Details of the Target
General Information of Target
| Target ID | LDTP02165 | |||||
|---|---|---|---|---|---|---|
| Target Name | Tropomyosin alpha-3 chain (TPM3) | |||||
| Gene Name | TPM3 | |||||
| Gene ID | 7170 | |||||
| Synonyms |
Tropomyosin alpha-3 chain; Gamma-tropomyosin; Tropomyosin-3; Tropomyosin-5; hTM5 |
|||||
| 3D Structure | ||||||
| Sequence |
MMEAIKKKMQMLKLDKENALDRAEQAEAEQKQAEERSKQLEDELAAMQKKLKGTEDELDK
YSEALKDAQEKLELAEKKAADAEAEVASLNRRIQLVEEELDRAQERLATALQKLEEAEKA ADESERGMKVIENRALKDEEKMELQEIQLKEAKHIAEEADRKYEEVARKLVIIEGDLERT EERAELAESKCSELEEELKNVTNNLKSLEAQAEKYSQKEDKYEEEIKILTDKLKEAETRA EFAERSVAKLEKTIDDLEDELYAQKLKYKAISEELDHALNDMTSI |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Tropomyosin family
|
|||||
| Subcellular location |
Cytoplasm, cytoskeleton
|
|||||
| Function |
Binds to actin filaments in muscle and non-muscle cells. Plays a central role, in association with the troponin complex, in the calcium dependent regulation of vertebrate striated muscle contraction. Smooth muscle contraction is regulated by interaction with caldesmon. In non-muscle cells is implicated in stabilizing cytoskeleton actin filaments.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
P3 Probe Info |
 |
10.00 | LDD0450 | [2] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [3] | |
|
ONAyne Probe Info |
 |
K137(1.06); K150(0.78); K232(3.09) | LDD0274 | [4] | |
|
STPyne Probe Info |
 |
K137(6.94); K150(9.38); K169(3.29); K218(10.00) | LDD0277 | [4] | |
|
OPA-S-S-alkyne Probe Info |
 |
K119(3.65); K153(4.13); K234(5.08); K129(7.91) | LDD3494 | [5] | |
|
Probe 1 Probe Info |
 |
Y163(48.39); Y222(101.52); Y262(29.49) | LDD3495 | [6] | |
|
DBIA Probe Info |
 |
C170(2.70) | LDD3313 | [7] | |
|
BTD Probe Info |
 |
C191(1.18) | LDD2089 | [8] | |
|
DA-P3 Probe Info |
 |
5.27 | LDD0181 | [9] | |
|
HHS-482 Probe Info |
 |
Y163(0.98) | LDD0285 | [10] | |
|
HHS-475 Probe Info |
 |
Y163(0.89) | LDD0264 | [11] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0231 | [12] | |
|
HHS-465 Probe Info |
 |
Y163(10.00) | LDD2237 | [13] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [14] | |
|
ATP probe Probe Info |
 |
K153(0.00); K162(0.00); K169(0.00); K141(0.00) | LDD0199 | [15] | |
|
CY-1 Probe Info |
 |
N.A. | LDD0246 | [16] | |
|
ATP probe Probe Info |
 |
K129(0.00); K249(0.00); K119(0.00); K169(0.00) | LDD0035 | [17] | |
|
AZ-9 Probe Info |
 |
D101(0.00); E98(0.00) | LDD0395 | [18] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [19] | |
|
SF Probe Info |
 |
Y163(0.00); K162(0.00); K232(0.00); K113(0.00) | LDD0028 | [20] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [21] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [22] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C087 Probe Info |
 |
7.73 | LDD1779 | [23] | |
|
C187 Probe Info |
 |
17.15 | LDD1865 | [23] | |
|
C210 Probe Info |
 |
38.85 | LDD1884 | [23] | |
|
C313 Probe Info |
 |
12.21 | LDD1980 | [23] | |
|
C354 Probe Info |
 |
6.96 | LDD2015 | [23] | |
|
C362 Probe Info |
 |
26.72 | LDD2023 | [23] | |
|
C391 Probe Info |
 |
12.73 | LDD2050 | [23] | |
|
FFF probe12 Probe Info |
 |
15.51 | LDD0473 | [24] | |
|
FFF probe13 Probe Info |
 |
14.76 | LDD0475 | [24] | |
|
FFF probe14 Probe Info |
 |
12.37 | LDD0477 | [24] | |
|
FFF probe2 Probe Info |
 |
13.96 | LDD0463 | [24] | |
|
FFF probe4 Probe Info |
 |
20.00 | LDD0466 | [24] | |
|
FFF probe9 Probe Info |
 |
7.12 | LDD0470 | [24] | |
|
Diazir Probe Info |
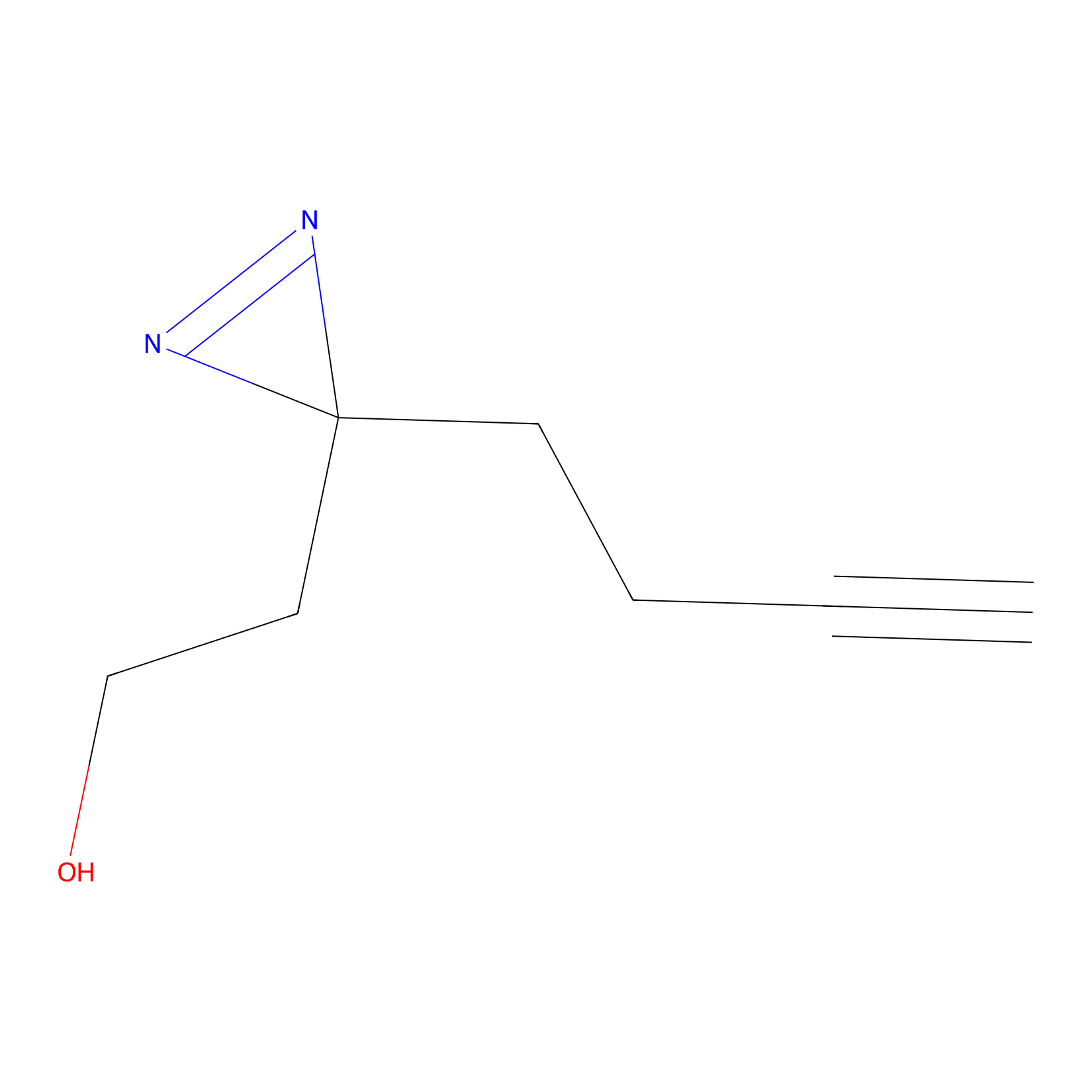 |
E85(0.00); E97(0.00); E98(0.00) | LDD0011 | [19] | |
|
OEA-DA Probe Info |
 |
4.36 | LDD0046 | [25] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C191(0.38) | LDD2132 | [8] |
| LDCM0156 | Aniline | NCI-H1299 | 11.50 | LDD0403 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C191(0.98) | LDD2091 | [8] |
| LDCM0116 | HHS-0101 | DM93 | Y163(0.89) | LDD0264 | [11] |
| LDCM0117 | HHS-0201 | DM93 | Y163(0.77) | LDD0265 | [11] |
| LDCM0118 | HHS-0301 | DM93 | Y163(0.89) | LDD0266 | [11] |
| LDCM0119 | HHS-0401 | DM93 | Y163(0.86) | LDD0267 | [11] |
| LDCM0120 | HHS-0701 | DM93 | Y163(0.74) | LDD0268 | [11] |
| LDCM0123 | JWB131 | DM93 | Y163(0.98) | LDD0285 | [10] |
| LDCM0124 | JWB142 | DM93 | Y163(0.88) | LDD0286 | [10] |
| LDCM0125 | JWB146 | DM93 | Y163(1.04) | LDD0287 | [10] |
| LDCM0126 | JWB150 | DM93 | Y163(4.39) | LDD0288 | [10] |
| LDCM0127 | JWB152 | DM93 | Y163(2.61) | LDD0289 | [10] |
| LDCM0128 | JWB198 | DM93 | Y163(1.92) | LDD0290 | [10] |
| LDCM0129 | JWB202 | DM93 | Y163(0.79) | LDD0291 | [10] |
| LDCM0130 | JWB211 | DM93 | Y163(1.14) | LDD0292 | [10] |
| LDCM0022 | KB02 | A101D | C170(2.83) | LDD2250 | [7] |
| LDCM0023 | KB03 | A101D | C170(2.94) | LDD2667 | [7] |
| LDCM0024 | KB05 | Hs 936.T | C170(2.70) | LDD3313 | [7] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0231 | [12] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C191(1.18) | LDD2089 | [8] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C191(0.90) | LDD2090 | [8] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C191(0.61) | LDD2115 | [8] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C191(0.82) | LDD2120 | [8] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C191(0.79) | LDD2128 | [8] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C191(0.43) | LDD2145 | [8] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C191(1.33) | LDD2147 | [8] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C191(0.69) | LDD2148 | [8] |
| LDCM0029 | Quercetin | HEK-293T | 5.27 | LDD0181 | [9] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| 26S proteasome regulatory subunit 8 (PSMC5) | AAA ATPase family | P62195 | |||
| Beta-secretase 2 (BACE2) | Peptidase A1 family | Q9Y5Z0 | |||
| Kallikrein-6 (KLK6) | Peptidase S1 family | Q92876 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nucleoporin p54 (NUP54) | NUP54 family | Q7Z3B4 | |||
| SNARE-associated protein Snapin (SNAPIN) | SNAPIN family | O95295 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Heat shock factor protein 2 (HSF2) | HSF family | Q03933 | |||
| THAP domain-containing protein 1 (THAP1) | THAP1 family | Q9NVV9 | |||
| THAP domain-containing protein 7 (THAP7) | . | Q9BT49 | |||
Other
References
