Details of the Target
General Information of Target
| Target ID | LDTP07605 | |||||
|---|---|---|---|---|---|---|
| Target Name | La-related protein 1 (LARP1) | |||||
| Gene Name | LARP1 | |||||
| Gene ID | 23367 | |||||
| Synonyms |
KIAA0731; LARP; La-related protein 1; La ribonucleoprotein domain family member 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MATQVEPLLPGGATLLQAEEHGGLVRKKPPPAPEGKGEPGPNDVRGGEPDGSARRPRPPC
AKPHKEGTGQQERESPRPLQLPGAEGPAISDGEEGGGEPGAGGGAAGAAGAGRRDFVEAP PPKVNPWTKNALPPVLTTVNGQSPPEHSAPAKVVRAAVPKQRKGSKVGDFGDAINWPTPG EIAHKSVQPQSHKPQPTRKLPPKKDMKEQEKGEGSDSKESPKTKSDESGEEKNGDEDCQR GGQKKKGNKHKWVPLQIDMKPEVPREKLASRPTRPPEPRHIPANRGEIKGSESATYVPVA PPTPAWQPEIKPEPAWHDQDETSSVKSDGAGGARASFRGRGRGRGRGRGRGRGGTRTHFD YQFGYRKFDGVEGPRTPKYMNNITYYFDNVSSTELYSVDQELLKDYIKRQIEYYFSVDNL ERDFFLRRKMDADGFLPITLIASFHRVQALTTDISLIFAALKDSKVVEIVDEKVRRREEP EKWPLPPIVDYSQTDFSQLLNCPEFVPRQHYQKETESAPGSPRAVTPVPTKTEEVSNLKT LPKGLSASLPDLDSENWIEVKKRPRPSPARPKKSEESRFSHLTSLPQQLPSQQLMSKDQD EQEELDFLFDEEMEQMDGRKNTFTAWSDEESDYEIDDRDVNKILIVTQTPHYMRRHPGGD RTGNHTSRAKMSAELAKVINDGLFYYEQDLWAEKFEPEYSQIKQEVENFKKVNMISREQF DTLTPEPPVDPNQEVPPGPPRFQQVPTDALANKLFGAPEPSTIARSLPTTVPESPNYRNT RTPRTPRTPQLKDSSQTSRFYPVVKEGRTLDAKMPRKRKTRHSSNPPLESHVGWVMDSRE HRPRTASISSSPSEGTPTVGSYGCTPQSLPKFQHPSHELLKENGFTQHVYHKYRRRCLNE RKRLGIGQSQEMNTLFRFWSFFLRDHFNKKMYEEFKQLALEDAKEGYRYGLECLFRYYSY GLEKKFRLDIFKDFQEETVKDYEAGQLYGLEKFWAFLKYSKAKNLDIDPKLQEYLGKFRR LEDFRVDPPMGEEGNHKRHSVVAGGGGGEGRKRCPSQSSSRPAAMISQPPTPPTGQPVRE DAKWTSQHSNTQTLGK |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
LARP family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
RNA-binding protein that regulates the translation of specific target mRNA species downstream of the mTORC1 complex, in function of growth signals and nutrient availability. Interacts on the one hand with the 3' poly-A tails that are present in all mRNA molecules, and on the other hand with the 7-methylguanosine cap structure of mRNAs containing a 5' terminal oligopyrimidine (5'TOP) motif, which is present in mRNAs encoding ribosomal proteins and several components of the translation machinery . The interaction with the 5' end of mRNAs containing a 5'TOP motif leads to translational repression by preventing the binding of EIF4G1. When mTORC1 is activated, LARP1 is phosphorylated and dissociates from the 5' untranslated region (UTR) of mRNA. Does not prevent binding of EIF4G1 to mRNAs that lack a 5'TOP motif. Interacts with the free 40S ribosome subunit and with ribosomes, both monosomes and polysomes. Under normal nutrient availability, interacts primarily with the 3' untranslated region (UTR) of mRNAs encoding ribosomal proteins and increases protein synthesis. Associates with actively translating ribosomes and stimulates translation of mRNAs containing a 5'TOP motif, thereby regulating protein synthesis, and as a consequence, cell growth and proliferation. Stabilizes mRNAs species with a 5'TOP motif, which is required to prevent apoptosis.; (Microbial infection) Positively regulates the replication of dengue virus (DENV).
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CCK81 | Deletion: p.T303PfsTer23 | DBIA Probe Info | |||
| FADU | SNV: p.V135A | . | |||
| HCT116 | Deletion: p.P145QfsTer10; p.T303PfsTer23 | DBIA Probe Info | |||
| HCT15 | SNV: p.H445Q; p.D981E | DBIA Probe Info | |||
| HT115 | SNV: p.R956Q | DBIA Probe Info | |||
| IM95 | Deletion: p.T303PfsTer23 | DBIA Probe Info | |||
| JURKAT | Deletion: p.D660TfsTer12 | Compound 10 Probe Info | |||
| KYSE410 | SNV: p.G242R | DBIA Probe Info | |||
| LS180 | SNV: p.W252C | DBIA Probe Info | |||
| MOLT4 | Deletion: p.T303PfsTer23 | IA-alkyne Probe Info | |||
| NCIH2286 | SNV: p.D610Y | DBIA Probe Info | |||
| RKO | SNV: p.G682D | DBIA Probe Info | |||
| RL952 | SNV: p.T780I | DBIA Probe Info | |||
| SKNSH | SNV: p.G103Ter | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
DAyne Probe Info |
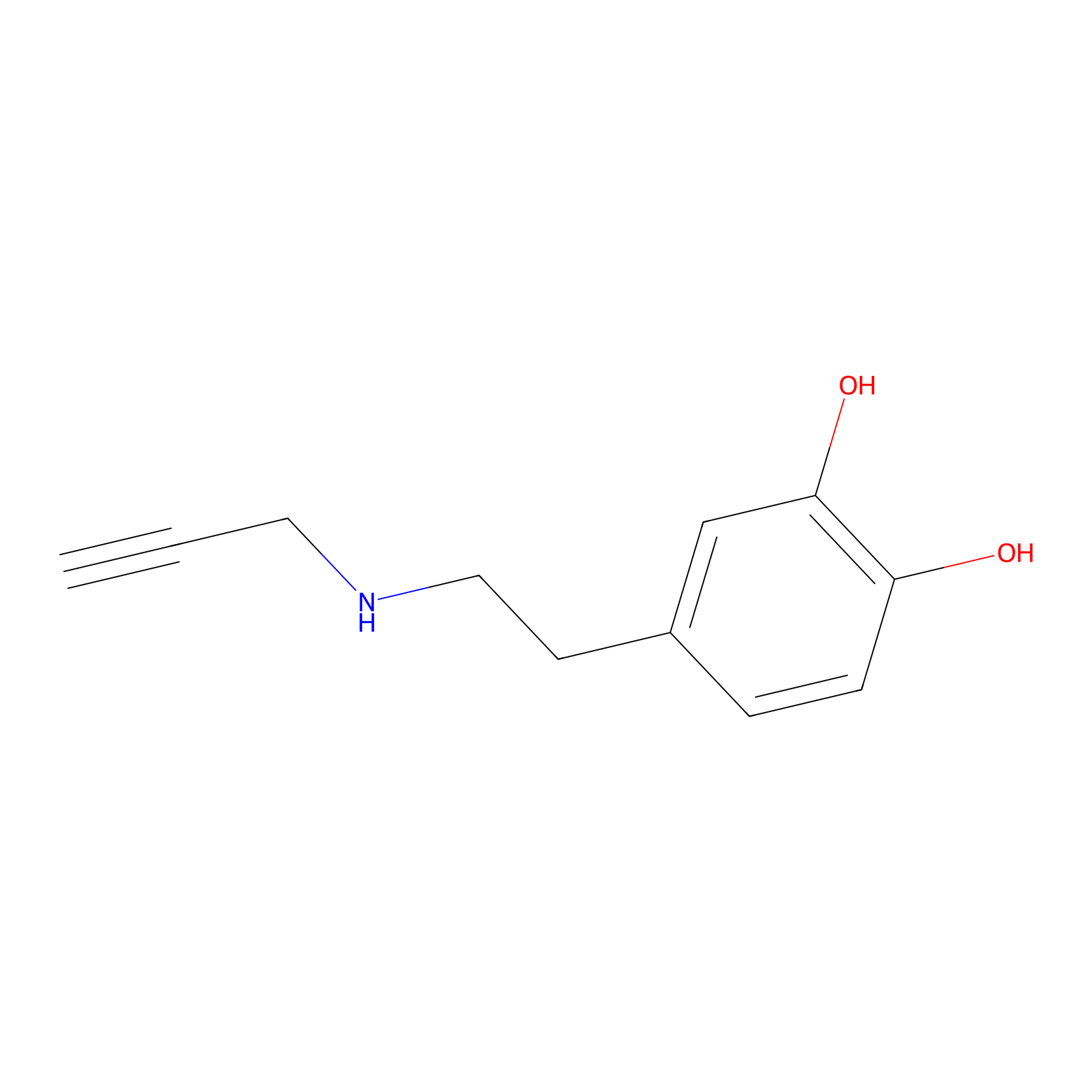 |
2.12 | LDD0261 | [2] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [3] | |
|
ONAyne Probe Info |
 |
K792(5.26) | LDD0274 | [4] | |
|
STPyne Probe Info |
 |
K1017(7.05); K160(5.88); K513(0.55); K531(10.00) | LDD0277 | [4] | |
|
Probe 1 Probe Info |
 |
Y777(5.19) | LDD3495 | [5] | |
|
BTD Probe Info |
 |
C897(1.50) | LDD1700 | [6] | |
|
Sulforaphane-probe2 Probe Info |
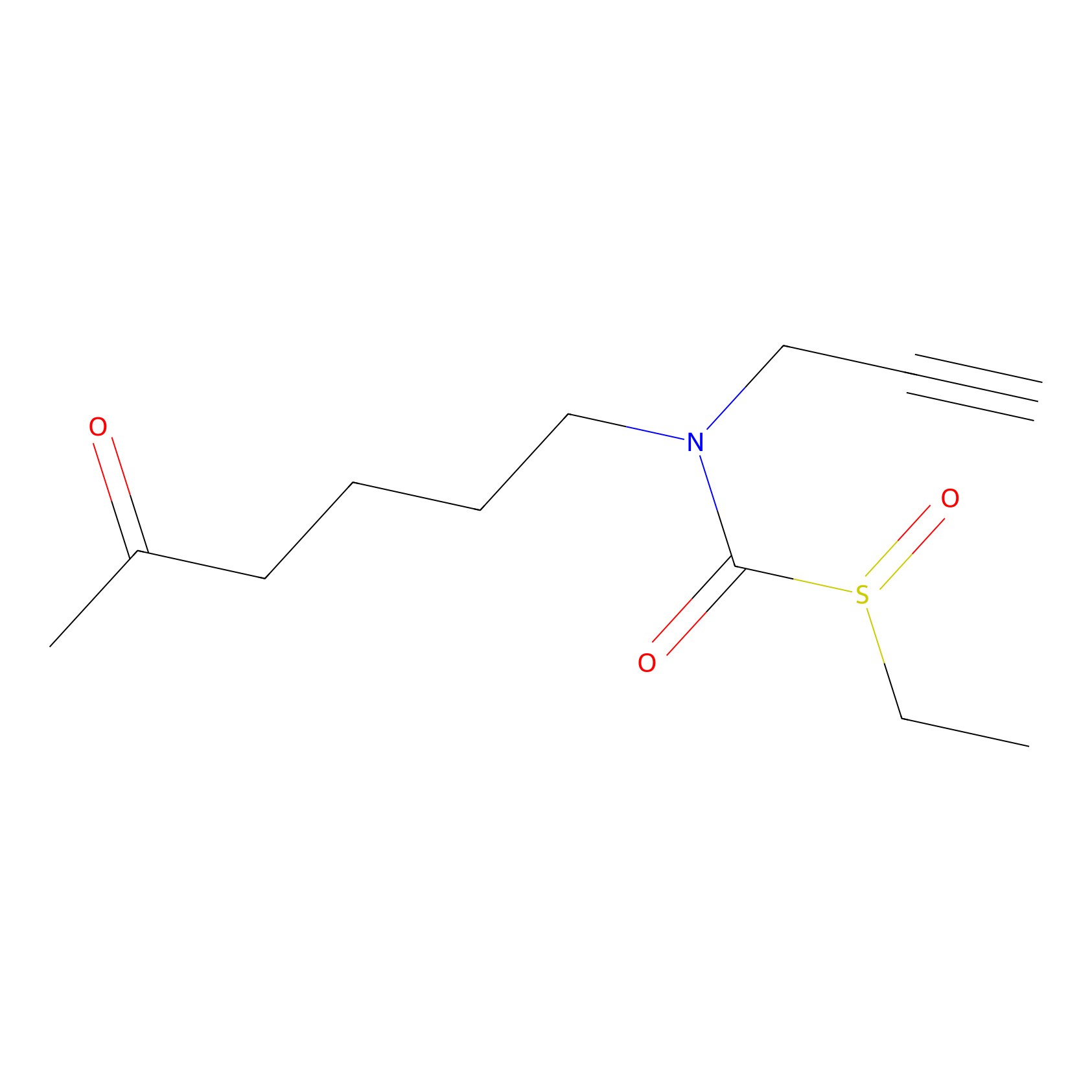 |
2.06 | LDD0042 | [7] | |
|
DA-P3 Probe Info |
 |
5.30 | LDD0181 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C238(3.79) | LDD0168 | [9] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [10] | |
|
DBIA Probe Info |
 |
C864(1.08); C238(1.04) | LDD0078 | [11] | |
|
5E-2FA Probe Info |
 |
H21(0.00); H192(0.00) | LDD2235 | [12] | |
|
ATP probe Probe Info |
 |
K1003(0.00); K367(0.00); K65(0.00); K260(0.00) | LDD0199 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C1054(0.00); C502(0.00); C238(0.00); C864(0.00) | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [15] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [16] | |
|
Lodoacetamide azide Probe Info |
 |
C1054(0.00); C502(0.00); C864(0.00); C953(0.00) | LDD0037 | [14] | |
|
IPM Probe Info |
 |
C1054(0.00); C864(0.00) | LDD0025 | [17] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [17] | |
|
NAIA_4 Probe Info |
 |
C502(0.00); C864(0.00) | LDD2226 | [18] | |
|
TFBX Probe Info |
 |
C1054(0.00); C864(0.00) | LDD0027 | [17] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [19] | |
|
Compound 10 Probe Info |
 |
C1054(0.00); C238(0.00); C864(0.00) | LDD2216 | [20] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [21] | |
|
PPMS Probe Info |
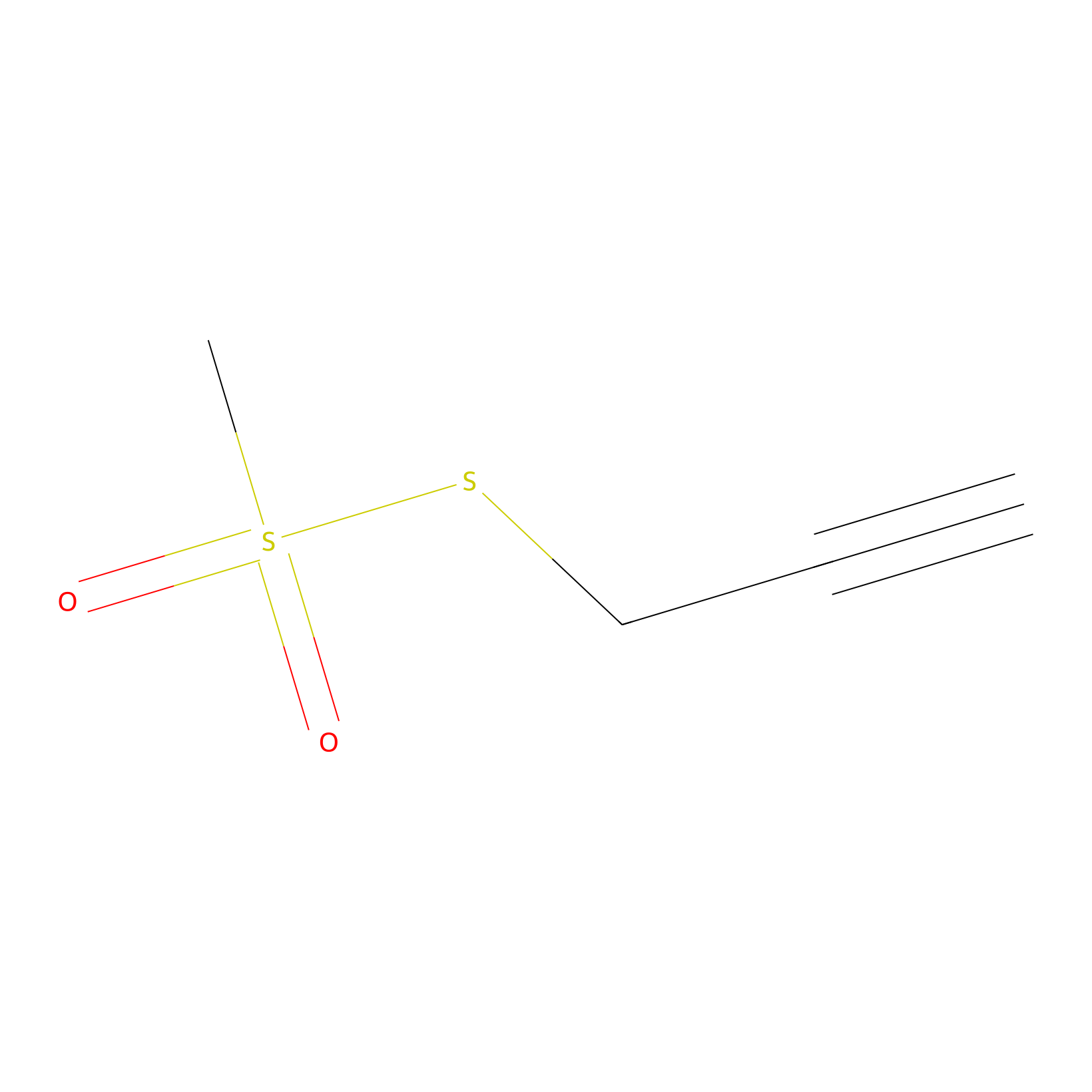 |
N.A. | LDD0008 | [19] | |
|
SF Probe Info |
 |
Y361(0.00); Y777(0.00); Y960(0.00); Y365(0.00) | LDD0028 | [22] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [19] | |
|
Phosphinate-6 Probe Info |
 |
C238(0.00); C60(0.00); C864(0.00) | LDD0018 | [23] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [24] | |
|
1c-yne Probe Info |
 |
K753(0.00); K429(0.00); K694(0.00) | LDD0228 | [25] | |
|
Acrolein Probe Info |
 |
H1039(0.00); C238(0.00) | LDD0217 | [26] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [26] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [27] | |
|
AOyne Probe Info |
 |
6.90 | LDD0443 | [28] | |
|
NAIA_5 Probe Info |
 |
C1054(0.00); C502(0.00); C953(0.00); C897(0.00) | LDD2223 | [18] | |
|
HHS-482 Probe Info |
 |
Y365(0.99) | LDD2239 | [29] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C191 Probe Info |
 |
9.58 | LDD1868 | [30] | |
|
FFF probe11 Probe Info |
 |
11.30 | LDD0471 | [31] | |
|
FFF probe13 Probe Info |
 |
14.73 | LDD0475 | [31] | |
|
FFF probe2 Probe Info |
 |
6.62 | LDD0463 | [31] | |
|
FFF probe3 Probe Info |
 |
18.47 | LDD0464 | [31] | |
|
FFF probe9 Probe Info |
 |
20.00 | LDD0470 | [31] | |
|
Staurosporine capture compound Probe Info |
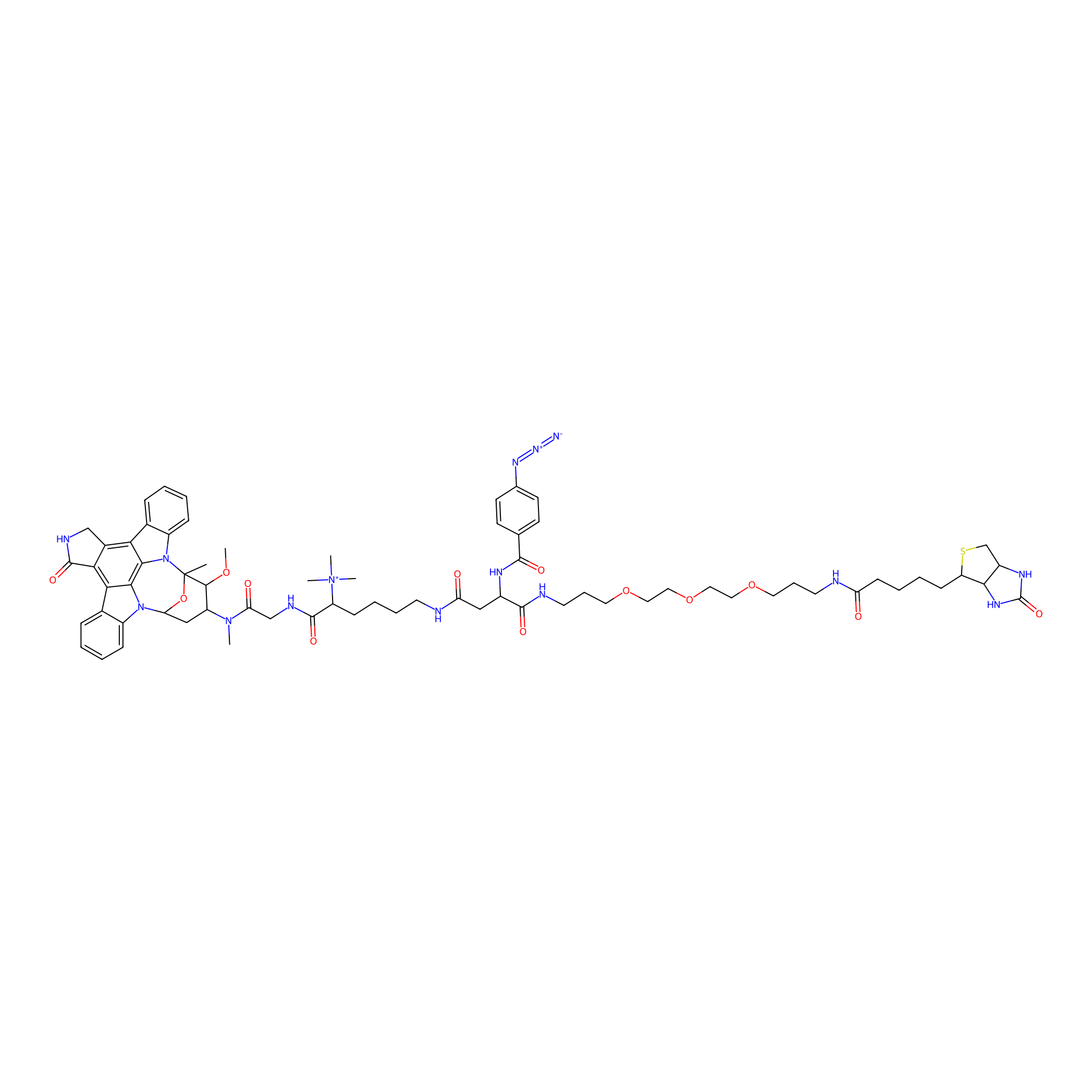 |
N.A. | LDD0083 | [32] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C1054(0.76) | LDD2142 | [6] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C1054(0.96) | LDD2112 | [6] |
| LDCM0025 | 4SU-RNA | HEK-293T | C238(3.79) | LDD0168 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C238(3.13) | LDD0169 | [9] |
| LDCM0214 | AC1 | HCT 116 | C238(0.85); C864(1.20); C953(1.18) | LDD0531 | [11] |
| LDCM0215 | AC10 | HCT 116 | C238(1.68); C864(1.21); C953(0.66) | LDD0532 | [11] |
| LDCM0216 | AC100 | HCT 116 | C238(0.96); C864(0.84) | LDD0533 | [11] |
| LDCM0217 | AC101 | HCT 116 | C238(1.08); C864(0.86) | LDD0534 | [11] |
| LDCM0218 | AC102 | HCT 116 | C238(0.85); C864(0.95) | LDD0535 | [11] |
| LDCM0219 | AC103 | HCT 116 | C238(1.45); C864(1.22) | LDD0536 | [11] |
| LDCM0220 | AC104 | HCT 116 | C238(0.91); C864(1.03) | LDD0537 | [11] |
| LDCM0221 | AC105 | HCT 116 | C238(1.45); C864(1.03) | LDD0538 | [11] |
| LDCM0222 | AC106 | HCT 116 | C238(1.56); C864(1.00) | LDD0539 | [11] |
| LDCM0223 | AC107 | HCT 116 | C238(1.38); C864(1.25) | LDD0540 | [11] |
| LDCM0224 | AC108 | HCT 116 | C238(1.23); C864(0.91) | LDD0541 | [11] |
| LDCM0225 | AC109 | HCT 116 | C238(0.81); C864(0.92) | LDD0542 | [11] |
| LDCM0226 | AC11 | HCT 116 | C238(1.30); C864(1.34); C953(0.63) | LDD0543 | [11] |
| LDCM0227 | AC110 | HCT 116 | C238(1.06); C864(0.97) | LDD0544 | [11] |
| LDCM0228 | AC111 | HCT 116 | C238(1.12); C864(0.92) | LDD0545 | [11] |
| LDCM0229 | AC112 | HCT 116 | C238(0.94); C864(1.12) | LDD0546 | [11] |
| LDCM0230 | AC113 | HCT 116 | C238(1.08); C864(1.00) | LDD0547 | [11] |
| LDCM0231 | AC114 | HCT 116 | C238(1.47); C864(1.15) | LDD0548 | [11] |
| LDCM0232 | AC115 | HCT 116 | C238(1.74); C864(1.33) | LDD0549 | [11] |
| LDCM0233 | AC116 | HCT 116 | C238(1.73); C864(1.42) | LDD0550 | [11] |
| LDCM0234 | AC117 | HCT 116 | C238(1.42); C864(1.15) | LDD0551 | [11] |
| LDCM0235 | AC118 | HCT 116 | C238(1.38); C864(1.13) | LDD0552 | [11] |
| LDCM0236 | AC119 | HCT 116 | C238(1.29); C864(1.18) | LDD0553 | [11] |
| LDCM0237 | AC12 | HCT 116 | C238(0.99); C864(1.07); C953(0.78) | LDD0554 | [11] |
| LDCM0238 | AC120 | HCT 116 | C238(0.83); C864(1.33) | LDD0555 | [11] |
| LDCM0239 | AC121 | HCT 116 | C238(1.53); C864(1.03) | LDD0556 | [11] |
| LDCM0240 | AC122 | HCT 116 | C238(1.20); C864(1.10) | LDD0557 | [11] |
| LDCM0241 | AC123 | HCT 116 | C238(1.35); C864(0.99) | LDD0558 | [11] |
| LDCM0242 | AC124 | HCT 116 | C238(1.07); C864(0.97) | LDD0559 | [11] |
| LDCM0243 | AC125 | HCT 116 | C238(1.28); C864(1.01) | LDD0560 | [11] |
| LDCM0244 | AC126 | HCT 116 | C238(1.79); C864(1.18) | LDD0561 | [11] |
| LDCM0245 | AC127 | HCT 116 | C238(1.95); C864(1.35) | LDD0562 | [11] |
| LDCM0246 | AC128 | HCT 116 | C238(1.52); C864(1.22); C953(0.82) | LDD0563 | [11] |
| LDCM0247 | AC129 | HCT 116 | C238(1.36); C864(0.94); C953(1.12) | LDD0564 | [11] |
| LDCM0249 | AC130 | HCT 116 | C238(2.09); C864(1.18); C953(0.87) | LDD0566 | [11] |
| LDCM0250 | AC131 | HCT 116 | C238(1.76); C864(0.96); C953(0.88) | LDD0567 | [11] |
| LDCM0251 | AC132 | HCT 116 | C238(1.45); C864(0.90); C953(1.04) | LDD0568 | [11] |
| LDCM0252 | AC133 | HCT 116 | C238(1.64); C864(0.98); C953(0.87) | LDD0569 | [11] |
| LDCM0253 | AC134 | HCT 116 | C238(2.00); C864(1.32); C953(0.81) | LDD0570 | [11] |
| LDCM0254 | AC135 | HCT 116 | C238(1.86); C864(1.03); C953(0.83) | LDD0571 | [11] |
| LDCM0255 | AC136 | HCT 116 | C238(1.05); C864(1.18); C953(0.95) | LDD0572 | [11] |
| LDCM0256 | AC137 | HCT 116 | C238(1.02); C864(1.03); C953(0.96) | LDD0573 | [11] |
| LDCM0257 | AC138 | HCT 116 | C238(1.58); C864(1.17); C953(0.80) | LDD0574 | [11] |
| LDCM0258 | AC139 | HCT 116 | C238(1.54); C864(1.08); C953(0.86) | LDD0575 | [11] |
| LDCM0259 | AC14 | HCT 116 | C238(1.69); C864(1.09); C953(0.72) | LDD0576 | [11] |
| LDCM0260 | AC140 | HCT 116 | C238(2.24); C864(1.23); C953(0.83) | LDD0577 | [11] |
| LDCM0261 | AC141 | HCT 116 | C238(1.58); C864(1.17); C953(0.81) | LDD0578 | [11] |
| LDCM0262 | AC142 | HCT 116 | C238(1.28); C864(0.91); C953(1.15) | LDD0579 | [11] |
| LDCM0263 | AC143 | HCT 116 | C953(0.95); C864(1.06); C238(1.17) | LDD0580 | [11] |
| LDCM0264 | AC144 | HCT 116 | C953(0.68); C864(1.31); C238(1.39) | LDD0581 | [11] |
| LDCM0265 | AC145 | HCT 116 | C953(0.83); C864(1.11); C238(1.13) | LDD0582 | [11] |
| LDCM0266 | AC146 | HCT 116 | C953(0.55); C864(1.39); C238(1.63) | LDD0583 | [11] |
| LDCM0267 | AC147 | HCT 116 | C953(0.55); C864(1.37); C238(1.40) | LDD0584 | [11] |
| LDCM0268 | AC148 | HCT 116 | C953(0.42); C238(2.04); C864(2.05) | LDD0585 | [11] |
| LDCM0269 | AC149 | HCT 116 | C953(0.50); C864(1.56); C238(1.76) | LDD0586 | [11] |
| LDCM0270 | AC15 | HCT 116 | C953(0.60); C864(1.21); C238(1.67) | LDD0587 | [11] |
| LDCM0271 | AC150 | HCT 116 | C864(1.13); C238(1.15); C953(1.18) | LDD0588 | [11] |
| LDCM0272 | AC151 | HCT 116 | C953(0.79); C238(1.21); C864(1.25) | LDD0589 | [11] |
| LDCM0273 | AC152 | HCT 116 | C953(0.51); C238(1.06); C864(1.36) | LDD0590 | [11] |
| LDCM0274 | AC153 | HCT 116 | C953(0.44); C864(2.23); C238(2.42) | LDD0591 | [11] |
| LDCM0621 | AC154 | HCT 116 | C238(1.68); C864(1.39); C953(0.84) | LDD2158 | [11] |
| LDCM0622 | AC155 | HCT 116 | C238(1.55); C864(1.42); C953(0.61) | LDD2159 | [11] |
| LDCM0623 | AC156 | HCT 116 | C238(1.24); C864(1.08); C953(0.81) | LDD2160 | [11] |
| LDCM0624 | AC157 | HCT 116 | C238(1.31); C864(1.22); C953(0.99) | LDD2161 | [11] |
| LDCM0276 | AC17 | HCT 116 | C238(0.95); C864(1.14) | LDD0593 | [11] |
| LDCM0277 | AC18 | HCT 116 | C238(0.64); C864(1.58) | LDD0594 | [11] |
| LDCM0278 | AC19 | HCT 116 | C864(1.18); C238(1.20) | LDD0595 | [11] |
| LDCM0279 | AC2 | HCT 116 | C238(0.98); C953(1.13); C864(1.18) | LDD0596 | [11] |
| LDCM0280 | AC20 | HCT 116 | C238(1.08); C864(1.22) | LDD0597 | [11] |
| LDCM0281 | AC21 | HCT 116 | C238(0.97); C864(1.21) | LDD0598 | [11] |
| LDCM0282 | AC22 | HCT 116 | C238(1.00); C864(1.08) | LDD0599 | [11] |
| LDCM0283 | AC23 | HCT 116 | C238(0.93); C864(1.17) | LDD0600 | [11] |
| LDCM0284 | AC24 | HCT 116 | C864(1.01); C238(1.31) | LDD0601 | [11] |
| LDCM0285 | AC25 | HCT 116 | C864(1.02); C238(2.03) | LDD0602 | [11] |
| LDCM0286 | AC26 | HCT 116 | C864(1.22); C238(2.23) | LDD0603 | [11] |
| LDCM0287 | AC27 | HCT 116 | C864(1.27); C238(2.09) | LDD0604 | [11] |
| LDCM0288 | AC28 | HCT 116 | C864(1.25); C238(1.69) | LDD0605 | [11] |
| LDCM0289 | AC29 | HCT 116 | C864(1.24); C238(2.62) | LDD0606 | [11] |
| LDCM0290 | AC3 | HCT 116 | C238(0.88); C953(1.06); C864(1.15) | LDD0607 | [11] |
| LDCM0291 | AC30 | HCT 116 | C864(1.42); C238(1.60) | LDD0608 | [11] |
| LDCM0292 | AC31 | HCT 116 | C864(1.15); C238(2.40) | LDD0609 | [11] |
| LDCM0293 | AC32 | HCT 116 | C864(1.50); C238(2.39) | LDD0610 | [11] |
| LDCM0294 | AC33 | HCT 116 | C864(1.34); C238(2.75) | LDD0611 | [11] |
| LDCM0295 | AC34 | HCT 116 | C864(1.71); C238(2.34) | LDD0612 | [11] |
| LDCM0296 | AC35 | HCT 116 | C238(0.95); C864(0.96); C953(1.43) | LDD0613 | [11] |
| LDCM0297 | AC36 | HCT 116 | C864(0.96); C953(1.16); C238(1.17) | LDD0614 | [11] |
| LDCM0298 | AC37 | HCT 116 | C864(0.97); C953(1.03); C238(1.22) | LDD0615 | [11] |
| LDCM0299 | AC38 | HCT 116 | C864(1.04); C238(1.08); C953(1.29) | LDD0616 | [11] |
| LDCM0300 | AC39 | HCT 116 | C238(1.04); C953(1.07); C864(1.15) | LDD0617 | [11] |
| LDCM0301 | AC4 | HCT 116 | C953(0.91); C238(1.00); C864(1.33) | LDD0618 | [11] |
| LDCM0302 | AC40 | HCT 116 | C953(0.88); C238(1.21); C864(1.29) | LDD0619 | [11] |
| LDCM0303 | AC41 | HCT 116 | C953(1.00); C864(1.10); C238(1.33) | LDD0620 | [11] |
| LDCM0304 | AC42 | HCT 116 | C953(1.03); C864(1.13); C238(1.18) | LDD0621 | [11] |
| LDCM0305 | AC43 | HCT 116 | C864(1.23); C953(1.25); C238(1.30) | LDD0622 | [11] |
| LDCM0306 | AC44 | HCT 116 | C953(1.02); C864(1.19); C238(1.24) | LDD0623 | [11] |
| LDCM0307 | AC45 | HCT 116 | C953(0.83); C864(1.28); C238(1.44) | LDD0624 | [11] |
| LDCM0308 | AC46 | HCT 116 | C864(1.04); C953(1.06); C238(1.24) | LDD0625 | [11] |
| LDCM0309 | AC47 | HCT 116 | C953(0.96); C864(0.99); C238(1.38) | LDD0626 | [11] |
| LDCM0310 | AC48 | HCT 116 | C864(1.00); C953(1.09); C238(1.29) | LDD0627 | [11] |
| LDCM0311 | AC49 | HCT 116 | C953(0.69); C238(1.50); C864(1.58) | LDD0628 | [11] |
| LDCM0312 | AC5 | HCT 116 | C238(0.85); C953(0.89); C864(1.32) | LDD0629 | [11] |
| LDCM0313 | AC50 | HCT 116 | C953(0.68); C864(1.37); C238(1.93) | LDD0630 | [11] |
| LDCM0314 | AC51 | HCT 116 | C864(0.90); C238(1.10); C953(1.17) | LDD0631 | [11] |
| LDCM0315 | AC52 | HCT 116 | C953(0.85); C864(1.03); C238(1.37) | LDD0632 | [11] |
| LDCM0316 | AC53 | HCT 116 | C953(0.86); C864(1.23); C238(1.46) | LDD0633 | [11] |
| LDCM0317 | AC54 | HCT 116 | C953(0.84); C238(1.19); C864(1.25) | LDD0634 | [11] |
| LDCM0318 | AC55 | HCT 116 | C953(0.84); C864(1.47); C238(1.91) | LDD0635 | [11] |
| LDCM0319 | AC56 | HCT 116 | C953(0.60); C864(2.28); C238(2.93) | LDD0636 | [11] |
| LDCM0320 | AC57 | HCT 116 | C953(0.69); C864(1.32); C238(1.71) | LDD0637 | [11] |
| LDCM0321 | AC58 | HCT 116 | C953(0.58); C864(1.53); C238(1.77) | LDD0638 | [11] |
| LDCM0322 | AC59 | HCT 116 | C953(0.59); C864(1.39); C238(1.56) | LDD0639 | [11] |
| LDCM0323 | AC6 | HCT 116 | C953(0.62); C864(1.35); C238(1.50) | LDD0640 | [11] |
| LDCM0324 | AC60 | HCT 116 | C953(0.70); C864(1.51); C238(1.86) | LDD0641 | [11] |
| LDCM0325 | AC61 | HCT 116 | C953(0.78); C864(1.27); C238(1.93) | LDD0642 | [11] |
| LDCM0326 | AC62 | HCT 116 | C953(0.54); C864(1.38); C238(2.03) | LDD0643 | [11] |
| LDCM0327 | AC63 | HCT 116 | C953(0.74); C238(1.39); C864(1.41) | LDD0644 | [11] |
| LDCM0328 | AC64 | HCT 116 | C953(0.60); C864(1.48); C238(1.73) | LDD0645 | [11] |
| LDCM0329 | AC65 | HCT 116 | C953(0.77); C864(1.30); C238(1.32) | LDD0646 | [11] |
| LDCM0330 | AC66 | HCT 116 | C953(0.82); C238(1.11); C864(1.33) | LDD0647 | [11] |
| LDCM0331 | AC67 | HCT 116 | C953(0.45); C238(1.26); C864(1.75) | LDD0648 | [11] |
| LDCM0332 | AC68 | HCT 116 | C953(1.03); C864(1.19); C238(1.24) | LDD0649 | [11] |
| LDCM0333 | AC69 | HCT 116 | C953(0.93); C864(1.22); C238(1.64) | LDD0650 | [11] |
| LDCM0334 | AC7 | HCT 116 | C953(0.76); C238(0.94); C864(1.02) | LDD0651 | [11] |
| LDCM0335 | AC70 | HCT 116 | C953(0.67); C864(1.22); C238(1.42) | LDD0652 | [11] |
| LDCM0336 | AC71 | HCT 116 | C953(1.11); C864(1.14); C238(1.23) | LDD0653 | [11] |
| LDCM0337 | AC72 | HCT 116 | C953(0.93); C864(1.16); C238(1.93) | LDD0654 | [11] |
| LDCM0338 | AC73 | HCT 116 | C953(0.57); C864(1.74); C238(2.40) | LDD0655 | [11] |
| LDCM0339 | AC74 | HCT 116 | C953(0.59); C864(1.57); C238(1.66) | LDD0656 | [11] |
| LDCM0340 | AC75 | HCT 116 | C953(0.51); C864(2.11); C238(2.85) | LDD0657 | [11] |
| LDCM0341 | AC76 | HCT 116 | C953(0.83); C864(1.27); C238(1.63) | LDD0658 | [11] |
| LDCM0342 | AC77 | HCT 116 | C953(0.76); C864(1.39); C238(2.10) | LDD0659 | [11] |
| LDCM0343 | AC78 | HCT 116 | C864(1.12); C953(1.21); C238(1.75) | LDD0660 | [11] |
| LDCM0344 | AC79 | HCT 116 | C953(1.02); C864(1.26); C238(1.57) | LDD0661 | [11] |
| LDCM0345 | AC8 | HCT 116 | C953(0.67); C238(0.93); C864(1.33) | LDD0662 | [11] |
| LDCM0346 | AC80 | HCT 116 | C953(0.84); C864(1.20); C238(1.57) | LDD0663 | [11] |
| LDCM0347 | AC81 | HCT 116 | C238(0.97); C864(1.16); C953(1.27) | LDD0664 | [11] |
| LDCM0348 | AC82 | HCT 116 | C953(0.57); C238(1.87); C864(2.03) | LDD0665 | [11] |
| LDCM0349 | AC83 | HCT 116 | C953(0.30); C238(1.85); C864(1.88) | LDD0666 | [11] |
| LDCM0350 | AC84 | HCT 116 | C953(0.30); C238(1.12); C864(1.73) | LDD0667 | [11] |
| LDCM0351 | AC85 | HCT 116 | C953(0.44); C238(0.90); C864(1.43) | LDD0668 | [11] |
| LDCM0352 | AC86 | HCT 116 | C953(0.40); C238(1.17); C864(1.35) | LDD0669 | [11] |
| LDCM0353 | AC87 | HCT 116 | C953(0.62); C238(0.88); C864(1.10) | LDD0670 | [11] |
| LDCM0354 | AC88 | HCT 116 | C953(0.35); C238(1.01); C864(1.31) | LDD0671 | [11] |
| LDCM0355 | AC89 | HCT 116 | C953(0.33); C864(1.45); C238(1.54) | LDD0672 | [11] |
| LDCM0357 | AC90 | HCT 116 | C953(0.93); C864(0.96); C238(1.09) | LDD0674 | [11] |
| LDCM0358 | AC91 | HCT 116 | C953(0.25); C864(1.78); C238(1.82) | LDD0675 | [11] |
| LDCM0359 | AC92 | HCT 116 | C953(0.30); C864(1.56); C238(1.74) | LDD0676 | [11] |
| LDCM0360 | AC93 | HCT 116 | C953(0.38); C238(0.94); C864(1.22) | LDD0677 | [11] |
| LDCM0361 | AC94 | HCT 116 | C953(0.59); C238(0.86); C864(1.33) | LDD0678 | [11] |
| LDCM0362 | AC95 | HCT 116 | C953(0.43); C864(1.16); C238(1.28) | LDD0679 | [11] |
| LDCM0363 | AC96 | HCT 116 | C953(0.31); C238(1.41); C864(1.44) | LDD0680 | [11] |
| LDCM0364 | AC97 | HCT 116 | C953(0.40); C238(1.18); C864(1.56) | LDD0681 | [11] |
| LDCM0365 | AC98 | HCT 116 | C864(1.57); C238(2.17) | LDD0682 | [11] |
| LDCM0366 | AC99 | HCT 116 | C238(0.82); C864(0.91) | LDD0683 | [11] |
| LDCM0248 | AKOS034007472 | HCT 116 | C238(1.37); C864(1.01); C953(0.91) | LDD0565 | [11] |
| LDCM0356 | AKOS034007680 | HCT 116 | C953(0.69); C864(1.12); C238(1.60) | LDD0673 | [11] |
| LDCM0275 | AKOS034007705 | HCT 116 | C953(0.41); C864(1.97); C238(2.66) | LDD0592 | [11] |
| LDCM0156 | Aniline | NCI-H1299 | N.A. | LDD0404 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C864(1.08); C238(1.04) | LDD0078 | [11] |
| LDCM0087 | Capsaicin | HEK-293T | 6.66 | LDD0185 | [8] |
| LDCM0630 | CCW28-3 | 231MFP | C864(1.71) | LDD2214 | [33] |
| LDCM0108 | Chloroacetamide | HeLa | H1039(0.00); H192(0.00); C238(0.00); H651(0.00) | LDD0222 | [26] |
| LDCM0632 | CL-Sc | Hep-G2 | C238(1.24); C238(0.98); C1054(0.65) | LDD2227 | [18] |
| LDCM0367 | CL1 | HCT 116 | C864(1.00); C238(1.08); C953(1.17) | LDD0684 | [11] |
| LDCM0368 | CL10 | HCT 116 | C953(0.68); C864(1.26); C238(1.89) | LDD0685 | [11] |
| LDCM0369 | CL100 | HCT 116 | C953(0.96); C238(1.04); C864(1.27) | LDD0686 | [11] |
| LDCM0370 | CL101 | HCT 116 | C953(0.55); C864(1.15); C238(1.42) | LDD0687 | [11] |
| LDCM0371 | CL102 | HCT 116 | C953(0.86); C864(0.94); C238(1.26) | LDD0688 | [11] |
| LDCM0372 | CL103 | HCT 116 | C953(0.73); C864(0.98); C238(1.15) | LDD0689 | [11] |
| LDCM0373 | CL104 | HCT 116 | C953(0.78); C864(1.16); C238(1.27) | LDD0690 | [11] |
| LDCM0374 | CL105 | HCT 116 | C238(1.00); C864(1.51) | LDD0691 | [11] |
| LDCM0375 | CL106 | HCT 116 | C238(1.12); C864(1.91) | LDD0692 | [11] |
| LDCM0376 | CL107 | HCT 116 | C238(1.14); C864(1.54) | LDD0693 | [11] |
| LDCM0377 | CL108 | HCT 116 | C238(0.93); C864(1.82) | LDD0694 | [11] |
| LDCM0378 | CL109 | HCT 116 | C238(1.23); C864(1.38) | LDD0695 | [11] |
| LDCM0379 | CL11 | HCT 116 | C953(0.62); C864(1.32); C238(2.17) | LDD0696 | [11] |
| LDCM0380 | CL110 | HCT 116 | C238(1.38); C864(1.67) | LDD0697 | [11] |
| LDCM0381 | CL111 | HCT 116 | C238(0.92); C864(1.40) | LDD0698 | [11] |
| LDCM0382 | CL112 | HCT 116 | C864(1.18); C238(1.69) | LDD0699 | [11] |
| LDCM0383 | CL113 | HCT 116 | C864(1.45); C238(2.04) | LDD0700 | [11] |
| LDCM0384 | CL114 | HCT 116 | C864(1.23); C238(2.27) | LDD0701 | [11] |
| LDCM0385 | CL115 | HCT 116 | C864(1.28); C238(2.10) | LDD0702 | [11] |
| LDCM0386 | CL116 | HCT 116 | C864(1.19); C238(2.32) | LDD0703 | [11] |
| LDCM0387 | CL117 | HCT 116 | C953(0.69); C238(1.63); C864(1.91) | LDD0704 | [11] |
| LDCM0388 | CL118 | HCT 116 | C864(1.11); C953(1.26); C238(1.34) | LDD0705 | [11] |
| LDCM0389 | CL119 | HCT 116 | C953(0.94); C864(1.24); C238(1.87) | LDD0706 | [11] |
| LDCM0390 | CL12 | HCT 116 | C953(0.62); C864(1.41); C238(2.86) | LDD0707 | [11] |
| LDCM0391 | CL120 | HCT 116 | C864(1.12); C953(1.15); C238(1.39) | LDD0708 | [11] |
| LDCM0392 | CL121 | HCT 116 | C953(0.78); C238(1.02); C864(1.06) | LDD0709 | [11] |
| LDCM0393 | CL122 | HCT 116 | C953(0.93); C864(1.04); C238(1.27) | LDD0710 | [11] |
| LDCM0394 | CL123 | HCT 116 | C953(0.73); C238(1.44); C864(1.54) | LDD0711 | [11] |
| LDCM0395 | CL124 | HCT 116 | C953(0.80); C864(1.47); C238(1.70) | LDD0712 | [11] |
| LDCM0396 | CL125 | HCT 116 | C953(1.01); C238(1.06); C864(1.14) | LDD0713 | [11] |
| LDCM0397 | CL126 | HCT 116 | C953(0.65); C238(1.04); C864(1.47) | LDD0714 | [11] |
| LDCM0398 | CL127 | HCT 116 | C953(0.78); C864(1.05); C238(1.24) | LDD0715 | [11] |
| LDCM0399 | CL128 | HCT 116 | C953(0.62); C864(1.51); C238(1.97) | LDD0716 | [11] |
| LDCM0400 | CL13 | HCT 116 | C953(0.69); C864(1.21); C238(1.59) | LDD0717 | [11] |
| LDCM0401 | CL14 | HCT 116 | C953(0.89); C864(1.05); C238(1.32) | LDD0718 | [11] |
| LDCM0402 | CL15 | HCT 116 | C953(0.53); C864(1.18); C238(1.34) | LDD0719 | [11] |
| LDCM0403 | CL16 | HCT 116 | C953(0.72); C238(0.94); C864(1.12) | LDD0720 | [11] |
| LDCM0404 | CL17 | HCT 116 | C953(0.58); C238(1.07); C864(1.16) | LDD0721 | [11] |
| LDCM0405 | CL18 | HCT 116 | C953(0.54); C864(1.08); C238(1.63) | LDD0722 | [11] |
| LDCM0406 | CL19 | HCT 116 | C953(0.57); C864(1.13); C238(1.33) | LDD0723 | [11] |
| LDCM0407 | CL2 | HCT 116 | C864(0.92); C238(1.02); C953(1.16) | LDD0724 | [11] |
| LDCM0408 | CL20 | HCT 116 | C953(0.51); C864(1.23); C238(1.60) | LDD0725 | [11] |
| LDCM0409 | CL21 | HCT 116 | C953(0.44); C238(1.56); C864(1.88) | LDD0726 | [11] |
| LDCM0410 | CL22 | HCT 116 | C953(0.29); C864(2.10); C238(2.10) | LDD0727 | [11] |
| LDCM0411 | CL23 | HCT 116 | C953(0.46); C864(1.10); C238(1.65) | LDD0728 | [11] |
| LDCM0412 | CL24 | HCT 116 | C953(0.39); C864(1.35); C238(1.85) | LDD0729 | [11] |
| LDCM0413 | CL25 | HCT 116 | C238(1.79); C864(1.50); C953(0.48) | LDD0730 | [11] |
| LDCM0414 | CL26 | HCT 116 | C238(1.64); C864(1.09); C953(0.50) | LDD0731 | [11] |
| LDCM0415 | CL27 | HCT 116 | C238(1.82); C864(1.09); C953(0.51) | LDD0732 | [11] |
| LDCM0416 | CL28 | HCT 116 | C238(1.90); C864(1.24); C953(0.51) | LDD0733 | [11] |
| LDCM0417 | CL29 | HCT 116 | C238(2.25); C864(1.35); C953(0.44) | LDD0734 | [11] |
| LDCM0418 | CL3 | HCT 116 | C238(1.09); C864(0.87); C953(0.98) | LDD0735 | [11] |
| LDCM0419 | CL30 | HCT 116 | C238(1.34); C864(1.02); C953(0.52) | LDD0736 | [11] |
| LDCM0420 | CL31 | HCT 116 | C238(1.28); C864(1.19); C953(0.64) | LDD0737 | [11] |
| LDCM0421 | CL32 | HCT 116 | C238(1.36); C864(1.19); C953(0.73) | LDD0738 | [11] |
| LDCM0422 | CL33 | HCT 116 | C238(1.35); C864(1.17); C953(0.55) | LDD0739 | [11] |
| LDCM0423 | CL34 | HCT 116 | C238(1.48); C864(1.52); C953(0.48) | LDD0740 | [11] |
| LDCM0424 | CL35 | HCT 116 | C238(1.51); C864(1.50); C953(0.52) | LDD0741 | [11] |
| LDCM0425 | CL36 | HCT 116 | C238(1.47); C864(1.42); C953(0.61) | LDD0742 | [11] |
| LDCM0426 | CL37 | HCT 116 | C238(2.09); C864(1.65); C953(0.44) | LDD0743 | [11] |
| LDCM0428 | CL39 | HCT 116 | C238(1.55); C864(1.63); C953(0.45) | LDD0745 | [11] |
| LDCM0429 | CL4 | HCT 116 | C238(1.23); C864(0.85); C953(1.02) | LDD0746 | [11] |
| LDCM0430 | CL40 | HCT 116 | C238(1.47); C864(1.45); C953(0.54) | LDD0747 | [11] |
| LDCM0431 | CL41 | HCT 116 | C238(1.58); C864(1.31); C953(0.54) | LDD0748 | [11] |
| LDCM0432 | CL42 | HCT 116 | C238(1.78); C864(3.35); C953(0.37) | LDD0749 | [11] |
| LDCM0433 | CL43 | HCT 116 | C238(1.71); C864(1.62); C953(0.46) | LDD0750 | [11] |
| LDCM0434 | CL44 | HCT 116 | C238(1.57); C864(1.42); C953(0.58) | LDD0751 | [11] |
| LDCM0435 | CL45 | HCT 116 | C238(1.51); C864(1.62); C953(0.52) | LDD0752 | [11] |
| LDCM0436 | CL46 | HCT 116 | C238(1.21); C864(0.82); C953(1.35) | LDD0753 | [11] |
| LDCM0437 | CL47 | HCT 116 | C238(1.30); C864(0.87); C953(0.93) | LDD0754 | [11] |
| LDCM0438 | CL48 | HCT 116 | C238(1.20); C864(0.70); C953(1.31) | LDD0755 | [11] |
| LDCM0439 | CL49 | HCT 116 | C238(0.95); C864(0.80); C953(0.96) | LDD0756 | [11] |
| LDCM0440 | CL5 | HCT 116 | C238(1.41); C864(0.91); C953(0.91) | LDD0757 | [11] |
| LDCM0441 | CL50 | HCT 116 | C238(1.17); C864(0.84); C953(1.43) | LDD0758 | [11] |
| LDCM0442 | CL51 | HCT 116 | C238(1.33); C864(0.80); C953(0.95) | LDD0759 | [11] |
| LDCM0443 | CL52 | HCT 116 | C238(1.59); C864(0.82); C953(0.85) | LDD0760 | [11] |
| LDCM0444 | CL53 | HCT 116 | C238(1.03); C864(0.89); C953(1.82) | LDD0761 | [11] |
| LDCM0445 | CL54 | HCT 116 | C238(1.21); C864(0.86); C953(1.09) | LDD0762 | [11] |
| LDCM0446 | CL55 | HCT 116 | C238(1.08); C864(0.72); C953(1.03) | LDD0763 | [11] |
| LDCM0447 | CL56 | HCT 116 | C238(1.06); C864(0.88); C953(1.06) | LDD0764 | [11] |
| LDCM0448 | CL57 | HCT 116 | C238(1.21); C864(0.85); C953(1.35) | LDD0765 | [11] |
| LDCM0449 | CL58 | HCT 116 | C238(1.14); C864(0.73); C953(1.50) | LDD0766 | [11] |
| LDCM0450 | CL59 | HCT 116 | C238(1.33); C864(0.87); C953(1.05) | LDD0767 | [11] |
| LDCM0451 | CL6 | HCT 116 | C238(1.21); C864(1.01); C953(0.85) | LDD0768 | [11] |
| LDCM0452 | CL60 | HCT 116 | C238(1.60); C864(0.78); C953(0.97) | LDD0769 | [11] |
| LDCM0453 | CL61 | HCT 116 | C238(1.20); C864(1.02) | LDD0770 | [11] |
| LDCM0454 | CL62 | HCT 116 | C238(1.07); C864(1.12) | LDD0771 | [11] |
| LDCM0455 | CL63 | HCT 116 | C238(1.34); C864(1.22) | LDD0772 | [11] |
| LDCM0456 | CL64 | HCT 116 | C238(1.37); C864(1.55) | LDD0773 | [11] |
| LDCM0457 | CL65 | HCT 116 | C238(0.94); C864(1.23) | LDD0774 | [11] |
| LDCM0458 | CL66 | HCT 116 | C238(1.43); C864(1.71) | LDD0775 | [11] |
| LDCM0459 | CL67 | HCT 116 | C238(1.22); C864(1.48) | LDD0776 | [11] |
| LDCM0460 | CL68 | HCT 116 | C238(1.50); C864(1.23) | LDD0777 | [11] |
| LDCM0461 | CL69 | HCT 116 | C238(1.58); C864(1.07) | LDD0778 | [11] |
| LDCM0462 | CL7 | HCT 116 | C238(1.59); C864(1.30); C953(0.77) | LDD0779 | [11] |
| LDCM0463 | CL70 | HCT 116 | C238(1.29); C864(1.20) | LDD0780 | [11] |
| LDCM0464 | CL71 | HCT 116 | C238(1.27); C864(1.36) | LDD0781 | [11] |
| LDCM0465 | CL72 | HCT 116 | C238(1.40); C864(1.18) | LDD0782 | [11] |
| LDCM0466 | CL73 | HCT 116 | C238(1.48); C864(1.30) | LDD0783 | [11] |
| LDCM0467 | CL74 | HCT 116 | C238(1.82); C864(1.30) | LDD0784 | [11] |
| LDCM0469 | CL76 | HCT 116 | C238(1.16); C864(1.11); C953(1.06) | LDD0786 | [11] |
| LDCM0470 | CL77 | HCT 116 | C238(1.13); C864(1.21); C953(0.82) | LDD0787 | [11] |
| LDCM0471 | CL78 | HCT 116 | C238(1.18); C864(1.20); C953(0.86) | LDD0788 | [11] |
| LDCM0472 | CL79 | HCT 116 | C238(1.62); C864(1.25); C953(0.65) | LDD0789 | [11] |
| LDCM0473 | CL8 | HCT 116 | C238(2.44); C864(1.70); C953(0.61) | LDD0790 | [11] |
| LDCM0474 | CL80 | HCT 116 | C238(0.90); C864(0.99); C953(1.05) | LDD0791 | [11] |
| LDCM0475 | CL81 | HCT 116 | C238(1.43); C864(1.12); C953(0.80) | LDD0792 | [11] |
| LDCM0476 | CL82 | HCT 116 | C238(1.76); C864(1.53); C953(0.56) | LDD0793 | [11] |
| LDCM0477 | CL83 | HCT 116 | C238(1.54); C864(1.64); C953(0.64) | LDD0794 | [11] |
| LDCM0478 | CL84 | HCT 116 | C238(1.86); C864(2.11); C953(0.50) | LDD0795 | [11] |
| LDCM0479 | CL85 | HCT 116 | C238(1.35); C864(1.34); C953(0.84) | LDD0796 | [11] |
| LDCM0480 | CL86 | HCT 116 | C238(0.86); C864(1.15); C953(1.08) | LDD0797 | [11] |
| LDCM0481 | CL87 | HCT 116 | C238(1.23); C864(1.21); C953(0.96) | LDD0798 | [11] |
| LDCM0482 | CL88 | HCT 116 | C238(1.56); C864(1.30); C953(0.75) | LDD0799 | [11] |
| LDCM0483 | CL89 | HCT 116 | C238(1.82); C864(2.31); C953(0.51) | LDD0800 | [11] |
| LDCM0484 | CL9 | HCT 116 | C238(1.38); C864(1.12); C953(0.82) | LDD0801 | [11] |
| LDCM0485 | CL90 | HCT 116 | C238(0.85); C864(1.00); C953(1.10) | LDD0802 | [11] |
| LDCM0486 | CL91 | HCT 116 | C238(1.28); C864(1.22); C953(1.07) | LDD0803 | [11] |
| LDCM0487 | CL92 | HCT 116 | C238(0.89); C864(1.19); C953(1.08) | LDD0804 | [11] |
| LDCM0488 | CL93 | HCT 116 | C238(1.21); C864(1.05); C953(1.14) | LDD0805 | [11] |
| LDCM0489 | CL94 | HCT 116 | C238(1.25); C864(1.42); C953(0.99) | LDD0806 | [11] |
| LDCM0490 | CL95 | HCT 116 | C238(1.25); C864(1.47); C953(0.93) | LDD0807 | [11] |
| LDCM0491 | CL96 | HCT 116 | C238(1.25); C864(1.25); C953(0.90) | LDD0808 | [11] |
| LDCM0492 | CL97 | HCT 116 | C238(0.98); C864(1.29); C953(1.02) | LDD0809 | [11] |
| LDCM0493 | CL98 | HCT 116 | C238(1.05); C864(1.27); C953(0.94) | LDD0810 | [11] |
| LDCM0494 | CL99 | HCT 116 | C238(1.00); C864(1.33); C953(0.97) | LDD0811 | [11] |
| LDCM0634 | CY-0357 | Hep-G2 | C864(0.48) | LDD2228 | [18] |
| LDCM0495 | E2913 | HEK-293T | C1054(1.16); C238(1.05); C60(1.14); C864(0.79) | LDD1698 | [34] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C1054(1.31); C864(1.00) | LDD1702 | [6] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 7.52 | LDD0183 | [8] |
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [10] |
| LDCM0625 | F8 | Ramos | C953(0.54); C1054(1.63); C238(1.62); C864(1.06) | LDD2187 | [35] |
| LDCM0572 | Fragment10 | Ramos | C953(0.52); C1054(2.53); C238(1.08); C864(1.00) | LDD2189 | [35] |
| LDCM0573 | Fragment11 | Ramos | C953(2.21); C1054(1.18); C238(20.00); C864(0.33) | LDD2190 | [35] |
| LDCM0574 | Fragment12 | Ramos | C953(0.57); C1054(1.84); C238(0.72); C864(0.75) | LDD2191 | [35] |
| LDCM0575 | Fragment13 | Ramos | C953(0.89); C1054(1.41); C238(0.86); C864(1.12) | LDD2192 | [35] |
| LDCM0576 | Fragment14 | Ramos | C953(0.67); C1054(0.79); C238(0.88); C864(0.86) | LDD2193 | [35] |
| LDCM0579 | Fragment20 | Ramos | C953(0.45); C1054(1.33); C238(0.74); C864(0.91) | LDD2194 | [35] |
| LDCM0580 | Fragment21 | Ramos | C953(0.81); C1054(0.99); C238(0.99); C864(0.79) | LDD2195 | [35] |
| LDCM0582 | Fragment23 | Ramos | C953(1.06); C1054(1.13); C238(0.91); C864(1.02) | LDD2196 | [35] |
| LDCM0578 | Fragment27 | Ramos | C953(0.94); C1054(1.35); C238(1.05); C864(1.12) | LDD2197 | [35] |
| LDCM0586 | Fragment28 | Ramos | C953(0.72); C1054(0.73); C238(0.73); C864(1.00) | LDD2198 | [35] |
| LDCM0588 | Fragment30 | Ramos | C953(0.74); C1054(1.92); C238(1.22); C864(1.44) | LDD2199 | [35] |
| LDCM0589 | Fragment31 | Ramos | C953(0.93); C1054(1.27); C238(1.13); C864(0.88) | LDD2200 | [35] |
| LDCM0590 | Fragment32 | Ramos | C953(0.40); C1054(3.66); C238(1.94); C864(1.08) | LDD2201 | [35] |
| LDCM0468 | Fragment33 | HCT 116 | C238(1.63); C864(1.11) | LDD0785 | [11] |
| LDCM0596 | Fragment38 | Ramos | C953(1.14); C1054(1.34); C238(0.74); C864(0.64) | LDD2203 | [35] |
| LDCM0566 | Fragment4 | Ramos | C953(0.52); C1054(1.01); C238(0.64); C864(1.19) | LDD2184 | [35] |
| LDCM0427 | Fragment51 | HCT 116 | C238(1.98); C864(2.33); C953(0.38) | LDD0744 | [11] |
| LDCM0610 | Fragment52 | Ramos | C953(1.14); C1054(2.91); C238(1.01); C864(1.34) | LDD2204 | [35] |
| LDCM0614 | Fragment56 | Ramos | C953(0.88); C1054(1.91); C238(1.18); C864(1.09) | LDD2205 | [35] |
| LDCM0569 | Fragment7 | Ramos | C953(0.59); C1054(1.16); C238(0.73); C864(1.14) | LDD2186 | [35] |
| LDCM0571 | Fragment9 | Ramos | C953(0.44); C1054(1.55); C238(0.74); C864(1.02) | LDD2188 | [35] |
| LDCM0107 | IAA | HeLa | H1039(0.00); H1088(0.00); H192(0.00); H581(0.00) | LDD0221 | [26] |
| LDCM0022 | KB02 | HCT 116 | C864(1.87) | LDD0080 | [11] |
| LDCM0023 | KB03 | HCT 116 | C864(1.19) | LDD0081 | [11] |
| LDCM0024 | KB05 | HCT 116 | C864(2.00) | LDD0082 | [11] |
| LDCM0030 | Luteolin | HEK-293T | 5.67 | LDD0182 | [8] |
| LDCM0109 | NEM | HeLa | H1039(0.00); H1088(0.00); H651(0.00); H581(0.00) | LDD0223 | [26] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C1054(0.65) | LDD2092 | [6] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C1054(1.05) | LDD2100 | [6] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C1054(0.79) | LDD2104 | [6] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C1054(1.13) | LDD2105 | [6] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C1054(0.63) | LDD2106 | [6] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C1054(0.96) | LDD2108 | [6] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C1054(1.16) | LDD2110 | [6] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C1054(0.60) | LDD2114 | [6] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C897(1.50) | LDD1700 | [6] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C864(0.58) | LDD2207 | [36] |
| LDCM0029 | Quercetin | HEK-293T | 5.30 | LDD0181 | [8] |
| LDCM0131 | RA190 | MM1.R | C864(1.77) | LDD0304 | [37] |
| LDCM0019 | Staurosporine | Hep-G2 | N.A. | LDD0083 | [32] |
| LDCM0003 | Sulforaphane | MCF-7 | 2.06 | LDD0042 | [7] |
| LDCM0021 | THZ1 | HCT 116 | C864(0.97); C238(1.18) | LDD2173 | [11] |
The Interaction Atlas With This Target
References
