GAPNPAQVPASSDSPSQQPKLQMRVPAVRLKTPTKPPCQ |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Sorting nexin family
|
|||||
| Subcellular location |
Early endosome membrane
|
|||||
| Function |
Involved in the retrograde transport from endosome to plasma membrane, a trafficking pathway that promotes the recycling of internalized transmembrane proteins. Following internalization, endocytosed transmembrane proteins are delivered to early endosomes and recycled to the plasma membrane instead of being degraded in lysosomes. SNX27 specifically binds and directs sorting of a subset of transmembrane proteins containing a PDZ-binding motif at the C-terminus: following interaction with target transmembrane proteins, associates with the retromer complex, preventing entry into the lysosomal pathway, and promotes retromer-tubule based plasma membrane recycling. SNX27 also binds with the WASH complex. Interacts with membranes containing phosphatidylinositol-3-phosphate (PtdIns(3P)). May participate in establishment of natural killer cell polarity. Recruits CYTIP to early endosomes.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| COLO800 | SNV: p.K97Q | DBIA Probe Info | |||
| HT | SNV: p.L78R | . | |||
| NOMO1 | SNV: p.V43G | DBIA Probe Info | |||
| WM2664 | Insertion: p.Y197VfsTer58 | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K222(10.00) | LDD0277 | [2] | |
|
Probe 1 Probe Info |
 |
Y242(49.28); Y269(8.15); Y400(51.63) | LDD3495 | [3] | |
|
BTD Probe Info |
 |
C519(2.42) | LDD2090 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C519(2.81) | LDD0169 | [5] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [6] | |
|
DBIA Probe Info |
 |
C30(1.16) | LDD0531 | [7] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [8] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C513(0.00); C456(0.00); C354(0.00); C30(0.00) | LDD0038 | [9] | |
|
IA-alkyne Probe Info |
 |
C513(0.00); C354(0.00); C30(0.00) | LDD0036 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
C513(0.00); C354(0.00); C456(0.00); C30(0.00) | LDD0037 | [9] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [10] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [11] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [12] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [12] | |
|
IPM Probe Info |
 |
C519(0.00); C30(0.00) | LDD0005 | [13] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [14] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [15] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [15] | |
|
AOyne Probe Info |
 |
13.20 | LDD0443 | [16] | |
|
NAIA_5 Probe Info |
 |
C513(0.00); C30(0.00) | LDD2223 | [17] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C017 Probe Info |
 |
5.54 | LDD1725 | [18] | |
|
C106 Probe Info |
 |
18.38 | LDD1793 | [18] | |
|
C108 Probe Info |
 |
33.59 | LDD1795 | [18] | |
|
C198 Probe Info |
 |
21.26 | LDD1874 | [18] | |
|
C201 Probe Info |
 |
26.35 | LDD1877 | [18] | |
|
C293 Probe Info |
 |
17.03 | LDD1963 | [18] | |
|
C314 Probe Info |
 |
15.78 | LDD1981 | [18] | |
|
C343 Probe Info |
 |
13.55 | LDD2005 | [18] | |
|
C350 Probe Info |
 |
34.30 | LDD2011 | [18] | |
|
C424 Probe Info |
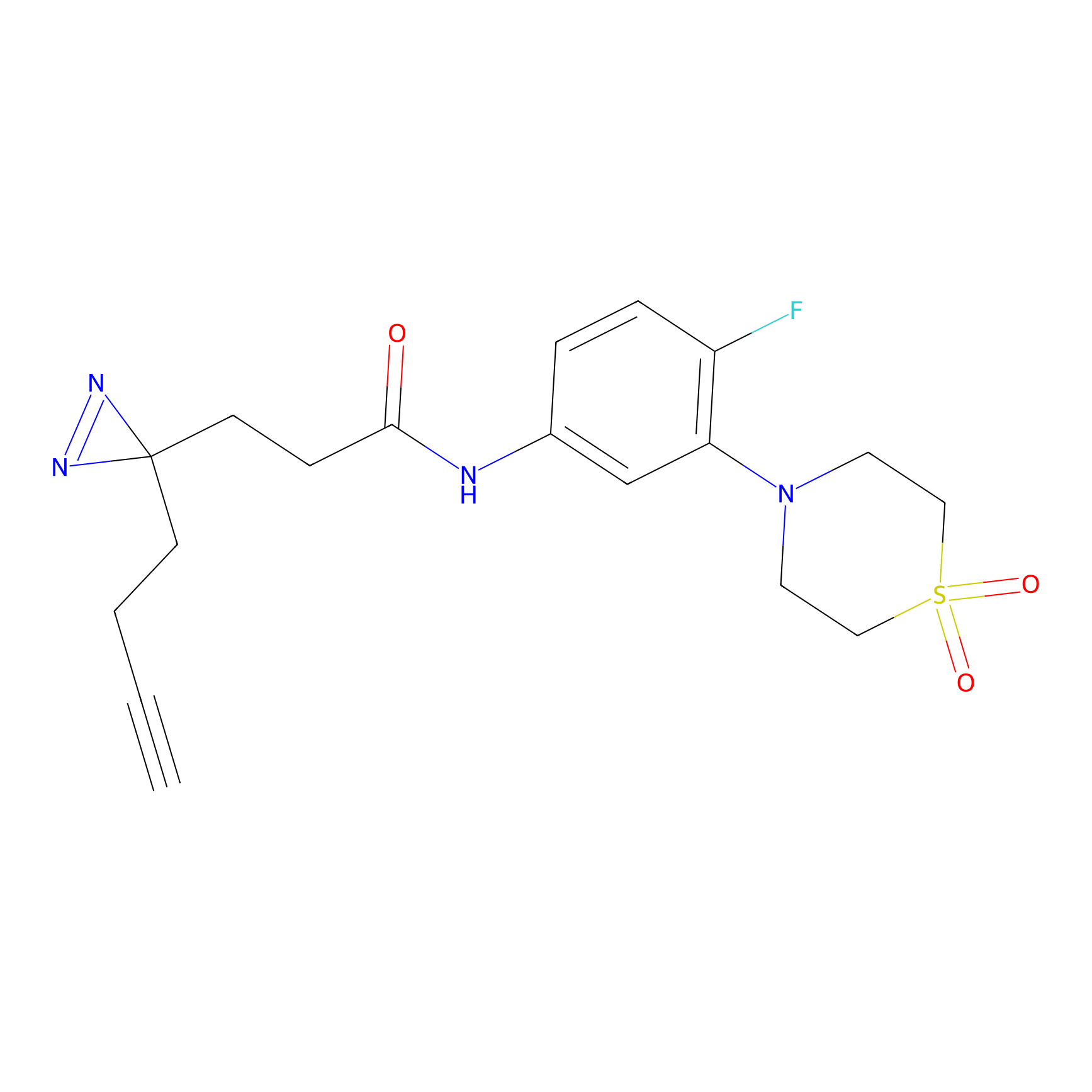 |
10.27 | LDD2079 | [18] | |
|
FFF probe11 Probe Info |
 |
7.99 | LDD0471 | [19] | |
|
FFF probe12 Probe Info |
 |
12.52 | LDD0473 | [19] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [19] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [19] | |
|
FFF probe2 Probe Info |
 |
10.55 | LDD0463 | [19] | |
|
OEA-DA Probe Info |
 |
5.79 | LDD0046 | [20] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C519(1.45) | LDD2117 | [4] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C519(2.81) | LDD0169 | [5] |
| LDCM0214 | AC1 | HCT 116 | C30(1.16) | LDD0531 | [7] |
| LDCM0215 | AC10 | HEK-293T | C354(1.33); C432(0.80); C487(1.42); C30(1.32) | LDD0813 | [7] |
| LDCM0216 | AC100 | HCT 116 | C30(1.25); C421(0.80); C432(0.85); C434(0.94) | LDD0533 | [7] |
| LDCM0217 | AC101 | HCT 116 | C30(0.90); C421(0.84); C432(0.77); C434(1.17) | LDD0534 | [7] |
| LDCM0218 | AC102 | HCT 116 | C30(0.46); C421(0.87); C432(0.87); C434(1.00) | LDD0535 | [7] |
| LDCM0219 | AC103 | HCT 116 | C30(1.12); C421(0.98); C432(0.94); C434(1.17) | LDD0536 | [7] |
| LDCM0220 | AC104 | HCT 116 | C30(0.78); C421(0.81); C432(0.79); C434(0.93) | LDD0537 | [7] |
| LDCM0221 | AC105 | HCT 116 | C30(1.30); C421(0.84); C432(0.78); C434(1.04) | LDD0538 | [7] |
| LDCM0222 | AC106 | HCT 116 | C30(1.58); C421(0.80); C432(0.71); C434(1.15) | LDD0539 | [7] |
| LDCM0223 | AC107 | HCT 116 | C30(1.15); C421(0.77); C432(0.79); C434(0.95) | LDD0540 | [7] |
| LDCM0224 | AC108 | HCT 116 | C30(0.86); C421(0.71); C432(0.68); C434(0.93) | LDD0541 | [7] |
| LDCM0225 | AC109 | HCT 116 | C30(0.69); C421(0.80); C432(0.74); C434(0.93) | LDD0542 | [7] |
| LDCM0226 | AC11 | HEK-293T | C354(1.42); C432(0.75); C487(1.79); C30(1.20) | LDD0824 | [7] |
| LDCM0227 | AC110 | HCT 116 | C30(0.56); C421(0.88); C432(0.91); C434(1.06) | LDD0544 | [7] |
| LDCM0228 | AC111 | HCT 116 | C30(0.90); C421(0.82); C432(0.71); C434(1.22) | LDD0545 | [7] |
| LDCM0229 | AC112 | HCT 116 | C30(0.80); C421(0.62); C432(0.74); C434(0.72) | LDD0546 | [7] |
| LDCM0230 | AC113 | HCT 116 | C30(1.28); C421(0.80); C432(0.87); C434(0.87) | LDD0547 | [7] |
| LDCM0231 | AC114 | HCT 116 | C30(2.12); C421(0.92); C432(0.83); C434(1.25) | LDD0548 | [7] |
| LDCM0232 | AC115 | HCT 116 | C30(1.58); C421(1.05); C432(0.83); C434(0.96) | LDD0549 | [7] |
| LDCM0233 | AC116 | HCT 116 | C30(1.64); C421(0.66); C432(0.70); C434(0.71) | LDD0550 | [7] |
| LDCM0234 | AC117 | HCT 116 | C30(1.44); C421(0.92); C432(0.78); C434(0.83) | LDD0551 | [7] |
| LDCM0235 | AC118 | HCT 116 | C30(1.73); C421(0.74); C432(0.80); C434(0.80) | LDD0552 | [7] |
| LDCM0236 | AC119 | HCT 116 | C30(1.44); C421(0.82); C432(0.84); C434(0.72) | LDD0553 | [7] |
| LDCM0237 | AC12 | HEK-293T | C354(1.56); C432(0.71); C487(2.70); C30(1.81) | LDD0835 | [7] |
| LDCM0238 | AC120 | HCT 116 | C30(0.71); C421(0.99); C432(1.16); C434(0.97) | LDD0555 | [7] |
| LDCM0239 | AC121 | HCT 116 | C30(1.17); C421(1.00); C432(1.07); C434(1.07) | LDD0556 | [7] |
| LDCM0240 | AC122 | HCT 116 | C30(1.56); C421(0.70); C432(0.74); C434(0.83) | LDD0557 | [7] |
| LDCM0241 | AC123 | HCT 116 | C30(1.50); C421(0.88); C432(0.88); C434(0.79) | LDD0558 | [7] |
| LDCM0242 | AC124 | HCT 116 | C30(0.97); C421(0.94); C432(1.03); C434(0.97) | LDD0559 | [7] |
| LDCM0243 | AC125 | HCT 116 | C30(1.41); C421(0.95); C432(0.94); C434(0.92) | LDD0560 | [7] |
| LDCM0244 | AC126 | HCT 116 | C30(1.35); C421(1.36); C432(1.09); C434(1.29) | LDD0561 | [7] |
| LDCM0245 | AC127 | HCT 116 | C30(2.49); C421(0.87); C432(0.93); C434(1.00) | LDD0562 | [7] |
| LDCM0246 | AC128 | HCT 116 | C432(2.28); C434(2.28) | LDD0563 | [7] |
| LDCM0247 | AC129 | HCT 116 | C432(2.56); C434(2.56) | LDD0564 | [7] |
| LDCM0249 | AC130 | HCT 116 | C432(1.40); C434(1.40) | LDD0566 | [7] |
| LDCM0250 | AC131 | HCT 116 | C432(1.63); C434(1.63) | LDD0567 | [7] |
| LDCM0251 | AC132 | HCT 116 | C432(0.92); C434(0.92) | LDD0568 | [7] |
| LDCM0252 | AC133 | HCT 116 | C432(1.52); C434(1.52) | LDD0569 | [7] |
| LDCM0253 | AC134 | HCT 116 | C432(1.76); C434(1.76) | LDD0570 | [7] |
| LDCM0254 | AC135 | HCT 116 | C432(1.58); C434(1.58) | LDD0571 | [7] |
| LDCM0255 | AC136 | HCT 116 | C432(1.36); C434(1.36) | LDD0572 | [7] |
| LDCM0256 | AC137 | HCT 116 | C432(1.53); C434(1.53) | LDD0573 | [7] |
| LDCM0257 | AC138 | HCT 116 | C432(1.72); C434(1.72) | LDD0574 | [7] |
| LDCM0258 | AC139 | HCT 116 | C432(1.29); C434(1.29) | LDD0575 | [7] |
| LDCM0259 | AC14 | HEK-293T | C354(1.26); C432(0.46); C487(1.69); C30(1.36) | LDD0857 | [7] |
| LDCM0260 | AC140 | HCT 116 | C432(1.42); C434(1.42) | LDD0577 | [7] |
| LDCM0261 | AC141 | HCT 116 | C432(1.14); C434(1.14) | LDD0578 | [7] |
| LDCM0262 | AC142 | HCT 116 | C432(1.34); C434(1.34) | LDD0579 | [7] |
| LDCM0263 | AC143 | HEK-293T | C354(0.99); C432(0.90); C487(1.05); C30(0.93) | LDD0861 | [7] |
| LDCM0264 | AC144 | HEK-293T | C354(1.50); C432(1.13); C487(1.16); C30(1.34) | LDD0862 | [7] |
| LDCM0265 | AC145 | HEK-293T | C354(1.40); C432(1.16); C487(1.14); C30(1.12) | LDD0863 | [7] |
| LDCM0266 | AC146 | HEK-293T | C354(1.29); C432(1.18); C487(1.06); C30(1.11) | LDD0864 | [7] |
| LDCM0267 | AC147 | HEK-293T | C354(1.33); C432(0.95); C487(1.37); C30(1.28) | LDD0865 | [7] |
| LDCM0268 | AC148 | HEK-293T | C354(0.92); C432(1.03); C487(1.07); C30(1.10) | LDD0866 | [7] |
| LDCM0269 | AC149 | HEK-293T | C354(0.97); C432(1.15); C487(1.11); C30(1.29) | LDD0867 | [7] |
| LDCM0270 | AC15 | HEK-293T | C354(1.56); C432(0.74); C487(2.74); C30(1.60) | LDD0868 | [7] |
| LDCM0271 | AC150 | HEK-293T | C354(1.05); C432(1.15); C487(0.92); C30(1.12) | LDD0869 | [7] |
| LDCM0272 | AC151 | HEK-293T | C354(1.26); C432(1.04); C487(1.09); C30(0.91) | LDD0870 | [7] |
| LDCM0273 | AC152 | HEK-293T | C354(1.14); C432(1.08); C487(1.01); C30(1.25) | LDD0871 | [7] |
| LDCM0274 | AC153 | HEK-293T | C354(1.07); C432(1.19); C487(1.08); C30(0.69) | LDD0872 | [7] |
| LDCM0621 | AC154 | HEK-293T | C354(0.98); C432(1.10); C487(0.91); C30(0.69) | LDD2162 | [7] |
| LDCM0622 | AC155 | HEK-293T | C354(1.42); C432(0.92); C487(1.14); C30(0.79) | LDD2163 | [7] |
| LDCM0623 | AC156 | HEK-293T | C354(1.24); C432(0.89); C487(1.43); C30(1.68) | LDD2164 | [7] |
| LDCM0624 | AC157 | HEK-293T | C354(1.67); C432(1.41); C487(1.06); C30(1.43) | LDD2165 | [7] |
| LDCM0276 | AC17 | HEK-293T | C354(1.63); C432(1.04); C421(1.04); C434(1.26) | LDD0874 | [7] |
| LDCM0277 | AC18 | HEK-293T | C354(1.57); C432(0.67); C421(0.67); C434(0.76) | LDD0875 | [7] |
| LDCM0278 | AC19 | HEK-293T | C354(1.51); C432(0.93); C421(0.93); C434(1.19) | LDD0876 | [7] |
| LDCM0279 | AC2 | HCT 116 | C30(1.04) | LDD0596 | [7] |
| LDCM0280 | AC20 | HEK-293T | C354(1.40); C432(0.57); C421(0.57); C434(0.68) | LDD0878 | [7] |
| LDCM0281 | AC21 | HEK-293T | C354(2.71); C432(0.71); C421(0.71); C434(0.95) | LDD0879 | [7] |
| LDCM0282 | AC22 | HEK-293T | C354(1.63); C432(0.58); C421(0.58); C434(0.70) | LDD0880 | [7] |
| LDCM0283 | AC23 | HEK-293T | C354(2.06); C432(0.62); C421(0.62); C434(0.75) | LDD0881 | [7] |
| LDCM0284 | AC24 | HEK-293T | C354(1.15); C432(0.96); C421(0.96); C434(1.31) | LDD0882 | [7] |
| LDCM0285 | AC25 | HCT 116 | C421(0.92); C432(0.92); C434(0.92) | LDD0602 | [7] |
| LDCM0286 | AC26 | HCT 116 | C421(0.46); C432(0.46); C434(0.46) | LDD0603 | [7] |
| LDCM0287 | AC27 | HCT 116 | C421(0.58); C432(0.58); C434(0.58) | LDD0604 | [7] |
| LDCM0288 | AC28 | HCT 116 | C421(0.44); C432(0.44); C434(0.44) | LDD0605 | [7] |
| LDCM0289 | AC29 | HCT 116 | C421(0.50); C432(0.50); C434(0.50) | LDD0606 | [7] |
| LDCM0290 | AC3 | HCT 116 | C30(1.65) | LDD0607 | [7] |
| LDCM0291 | AC30 | HCT 116 | C421(0.42); C432(0.42); C434(0.42) | LDD0608 | [7] |
| LDCM0292 | AC31 | HCT 116 | C421(0.59); C432(0.59); C434(0.59) | LDD0609 | [7] |
| LDCM0293 | AC32 | HCT 116 | C421(0.30); C432(0.30); C434(0.30) | LDD0610 | [7] |
| LDCM0294 | AC33 | HCT 116 | C421(0.91); C432(0.91); C434(0.91) | LDD0611 | [7] |
| LDCM0295 | AC34 | HCT 116 | C421(2.12); C432(2.12); C434(2.12) | LDD0612 | [7] |
| LDCM0296 | AC35 | HCT 116 | C421(1.48); C432(1.48) | LDD0613 | [7] |
| LDCM0297 | AC36 | HCT 116 | C421(0.78); C432(0.78) | LDD0614 | [7] |
| LDCM0298 | AC37 | HCT 116 | C421(0.72); C432(0.72) | LDD0615 | [7] |
| LDCM0299 | AC38 | HCT 116 | C421(1.28); C432(1.28) | LDD0616 | [7] |
| LDCM0300 | AC39 | HCT 116 | C421(1.26); C432(1.26) | LDD0617 | [7] |
| LDCM0301 | AC4 | HCT 116 | C30(1.17) | LDD0618 | [7] |
| LDCM0302 | AC40 | HCT 116 | C421(1.35); C432(1.35) | LDD0619 | [7] |
| LDCM0303 | AC41 | HCT 116 | C421(0.75); C432(0.75) | LDD0620 | [7] |
| LDCM0304 | AC42 | HCT 116 | C421(0.92); C432(0.92) | LDD0621 | [7] |
| LDCM0305 | AC43 | HCT 116 | C421(0.54); C432(0.54) | LDD0622 | [7] |
| LDCM0306 | AC44 | HCT 116 | C421(1.41); C432(1.41) | LDD0623 | [7] |
| LDCM0307 | AC45 | HCT 116 | C421(0.96); C432(0.96) | LDD0624 | [7] |
| LDCM0308 | AC46 | HCT 116 | C421(0.55); C432(0.55); C434(0.55); C30(0.91) | LDD0625 | [7] |
| LDCM0309 | AC47 | HCT 116 | C421(0.51); C432(0.51); C434(0.51); C30(0.89) | LDD0626 | [7] |
| LDCM0310 | AC48 | HCT 116 | C421(0.61); C432(0.61); C434(0.61); C30(0.91) | LDD0627 | [7] |
| LDCM0311 | AC49 | HCT 116 | C421(0.46); C432(0.46); C434(0.46); C30(1.11) | LDD0628 | [7] |
| LDCM0312 | AC5 | HCT 116 | C30(1.03) | LDD0629 | [7] |
| LDCM0313 | AC50 | HCT 116 | C421(0.44); C432(0.44); C434(0.44); C30(1.99) | LDD0630 | [7] |
| LDCM0314 | AC51 | HCT 116 | C421(0.56); C432(0.56); C434(0.56); C30(0.74) | LDD0631 | [7] |
| LDCM0315 | AC52 | HCT 116 | C421(0.60); C432(0.60); C434(0.60); C30(1.13) | LDD0632 | [7] |
| LDCM0316 | AC53 | HCT 116 | C421(0.79); C432(0.79); C434(0.79); C30(1.52) | LDD0633 | [7] |
| LDCM0317 | AC54 | HCT 116 | C421(0.74); C432(0.74); C434(0.74); C30(0.95) | LDD0634 | [7] |
| LDCM0318 | AC55 | HCT 116 | C421(0.67); C432(0.67); C434(0.67); C30(1.30) | LDD0635 | [7] |
| LDCM0319 | AC56 | HCT 116 | C421(0.53); C432(0.53); C434(0.53); C30(1.89) | LDD0636 | [7] |
| LDCM0320 | AC57 | HCT 116 | C421(1.32); C432(1.32); C434(1.32) | LDD0637 | [7] |
| LDCM0321 | AC58 | HCT 116 | C421(1.40); C432(1.40); C434(1.40) | LDD0638 | [7] |
| LDCM0322 | AC59 | HCT 116 | C421(1.37); C432(1.37); C434(1.37) | LDD0639 | [7] |
| LDCM0323 | AC6 | HEK-293T | C354(2.51); C432(0.96); C487(1.29); C30(1.04) | LDD0921 | [7] |
| LDCM0324 | AC60 | HCT 116 | C421(1.87); C432(1.87); C434(1.87) | LDD0641 | [7] |
| LDCM0325 | AC61 | HCT 116 | C421(0.86); C432(0.86); C434(0.86) | LDD0642 | [7] |
| LDCM0326 | AC62 | HCT 116 | C421(1.36); C432(1.36); C434(1.36) | LDD0643 | [7] |
| LDCM0327 | AC63 | HCT 116 | C421(1.72); C432(1.72); C434(1.72) | LDD0644 | [7] |
| LDCM0328 | AC64 | HCT 116 | C421(1.44); C432(1.44); C434(1.44) | LDD0645 | [7] |
| LDCM0329 | AC65 | HCT 116 | C421(0.60); C432(0.60); C434(0.60) | LDD0646 | [7] |
| LDCM0330 | AC66 | HCT 116 | C421(0.68); C432(0.68); C434(0.68) | LDD0647 | [7] |
| LDCM0331 | AC67 | HCT 116 | C421(1.83); C432(1.83); C434(1.83) | LDD0648 | [7] |
| LDCM0332 | AC68 | HCT 116 | C434(1.16); C421(1.17); C432(1.17); C30(1.21) | LDD0649 | [7] |
| LDCM0333 | AC69 | HCT 116 | C30(0.77); C434(1.04); C421(1.05); C432(1.05) | LDD0650 | [7] |
| LDCM0334 | AC7 | HEK-293T | C354(1.57); C432(1.11); C487(1.20); C30(0.97) | LDD0932 | [7] |
| LDCM0335 | AC70 | HCT 116 | C30(0.78); C434(0.88); C421(0.91); C432(0.91) | LDD0652 | [7] |
| LDCM0336 | AC71 | HCT 116 | C30(0.85); C421(1.37); C432(1.37); C434(1.43) | LDD0653 | [7] |
| LDCM0337 | AC72 | HCT 116 | C30(0.85); C434(1.01); C421(1.03); C432(1.03) | LDD0654 | [7] |
| LDCM0338 | AC73 | HCT 116 | C30(0.72); C434(0.76); C421(0.77); C432(0.77) | LDD0655 | [7] |
| LDCM0339 | AC74 | HCT 116 | C30(1.03); C421(1.14); C432(1.14); C434(1.17) | LDD0656 | [7] |
| LDCM0340 | AC75 | HCT 116 | C421(0.69); C432(0.69); C434(0.72); C30(1.20) | LDD0657 | [7] |
| LDCM0341 | AC76 | HCT 116 | C30(1.10); C421(1.23); C432(1.23); C434(1.34) | LDD0658 | [7] |
| LDCM0342 | AC77 | HCT 116 | C30(1.10); C434(1.26); C421(1.28); C432(1.28) | LDD0659 | [7] |
| LDCM0343 | AC78 | HCT 116 | C30(0.96); C421(1.02); C432(1.02); C434(1.04) | LDD0660 | [7] |
| LDCM0344 | AC79 | HCT 116 | C30(0.78); C421(1.02); C432(1.02); C434(1.06) | LDD0661 | [7] |
| LDCM0345 | AC8 | HEK-293T | C354(2.95); C432(1.27); C487(1.09); C30(1.19) | LDD0943 | [7] |
| LDCM0346 | AC80 | HCT 116 | C421(0.97); C432(0.97); C434(1.00); C30(1.13) | LDD0663 | [7] |
| LDCM0347 | AC81 | HCT 116 | C30(0.86); C421(1.01); C432(1.01); C434(1.04) | LDD0664 | [7] |
| LDCM0348 | AC82 | HCT 116 | C30(0.81); C421(1.15); C432(1.15); C434(1.28) | LDD0665 | [7] |
| LDCM0349 | AC83 | HCT 116 | C30(1.58) | LDD0666 | [7] |
| LDCM0350 | AC84 | HCT 116 | C30(1.18) | LDD0667 | [7] |
| LDCM0351 | AC85 | HCT 116 | C30(0.96) | LDD0668 | [7] |
| LDCM0352 | AC86 | HCT 116 | C30(1.00) | LDD0669 | [7] |
| LDCM0353 | AC87 | HCT 116 | C30(0.94) | LDD0670 | [7] |
| LDCM0354 | AC88 | HCT 116 | C30(0.80) | LDD0671 | [7] |
| LDCM0355 | AC89 | HCT 116 | C30(1.67) | LDD0672 | [7] |
| LDCM0357 | AC90 | HCT 116 | C30(0.53) | LDD0674 | [7] |
| LDCM0358 | AC91 | HCT 116 | C30(1.92) | LDD0675 | [7] |
| LDCM0359 | AC92 | HCT 116 | C30(1.55) | LDD0676 | [7] |
| LDCM0360 | AC93 | HCT 116 | C30(0.67) | LDD0677 | [7] |
| LDCM0361 | AC94 | HCT 116 | C30(0.78) | LDD0678 | [7] |
| LDCM0362 | AC95 | HCT 116 | C30(1.12) | LDD0679 | [7] |
| LDCM0363 | AC96 | HCT 116 | C30(1.10) | LDD0680 | [7] |
| LDCM0364 | AC97 | HCT 116 | C30(1.20) | LDD0681 | [7] |
| LDCM0365 | AC98 | HCT 116 | C432(0.83); C421(0.85); C434(1.08); C30(1.36) | LDD0682 | [7] |
| LDCM0366 | AC99 | HCT 116 | C30(0.60); C421(0.89); C432(0.91); C434(1.09) | LDD0683 | [7] |
| LDCM0248 | AKOS034007472 | HEK-293T | C354(1.71); C432(0.78); C487(1.16); C30(1.31) | LDD0846 | [7] |
| LDCM0356 | AKOS034007680 | HEK-293T | C354(1.48); C432(0.62); C487(1.55); C30(1.43) | LDD0954 | [7] |
| LDCM0275 | AKOS034007705 | HEK-293T | C354(1.79); C432(0.48); C487(1.30); C30(1.30) | LDD0873 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | 11.62 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [15] |
| LDCM0632 | CL-Sc | Hep-G2 | C30(0.55) | LDD2227 | [17] |
| LDCM0367 | CL1 | HEK-293T | C432(0.91); C487(0.96); C30(3.28); C421(1.02) | LDD0965 | [7] |
| LDCM0368 | CL10 | HEK-293T | C432(1.19); C487(1.30); C30(1.56); C421(0.63) | LDD0966 | [7] |
| LDCM0369 | CL100 | HCT 116 | C30(1.14) | LDD0686 | [7] |
| LDCM0370 | CL101 | HEK-293T | C354(1.70); C432(0.86); C487(1.46); C30(1.42) | LDD0968 | [7] |
| LDCM0371 | CL102 | HEK-293T | C354(1.77); C432(0.71); C487(1.60); C30(1.65) | LDD0969 | [7] |
| LDCM0372 | CL103 | HEK-293T | C354(2.03); C432(0.98); C487(1.39); C30(1.65) | LDD0970 | [7] |
| LDCM0373 | CL104 | HEK-293T | C354(2.02); C432(1.80); C487(1.58); C30(1.11) | LDD0971 | [7] |
| LDCM0374 | CL105 | HEK-293T | C354(1.94); C432(0.58); C421(0.58); C434(0.69) | LDD0972 | [7] |
| LDCM0375 | CL106 | HEK-293T | C354(1.01); C432(0.71); C421(0.71); C434(0.99) | LDD0973 | [7] |
| LDCM0376 | CL107 | HEK-293T | C354(1.60); C432(0.59); C421(0.59); C434(0.83) | LDD0974 | [7] |
| LDCM0377 | CL108 | HEK-293T | C354(1.14); C432(1.15); C421(1.15); C434(1.57) | LDD0975 | [7] |
| LDCM0378 | CL109 | HEK-293T | C354(1.34); C432(0.59); C421(0.59); C434(0.73) | LDD0976 | [7] |
| LDCM0379 | CL11 | HEK-293T | C432(1.10); C487(1.51); C30(2.15); C421(0.68) | LDD0977 | [7] |
| LDCM0380 | CL110 | HEK-293T | C354(1.87); C432(0.62); C421(0.62); C434(0.76) | LDD0978 | [7] |
| LDCM0381 | CL111 | HEK-293T | C354(1.50); C432(0.53); C421(0.53); C434(0.64) | LDD0979 | [7] |
| LDCM0382 | CL112 | HCT 116 | C421(0.85); C432(0.85); C434(0.85) | LDD0699 | [7] |
| LDCM0383 | CL113 | HCT 116 | C421(0.65); C432(0.65); C434(0.65) | LDD0700 | [7] |
| LDCM0384 | CL114 | HCT 116 | C421(1.43); C432(1.43); C434(1.43) | LDD0701 | [7] |
| LDCM0385 | CL115 | HCT 116 | C421(1.68); C432(1.68); C434(1.68) | LDD0702 | [7] |
| LDCM0386 | CL116 | HCT 116 | C421(1.05); C432(1.05); C434(1.05) | LDD0703 | [7] |
| LDCM0387 | CL117 | HCT 116 | C421(1.77); C432(1.77) | LDD0704 | [7] |
| LDCM0388 | CL118 | HCT 116 | C421(0.69); C432(0.69) | LDD0705 | [7] |
| LDCM0389 | CL119 | HCT 116 | C421(0.68); C432(0.68) | LDD0706 | [7] |
| LDCM0390 | CL12 | HEK-293T | C432(1.09); C487(1.12); C30(1.09); C421(0.68) | LDD0988 | [7] |
| LDCM0391 | CL120 | HCT 116 | C421(0.73); C432(0.73) | LDD0708 | [7] |
| LDCM0392 | CL121 | HCT 116 | C30(0.73); C421(0.94); C432(0.94); C434(0.94) | LDD0709 | [7] |
| LDCM0393 | CL122 | HCT 116 | C421(0.80); C432(0.80); C434(0.80); C30(1.16) | LDD0710 | [7] |
| LDCM0394 | CL123 | HCT 116 | C421(0.72); C432(0.72); C434(0.72); C30(1.37) | LDD0711 | [7] |
| LDCM0395 | CL124 | HCT 116 | C421(0.54); C432(0.54); C434(0.54); C30(1.58) | LDD0712 | [7] |
| LDCM0396 | CL125 | HCT 116 | C421(0.72); C432(0.72); C434(0.72) | LDD0713 | [7] |
| LDCM0397 | CL126 | HCT 116 | C421(0.73); C432(0.73); C434(0.73) | LDD0714 | [7] |
| LDCM0398 | CL127 | HCT 116 | C421(1.07); C432(1.07); C434(1.07) | LDD0715 | [7] |
| LDCM0399 | CL128 | HCT 116 | C421(1.19); C432(1.19); C434(1.19) | LDD0716 | [7] |
| LDCM0400 | CL13 | HEK-293T | C432(1.29); C487(0.92); C30(1.34); C421(1.20) | LDD0998 | [7] |
| LDCM0401 | CL14 | HEK-293T | C432(1.00); C487(0.86); C30(1.98); C421(1.30) | LDD0999 | [7] |
| LDCM0402 | CL15 | HEK-293T | C432(1.10); C487(1.48); C30(1.41); C421(1.03) | LDD1000 | [7] |
| LDCM0403 | CL16 | HEK-293T | C432(1.22); C421(1.22); C434(1.22) | LDD1001 | [7] |
| LDCM0404 | CL17 | HCT 116 | C421(0.67); C432(0.67); C434(0.67); C30(1.56) | LDD0721 | [7] |
| LDCM0405 | CL18 | HCT 116 | C421(0.86); C432(0.86); C434(0.86); C30(1.27) | LDD0722 | [7] |
| LDCM0406 | CL19 | HCT 116 | C421(0.84); C432(0.84); C434(0.84); C30(1.18) | LDD0723 | [7] |
| LDCM0407 | CL2 | HEK-293T | C432(1.03); C487(1.24); C30(1.49); C421(0.93) | LDD1005 | [7] |
| LDCM0408 | CL20 | HCT 116 | C421(0.65); C432(0.65); C434(0.65); C30(1.28) | LDD0725 | [7] |
| LDCM0409 | CL21 | HCT 116 | C421(0.85); C432(0.85); C434(0.85); C30(1.54) | LDD0726 | [7] |
| LDCM0410 | CL22 | HCT 116 | C30(0.91); C421(1.04); C432(1.04); C434(1.04) | LDD0727 | [7] |
| LDCM0411 | CL23 | HCT 116 | C421(0.67); C432(0.67); C434(0.67); C30(1.09) | LDD0728 | [7] |
| LDCM0412 | CL24 | HCT 116 | C30(1.31); C421(1.02); C432(1.02); C434(1.02) | LDD0729 | [7] |
| LDCM0413 | CL25 | HCT 116 | C30(1.09); C421(0.87); C432(0.87); C434(0.87) | LDD0730 | [7] |
| LDCM0414 | CL26 | HCT 116 | C30(1.57); C421(1.00); C432(1.00); C434(1.00) | LDD0731 | [7] |
| LDCM0415 | CL27 | HCT 116 | C30(1.18); C421(1.06); C432(1.06); C434(1.06) | LDD0732 | [7] |
| LDCM0416 | CL28 | HCT 116 | C30(1.55); C421(0.69); C432(0.69); C434(0.69) | LDD0733 | [7] |
| LDCM0417 | CL29 | HCT 116 | C30(1.37); C421(1.05); C432(1.05); C434(1.05) | LDD0734 | [7] |
| LDCM0418 | CL3 | HEK-293T | C432(1.01); C487(1.01); C30(1.03); C421(0.95) | LDD1016 | [7] |
| LDCM0419 | CL30 | HCT 116 | C30(0.80); C421(0.95); C432(0.95); C434(0.95) | LDD0736 | [7] |
| LDCM0420 | CL31 | HCT 116 | C30(1.05); C421(0.81); C432(0.81); C434(0.81) | LDD0737 | [7] |
| LDCM0421 | CL32 | HCT 116 | C30(0.75) | LDD0738 | [7] |
| LDCM0422 | CL33 | HCT 116 | C30(0.95) | LDD0739 | [7] |
| LDCM0423 | CL34 | HCT 116 | C30(0.89) | LDD0740 | [7] |
| LDCM0424 | CL35 | HCT 116 | C30(1.48) | LDD0741 | [7] |
| LDCM0425 | CL36 | HCT 116 | C30(1.44) | LDD0742 | [7] |
| LDCM0426 | CL37 | HCT 116 | C30(1.83) | LDD0743 | [7] |
| LDCM0428 | CL39 | HCT 116 | C30(0.93) | LDD0745 | [7] |
| LDCM0429 | CL4 | HEK-293T | C432(1.36); C487(1.25); C30(0.93); C421(1.16) | LDD1027 | [7] |
| LDCM0430 | CL40 | HCT 116 | C30(1.18) | LDD0747 | [7] |
| LDCM0431 | CL41 | HCT 116 | C30(0.88) | LDD0748 | [7] |
| LDCM0432 | CL42 | HCT 116 | C30(1.45) | LDD0749 | [7] |
| LDCM0433 | CL43 | HCT 116 | C30(0.69) | LDD0750 | [7] |
| LDCM0434 | CL44 | HCT 116 | C30(0.79) | LDD0751 | [7] |
| LDCM0435 | CL45 | HCT 116 | C30(0.71) | LDD0752 | [7] |
| LDCM0436 | CL46 | HCT 116 | C421(3.08); C432(3.08); C434(3.08) | LDD0753 | [7] |
| LDCM0437 | CL47 | HCT 116 | C421(0.83); C432(0.83); C434(0.83) | LDD0754 | [7] |
| LDCM0438 | CL48 | HCT 116 | C421(1.37); C432(1.37); C434(1.37) | LDD0755 | [7] |
| LDCM0439 | CL49 | HCT 116 | C421(0.90); C432(0.90); C434(0.90) | LDD0756 | [7] |
| LDCM0440 | CL5 | HEK-293T | C432(0.96); C487(1.32); C30(0.91); C421(0.86) | LDD1038 | [7] |
| LDCM0441 | CL50 | HCT 116 | C421(1.06); C432(1.06); C434(1.06) | LDD0758 | [7] |
| LDCM0442 | CL51 | HCT 116 | C421(1.15); C432(1.15); C434(1.15) | LDD0759 | [7] |
| LDCM0443 | CL52 | HCT 116 | C421(1.37); C432(1.37); C434(1.37) | LDD0760 | [7] |
| LDCM0444 | CL53 | HCT 116 | C421(1.25); C432(1.25); C434(1.25) | LDD0761 | [7] |
| LDCM0445 | CL54 | HCT 116 | C421(1.54); C432(1.54); C434(1.54) | LDD0762 | [7] |
| LDCM0446 | CL55 | HCT 116 | C421(2.24); C432(2.24); C434(2.24) | LDD0763 | [7] |
| LDCM0447 | CL56 | HCT 116 | C421(1.50); C432(1.50); C434(1.50) | LDD0764 | [7] |
| LDCM0448 | CL57 | HCT 116 | C421(1.79); C432(1.79); C434(1.79) | LDD0765 | [7] |
| LDCM0449 | CL58 | HCT 116 | C421(1.03); C432(1.03); C434(1.03) | LDD0766 | [7] |
| LDCM0450 | CL59 | HCT 116 | C421(1.48); C432(1.48); C434(1.48) | LDD0767 | [7] |
| LDCM0451 | CL6 | HEK-293T | C432(1.05); C487(0.97); C30(1.15); C421(1.17) | LDD1049 | [7] |
| LDCM0452 | CL60 | HCT 116 | C421(1.49); C432(1.49); C434(1.49) | LDD0769 | [7] |
| LDCM0453 | CL61 | HEK-293T | C354(1.25); C432(1.17); C487(0.81); C421(1.09) | LDD1051 | [7] |
| LDCM0454 | CL62 | HEK-293T | C354(1.26); C432(0.79); C487(1.16); C421(0.88) | LDD1052 | [7] |
| LDCM0455 | CL63 | HEK-293T | C354(1.57); C432(1.24); C487(1.10); C421(0.91) | LDD1053 | [7] |
| LDCM0456 | CL64 | HEK-293T | C354(0.96); C432(1.06); C487(0.91); C421(1.09) | LDD1054 | [7] |
| LDCM0457 | CL65 | HEK-293T | C354(1.41); C432(0.65); C487(1.22); C421(0.92) | LDD1055 | [7] |
| LDCM0458 | CL66 | HEK-293T | C354(1.17); C432(0.97); C487(0.94); C421(1.29) | LDD1056 | [7] |
| LDCM0459 | CL67 | HEK-293T | C354(0.86); C432(1.29); C487(1.09); C421(1.09) | LDD1057 | [7] |
| LDCM0460 | CL68 | HEK-293T | C354(1.07); C432(0.59); C487(1.27); C421(0.85) | LDD1058 | [7] |
| LDCM0461 | CL69 | HEK-293T | C354(1.33); C432(0.87); C487(0.78); C421(0.90) | LDD1059 | [7] |
| LDCM0462 | CL7 | HEK-293T | C432(1.20); C487(1.05); C30(1.41); C421(1.16) | LDD1060 | [7] |
| LDCM0463 | CL70 | HEK-293T | C354(0.84); C432(0.86); C487(1.44); C421(1.07) | LDD1061 | [7] |
| LDCM0464 | CL71 | HEK-293T | C354(1.31); C432(0.98); C487(1.20); C421(0.93) | LDD1062 | [7] |
| LDCM0465 | CL72 | HEK-293T | C354(1.21); C432(0.86); C487(1.01); C421(0.85) | LDD1063 | [7] |
| LDCM0466 | CL73 | HEK-293T | C354(0.87); C432(0.86); C487(1.29); C421(1.10) | LDD1064 | [7] |
| LDCM0467 | CL74 | HEK-293T | C354(1.39); C432(1.39); C487(1.32); C421(1.05) | LDD1065 | [7] |
| LDCM0469 | CL76 | HEK-293T | C354(1.22); C432(0.84); C421(0.81); C434(0.84) | LDD1067 | [7] |
| LDCM0470 | CL77 | HEK-293T | C354(1.20); C432(0.75); C421(0.86); C434(0.75) | LDD1068 | [7] |
| LDCM0471 | CL78 | HEK-293T | C354(1.79); C432(0.86); C421(1.16); C434(0.86) | LDD1069 | [7] |
| LDCM0472 | CL79 | HEK-293T | C354(1.23); C432(0.86); C421(0.83); C434(0.86) | LDD1070 | [7] |
| LDCM0473 | CL8 | HEK-293T | C432(1.15); C30(0.86); C421(0.85); C434(1.22) | LDD1071 | [7] |
| LDCM0474 | CL80 | HEK-293T | C354(1.20); C432(0.87); C421(0.92); C434(0.87) | LDD1072 | [7] |
| LDCM0475 | CL81 | HEK-293T | C354(1.28); C432(0.85); C421(0.87); C434(0.85) | LDD1073 | [7] |
| LDCM0476 | CL82 | HEK-293T | C354(0.86); C432(0.82); C421(0.83); C434(0.82) | LDD1074 | [7] |
| LDCM0477 | CL83 | HEK-293T | C354(0.92); C432(0.87); C421(0.92); C434(0.87) | LDD1075 | [7] |
| LDCM0478 | CL84 | HEK-293T | C354(1.59); C432(0.88); C421(1.33); C434(0.88) | LDD1076 | [7] |
| LDCM0479 | CL85 | HEK-293T | C354(1.84); C432(0.76); C421(0.79); C434(0.76) | LDD1077 | [7] |
| LDCM0480 | CL86 | HEK-293T | C354(1.47); C432(0.74); C421(0.95); C434(0.74) | LDD1078 | [7] |
| LDCM0481 | CL87 | HEK-293T | C354(1.42); C432(0.63); C421(0.75); C434(0.63) | LDD1079 | [7] |
| LDCM0482 | CL88 | HEK-293T | C354(1.37); C432(0.74); C421(0.92); C434(0.74) | LDD1080 | [7] |
| LDCM0483 | CL89 | HEK-293T | C354(1.40); C432(0.76); C421(0.86); C434(0.76) | LDD1081 | [7] |
| LDCM0484 | CL9 | HEK-293T | C432(1.22); C30(1.53); C421(0.76); C434(0.97) | LDD1082 | [7] |
| LDCM0485 | CL90 | HEK-293T | C354(1.07); C432(0.76); C421(0.83); C434(0.76) | LDD1083 | [7] |
| LDCM0486 | CL91 | HCT 116 | C30(0.80) | LDD0803 | [7] |
| LDCM0487 | CL92 | HCT 116 | C30(0.97) | LDD0804 | [7] |
| LDCM0488 | CL93 | HCT 116 | C30(0.96) | LDD0805 | [7] |
| LDCM0489 | CL94 | HCT 116 | C30(1.84) | LDD0806 | [7] |
| LDCM0490 | CL95 | HCT 116 | C30(1.29) | LDD0807 | [7] |
| LDCM0491 | CL96 | HCT 116 | C30(1.29) | LDD0808 | [7] |
| LDCM0492 | CL97 | HCT 116 | C30(1.10) | LDD0809 | [7] |
| LDCM0493 | CL98 | HCT 116 | C30(1.02) | LDD0810 | [7] |
| LDCM0494 | CL99 | HCT 116 | C30(1.09) | LDD0811 | [7] |
| LDCM0495 | E2913 | HEK-293T | C30(0.72); C456(1.13); C513(1.02) | LDD1698 | [21] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C519(0.95) | LDD1702 | [4] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [6] |
| LDCM0625 | F8 | Ramos | C30(1.70) | LDD2187 | [22] |
| LDCM0572 | Fragment10 | Ramos | C30(1.75) | LDD2189 | [22] |
| LDCM0574 | Fragment12 | Ramos | C30(0.49) | LDD2191 | [22] |
| LDCM0575 | Fragment13 | Ramos | C30(0.75) | LDD2192 | [22] |
| LDCM0576 | Fragment14 | Ramos | C30(0.58) | LDD2193 | [22] |
| LDCM0579 | Fragment20 | Ramos | C30(0.65) | LDD2194 | [22] |
| LDCM0580 | Fragment21 | Ramos | C30(0.86) | LDD2195 | [22] |
| LDCM0582 | Fragment23 | Ramos | C30(0.76) | LDD2196 | [22] |
| LDCM0578 | Fragment27 | Ramos | C30(0.74) | LDD2197 | [22] |
| LDCM0586 | Fragment28 | Ramos | C30(0.89) | LDD2198 | [22] |
| LDCM0588 | Fragment30 | Ramos | C30(0.88) | LDD2199 | [22] |
| LDCM0589 | Fragment31 | Ramos | C30(0.57) | LDD2200 | [22] |
| LDCM0590 | Fragment32 | Ramos | C30(1.57) | LDD2201 | [22] |
| LDCM0468 | Fragment33 | HEK-293T | C354(1.46); C432(1.46); C487(1.48); C421(1.27) | LDD1066 | [7] |
| LDCM0596 | Fragment38 | Ramos | C30(0.71) | LDD2203 | [22] |
| LDCM0566 | Fragment4 | Ramos | C30(0.73) | LDD2184 | [22] |
| LDCM0427 | Fragment51 | HCT 116 | C30(1.70) | LDD0744 | [7] |
| LDCM0610 | Fragment52 | Ramos | C30(0.88) | LDD2204 | [22] |
| LDCM0614 | Fragment56 | Ramos | C30(1.17) | LDD2205 | [22] |
| LDCM0569 | Fragment7 | Ramos | C30(0.98) | LDD2186 | [22] |
| LDCM0571 | Fragment9 | Ramos | C30(0.82) | LDD2188 | [22] |
| LDCM0022 | KB02 | HEK-293T | C30(0.92); C354(0.96); C513(1.07); C434(1.27) | LDD1492 | [21] |
| LDCM0023 | KB03 | HEK-293T | C30(0.92); C354(1.01); C513(1.01); C434(1.09) | LDD1497 | [21] |
| LDCM0024 | KB05 | COLO792 | C513(1.97); C354(1.64) | LDD3310 | [23] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C519(2.42) | LDD2090 | [4] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C519(1.20) | LDD2107 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C519(0.61) | LDD2109 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C519(0.82) | LDD2125 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C519(0.83) | LDD2127 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C519(1.00) | LDD2136 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C519(0.96) | LDD2137 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C519(1.08) | LDD2140 | [4] |
The Interaction Atlas With This Target
References
