Details of the Target
General Information of Target
| Target ID | LDTP01419 | |||||
|---|---|---|---|---|---|---|
| Target Name | Retinal dehydrogenase 2 (ALDH1A2) | |||||
| Gene Name | ALDH1A2 | |||||
| Gene ID | 8854 | |||||
| Synonyms |
RALDH2; Retinal dehydrogenase 2; RALDH 2; RalDH2; EC 1.2.1.36; Aldehyde dehydrogenase family 1 member A2; ALDH1A2; Retinaldehyde-specific dehydrogenase type 2; RALDH(II) |
|||||
| 3D Structure | ||||||
| Sequence |
MTSSKIEMPGEVKADPAALMASLHLLPSPTPNLEIKYTKIFINNEWQNSESGRVFPVYNP
ATGEQVCEVQEADKADIDKAVQAARLAFSLGSVWRRMDASERGRLLDKLADLVERDRAVL ATMESLNGGKPFLQAFYVDLQGVIKTFRYYAGWADKIHGMTIPVDGDYFTFTRHEPIGVC GQIIPWNFPLLMFAWKIAPALCCGNTVVIKPAEQTPLSALYMGALIKEAGFPPGVINILP GYGPTAGAAIASHIGIDKIAFTGSTEVGKLIQEAAGRSNLKRVTLELGGKSPNIIFADAD LDYAVEQAHQGVFFNQGQCCTAGSRIFVEESIYEEFVRRSVERAKRRVVGSPFDPTTEQG PQIDKKQYNKILELIQSGVAEGAKLECGGKGLGRKGFFIEPTVFSNVTDDMRIAKEEIFG PVQEILRFKTMDEVIERANNSDFGLVAAVFTNDINKALTVSSAMQAGTVWINCYNALNAQ SPFGGFKMSGNGREMGEFGLREYSEVKTVTVKIPQKNS |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Aldehyde dehydrogenase family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Catalyzes the NAD-dependent oxidation of aldehyde substrates, such as all-trans-retinal and all-trans-13,14-dihydroretinal, to their corresponding carboxylic acids, all-trans-retinoate and all-trans-13,14-dihydroretinoate, respectively. Retinoate signaling is critical for the transcriptional control of many genes, for instance it is crucial for initiation of meiosis in both male and female (Probable). Recognizes retinal as substrate, both in its free form and when bound to cellular retinol-binding protein. Can metabolize octanal and decanal, but has only very low activity with benzaldehyde, acetaldehyde and propanal. Displays complete lack of activity with citral.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
A-EBA Probe Info |
 |
4.08 | LDD0215 | [1] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [2] | |
|
AZ-9 Probe Info |
 |
E286(1.00) | LDD2208 | [3] | |
|
P11 Probe Info |
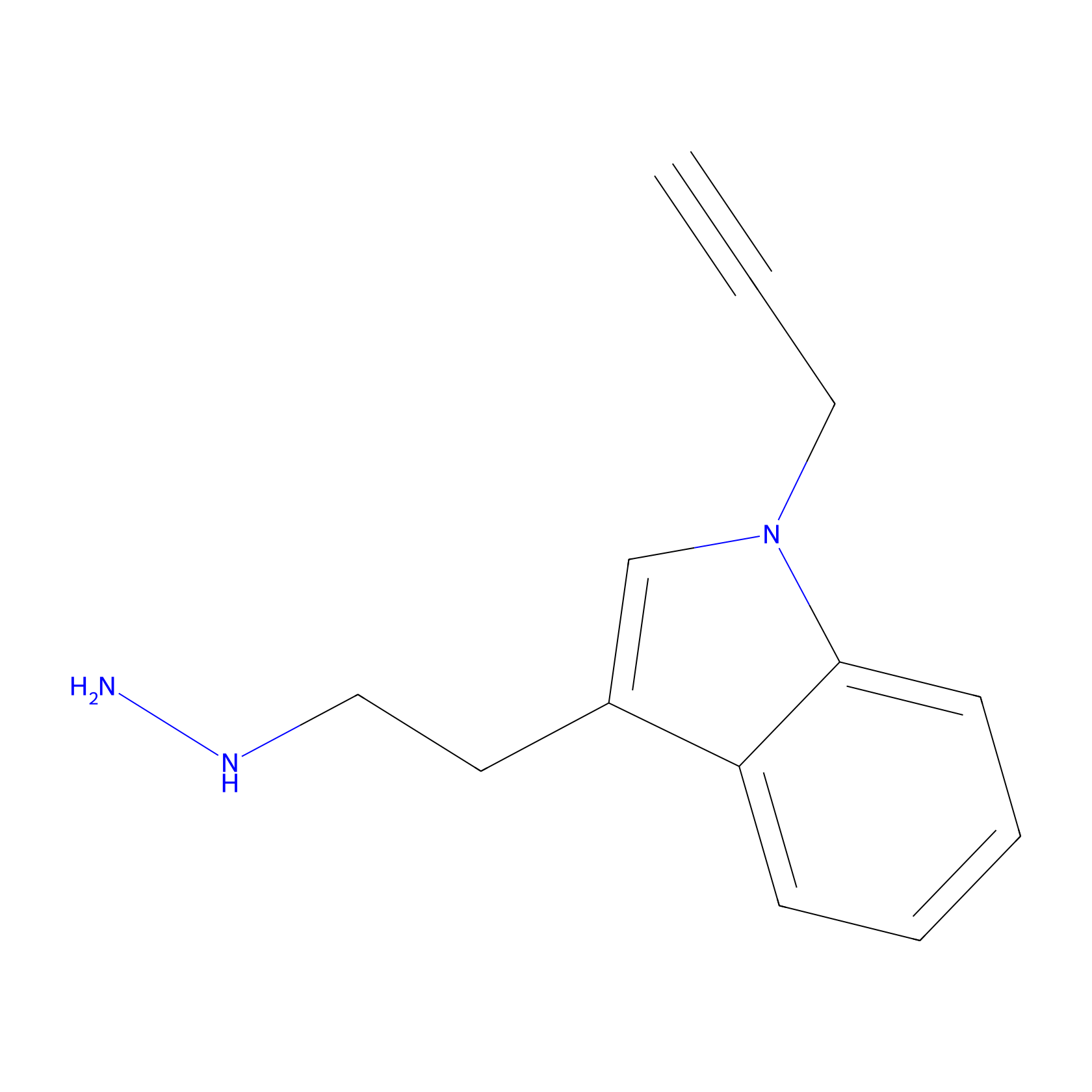 |
20.00 | LDD0201 | [4] | |
|
Alkylaryl probe 3 Probe Info |
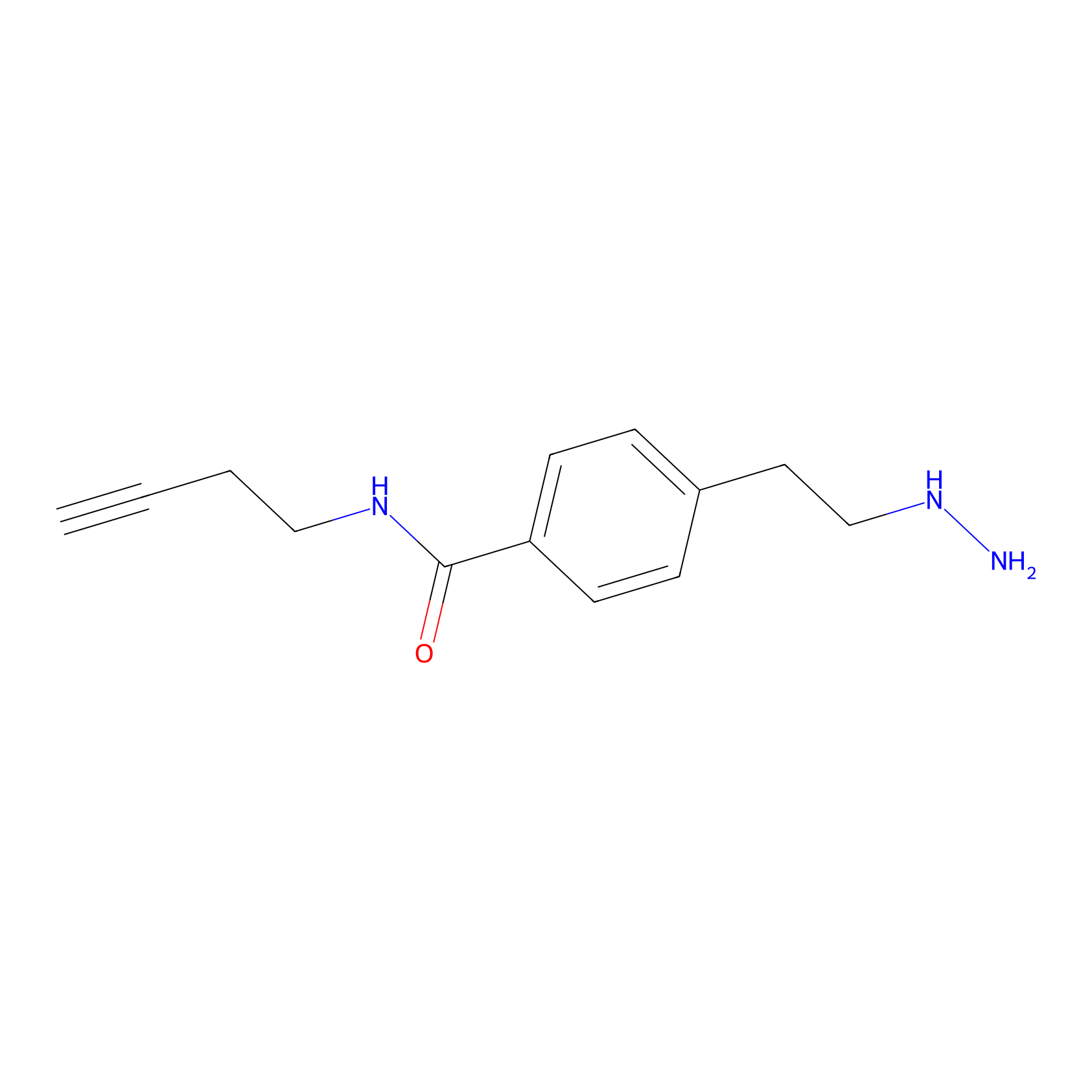 |
3.13 | LDD0282 | [5] | |
|
Johansson_61 Probe Info |
 |
_(10.81) | LDD1487 | [6] | |
|
DBIA Probe Info |
 |
C387(8.30) | LDD0209 | [7] | |
|
IA-alkyne Probe Info |
 |
C202(0.00); C67(0.00); C180(0.00) | LDD0165 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [9] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [10] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [11] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [11] | |
|
AOyne Probe Info |
 |
14.60 | LDD0443 | [12] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C024 Probe Info |
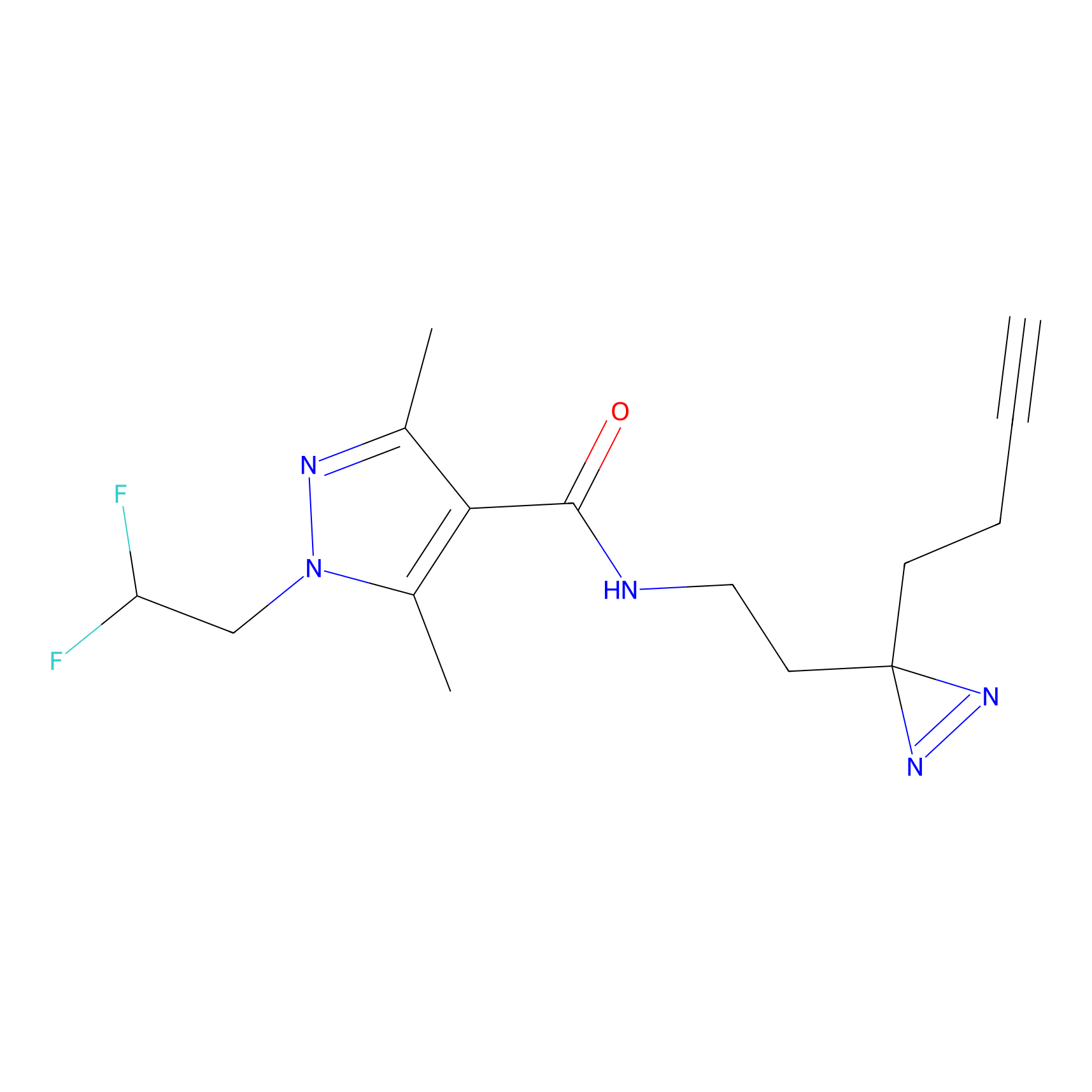 |
5.46 | LDD1730 | [13] | |
|
C138 Probe Info |
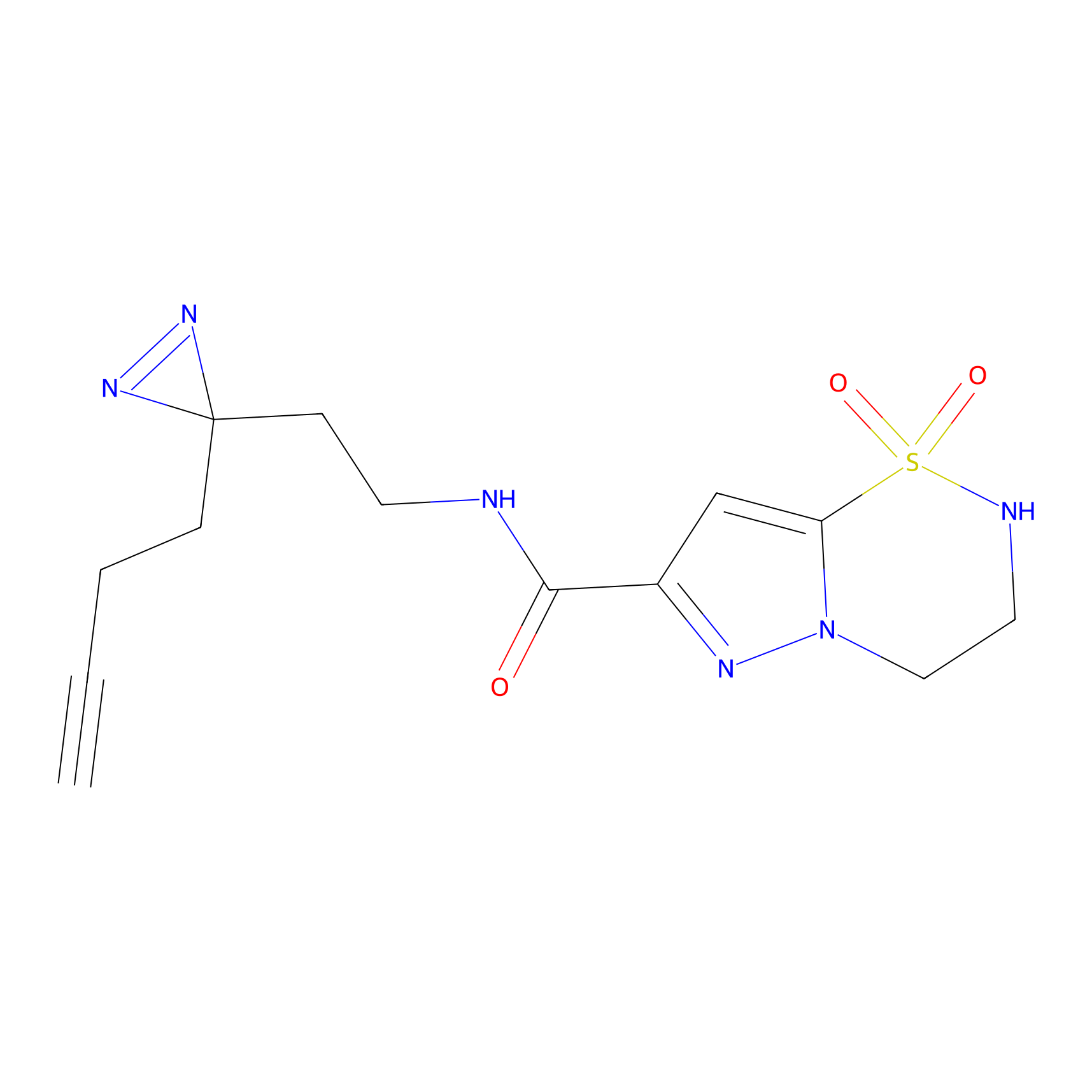 |
5.58 | LDD1820 | [13] | |
|
C145 Probe Info |
 |
10.34 | LDD1827 | [13] | |
|
C147 Probe Info |
 |
11.24 | LDD1829 | [13] | |
|
C234 Probe Info |
 |
5.58 | LDD1907 | [13] | |
|
C347 Probe Info |
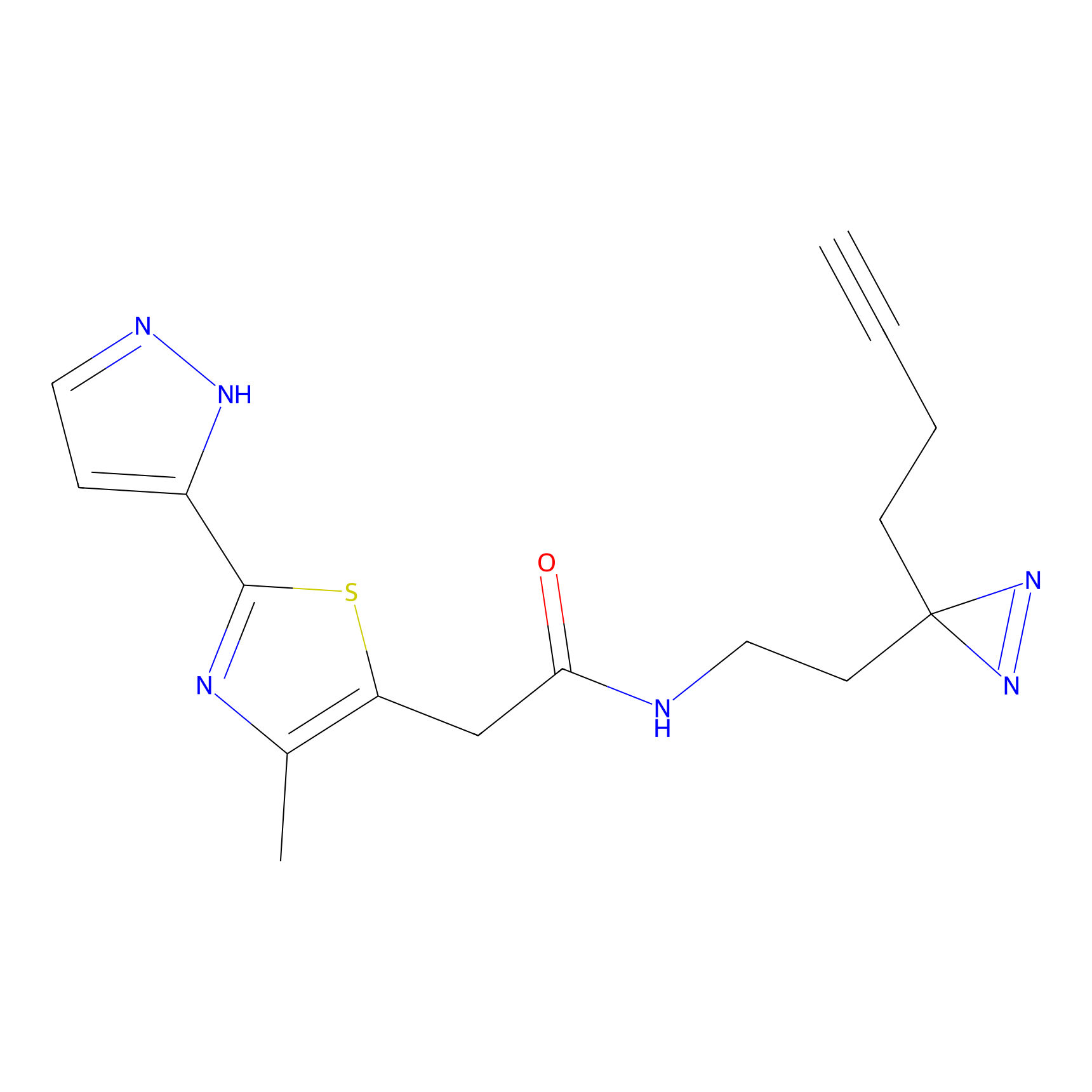 |
5.03 | LDD2008 | [13] | |
|
C353 Probe Info |
 |
15.24 | LDD2014 | [13] | |
|
C354 Probe Info |
 |
12.04 | LDD2015 | [13] | |
|
C355 Probe Info |
 |
22.63 | LDD2016 | [13] | |
|
C361 Probe Info |
 |
32.90 | LDD2022 | [13] | |
|
C430 Probe Info |
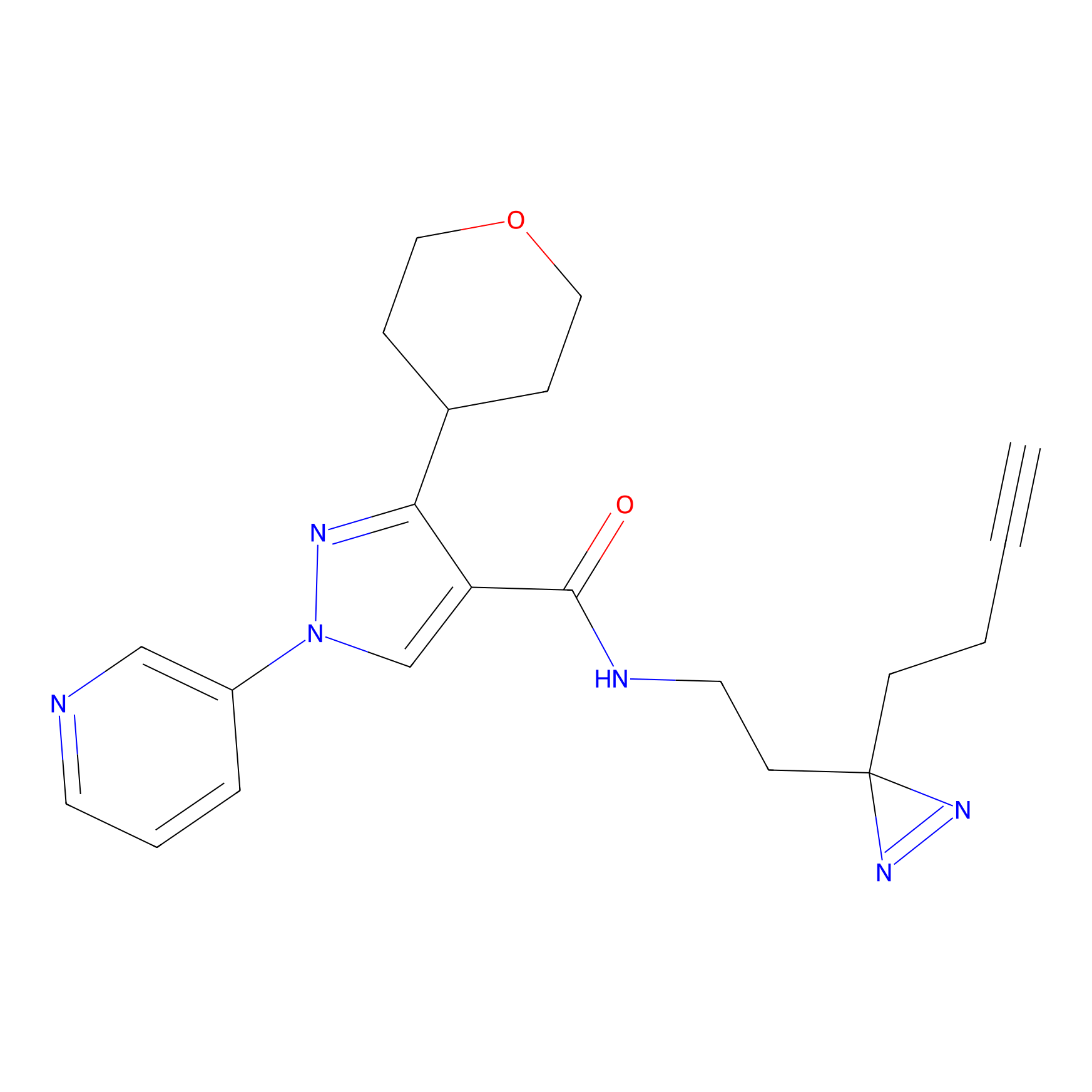 |
9.99 | LDD2085 | [13] | |
|
C432 Probe Info |
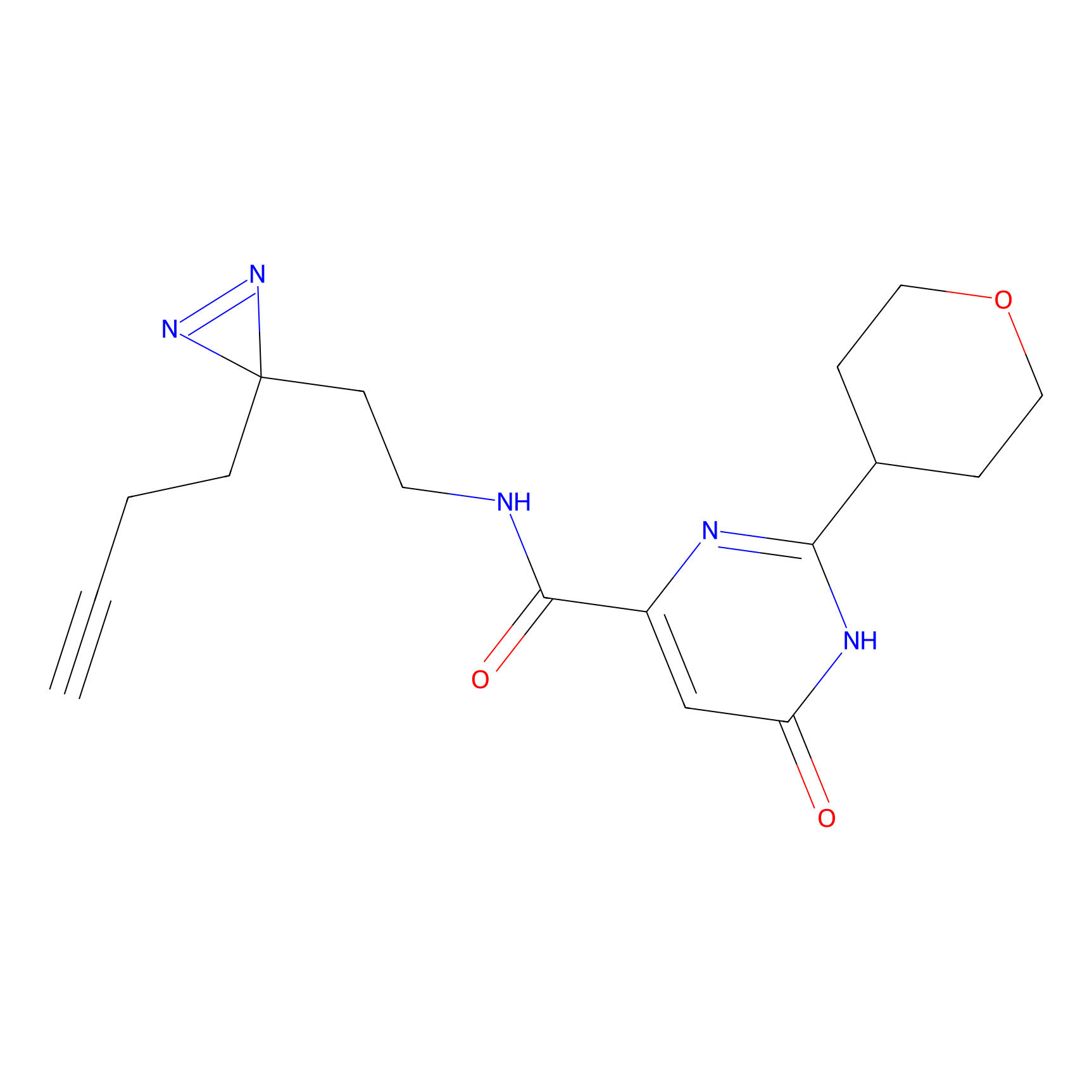 |
8.94 | LDD2087 | [13] | |
|
C433 Probe Info |
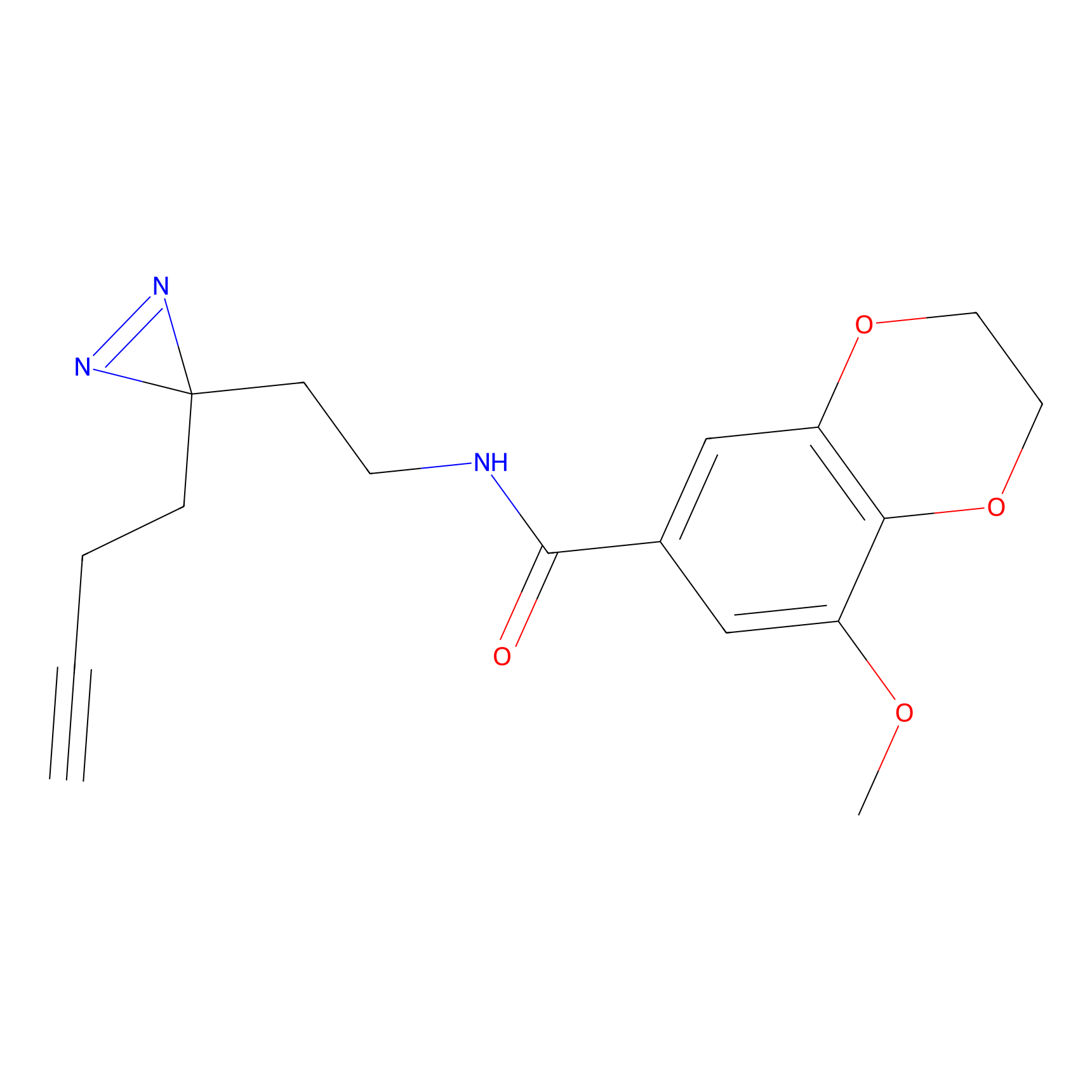 |
8.51 | LDD2088 | [13] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [14] | |
|
AEA-DA Probe Info |
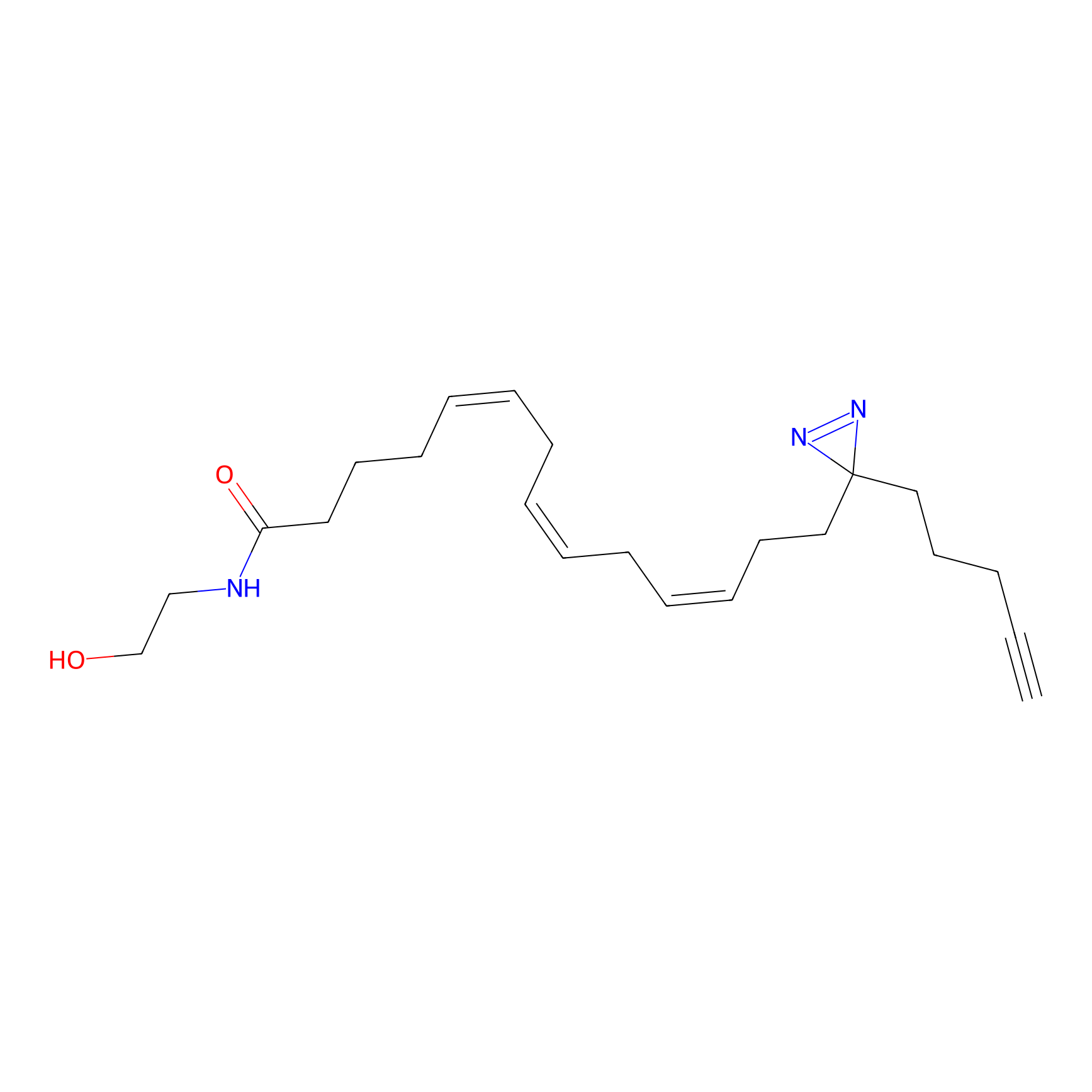 |
3.29 | LDD0333 | [15] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C320(1.09) | LDD1507 | [16] |
| LDCM0215 | AC10 | HEK-293T | C320(1.02) | LDD1508 | [16] |
| LDCM0226 | AC11 | HEK-293T | C320(1.11) | LDD1509 | [16] |
| LDCM0237 | AC12 | HEK-293T | C320(1.08) | LDD1510 | [16] |
| LDCM0259 | AC14 | HEK-293T | C320(0.95) | LDD1512 | [16] |
| LDCM0270 | AC15 | HEK-293T | C320(0.86) | LDD1513 | [16] |
| LDCM0276 | AC17 | HEK-293T | C320(1.10) | LDD1515 | [16] |
| LDCM0277 | AC18 | HEK-293T | C320(1.34) | LDD1516 | [16] |
| LDCM0278 | AC19 | HEK-293T | C320(1.02) | LDD1517 | [16] |
| LDCM0279 | AC2 | HEK-293T | C320(0.92) | LDD1518 | [16] |
| LDCM0280 | AC20 | HEK-293T | C320(0.93) | LDD1519 | [16] |
| LDCM0281 | AC21 | HEK-293T | C473(1.03); C320(0.90) | LDD1520 | [16] |
| LDCM0282 | AC22 | HEK-293T | C320(0.91) | LDD1521 | [16] |
| LDCM0283 | AC23 | HEK-293T | C320(1.04) | LDD1522 | [16] |
| LDCM0284 | AC24 | HEK-293T | C67(1.25) | LDD1523 | [16] |
| LDCM0285 | AC25 | HEK-293T | C320(1.07) | LDD1524 | [16] |
| LDCM0286 | AC26 | HEK-293T | C320(1.07) | LDD1525 | [16] |
| LDCM0287 | AC27 | HEK-293T | C320(1.17) | LDD1526 | [16] |
| LDCM0288 | AC28 | HEK-293T | C320(1.41) | LDD1527 | [16] |
| LDCM0289 | AC29 | HEK-293T | C473(0.98); C320(0.96) | LDD1528 | [16] |
| LDCM0290 | AC3 | HEK-293T | C320(1.36) | LDD1529 | [16] |
| LDCM0291 | AC30 | HEK-293T | C320(1.10) | LDD1530 | [16] |
| LDCM0292 | AC31 | HEK-293T | C320(0.99) | LDD1531 | [16] |
| LDCM0293 | AC32 | HEK-293T | C67(1.11) | LDD1532 | [16] |
| LDCM0294 | AC33 | HEK-293T | C320(0.96) | LDD1533 | [16] |
| LDCM0295 | AC34 | HEK-293T | C320(0.99) | LDD1534 | [16] |
| LDCM0296 | AC35 | HEK-293T | C320(1.33) | LDD1535 | [16] |
| LDCM0297 | AC36 | HEK-293T | C320(0.86) | LDD1536 | [16] |
| LDCM0298 | AC37 | HEK-293T | C473(0.86); C320(1.05) | LDD1537 | [16] |
| LDCM0299 | AC38 | HEK-293T | C320(1.07) | LDD1538 | [16] |
| LDCM0300 | AC39 | HEK-293T | C320(1.07) | LDD1539 | [16] |
| LDCM0301 | AC4 | HEK-293T | C320(0.89) | LDD1540 | [16] |
| LDCM0302 | AC40 | HEK-293T | C67(1.08) | LDD1541 | [16] |
| LDCM0303 | AC41 | HEK-293T | C320(1.17) | LDD1542 | [16] |
| LDCM0304 | AC42 | HEK-293T | C320(1.07) | LDD1543 | [16] |
| LDCM0305 | AC43 | HEK-293T | C320(1.12) | LDD1544 | [16] |
| LDCM0306 | AC44 | HEK-293T | C320(1.06) | LDD1545 | [16] |
| LDCM0307 | AC45 | HEK-293T | C473(0.86); C320(0.97) | LDD1546 | [16] |
| LDCM0308 | AC46 | HEK-293T | C320(1.32) | LDD1547 | [16] |
| LDCM0309 | AC47 | HEK-293T | C320(0.84) | LDD1548 | [16] |
| LDCM0310 | AC48 | HEK-293T | C67(1.05) | LDD1549 | [16] |
| LDCM0311 | AC49 | HEK-293T | C320(1.07) | LDD1550 | [16] |
| LDCM0312 | AC5 | HEK-293T | C473(0.75); C320(0.87) | LDD1551 | [16] |
| LDCM0313 | AC50 | HEK-293T | C320(1.01) | LDD1552 | [16] |
| LDCM0314 | AC51 | HEK-293T | C320(1.54) | LDD1553 | [16] |
| LDCM0315 | AC52 | HEK-293T | C320(0.86) | LDD1554 | [16] |
| LDCM0316 | AC53 | HEK-293T | C473(0.81); C320(0.92) | LDD1555 | [16] |
| LDCM0317 | AC54 | HEK-293T | C320(0.96) | LDD1556 | [16] |
| LDCM0318 | AC55 | HEK-293T | C320(1.10) | LDD1557 | [16] |
| LDCM0319 | AC56 | HEK-293T | C67(1.04) | LDD1558 | [16] |
| LDCM0320 | AC57 | HEK-293T | C320(1.10) | LDD1559 | [16] |
| LDCM0321 | AC58 | HEK-293T | C320(0.94) | LDD1560 | [16] |
| LDCM0322 | AC59 | HEK-293T | C320(1.58) | LDD1561 | [16] |
| LDCM0323 | AC6 | HEK-293T | C320(1.31) | LDD1562 | [16] |
| LDCM0324 | AC60 | HEK-293T | C320(1.31) | LDD1563 | [16] |
| LDCM0325 | AC61 | HEK-293T | C473(0.99); C320(0.88) | LDD1564 | [16] |
| LDCM0326 | AC62 | HEK-293T | C320(1.05) | LDD1565 | [16] |
| LDCM0327 | AC63 | HEK-293T | C320(1.22) | LDD1566 | [16] |
| LDCM0328 | AC64 | HEK-293T | C67(1.01) | LDD1567 | [16] |
| LDCM0334 | AC7 | HEK-293T | C320(0.86) | LDD1568 | [16] |
| LDCM0345 | AC8 | HEK-293T | C67(1.18) | LDD1569 | [16] |
| LDCM0248 | AKOS034007472 | HEK-293T | C473(0.93); C320(0.96) | LDD1511 | [16] |
| LDCM0356 | AKOS034007680 | HEK-293T | C320(1.19) | LDD1570 | [16] |
| LDCM0275 | AKOS034007705 | HEK-293T | C67(1.03) | LDD1514 | [16] |
| LDCM0088 | C45 | HEK-293T | 20.00 | LDD0201 | [4] |
| LDCM0368 | CL10 | HEK-293T | C320(2.44) | LDD1572 | [16] |
| LDCM0379 | CL11 | HEK-293T | C320(1.56) | LDD1583 | [16] |
| LDCM0390 | CL12 | HEK-293T | C67(1.15) | LDD1594 | [16] |
| LDCM0404 | CL17 | HEK-293T | C320(1.59) | LDD1608 | [16] |
| LDCM0405 | CL18 | HEK-293T | C320(1.56) | LDD1609 | [16] |
| LDCM0406 | CL19 | HEK-293T | C320(1.98) | LDD1610 | [16] |
| LDCM0408 | CL20 | HEK-293T | C320(1.18) | LDD1612 | [16] |
| LDCM0409 | CL21 | HEK-293T | C473(0.76); C320(1.12) | LDD1613 | [16] |
| LDCM0410 | CL22 | HEK-293T | C320(2.32) | LDD1614 | [16] |
| LDCM0411 | CL23 | HEK-293T | C320(1.31) | LDD1615 | [16] |
| LDCM0412 | CL24 | HEK-293T | C67(1.00) | LDD1616 | [16] |
| LDCM0417 | CL29 | HEK-293T | C320(0.94) | LDD1621 | [16] |
| LDCM0419 | CL30 | HEK-293T | C320(1.03) | LDD1623 | [16] |
| LDCM0420 | CL31 | HEK-293T | C320(2.07) | LDD1624 | [16] |
| LDCM0421 | CL32 | HEK-293T | C320(1.53) | LDD1625 | [16] |
| LDCM0422 | CL33 | HEK-293T | C473(0.87); C320(1.00) | LDD1626 | [16] |
| LDCM0423 | CL34 | HEK-293T | C320(1.75) | LDD1627 | [16] |
| LDCM0424 | CL35 | HEK-293T | C320(1.46) | LDD1628 | [16] |
| LDCM0425 | CL36 | HEK-293T | C67(1.12) | LDD1629 | [16] |
| LDCM0431 | CL41 | HEK-293T | C320(1.91) | LDD1635 | [16] |
| LDCM0432 | CL42 | HEK-293T | C320(1.58) | LDD1636 | [16] |
| LDCM0433 | CL43 | HEK-293T | C320(1.54) | LDD1637 | [16] |
| LDCM0434 | CL44 | HEK-293T | C320(1.46) | LDD1638 | [16] |
| LDCM0435 | CL45 | HEK-293T | C473(0.66); C320(0.86) | LDD1639 | [16] |
| LDCM0436 | CL46 | HEK-293T | C320(1.79) | LDD1640 | [16] |
| LDCM0437 | CL47 | HEK-293T | C320(1.90) | LDD1641 | [16] |
| LDCM0438 | CL48 | HEK-293T | C67(1.21) | LDD1642 | [16] |
| LDCM0440 | CL5 | HEK-293T | C320(1.83) | LDD1644 | [16] |
| LDCM0444 | CL53 | HEK-293T | C320(1.60) | LDD1647 | [16] |
| LDCM0445 | CL54 | HEK-293T | C320(1.20) | LDD1648 | [16] |
| LDCM0446 | CL55 | HEK-293T | C320(3.03) | LDD1649 | [16] |
| LDCM0447 | CL56 | HEK-293T | C320(1.39) | LDD1650 | [16] |
| LDCM0448 | CL57 | HEK-293T | C473(0.72); C320(0.87) | LDD1651 | [16] |
| LDCM0449 | CL58 | HEK-293T | C320(1.81) | LDD1652 | [16] |
| LDCM0450 | CL59 | HEK-293T | C320(1.45) | LDD1653 | [16] |
| LDCM0451 | CL6 | HEK-293T | C320(1.53) | LDD1654 | [16] |
| LDCM0452 | CL60 | HEK-293T | C67(1.17) | LDD1655 | [16] |
| LDCM0457 | CL65 | HEK-293T | C320(1.18) | LDD1660 | [16] |
| LDCM0458 | CL66 | HEK-293T | C320(1.63) | LDD1661 | [16] |
| LDCM0459 | CL67 | HEK-293T | C320(1.41) | LDD1662 | [16] |
| LDCM0460 | CL68 | HEK-293T | C320(1.22) | LDD1663 | [16] |
| LDCM0461 | CL69 | HEK-293T | C473(0.93); C320(1.04) | LDD1664 | [16] |
| LDCM0462 | CL7 | HEK-293T | C320(1.54) | LDD1665 | [16] |
| LDCM0463 | CL70 | HEK-293T | C320(1.91) | LDD1666 | [16] |
| LDCM0464 | CL71 | HEK-293T | C320(1.14) | LDD1667 | [16] |
| LDCM0465 | CL72 | HEK-293T | C67(1.10) | LDD1668 | [16] |
| LDCM0470 | CL77 | HEK-293T | C320(1.62) | LDD1673 | [16] |
| LDCM0471 | CL78 | HEK-293T | C320(1.53) | LDD1674 | [16] |
| LDCM0472 | CL79 | HEK-293T | C320(1.70) | LDD1675 | [16] |
| LDCM0473 | CL8 | HEK-293T | C320(1.39) | LDD1676 | [16] |
| LDCM0474 | CL80 | HEK-293T | C320(0.94) | LDD1677 | [16] |
| LDCM0475 | CL81 | HEK-293T | C473(0.90); C320(1.34) | LDD1678 | [16] |
| LDCM0476 | CL82 | HEK-293T | C320(1.80) | LDD1679 | [16] |
| LDCM0477 | CL83 | HEK-293T | C320(1.97) | LDD1680 | [16] |
| LDCM0478 | CL84 | HEK-293T | C67(0.87) | LDD1681 | [16] |
| LDCM0483 | CL89 | HEK-293T | C320(1.83) | LDD1686 | [16] |
| LDCM0484 | CL9 | HEK-293T | C473(0.84); C320(1.17) | LDD1687 | [16] |
| LDCM0485 | CL90 | HEK-293T | C320(1.65) | LDD1688 | [16] |
| LDCM0486 | CL91 | HEK-293T | C320(1.22) | LDD1689 | [16] |
| LDCM0487 | CL92 | HEK-293T | C320(1.38) | LDD1690 | [16] |
| LDCM0488 | CL93 | HEK-293T | C473(0.95); C320(1.03) | LDD1691 | [16] |
| LDCM0489 | CL94 | HEK-293T | C320(1.13) | LDD1692 | [16] |
| LDCM0490 | CL95 | HEK-293T | C320(1.13) | LDD1693 | [16] |
| LDCM0491 | CL96 | HEK-293T | C67(1.15) | LDD1694 | [16] |
| LDCM0615 | Fragment63-R | Jurkat | _(10.81) | LDD1487 | [6] |
| LDCM0617 | Fragment63-S | Jurkat | _(20.00) | LDD1490 | [6] |
| LDCM0023 | KB03 | Jurkat | C387(8.30) | LDD0209 | [7] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C319(1.12) | LDD2207 | [17] |
| LDCM0099 | Phenelzine | HEK-293T | 3.13 | LDD0282 | [5] |
| LDCM0136 | SR-4559 | HEK-293T | 3.29 | LDD0333 | [15] |
The Interaction Atlas With This Target
References
