Details of the Target
General Information of Target
| Target ID | LDTP11161 | |||||
|---|---|---|---|---|---|---|
| Target Name | Extended synaptotagmin-1 (ESYT1) | |||||
| Gene Name | ESYT1 | |||||
| Gene ID | 23344 | |||||
| Synonyms |
FAM62A; KIAA0747; MBC2; Extended synaptotagmin-1; E-Syt1; Membrane-bound C2 domain-containing protein |
|||||
| 3D Structure | ||||||
| Sequence |
MASRWQNMGTSVRRRSLQHQEQLEDSKELQPVVSHQETSVGALGSLCRQFQRRLPLRAVN
LNLRAGPSWKRLETPEPGQQGLQAAARSAKSALGAVSQRIQESCQSGTKWLVETQVKARR RKRGAQKGSGSPTHSLSQKSTRLSGAAPAHSAADPWEKEHHRLSVRMGSHAHPLRRSRRE AAFRSPYSSTEPLCSPSESDSDLEPVGAGIQHLQKLSQELDEAIMAEERKQALSDRQGFI LKDVYASP |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Extended synaptotagmin family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Binds glycerophospholipids in a barrel-like domain and may play a role in cellular lipid transport. Binds calcium (via the C2 domains) and translocates to sites of contact between the endoplasmic reticulum and the cell membrane in response to increased cytosolic calcium levels. Helps tether the endoplasmic reticulum to the cell membrane and promotes the formation of appositions between the endoplasmic reticulum and the cell membrane.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CAL120 | Deletion: p.S12_P13del | DBIA Probe Info | |||
| HCT116 | Deletion: p.R771VfsTer7 | . | |||
| HT | SNV: p.L1033P | DBIA Probe Info | |||
| HT3 | SNV: p.C734G | DBIA Probe Info | |||
| JURKAT | SNV: p.Q381Ter | Compound 10 Probe Info | |||
| L428 | SNV: p.A528V | DBIA Probe Info | |||
| MDAMB453 | SNV: p.S581C; p.L852F | DBIA Probe Info | |||
| MM1S | SNV: p.T739S | . | |||
| MOLT4 | SNV: p.Q903H | IA-alkyne Probe Info | |||
| SBC5 | SNV: p.A614S | DBIA Probe Info | |||
| SNU1 | SNV: p.S935R | DBIA Probe Info | |||
| TOV21G | SNV: p.A630V | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
13.15 | LDD0402 | [1] | |
|
A-EBA Probe Info |
 |
2.70 | LDD0215 | [2] | |
|
DA-P2 Probe Info |
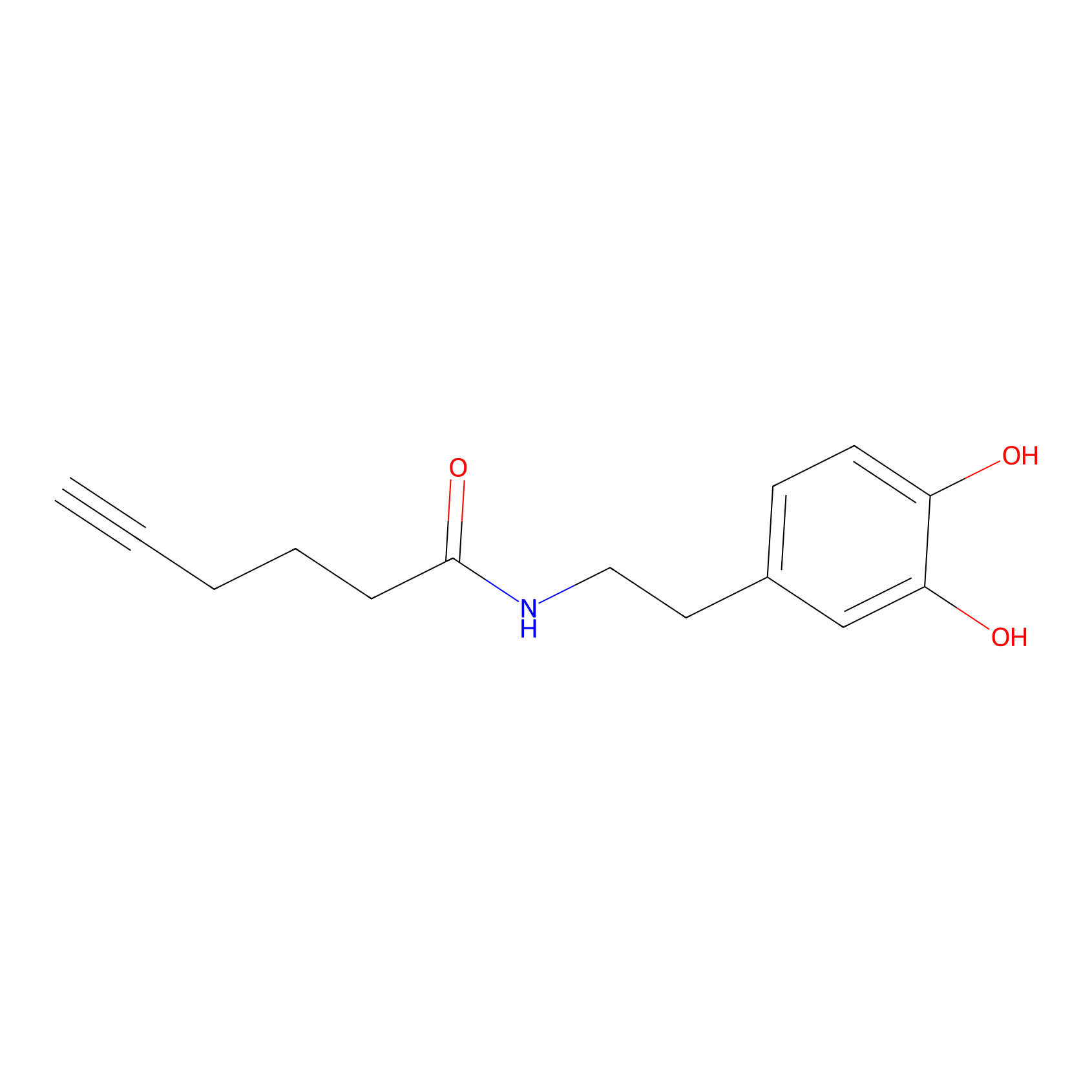 |
4.99 | LDD0348 | [3] | |
|
HRP Probe Info |
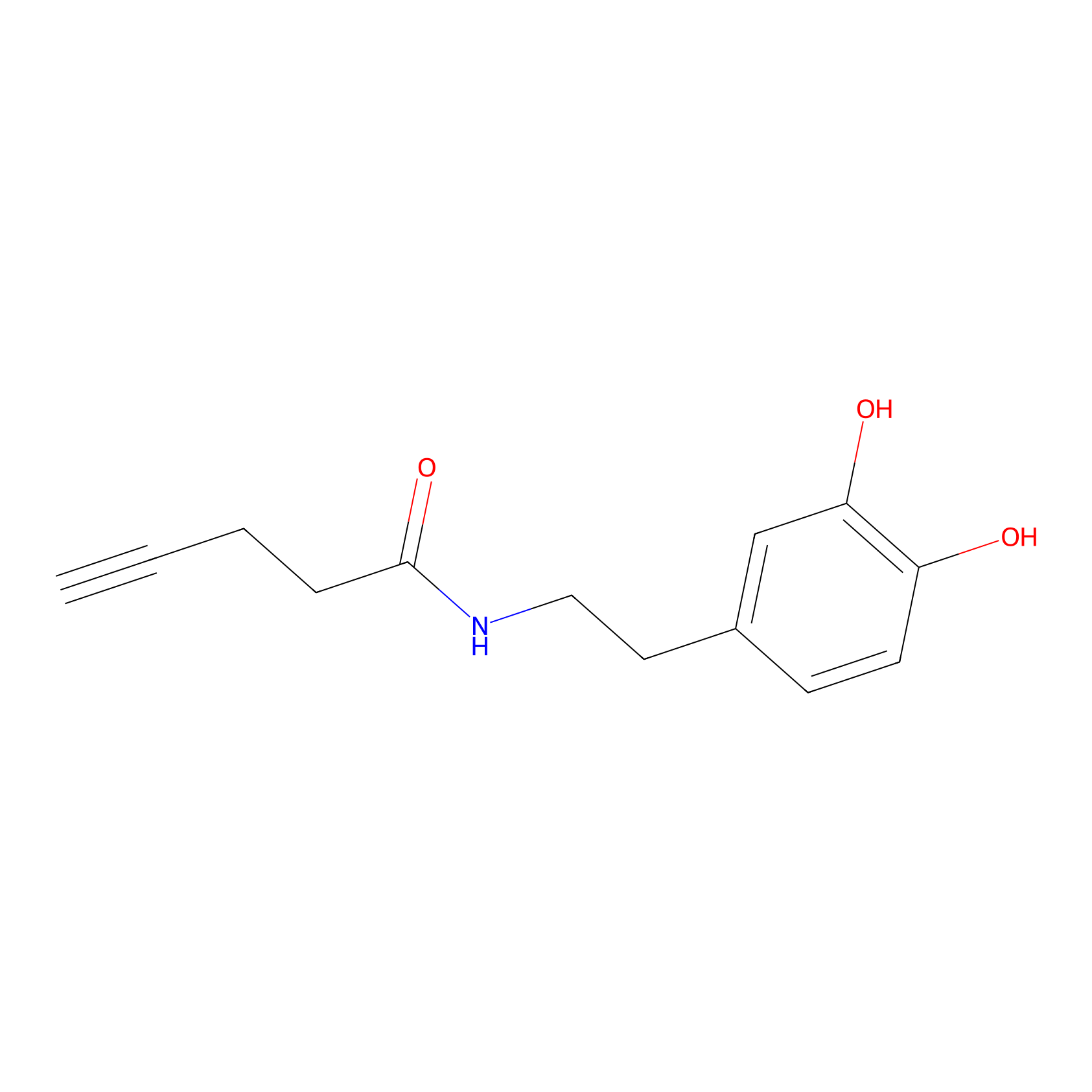 |
4.02 | LDD0347 | [3] | |
|
TH211 Probe Info |
 |
Y121(12.22); Y518(20.00); Y803(8.50); Y822(6.85) | LDD0260 | [4] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [5] | |
|
ONAyne Probe Info |
 |
K197(0.58); K344(10.00); K662(8.33) | LDD0274 | [6] | |
|
STPyne Probe Info |
 |
K1018(8.33); K1054(10.00); K1060(6.14); K118(10.00) | LDD0277 | [6] | |
|
OPA-S-S-alkyne Probe Info |
 |
K197(5.31) | LDD3494 | [7] | |
|
DBIA Probe Info |
 |
C532(2.36) | LDD3311 | [8] | |
|
P12 Probe Info |
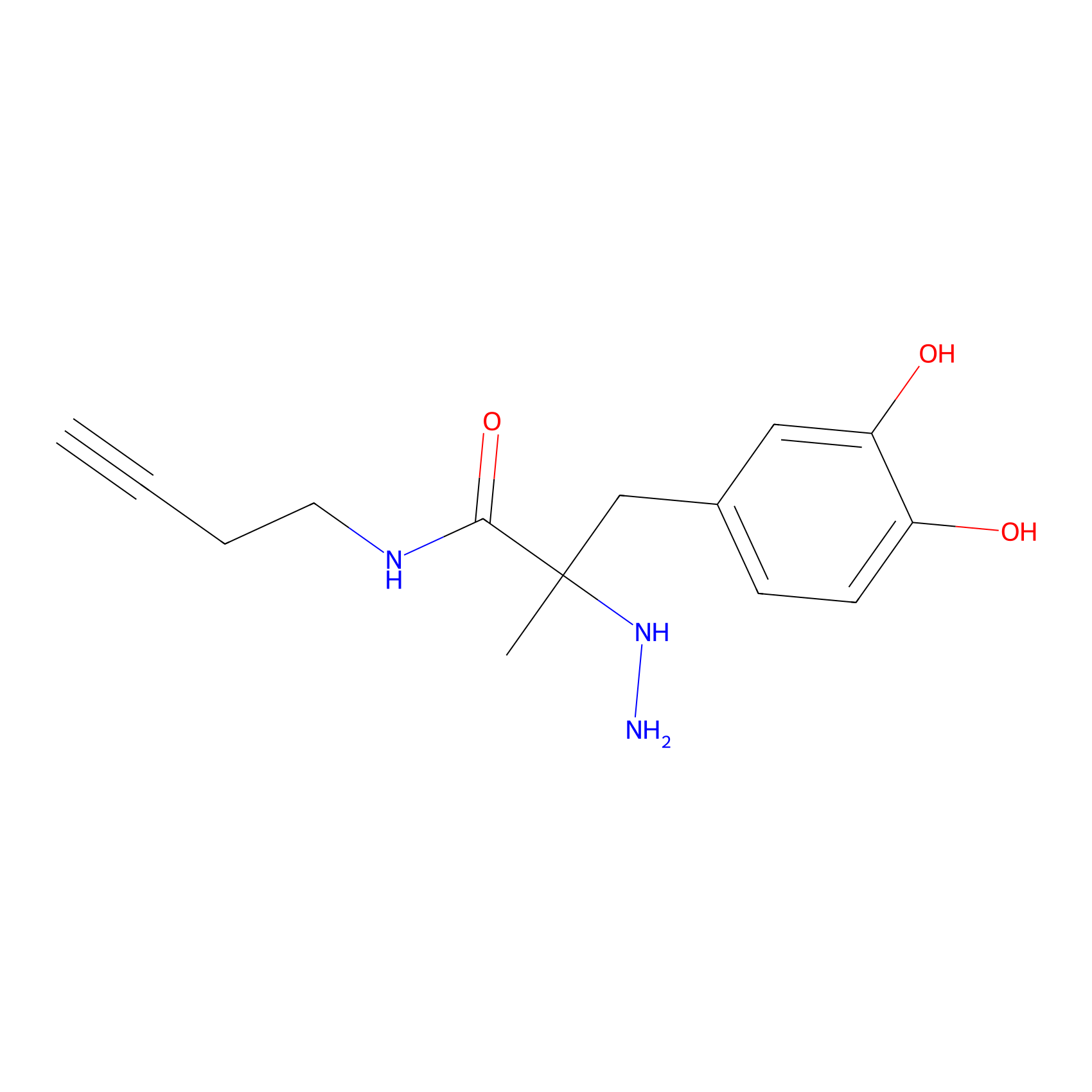 |
10.62 | LDD0202 | [9] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [10] | |
|
BTD Probe Info |
 |
C370(1.90) | LDD1700 | [11] | |
|
Sulforaphane-probe2 Probe Info |
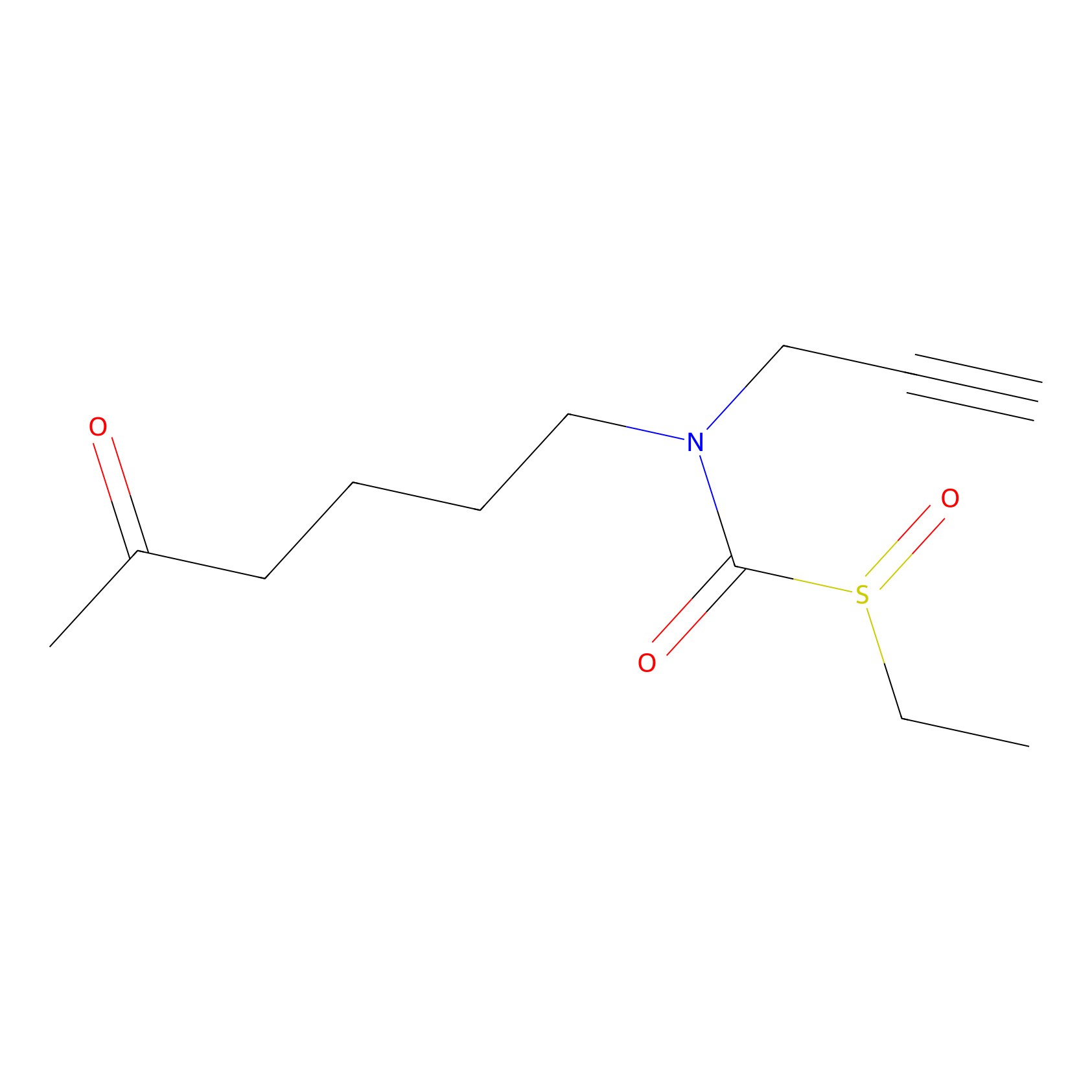 |
1.36 | LDD0160 | [12] | |
|
DA-P3 Probe Info |
 |
7.18 | LDD0179 | [3] | |
|
AHL-Pu-1 Probe Info |
 |
C635(4.35); C611(3.59); C604(4.21) | LDD0171 | [13] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [14] | |
|
ATP probe Probe Info |
 |
K1060(0.00); K662(0.00); K197(0.00); K671(0.00) | LDD0199 | [15] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C522(0.00); C890(0.00); C635(0.00); C370(0.00) | LDD0038 | [16] | |
|
IA-alkyne Probe Info |
 |
C522(0.00); C635(0.00) | LDD0032 | [17] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [18] | |
|
Lodoacetamide azide Probe Info |
 |
C522(0.00); C890(0.00); C995(0.00); C370(0.00) | LDD0037 | [16] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [19] | |
|
TFBX Probe Info |
 |
C370(0.00); C995(0.00) | LDD0027 | [19] | |
|
WYneN Probe Info |
 |
C370(0.00); C635(0.00) | LDD0021 | [20] | |
|
aHNE Probe Info |
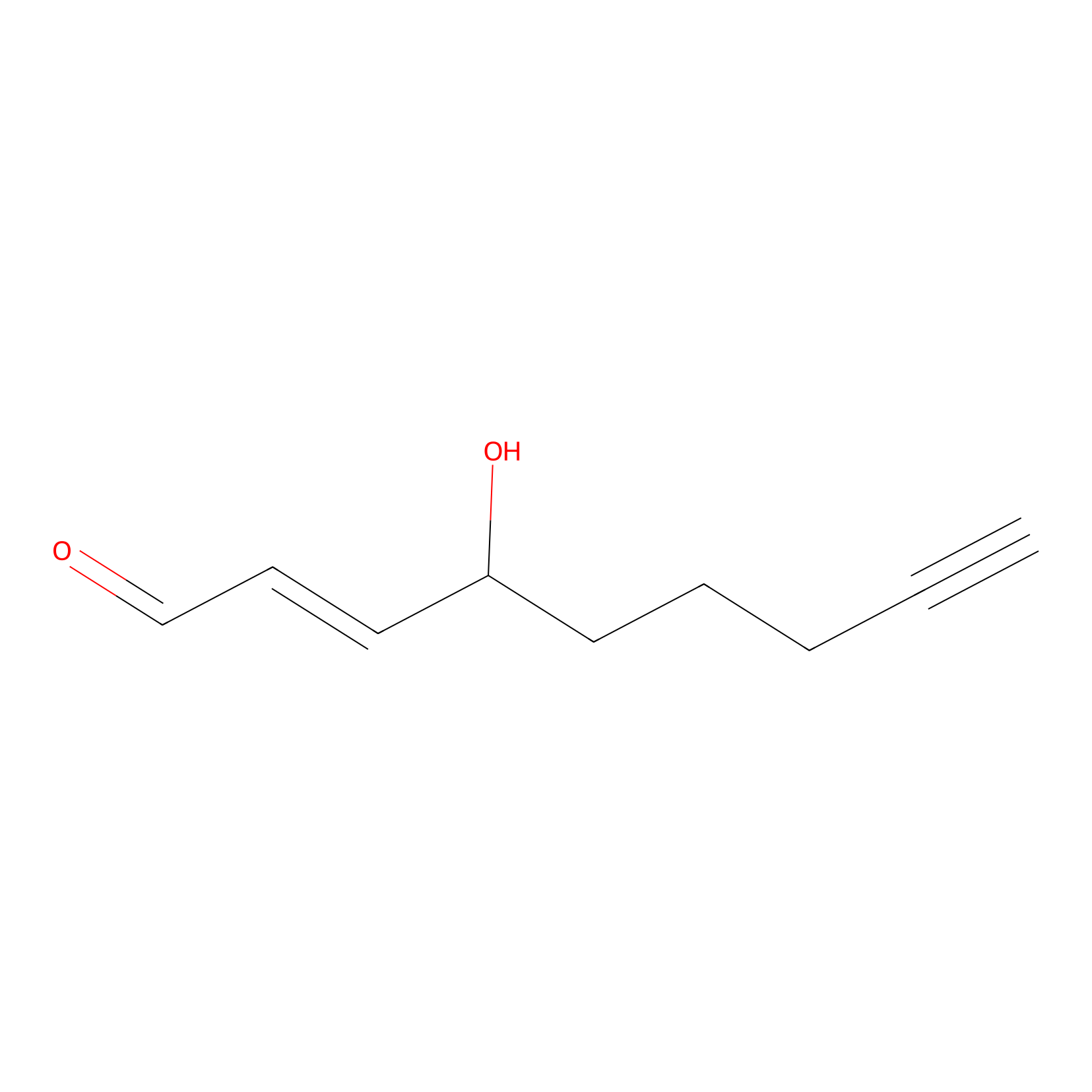 |
N.A. | LDD0001 | [20] | |
|
Compound 10 Probe Info |
 |
C370(0.00); C522(0.00) | LDD2216 | [21] | |
|
Compound 11 Probe Info |
 |
C370(0.00); C522(0.00) | LDD2213 | [21] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [20] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [22] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [23] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [20] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [24] | |
|
Ox-W18 Probe Info |
 |
W750(0.00); W1043(0.00) | LDD2175 | [25] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [26] | |
|
Acrolein Probe Info |
 |
C522(0.00); C635(0.00); H652(0.00) | LDD0217 | [27] | |
|
Crotonaldehyde Probe Info |
 |
C522(0.00); C370(0.00); C995(0.00) | LDD0219 | [27] | |
|
Methacrolein Probe Info |
 |
C370(0.00); C995(0.00) | LDD0218 | [27] | |
|
W1 Probe Info |
 |
C370(0.00); C522(0.00) | LDD0236 | [28] | |
|
AOyne Probe Info |
 |
5.00 | LDD0443 | [29] | |
|
NAIA_5 Probe Info |
 |
C522(0.00); C890(0.00); C995(0.00); C370(0.00) | LDD2223 | [30] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [31] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe11 Probe Info |
 |
10.26 | LDD0471 | [32] | |
|
FFF probe12 Probe Info |
 |
12.37 | LDD0473 | [32] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [32] | |
|
FFF probe14 Probe Info |
 |
12.87 | LDD0477 | [32] | |
|
FFF probe2 Probe Info |
 |
17.00 | LDD0463 | [32] | |
|
FFF probe3 Probe Info |
 |
15.90 | LDD0464 | [32] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [33] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [33] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [34] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0072 | [35] | |
|
OEA-DA Probe Info |
 |
5.10 | LDD0046 | [36] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C370(0.85); C995(1.17) | LDD2117 | [11] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C370(1.36) | LDD2152 | [11] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C635(4.35); C611(3.59); C604(4.21) | LDD0171 | [13] |
| LDCM0214 | AC1 | HEK-293T | C635(0.91); C370(0.90); C995(1.04); C522(0.98) | LDD1507 | [37] |
| LDCM0215 | AC10 | HEK-293T | C370(0.82); C995(0.96); C522(0.98); C890(1.00) | LDD1508 | [37] |
| LDCM0226 | AC11 | HEK-293T | C635(0.99); C370(1.09); C995(0.95); C522(1.00) | LDD1509 | [37] |
| LDCM0237 | AC12 | HEK-293T | C635(0.98); C370(0.98); C995(1.07); C522(0.97) | LDD1510 | [37] |
| LDCM0259 | AC14 | HEK-293T | C635(1.06); C370(0.93); C995(1.05); C522(0.98) | LDD1512 | [37] |
| LDCM0270 | AC15 | HEK-293T | C635(0.97); C370(0.79); C995(1.04); C522(0.98) | LDD1513 | [37] |
| LDCM0276 | AC17 | HEK-293T | C635(0.95); C370(0.84); C995(1.05); C522(0.95) | LDD1515 | [37] |
| LDCM0277 | AC18 | HEK-293T | C370(0.95); C995(0.97); C522(0.92); C890(0.80) | LDD1516 | [37] |
| LDCM0278 | AC19 | HEK-293T | C635(0.88); C370(0.96); C995(0.99); C522(1.05) | LDD1517 | [37] |
| LDCM0279 | AC2 | HEK-293T | C370(1.04); C995(0.98); C522(1.06); C890(1.18) | LDD1518 | [37] |
| LDCM0280 | AC20 | HEK-293T | C635(1.04); C370(0.95); C995(0.97); C522(1.01) | LDD1519 | [37] |
| LDCM0281 | AC21 | HEK-293T | C635(0.86); C370(0.81); C995(1.00); C522(1.03) | LDD1520 | [37] |
| LDCM0282 | AC22 | HEK-293T | C635(1.07); C370(0.97); C995(1.05); C522(0.94) | LDD1521 | [37] |
| LDCM0283 | AC23 | HEK-293T | C635(1.05); C370(0.85); C995(0.99); C522(0.98) | LDD1522 | [37] |
| LDCM0284 | AC24 | HEK-293T | C635(1.01); C370(0.99); C995(1.01); C522(1.05) | LDD1523 | [37] |
| LDCM0285 | AC25 | HEK-293T | C635(1.03); C370(0.86); C995(1.07); C522(1.01) | LDD1524 | [37] |
| LDCM0286 | AC26 | HEK-293T | C370(0.92); C995(1.02); C522(1.08); C890(0.79) | LDD1525 | [37] |
| LDCM0287 | AC27 | HEK-293T | C635(1.01); C370(1.05); C995(0.86); C522(1.00) | LDD1526 | [37] |
| LDCM0288 | AC28 | HEK-293T | C635(0.89); C370(1.01); C995(0.96); C522(0.95) | LDD1527 | [37] |
| LDCM0289 | AC29 | HEK-293T | C635(1.00); C370(0.84); C995(1.03); C522(0.94) | LDD1528 | [37] |
| LDCM0290 | AC3 | HEK-293T | C635(0.98); C370(1.08); C995(0.85); C522(1.08) | LDD1529 | [37] |
| LDCM0291 | AC30 | HEK-293T | C635(1.01); C370(1.03); C995(1.04); C522(1.01) | LDD1530 | [37] |
| LDCM0292 | AC31 | HEK-293T | C635(0.91); C370(0.87); C995(1.02); C522(0.95) | LDD1531 | [37] |
| LDCM0293 | AC32 | HEK-293T | C635(1.09); C370(1.02); C995(1.00); C522(1.13) | LDD1532 | [37] |
| LDCM0294 | AC33 | HEK-293T | C635(1.08); C370(0.89); C995(1.07); C522(1.03) | LDD1533 | [37] |
| LDCM0295 | AC34 | HEK-293T | C370(0.89); C995(0.96); C522(0.99); C890(0.99) | LDD1534 | [37] |
| LDCM0296 | AC35 | HEK-293T | C635(1.13); C370(1.00); C995(0.99); C522(1.05) | LDD1535 | [37] |
| LDCM0297 | AC36 | HEK-293T | C635(1.17); C370(0.95); C995(0.95); C522(1.07) | LDD1536 | [37] |
| LDCM0298 | AC37 | HEK-293T | C635(1.05); C370(0.86); C995(0.99); C522(1.06) | LDD1537 | [37] |
| LDCM0299 | AC38 | HEK-293T | C635(0.97); C370(0.93); C995(1.00); C522(0.96) | LDD1538 | [37] |
| LDCM0300 | AC39 | HEK-293T | C635(0.88); C370(0.79); C995(1.01); C522(1.03) | LDD1539 | [37] |
| LDCM0301 | AC4 | HEK-293T | C635(0.96); C370(1.07); C995(1.06); C522(1.01) | LDD1540 | [37] |
| LDCM0302 | AC40 | HEK-293T | C635(1.01); C370(1.00); C995(1.02); C522(0.99) | LDD1541 | [37] |
| LDCM0303 | AC41 | HEK-293T | C635(1.06); C370(0.83); C995(1.03); C522(0.92) | LDD1542 | [37] |
| LDCM0304 | AC42 | HEK-293T | C370(0.87); C995(1.01); C522(0.92); C890(1.02) | LDD1543 | [37] |
| LDCM0305 | AC43 | HEK-293T | C635(1.09); C370(1.08); C995(0.94); C522(1.03) | LDD1544 | [37] |
| LDCM0306 | AC44 | HEK-293T | C635(0.84); C370(0.97); C995(0.98); C522(1.16) | LDD1545 | [37] |
| LDCM0307 | AC45 | HEK-293T | C635(1.08); C370(0.76); C995(1.04); C522(0.96) | LDD1546 | [37] |
| LDCM0308 | AC46 | HEK-293T | C635(0.96); C370(0.93); C995(1.01); C522(1.04) | LDD1547 | [37] |
| LDCM0309 | AC47 | HEK-293T | C635(0.97); C370(0.79); C995(1.00); C522(1.05) | LDD1548 | [37] |
| LDCM0310 | AC48 | HEK-293T | C635(1.07); C370(0.99); C995(1.05); C522(1.01) | LDD1549 | [37] |
| LDCM0311 | AC49 | HEK-293T | C635(0.81); C370(0.70); C995(1.03); C522(1.03) | LDD1550 | [37] |
| LDCM0312 | AC5 | HEK-293T | C635(0.95); C370(1.02); C995(0.96); C522(0.94) | LDD1551 | [37] |
| LDCM0313 | AC50 | HEK-293T | C370(0.84); C995(0.98); C522(1.05); C890(0.95) | LDD1552 | [37] |
| LDCM0314 | AC51 | HEK-293T | C635(0.95); C370(1.06); C995(0.89); C522(1.07) | LDD1553 | [37] |
| LDCM0315 | AC52 | HEK-293T | C635(0.91); C370(0.97); C995(0.98); C522(0.96) | LDD1554 | [37] |
| LDCM0316 | AC53 | HEK-293T | C635(0.93); C370(0.70); C995(0.99); C522(0.96) | LDD1555 | [37] |
| LDCM0317 | AC54 | HEK-293T | C635(0.95); C370(0.93); C995(0.89); C522(1.01) | LDD1556 | [37] |
| LDCM0318 | AC55 | HEK-293T | C635(0.80); C370(0.68); C995(0.96); C522(1.08) | LDD1557 | [37] |
| LDCM0319 | AC56 | HEK-293T | C635(0.98); C370(1.01); C995(0.96); C522(0.99) | LDD1558 | [37] |
| LDCM0320 | AC57 | HEK-293T | C635(0.86); C370(0.96); C995(0.95); C522(1.09) | LDD1559 | [37] |
| LDCM0321 | AC58 | HEK-293T | C370(1.01); C995(0.88); C522(1.00); C890(0.86) | LDD1560 | [37] |
| LDCM0322 | AC59 | HEK-293T | C635(0.92); C370(1.09); C995(0.91); C522(1.04) | LDD1561 | [37] |
| LDCM0323 | AC6 | HEK-293T | C635(1.08); C370(0.98); C995(0.99); C522(1.06) | LDD1562 | [37] |
| LDCM0324 | AC60 | HEK-293T | C635(0.93); C370(1.04); C995(0.97); C522(0.98) | LDD1563 | [37] |
| LDCM0325 | AC61 | HEK-293T | C635(1.00); C370(0.93); C995(0.88); C522(0.99) | LDD1564 | [37] |
| LDCM0326 | AC62 | HEK-293T | C635(0.86); C370(1.01); C995(0.89); C522(0.99) | LDD1565 | [37] |
| LDCM0327 | AC63 | HEK-293T | C635(0.92); C370(1.06); C995(0.95); C522(1.06) | LDD1566 | [37] |
| LDCM0328 | AC64 | HEK-293T | C635(0.98); C370(1.01); C995(0.85); C522(0.97) | LDD1567 | [37] |
| LDCM0334 | AC7 | HEK-293T | C635(0.86); C370(0.95); C995(0.94); C522(1.04) | LDD1568 | [37] |
| LDCM0345 | AC8 | HEK-293T | C635(1.10); C370(0.96); C995(0.92); C522(1.03) | LDD1569 | [37] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C995(0.71) | LDD2113 | [11] |
| LDCM0248 | AKOS034007472 | HEK-293T | C635(0.94); C370(0.78); C995(0.97); C522(0.96) | LDD1511 | [37] |
| LDCM0356 | AKOS034007680 | HEK-293T | C635(0.95); C370(0.84); C995(1.05); C522(0.95) | LDD1570 | [37] |
| LDCM0275 | AKOS034007705 | HEK-293T | C635(1.02); C370(0.95); C995(0.99); C522(1.01) | LDD1514 | [37] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C370(0.58) | LDD2091 | [11] |
| LDCM0088 | C45 | HEK-293T | 10.62 | LDD0202 | [9] |
| LDCM0087 | Capsaicin | HEK-293T | 14.41 | LDD0185 | [3] |
| LDCM0630 | CCW28-3 | 231MFP | C611(1.45) | LDD2214 | [38] |
| LDCM0108 | Chloroacetamide | HeLa | C522(0.00); C995(0.00); H652(0.00); K855(0.00) | LDD0222 | [27] |
| LDCM0367 | CL1 | HEK-293T | C635(0.97); C370(0.92); C995(1.06); C522(1.25) | LDD1571 | [37] |
| LDCM0368 | CL10 | HEK-293T | C635(1.33); C370(0.79); C995(1.61); C522(1.06) | LDD1572 | [37] |
| LDCM0369 | CL100 | HEK-293T | C635(1.07); C370(0.96); C995(1.03); C522(1.00) | LDD1573 | [37] |
| LDCM0370 | CL101 | HEK-293T | C635(0.94); C370(0.96); C995(0.91); C522(1.12) | LDD1574 | [37] |
| LDCM0371 | CL102 | HEK-293T | C370(0.97); C995(1.05); C522(1.02); C611(1.19) | LDD1575 | [37] |
| LDCM0372 | CL103 | HEK-293T | C635(1.05); C370(0.76); C995(1.01); C522(1.03) | LDD1576 | [37] |
| LDCM0373 | CL104 | HEK-293T | C635(0.98); C370(0.80); C995(1.07); C522(1.01) | LDD1577 | [37] |
| LDCM0374 | CL105 | HEK-293T | C635(0.97); C370(1.04); C995(0.94); C522(1.14) | LDD1578 | [37] |
| LDCM0375 | CL106 | HEK-293T | C370(0.99); C995(0.93); C522(1.04); C611(1.37) | LDD1579 | [37] |
| LDCM0376 | CL107 | HEK-293T | C635(0.89); C370(0.73); C995(1.05); C522(0.98) | LDD1580 | [37] |
| LDCM0377 | CL108 | HEK-293T | C635(0.99); C370(0.78); C995(1.03); C522(0.96) | LDD1581 | [37] |
| LDCM0378 | CL109 | HEK-293T | C635(0.99); C370(1.05); C995(1.03); C522(1.10) | LDD1582 | [37] |
| LDCM0379 | CL11 | HEK-293T | C635(0.99); C370(1.02); C995(1.48); C522(1.26) | LDD1583 | [37] |
| LDCM0380 | CL110 | HEK-293T | C370(0.97); C995(1.22); C522(1.13); C611(0.83) | LDD1584 | [37] |
| LDCM0381 | CL111 | HEK-293T | C635(1.11); C370(0.87); C995(1.09); C522(1.04) | LDD1585 | [37] |
| LDCM0382 | CL112 | HEK-293T | C635(1.06); C370(0.81); C995(1.10); C522(1.03) | LDD1586 | [37] |
| LDCM0383 | CL113 | HEK-293T | C635(0.92); C370(1.00); C995(0.93); C522(1.02) | LDD1587 | [37] |
| LDCM0384 | CL114 | HEK-293T | C370(1.04); C995(1.04); C522(1.04); C611(1.69) | LDD1588 | [37] |
| LDCM0385 | CL115 | HEK-293T | C635(1.08); C370(0.82); C995(1.05); C522(1.02) | LDD1589 | [37] |
| LDCM0386 | CL116 | HEK-293T | C635(1.04); C370(0.82); C995(1.02); C522(0.97) | LDD1590 | [37] |
| LDCM0387 | CL117 | HEK-293T | C635(0.84); C370(1.02); C995(0.95); C522(0.96) | LDD1591 | [37] |
| LDCM0388 | CL118 | HEK-293T | C370(0.91); C995(0.95); C522(1.07); C611(0.97) | LDD1592 | [37] |
| LDCM0389 | CL119 | HEK-293T | C635(1.01); C370(0.76); C995(1.04); C522(1.02) | LDD1593 | [37] |
| LDCM0390 | CL12 | HEK-293T | C635(1.31); C370(0.87); C995(1.39); C522(1.03) | LDD1594 | [37] |
| LDCM0391 | CL120 | HEK-293T | C635(1.04); C370(0.92); C995(1.13); C522(1.03) | LDD1595 | [37] |
| LDCM0392 | CL121 | HEK-293T | C635(0.80); C370(1.10); C995(0.94); C522(1.08) | LDD1596 | [37] |
| LDCM0393 | CL122 | HEK-293T | C370(0.91); C995(0.92); C522(1.02); C611(0.91) | LDD1597 | [37] |
| LDCM0394 | CL123 | HEK-293T | C635(0.80); C370(0.72); C995(0.97); C522(1.09) | LDD1598 | [37] |
| LDCM0395 | CL124 | HEK-293T | C635(0.99); C370(0.61); C995(0.94); C522(1.06) | LDD1599 | [37] |
| LDCM0396 | CL125 | HEK-293T | C635(0.59); C370(1.03); C995(0.77); C522(1.11) | LDD1600 | [37] |
| LDCM0397 | CL126 | HEK-293T | C370(1.05); C995(0.82); C522(1.01); C611(1.29) | LDD1601 | [37] |
| LDCM0398 | CL127 | HEK-293T | C635(0.78); C370(1.04); C995(0.88); C522(1.10) | LDD1602 | [37] |
| LDCM0399 | CL128 | HEK-293T | C635(1.01); C370(0.99); C995(0.88); C522(0.92) | LDD1603 | [37] |
| LDCM0400 | CL13 | HEK-293T | C635(0.91); C370(0.91); C995(0.99); C522(0.95) | LDD1604 | [37] |
| LDCM0401 | CL14 | HEK-293T | C370(1.01); C995(0.92); C522(0.98); C611(1.14) | LDD1605 | [37] |
| LDCM0402 | CL15 | HEK-293T | C635(0.90); C370(0.77); C995(1.02); C522(1.17) | LDD1606 | [37] |
| LDCM0403 | CL16 | HEK-293T | C635(1.04); C370(0.85); C995(0.95); C522(0.97) | LDD1607 | [37] |
| LDCM0404 | CL17 | HEK-293T | C635(0.85); C370(0.82); C995(1.15); C522(1.13) | LDD1608 | [37] |
| LDCM0405 | CL18 | HEK-293T | C370(0.80); C995(0.95); C522(1.04); C890(1.08) | LDD1609 | [37] |
| LDCM0406 | CL19 | HEK-293T | C635(1.02); C370(0.94); C995(0.83); C522(1.07) | LDD1610 | [37] |
| LDCM0407 | CL2 | HEK-293T | C370(0.98); C995(0.95); C522(1.01); C611(1.14) | LDD1611 | [37] |
| LDCM0408 | CL20 | HEK-293T | C635(0.98); C370(0.85); C995(1.21); C522(1.13) | LDD1612 | [37] |
| LDCM0409 | CL21 | HEK-293T | C635(1.00); C370(0.76); C995(1.28); C522(1.21) | LDD1613 | [37] |
| LDCM0410 | CL22 | HEK-293T | C635(1.07); C370(0.78); C995(1.48); C522(0.97) | LDD1614 | [37] |
| LDCM0411 | CL23 | HEK-293T | C635(1.04); C370(0.81); C995(1.38); C522(1.05) | LDD1615 | [37] |
| LDCM0412 | CL24 | HEK-293T | C635(1.16); C370(0.86); C995(1.28); C522(1.05) | LDD1616 | [37] |
| LDCM0413 | CL25 | HEK-293T | C635(0.73); C370(1.09); C995(0.93); C522(1.00) | LDD1617 | [37] |
| LDCM0414 | CL26 | HEK-293T | C370(0.99); C995(0.96); C522(0.93); C611(1.19) | LDD1618 | [37] |
| LDCM0415 | CL27 | HEK-293T | C635(0.92); C370(0.80); C995(1.08); C522(1.00) | LDD1619 | [37] |
| LDCM0416 | CL28 | HEK-293T | C635(1.01); C370(0.76); C995(1.02); C522(1.16) | LDD1620 | [37] |
| LDCM0417 | CL29 | HEK-293T | C635(0.94); C370(0.82); C995(1.10); C522(0.91) | LDD1621 | [37] |
| LDCM0418 | CL3 | HEK-293T | C635(0.99); C370(0.94); C995(1.02); C522(1.10) | LDD1622 | [37] |
| LDCM0419 | CL30 | HEK-293T | C370(0.84); C995(1.07); C522(0.99); C890(1.08) | LDD1623 | [37] |
| LDCM0420 | CL31 | HEK-293T | C635(0.87); C370(0.99); C995(1.04); C522(1.05) | LDD1624 | [37] |
| LDCM0421 | CL32 | HEK-293T | C635(0.82); C370(0.94); C995(1.09); C522(1.20) | LDD1625 | [37] |
| LDCM0422 | CL33 | HEK-293T | C635(0.86); C370(0.78); C995(1.26); C522(1.15) | LDD1626 | [37] |
| LDCM0423 | CL34 | HEK-293T | C635(0.96); C370(0.83); C995(1.28); C522(1.13) | LDD1627 | [37] |
| LDCM0424 | CL35 | HEK-293T | C635(0.94); C370(0.80); C995(1.45); C522(1.00) | LDD1628 | [37] |
| LDCM0425 | CL36 | HEK-293T | C635(1.15); C370(0.91); C995(1.35); C522(1.08) | LDD1629 | [37] |
| LDCM0426 | CL37 | HEK-293T | C635(0.76); C370(1.00); C995(0.91); C522(1.19) | LDD1630 | [37] |
| LDCM0428 | CL39 | HEK-293T | C635(1.13); C370(0.90); C995(1.08); C522(1.08) | LDD1632 | [37] |
| LDCM0429 | CL4 | HEK-293T | C635(1.02); C370(1.02); C995(0.96); C522(0.97) | LDD1633 | [37] |
| LDCM0430 | CL40 | HEK-293T | C635(0.95); C370(1.00); C995(0.95); C522(0.92) | LDD1634 | [37] |
| LDCM0431 | CL41 | HEK-293T | C635(1.10); C370(0.91); C995(1.12); C522(1.00) | LDD1635 | [37] |
| LDCM0432 | CL42 | HEK-293T | C370(0.98); C995(0.97); C522(0.95); C890(1.20) | LDD1636 | [37] |
| LDCM0433 | CL43 | HEK-293T | C635(1.06); C370(1.12); C995(1.06); C522(1.02) | LDD1637 | [37] |
| LDCM0434 | CL44 | HEK-293T | C635(1.01); C370(0.94); C995(1.17); C522(0.93) | LDD1638 | [37] |
| LDCM0435 | CL45 | HEK-293T | C635(1.04); C370(0.76); C995(1.17); C522(1.08) | LDD1639 | [37] |
| LDCM0436 | CL46 | HEK-293T | C635(1.09); C370(0.87); C995(1.39); C522(1.07) | LDD1640 | [37] |
| LDCM0437 | CL47 | HEK-293T | C635(1.10); C370(0.91); C995(1.32); C522(1.12) | LDD1641 | [37] |
| LDCM0438 | CL48 | HEK-293T | C635(1.13); C370(0.89); C995(1.34); C522(1.01) | LDD1642 | [37] |
| LDCM0439 | CL49 | HEK-293T | C635(0.72); C370(1.05); C995(0.94); C522(0.96) | LDD1643 | [37] |
| LDCM0440 | CL5 | HEK-293T | C635(0.80); C370(0.94); C995(1.05); C522(1.00) | LDD1644 | [37] |
| LDCM0441 | CL50 | HEK-293T | C370(1.01); C995(1.02); C522(0.93); C611(0.90) | LDD1645 | [37] |
| LDCM0443 | CL52 | HEK-293T | C635(1.03); C370(0.92); C995(1.20); C522(1.00) | LDD1646 | [37] |
| LDCM0444 | CL53 | HEK-293T | C635(0.96); C370(0.93); C995(1.13); C522(1.11) | LDD1647 | [37] |
| LDCM0445 | CL54 | HEK-293T | C370(0.68); C995(1.04); C522(1.15); C890(0.77) | LDD1648 | [37] |
| LDCM0446 | CL55 | HEK-293T | C635(0.99); C370(0.96); C995(0.96); C522(1.13) | LDD1649 | [37] |
| LDCM0447 | CL56 | HEK-293T | C635(1.02); C370(0.94); C995(1.26); C522(1.11) | LDD1650 | [37] |
| LDCM0448 | CL57 | HEK-293T | C635(1.02); C370(0.81); C995(1.39); C522(1.12) | LDD1651 | [37] |
| LDCM0449 | CL58 | HEK-293T | C635(0.95); C370(0.86); C995(1.50); C522(1.05) | LDD1652 | [37] |
| LDCM0450 | CL59 | HEK-293T | C635(0.98); C370(0.77); C995(1.32); C522(1.11) | LDD1653 | [37] |
| LDCM0451 | CL6 | HEK-293T | C370(1.01); C995(0.98); C522(1.12); C890(1.06) | LDD1654 | [37] |
| LDCM0452 | CL60 | HEK-293T | C635(1.30); C370(0.89); C995(1.33); C522(1.20) | LDD1655 | [37] |
| LDCM0453 | CL61 | HEK-293T | C635(0.87); C370(1.02); C995(0.85); C522(0.97) | LDD1656 | [37] |
| LDCM0454 | CL62 | HEK-293T | C370(1.07); C995(0.91); C522(1.04); C611(0.89) | LDD1657 | [37] |
| LDCM0455 | CL63 | HEK-293T | C635(0.92); C370(0.88); C995(1.03); C522(0.94) | LDD1658 | [37] |
| LDCM0456 | CL64 | HEK-293T | C635(1.04); C370(0.89); C995(1.10); C522(1.17) | LDD1659 | [37] |
| LDCM0457 | CL65 | HEK-293T | C635(1.03); C370(0.85); C995(1.04); C522(0.91) | LDD1660 | [37] |
| LDCM0458 | CL66 | HEK-293T | C370(0.92); C995(0.99); C522(1.13); C890(1.12) | LDD1661 | [37] |
| LDCM0459 | CL67 | HEK-293T | C635(0.97); C370(1.12); C995(0.90); C522(1.01) | LDD1662 | [37] |
| LDCM0460 | CL68 | HEK-293T | C635(0.89); C370(0.97); C995(1.14); C522(1.20) | LDD1663 | [37] |
| LDCM0461 | CL69 | HEK-293T | C635(1.05); C370(0.69); C995(1.11); C522(1.17) | LDD1664 | [37] |
| LDCM0462 | CL7 | HEK-293T | C635(0.96); C370(0.93); C995(0.97); C522(1.06) | LDD1665 | [37] |
| LDCM0463 | CL70 | HEK-293T | C635(1.05); C370(0.86); C995(1.37); C522(0.96) | LDD1666 | [37] |
| LDCM0464 | CL71 | HEK-293T | C635(0.88); C370(0.72); C995(1.29); C522(1.07) | LDD1667 | [37] |
| LDCM0465 | CL72 | HEK-293T | C635(1.12); C370(0.89); C995(1.24); C522(0.99) | LDD1668 | [37] |
| LDCM0466 | CL73 | HEK-293T | C635(0.80); C370(1.00); C995(0.95); C522(1.05) | LDD1669 | [37] |
| LDCM0467 | CL74 | HEK-293T | C370(0.94); C995(0.90); C522(1.06); C611(0.87) | LDD1670 | [37] |
| LDCM0469 | CL76 | HEK-293T | C635(1.03); C370(0.88); C995(0.86); C522(0.96) | LDD1672 | [37] |
| LDCM0470 | CL77 | HEK-293T | C635(0.82); C370(0.84); C995(1.09); C522(1.02) | LDD1673 | [37] |
| LDCM0471 | CL78 | HEK-293T | C370(0.84); C995(1.01); C522(0.98); C890(0.96) | LDD1674 | [37] |
| LDCM0472 | CL79 | HEK-293T | C635(0.95); C370(1.02); C995(0.94); C522(1.09) | LDD1675 | [37] |
| LDCM0473 | CL8 | HEK-293T | C635(0.74); C370(0.87); C995(1.20); C522(1.09) | LDD1676 | [37] |
| LDCM0474 | CL80 | HEK-293T | C635(0.82); C370(0.91); C995(1.11); C522(1.00) | LDD1677 | [37] |
| LDCM0475 | CL81 | HEK-293T | C635(1.00); C370(0.62); C995(1.02); C522(1.01) | LDD1678 | [37] |
| LDCM0476 | CL82 | HEK-293T | C635(0.96); C370(0.90); C995(1.26); C522(1.12) | LDD1679 | [37] |
| LDCM0477 | CL83 | HEK-293T | C635(0.99); C370(0.85); C995(1.26); C522(1.04) | LDD1680 | [37] |
| LDCM0478 | CL84 | HEK-293T | C635(0.95); C370(0.91); C995(1.12); C522(1.26) | LDD1681 | [37] |
| LDCM0479 | CL85 | HEK-293T | C635(0.70); C370(1.03); C995(0.81); C522(0.99) | LDD1682 | [37] |
| LDCM0480 | CL86 | HEK-293T | C370(1.04); C995(0.76); C522(1.13); C611(1.99) | LDD1683 | [37] |
| LDCM0481 | CL87 | HEK-293T | C635(0.74); C370(1.03); C995(0.83); C522(1.08) | LDD1684 | [37] |
| LDCM0482 | CL88 | HEK-293T | C635(1.05); C370(1.04); C995(0.87); C522(1.01) | LDD1685 | [37] |
| LDCM0483 | CL89 | HEK-293T | C635(0.67); C370(0.94); C995(0.83); C522(1.02) | LDD1686 | [37] |
| LDCM0484 | CL9 | HEK-293T | C635(1.11); C370(0.99); C995(1.19); C522(1.00) | LDD1687 | [37] |
| LDCM0485 | CL90 | HEK-293T | C370(0.92); C995(1.11); C522(1.10); C890(1.19) | LDD1688 | [37] |
| LDCM0486 | CL91 | HEK-293T | C635(0.74); C370(1.03); C995(0.78); C522(1.08) | LDD1689 | [37] |
| LDCM0487 | CL92 | HEK-293T | C635(0.78); C370(0.96); C995(0.98); C522(1.08) | LDD1690 | [37] |
| LDCM0488 | CL93 | HEK-293T | C635(0.80); C370(0.79); C995(1.08); C522(1.04) | LDD1691 | [37] |
| LDCM0489 | CL94 | HEK-293T | C635(1.18); C370(0.88); C995(1.16); C522(1.08) | LDD1692 | [37] |
| LDCM0490 | CL95 | HEK-293T | C635(0.84); C370(0.97); C995(1.27); C522(1.00) | LDD1693 | [37] |
| LDCM0491 | CL96 | HEK-293T | C635(0.78); C370(0.90); C995(0.80); C522(1.12) | LDD1694 | [37] |
| LDCM0492 | CL97 | HEK-293T | C635(0.91); C370(1.01); C995(0.99); C522(1.27) | LDD1695 | [37] |
| LDCM0493 | CL98 | HEK-293T | C370(1.03); C995(1.04); C522(1.03); C611(1.02) | LDD1696 | [37] |
| LDCM0494 | CL99 | HEK-293T | C635(1.04); C370(1.11); C995(0.97); C522(1.04) | LDD1697 | [37] |
| LDCM0028 | Dobutamine | HEK-293T | 6.02 | LDD0180 | [3] |
| LDCM0027 | Dopamine | HEK-293T | 7.18 | LDD0179 | [3] |
| LDCM0495 | E2913 | HEK-293T | C635(1.13); C370(0.88); C995(1.05); C522(1.01) | LDD1698 | [37] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C995(2.33); C370(2.04); C635(1.22) | LDD1702 | [11] |
| LDCM0031 | Epigallocatechin gallate | HEK-293T | 5.87 | LDD0183 | [3] |
| LDCM0625 | F8 | Ramos | C635(2.84) | LDD2187 | [39] |
| LDCM0572 | Fragment10 | Ramos | C635(1.09) | LDD2189 | [39] |
| LDCM0573 | Fragment11 | Ramos | C522(3.87); C635(5.58) | LDD2190 | [39] |
| LDCM0574 | Fragment12 | Ramos | C635(0.46) | LDD2191 | [39] |
| LDCM0576 | Fragment14 | Ramos | C522(1.27); C635(0.55) | LDD2193 | [39] |
| LDCM0579 | Fragment20 | Ramos | C635(0.40) | LDD2194 | [39] |
| LDCM0580 | Fragment21 | Ramos | C635(0.59) | LDD2195 | [39] |
| LDCM0582 | Fragment23 | Ramos | C635(1.11) | LDD2196 | [39] |
| LDCM0578 | Fragment27 | Ramos | C635(0.93) | LDD2197 | [39] |
| LDCM0586 | Fragment28 | Ramos | C635(0.66) | LDD2198 | [39] |
| LDCM0590 | Fragment32 | Ramos | C635(0.87) | LDD2201 | [39] |
| LDCM0468 | Fragment33 | HEK-293T | C635(0.91); C370(0.80); C995(0.96); C522(1.01) | LDD1671 | [37] |
| LDCM0596 | Fragment38 | Ramos | C635(0.77) | LDD2203 | [39] |
| LDCM0427 | Fragment51 | HEK-293T | C370(1.04); C995(0.98); C522(0.96); C611(1.26) | LDD1631 | [37] |
| LDCM0610 | Fragment52 | Ramos | C635(1.24) | LDD2204 | [39] |
| LDCM0569 | Fragment7 | Ramos | C522(0.77); C635(0.65) | LDD2186 | [39] |
| LDCM0571 | Fragment9 | Ramos | C635(0.45) | LDD2188 | [39] |
| LDCM0107 | IAA | HeLa | C522(0.00); H652(0.00); H960(0.00) | LDD0221 | [27] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [10] |
| LDCM0022 | KB02 | HEK-293T | C635(1.26); C370(1.00); C995(1.01); C522(0.79) | LDD1492 | [37] |
| LDCM0023 | KB03 | HEK-293T | C635(0.99); C370(0.99); C995(0.96); C522(0.90) | LDD1497 | [37] |
| LDCM0024 | KB05 | G361 | C532(2.36) | LDD3311 | [8] |
| LDCM0030 | Luteolin | HEK-293T | 14.42 | LDD0182 | [3] |
| LDCM0109 | NEM | HeLa | H652(0.00); H857(0.00); H960(0.00) | LDD0223 | [27] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C635(0.42) | LDD2089 | [11] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C370(1.63) | LDD2090 | [11] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C370(0.86); C635(0.96) | LDD2093 | [11] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C635(1.26) | LDD2094 | [11] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C370(0.97) | LDD2099 | [11] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C370(0.77) | LDD2101 | [11] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C370(1.85) | LDD2105 | [11] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C635(0.74) | LDD2106 | [11] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C370(0.91); C635(1.15); C995(1.14) | LDD2107 | [11] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C370(0.66); C995(0.71) | LDD2109 | [11] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C370(0.90) | LDD2111 | [11] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C370(0.47) | LDD2115 | [11] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C635(0.90) | LDD2116 | [11] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C370(0.49) | LDD2118 | [11] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C370(0.97) | LDD2120 | [11] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C635(1.14) | LDD2122 | [11] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C370(0.73); C635(0.97); C995(1.34) | LDD2123 | [11] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C370(0.22); C635(1.30) | LDD2124 | [11] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C370(0.74); C995(1.07) | LDD2125 | [11] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C370(0.18) | LDD2126 | [11] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C370(0.96) | LDD2127 | [11] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C370(0.99); C635(0.86) | LDD2129 | [11] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C370(0.38) | LDD2134 | [11] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C370(1.13); C635(1.12) | LDD2135 | [11] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C370(1.15) | LDD2136 | [11] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C370(0.97) | LDD2137 | [11] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C370(1.90) | LDD1700 | [11] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C995(0.94) | LDD2140 | [11] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C370(0.34) | LDD2141 | [11] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C370(0.97) | LDD2143 | [11] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C370(2.57) | LDD2144 | [11] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C995(1.10) | LDD2146 | [11] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C370(0.45) | LDD2148 | [11] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C370(0.35) | LDD2149 | [11] |
| LDCM0032 | Oleacein | HEK-293T | 5.27 | LDD0184 | [3] |
| LDCM0029 | Quercetin | HEK-293T | 12.83 | LDD0181 | [3] |
| LDCM0003 | Sulforaphane | MDA-MB-231 | 1.36 | LDD0160 | [12] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
References
