Details of the Target
General Information of Target
| Target ID | LDTP07205 | |||||
|---|---|---|---|---|---|---|
| Target Name | Telomere-associated protein RIF1 (RIF1) | |||||
| Gene Name | RIF1 | |||||
| Gene ID | 55183 | |||||
| Synonyms |
Telomere-associated protein RIF1; Rap1-interacting factor 1 homolog |
|||||
| 3D Structure | ||||||
| Sequence |
MTARGQSPLAPLLETLEDPSASHGGQTDAYLTLTSRMTGEEGKEVITEIEKKLPRLYKVL
KTHISSQNSELSSAALQALGFCLYNPKITSELSEANALELLSKLNDTIKNSDKNVRTRAL WVISKQTFPSEVVGKMVSSIIDSLEILFNKGETHSAVVDFEALNVIVRLIEQAPIQMGEE AVRWAKLVIPLVVHSAQKVHLRGATALEMGMPLLLQKQQEIASITEQLMTTKLISELQKL FMSKNETYVLKLWPLFVKLLGRTLHRSGSFINSLLQLEELGFRSGAPMIKKIAFIAWKSL IDNFALNPDILCSAKRLKLLMQPLSSIHVRTETLALTKLEVWWYLLMRLGPHLPANFEQV CVPLIQSTISIDSNASPQGNSCHVATSPGLNPMTPVHKGASSPYGAPGTPRMNLSSNLGG MATIPSIQLLGLEMLLHFLLGPEALSFAKQNKLVLSLEPLEHPLISSPSFFSKHANTLIT AVHDSFVAVGKDAPDVVVSAIWKELISLVKSVTESGNKKEKPGSEVLTLLLKSLESIVKS EVFPVSKTLVLMEITIKGLPQKVLGSPAYQVANMDILNGTPALFLIQLIFNNFLECGVSD ERFFLSLESLVGCVLSGPTSPLAFSDSVLNVINQNAKQLENKEHLWKMWSVIVTPLTELI NQTNEVNQGDALEHNFSAIYGALTLPVNHIFSEQRFPVATMKTLLRTWSELYRAFARCAA LVATAEENLCCEELSSKIMSSLEDEGFSNLLFVDRIIYIITVMVDCIDFSPYNIKYQPKV KSPQRPSDWSKKKNEPLGKLTSLFKLIVKVIYSFHTLSFKEAHSDTLFTIGNSITGIISS VLGHISLPSMIRKIFATLTRPLALFYENSKLDEVPKVYSCLNNKLEKLLGEIIACLQFSY TGTYDSELLEQLSPLLCIIFLHKNKQIRKQSAQFWNATFAKVMMLVYPEELKPVLTQAKQ KFLLLLPGLETVEMMEESSGPYSDGTENSQLNVKISGMERKSNGKRDSFLAQTKNKKENM KPAAKLKLESSSLKVKGEILLEEEKSTDFVFIPPEGKDAKERILTDHQKEVLKTKRCDIP AMYNNLDVSQDTLFTQYSQEEPMEIPTLTRKPKEDSKMMITEEQMDSDIVIPQDVTEDCG MAEHLEKSSLSNNECGSLDKTSPEMSNSNNDERKKALISSRKTSTECASSTENSFVVSSS SVSNTTVAGTPPYPTSRRQTFITLEKFDGSENRPFSPSPLNNISSTVTVKNNQETMIKTD FLPKAKQREGTFSKSDSEKIVNGTKRSSRRAGKAEQTGNKRSKPLMRSEPEKNTEESVEG IVVLENNPPGLLNQTECVSDNQVHLSESTMEHDNTKLKAATVENAVLLETNTVEEKNVEI NLESKENTPPVVISADQMVNEDSQVQITPNQKTLRRSSRRRSEVVESTTESQDKENSHQK KERRKEEEKPLQKSPLHIKDDVLPKQKLIAEQTLQENLIEKGSNLHEKTLGETSANAETE QNKKKADPENIKSEGDGTQDIVDKSSEKLVRGRTRYQTRRASQGLLSSIENSESDSSEAK EEGSRKKRSGKWKNKSNESVDIQDQEEKVVKQECIKAENQSHDYKATSEEDVSIKSPICE KQDESNTVICQDSTVTSDLLQVPDDLPNVCEEKNETSKYAEYSFTSLPVPESNLRTRNAI KRLHKRDSFDNCSLGESSKIGISDISSLSEKTFQTLECQHKRSRRVRRSKGCDCCGEKSQ PQEKSLIGLKNTENNDVEISETKKADVQAPVSPSETSQANPYSEGQFLDEHHSVNFHLGL KEDNDTINDSLIVSETKSKENTMQESLPSGIVNFREEICDMDSSEAMSLESQESPNENFK TVGPCLGDSKNVSQESLETKEEKPEETPKMELSLENVTVEGNACKVTESNLEKAKTMELN VGNEASFHGQERTKTGISEEAAIEENKRNDDSEADTAKLNAKEVATEEFNSDISLSDNTT PVKLNAQTEISEQTAAGELDGGNDVSDLHSSEETNTKMKNNEEMMIGEAMAETGHDGETE NEGITTKTSKPDEAETNMLTAEMDNFVCDTVEMSTEEGIIDANKTETNTEYSKSEEKLDN NQMVMESDILQEDHHTSQKVEEPSQCLASGTAISELIIEDNNASPQKLRELDPSLVSAND SPSGMQTRCVWSPLASPSTSILKRGLKRSQEDEISSPVNKVRRVSFADPIYQAGLADDID RRCSIVRSHSSNSSPIGKSVKTSPTTQSKHNTTSAKGFLSPGSRSPKFKSSKKCLISEMA KESIPCPTESVYPPLVNCVAPVDIILPQITSNMWARGLGQLIRAKNIKTIGDLSTLTASE IKTLPIRSPKVSNVKKALRIYHEQQVKTRGLEEIPVFDISEKTVNGIENKSLSPDEERLV SDIIDPVALEIPLSKNLLAQISALALQLDSEDLHNYSGSQLFEMHEKLSCMANSVIKNLQ SRWRSPSHENSI |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
RIF1 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Key regulator of TP53BP1 that plays a key role in the repair of double-strand DNA breaks (DSBs) in response to DNA damage: acts by promoting non-homologous end joining (NHEJ)-mediated repair of DSBs. In response to DNA damage, interacts with ATM-phosphorylated TP53BP1. Interaction with TP53BP1 leads to dissociate the interaction between NUDT16L1/TIRR and TP53BP1, thereby unmasking the tandem Tudor-like domain of TP53BP1 and allowing recruitment to DNA DSBs. Once recruited to DSBs, RIF1 and TP53BP1 act by promoting NHEJ-mediated repair of DSBs. In the same time, RIF1 and TP53BP1 specifically counteract the function of BRCA1 by blocking DSBs resection via homologous recombination (HR) during G1 phase. Also required for immunoglobulin class-switch recombination (CSR) during antibody genesis, a process that involves the generation of DNA DSBs. Promotes NHEJ of dysfunctional telomeres.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CAMA1 | SNV: p.E1598K | . | |||
| CCK81 | SNV: p.C1734R | . | |||
| HCT116 | Insertion: p.E1442GfsTer4 | DBIA Probe Info | |||
| HCT15 | SNV: p.A293V; p.E1878K | . | |||
| HT115 | SNV: p.E1857Ter | . | |||
| ICC10 | SNV: p.K1312N | . | |||
| ICC106 | SNV: p.K1312N | . | |||
| ICC108 | SNV: p.K1312N | . | |||
| JHH7 | SNV: p.K449N | . | |||
| KYSE150 | SNV: p.S1089C | . | |||
| LNCaP clone FGC | SNV: p.P1455T | . | |||
| MCC13 | SNV: p.P410L Substitution: p.P1310Ter |
. | |||
| NCIH2172 | SNV: p.R785T | . | |||
| NCIH716 | SNV: p.S511A | . | |||
| OPM2 | SNV: p.S269G | . | |||
| REH | SNV: p.V193A | . | |||
| RKO | SNV: p.E1430G | . | |||
| RL | SNV: p.M739I | . | |||
| SNGM | SNV: p.Q1874H | . | |||
| SUPT1 | SNV: p.V841I | . | |||
| VMRCRCW | SNV: p.E1143G | . | |||
| WM1552C | SNV: p.E1885Ter | . | |||
| YSCCC | SNV: p.E727K; p.P1163S | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
A-EBA Probe Info |
 |
2.42 | LDD0215 | [2] | |
|
TH211 Probe Info |
 |
Y947(17.48); Y2361(10.03); Y404(5.99); Y2211(5.64) | LDD0257 | [3] | |
|
TH216 Probe Info |
 |
Y404(7.84) | LDD0259 | [3] | |
|
STPyne Probe Info |
 |
K1528(10.00) | LDD0277 | [4] | |
|
P12 Probe Info |
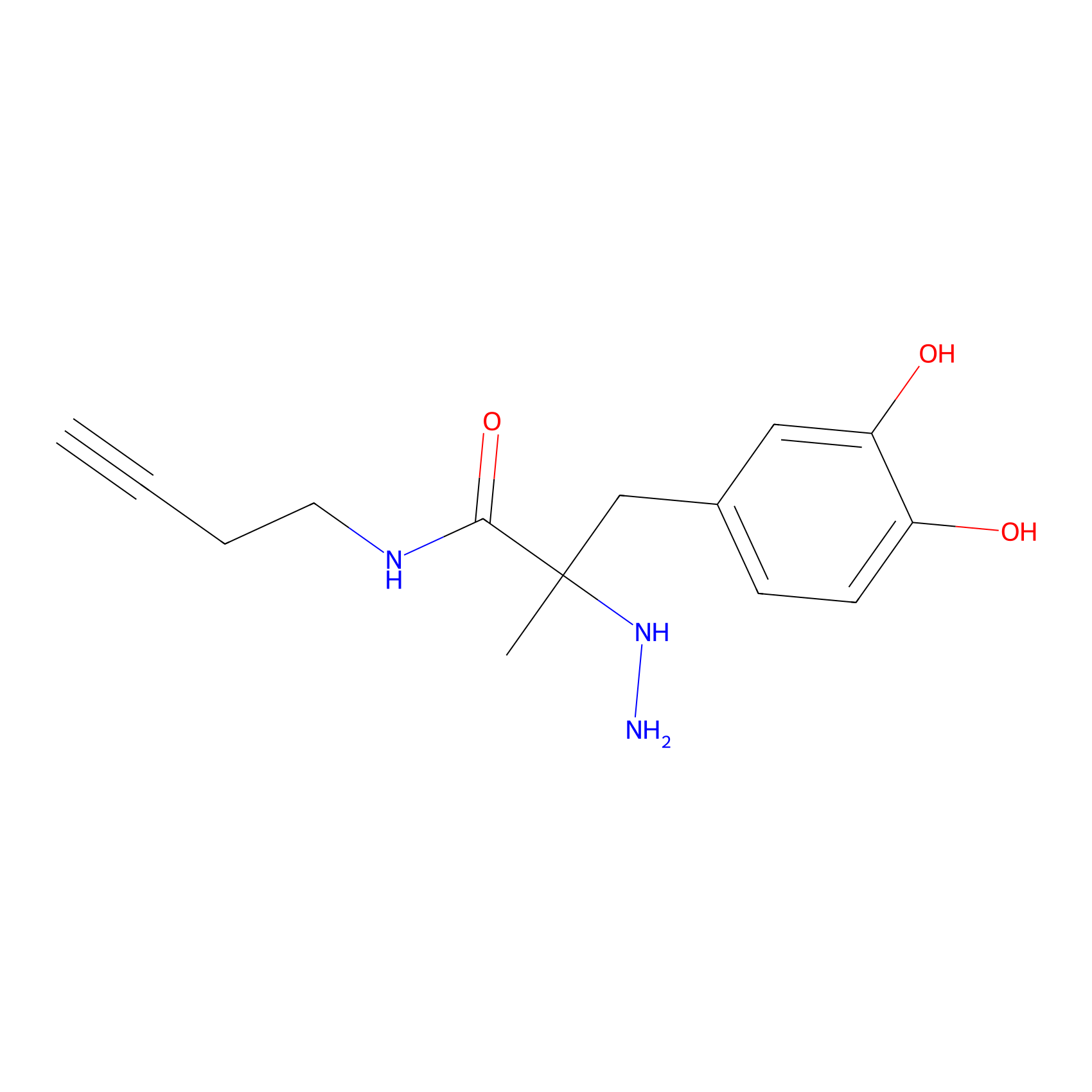 |
11.91 | LDD0202 | [5] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [6] | |
|
BTD Probe Info |
 |
C2274(0.64) | LDD2089 | [7] | |
|
DA-P3 Probe Info |
 |
4.57 | LDD0180 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C1155(10.64); C2298(5.06); C2274(2.03); C2169(2.00) | LDD0168 | [9] | |
|
Alkyne-RA190 Probe Info |
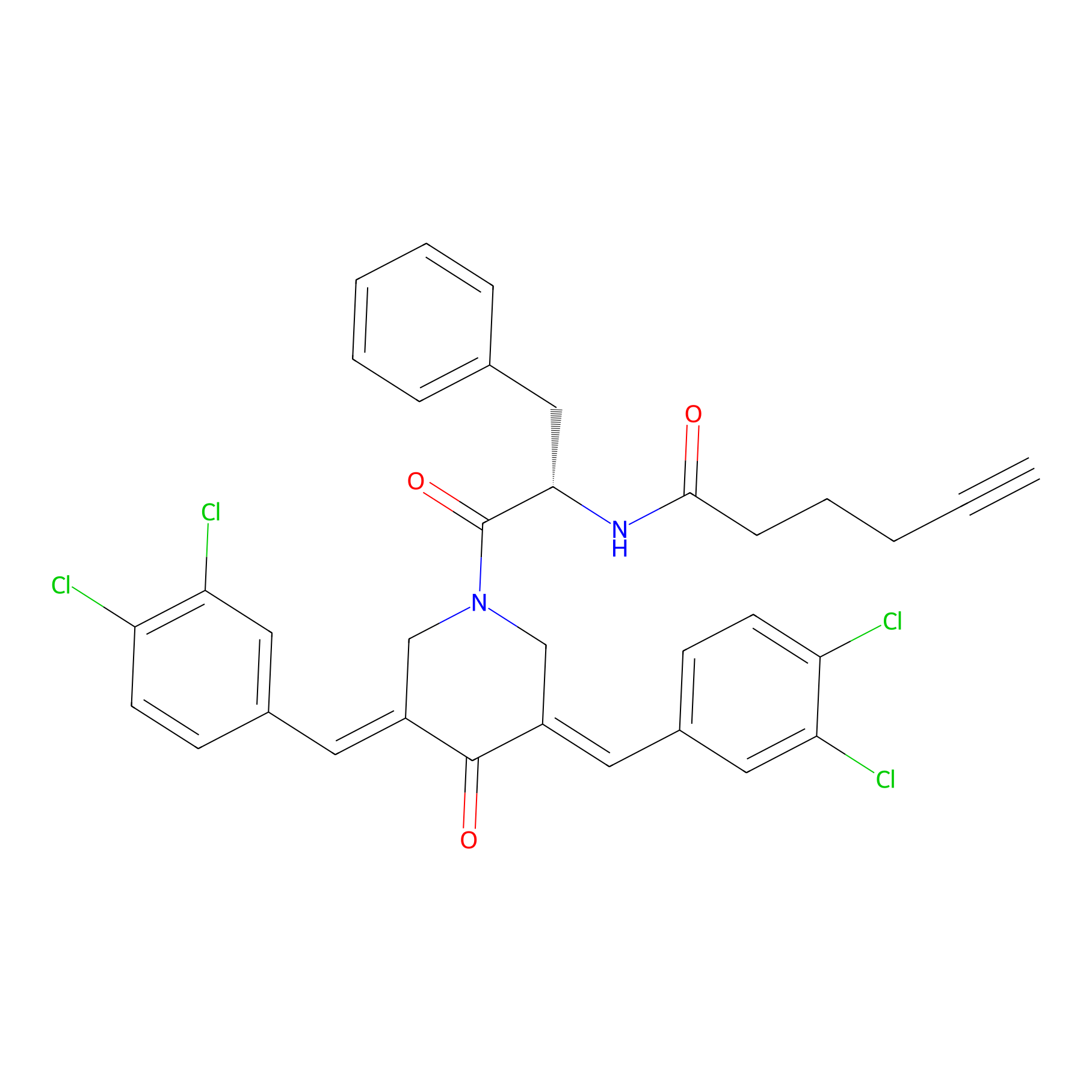 |
4.48 | LDD0299 | [10] | |
|
HHS-475 Probe Info |
 |
Y404(0.34) | LDD0264 | [11] | |
|
DBIA Probe Info |
 |
C880(1.25); C2450(1.10); C1718(1.02); C2169(1.00) | LDD0078 | [12] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [13] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C2274(0.00); C312(0.00); C82(0.00); C1692(0.00) | LDD0038 | [14] | |
|
IA-alkyne Probe Info |
 |
C1718(0.00); C2169(0.00); C312(0.00) | LDD0032 | [15] | |
|
Lodoacetamide azide Probe Info |
 |
C2169(0.00); C312(0.00); C1155(0.00); C1718(0.00) | LDD0037 | [14] | |
|
NAIA_4 Probe Info |
 |
C1692(0.00); C2169(0.00) | LDD2226 | [16] | |
|
TFBX Probe Info |
 |
C1718(0.00); C1692(0.00) | LDD0027 | [17] | |
|
WYneN Probe Info |
 |
C2169(0.00); C1155(0.00); C312(0.00) | LDD0021 | [18] | |
|
aHNE Probe Info |
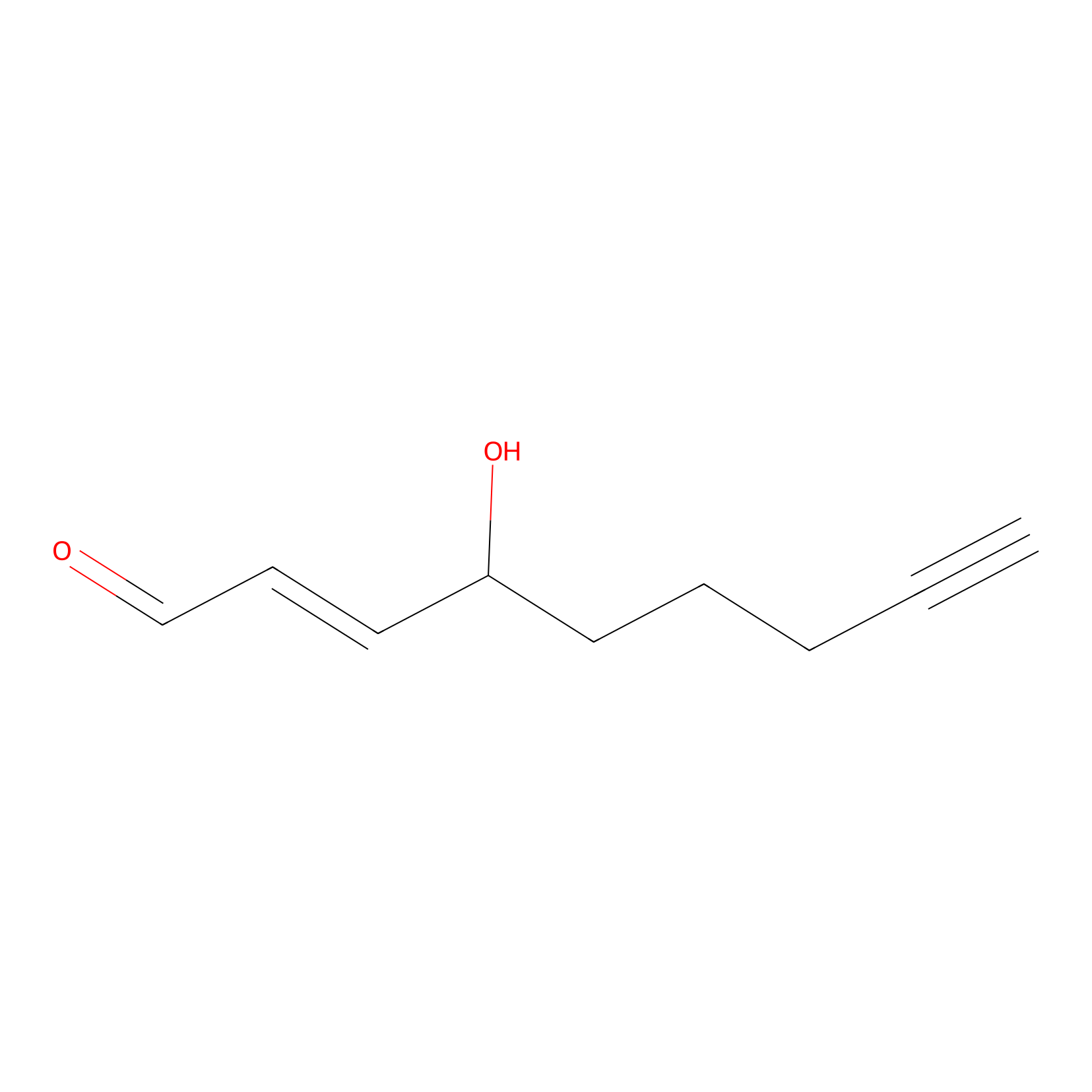 |
N.A. | LDD0001 | [18] | |
|
Compound 10 Probe Info |
 |
C1718(0.00); C2169(0.00); C2274(0.00); C312(0.00) | LDD2216 | [19] | |
|
Compound 11 Probe Info |
 |
C1692(0.00); C1718(0.00); C2169(0.00); C2274(0.00) | LDD2213 | [19] | |
|
ENE Probe Info |
 |
C2450(0.00); C1155(0.00); C880(0.00); C1865(0.00) | LDD0006 | [18] | |
|
IPM Probe Info |
 |
C312(0.00); C2169(0.00); C2274(0.00); C1718(0.00) | LDD0005 | [18] | |
|
PPMS Probe Info |
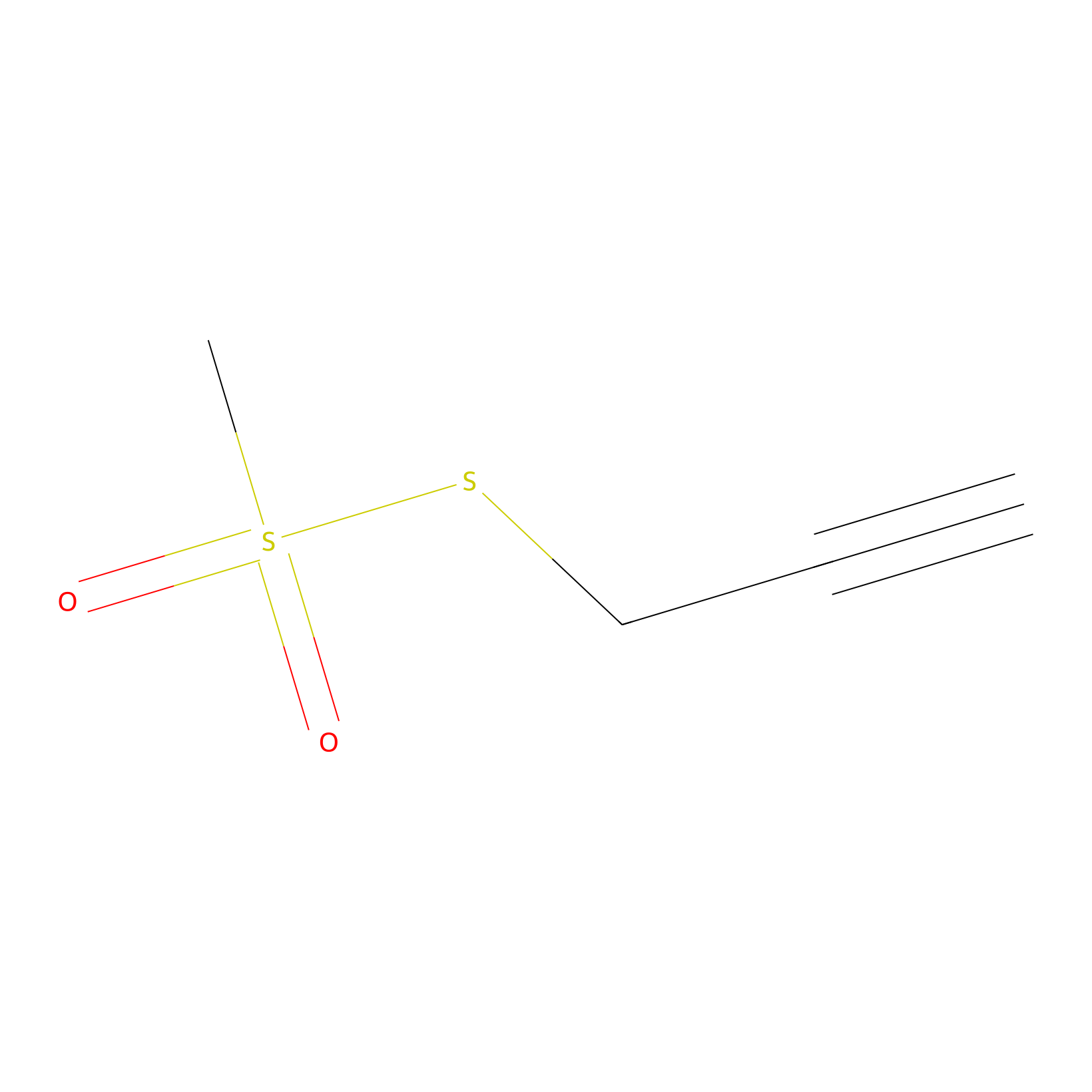 |
N.A. | LDD0008 | [18] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [20] | |
|
VSF Probe Info |
 |
C1904(0.00); C2126(0.00); C2169(0.00); C1619(0.00) | LDD0007 | [18] | |
|
Phosphinate-6 Probe Info |
 |
C1735(0.00); C2223(0.00); C312(0.00); C1734(0.00) | LDD0018 | [21] | |
|
1c-yne Probe Info |
 |
K1870(0.00); K125(0.00); K103(0.00) | LDD0228 | [22] | |
|
Acrolein Probe Info |
 |
C1718(0.00); C1692(0.00) | LDD0217 | [23] | |
|
Methacrolein Probe Info |
 |
C1155(0.00); C2450(0.00); C1692(0.00); C1718(0.00) | LDD0218 | [23] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [24] | |
|
AOyne Probe Info |
 |
7.90 | LDD0443 | [25] | |
|
NAIA_5 Probe Info |
 |
C2274(0.00); C2450(0.00); C1718(0.00) | LDD2223 | [16] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe3 Probe Info |
 |
14.67 | LDD0465 | [26] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C2450(0.92) | LDD2117 | [7] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C2274(1.52) | LDD2152 | [7] |
| LDCM0025 | 4SU-RNA | HEK-293T | C1155(10.64); C2298(5.06); C2274(2.03); C2169(2.00) | LDD0168 | [9] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C2286(13.13); C2298(16.55); C2126(2.27); C2068(6.21) | LDD0169 | [9] |
| LDCM0214 | AC1 | HCT 116 | C1732(0.88); C1155(0.87); C1734(0.92); C1735(0.88) | LDD0531 | [12] |
| LDCM0215 | AC10 | HCT 116 | C1732(0.94); C1155(0.78); C1630(1.42); C1650(1.42) | LDD0532 | [12] |
| LDCM0216 | AC100 | HCT 116 | C1155(0.97); C1692(1.05); C1865(0.97); C1904(0.75) | LDD0533 | [12] |
| LDCM0217 | AC101 | HCT 116 | C1155(1.01); C1692(1.17); C1865(1.13); C1904(0.94) | LDD0534 | [12] |
| LDCM0218 | AC102 | HCT 116 | C1155(1.00); C1692(0.90); C1865(1.04); C1904(0.84) | LDD0535 | [12] |
| LDCM0219 | AC103 | HCT 116 | C1155(1.18); C1692(1.04); C1865(1.21); C1904(0.96) | LDD0536 | [12] |
| LDCM0220 | AC104 | HCT 116 | C1155(1.07); C1692(0.78); C1865(1.29); C1904(1.03) | LDD0537 | [12] |
| LDCM0221 | AC105 | HCT 116 | C1155(1.18); C1692(1.39); C1865(1.52); C1904(0.94) | LDD0538 | [12] |
| LDCM0222 | AC106 | HCT 116 | C1155(1.13); C1692(1.33); C1865(1.21); C1904(0.99) | LDD0539 | [12] |
| LDCM0223 | AC107 | HCT 116 | C1155(0.97); C1692(1.04); C1865(1.22); C1904(0.84) | LDD0540 | [12] |
| LDCM0224 | AC108 | HCT 116 | C1155(0.88); C1692(1.13); C1865(0.90); C1904(0.77) | LDD0541 | [12] |
| LDCM0225 | AC109 | HCT 116 | C1155(0.95); C1692(0.91); C1865(0.97); C1904(0.86) | LDD0542 | [12] |
| LDCM0226 | AC11 | HCT 116 | C1732(1.19); C1155(0.84); C1630(1.65); C1650(1.65) | LDD0543 | [12] |
| LDCM0227 | AC110 | HCT 116 | C1155(1.11); C1692(1.15); C1865(1.08); C1904(0.96) | LDD0544 | [12] |
| LDCM0228 | AC111 | HCT 116 | C1155(1.00); C1692(1.14); C1865(1.14); C1904(1.14) | LDD0545 | [12] |
| LDCM0229 | AC112 | HCT 116 | C1155(1.04); C1692(1.12); C1865(1.14); C1904(1.15) | LDD0546 | [12] |
| LDCM0230 | AC113 | HCT 116 | C1732(0.78); C1155(1.00); C1734(0.91); C1735(1.09) | LDD0547 | [12] |
| LDCM0231 | AC114 | HCT 116 | C1732(0.85); C1155(1.20); C1734(0.96); C1735(1.13) | LDD0548 | [12] |
| LDCM0232 | AC115 | HCT 116 | C1732(0.98); C1155(1.40); C1734(1.02); C1735(1.36) | LDD0549 | [12] |
| LDCM0233 | AC116 | HCT 116 | C1732(1.05); C1155(1.33); C1734(0.96); C1735(1.24) | LDD0550 | [12] |
| LDCM0234 | AC117 | HCT 116 | C1732(1.33); C1155(1.09); C1734(0.96); C1735(1.38) | LDD0551 | [12] |
| LDCM0235 | AC118 | HCT 116 | C1732(1.18); C1155(1.14); C1734(1.00); C1735(0.94) | LDD0552 | [12] |
| LDCM0236 | AC119 | HCT 116 | C1732(1.17); C1155(1.14); C1734(1.10); C1735(1.20) | LDD0553 | [12] |
| LDCM0237 | AC12 | HCT 116 | C1732(0.99); C1155(0.78); C1630(1.09); C1650(1.09) | LDD0554 | [12] |
| LDCM0238 | AC120 | HCT 116 | C1732(0.91); C1155(1.10); C1734(1.17); C1735(1.25) | LDD0555 | [12] |
| LDCM0239 | AC121 | HCT 116 | C1732(1.29); C1155(0.99); C1734(0.87); C1735(1.00) | LDD0556 | [12] |
| LDCM0240 | AC122 | HCT 116 | C1732(0.93); C1155(1.01); C1734(1.05); C1735(1.11) | LDD0557 | [12] |
| LDCM0241 | AC123 | HCT 116 | C1732(1.48); C1155(1.06); C1734(0.85); C1735(0.94) | LDD0558 | [12] |
| LDCM0242 | AC124 | HCT 116 | C1732(1.15); C1155(1.04); C1734(0.89); C1735(0.97) | LDD0559 | [12] |
| LDCM0243 | AC125 | HCT 116 | C1732(0.88); C1155(1.01); C1734(1.00); C1735(1.13) | LDD0560 | [12] |
| LDCM0244 | AC126 | HCT 116 | C1732(0.90); C1155(1.18); C1734(0.92); C1735(1.38) | LDD0561 | [12] |
| LDCM0245 | AC127 | HCT 116 | C1732(1.21); C1155(1.27); C1734(1.00); C1735(1.37) | LDD0562 | [12] |
| LDCM0246 | AC128 | HCT 116 | C1732(1.13); C1155(0.55); C1734(0.36); C1735(1.10) | LDD0563 | [12] |
| LDCM0247 | AC129 | HCT 116 | C1732(0.72); C1155(0.67); C1734(0.33); C1735(0.78) | LDD0564 | [12] |
| LDCM0249 | AC130 | HCT 116 | C1732(1.11); C1155(0.73); C1734(0.41); C1735(0.84) | LDD0566 | [12] |
| LDCM0250 | AC131 | HCT 116 | C1732(0.67); C1155(0.68); C1734(0.45); C1735(0.71) | LDD0567 | [12] |
| LDCM0251 | AC132 | HCT 116 | C1732(1.00); C1155(0.80); C1734(0.50); C1735(0.87) | LDD0568 | [12] |
| LDCM0252 | AC133 | HCT 116 | C1732(0.85); C1155(0.82); C1734(0.45); C1735(0.84) | LDD0569 | [12] |
| LDCM0253 | AC134 | HCT 116 | C1732(1.11); C1155(0.72); C1734(0.54); C1735(0.89) | LDD0570 | [12] |
| LDCM0254 | AC135 | HCT 116 | C1732(0.97); C1155(0.73); C1734(0.49); C1735(0.93) | LDD0571 | [12] |
| LDCM0255 | AC136 | HCT 116 | C1732(0.90); C1155(0.92); C1734(0.51); C1735(0.75) | LDD0572 | [12] |
| LDCM0256 | AC137 | HCT 116 | C1732(1.01); C1155(0.66); C1734(0.56); C1735(0.56) | LDD0573 | [12] |
| LDCM0257 | AC138 | HCT 116 | C1732(1.12); C1155(0.82); C1734(0.54); C1735(0.71) | LDD0574 | [12] |
| LDCM0258 | AC139 | HCT 116 | C1732(0.92); C1155(0.77); C1734(0.56); C1735(0.62) | LDD0575 | [12] |
| LDCM0259 | AC14 | HCT 116 | C1732(0.88); C1155(0.99); C1630(1.58); C1650(1.58) | LDD0576 | [12] |
| LDCM0260 | AC140 | HCT 116 | C1732(0.93); C1155(0.87); C1734(0.61); C1735(0.73) | LDD0577 | [12] |
| LDCM0261 | AC141 | HCT 116 | C1732(1.08); C1155(0.95); C1734(0.63); C1735(0.82) | LDD0578 | [12] |
| LDCM0262 | AC142 | HCT 116 | C1732(1.09); C1155(0.73); C1734(0.49); C1735(0.62) | LDD0579 | [12] |
| LDCM0263 | AC143 | HCT 116 | C1732(0.97); C1734(0.97); C2450(1.10); C2169(1.10) | LDD0580 | [12] |
| LDCM0264 | AC144 | HCT 116 | C1734(0.93); C1732(1.04); C1155(1.04); C1865(1.20) | LDD0581 | [12] |
| LDCM0265 | AC145 | HCT 116 | C1732(0.99); C1734(0.99); C1155(1.07); C2169(1.08) | LDD0582 | [12] |
| LDCM0266 | AC146 | HCT 116 | C1732(0.89); C1155(0.98); C1734(1.00); C880(1.00) | LDD0583 | [12] |
| LDCM0267 | AC147 | HCT 116 | C1155(0.86); C1734(0.90); C1732(1.07); C880(1.22) | LDD0584 | [12] |
| LDCM0268 | AC148 | HCT 116 | C1732(0.95); C1155(1.00); C1734(1.05); C880(1.66) | LDD0585 | [12] |
| LDCM0269 | AC149 | HCT 116 | C1732(0.86); C1734(0.99); C1155(1.09); C880(1.32) | LDD0586 | [12] |
| LDCM0270 | AC15 | HCT 116 | C2450(0.84); C1865(0.86); C1734(0.86); C1155(0.90) | LDD0587 | [12] |
| LDCM0271 | AC150 | HCT 116 | C1155(0.93); C1734(0.95); C1732(0.96); C880(1.00) | LDD0588 | [12] |
| LDCM0272 | AC151 | HCT 116 | C1734(0.82); C1732(0.99); C1155(1.00); C2450(1.05) | LDD0589 | [12] |
| LDCM0273 | AC152 | HCT 116 | C1734(0.91); C1732(1.00); C1155(1.00); C1865(1.22) | LDD0590 | [12] |
| LDCM0274 | AC153 | HCT 116 | C1732(1.09); C1155(1.18); C1734(1.28); C2450(1.83) | LDD0591 | [12] |
| LDCM0621 | AC154 | HCT 116 | C1732(0.95); C1155(1.00); C1692(2.39); C1734(0.95) | LDD2158 | [12] |
| LDCM0622 | AC155 | HCT 116 | C1732(0.94); C1155(1.03); C1692(2.62); C1734(0.97) | LDD2159 | [12] |
| LDCM0623 | AC156 | HCT 116 | C1732(1.00); C1155(0.94); C1692(1.72); C1734(1.07) | LDD2160 | [12] |
| LDCM0624 | AC157 | HCT 116 | C1732(1.26); C1155(1.07); C1692(2.10); C1734(1.24) | LDD2161 | [12] |
| LDCM0276 | AC17 | HCT 116 | C2169(0.14); C1734(0.56); C1735(0.73); C2274(0.78) | LDD0593 | [12] |
| LDCM0277 | AC18 | HCT 116 | C2169(0.19); C1734(0.70); C1732(0.85); C1735(0.87) | LDD0594 | [12] |
| LDCM0278 | AC19 | HCT 116 | C2169(0.14); C1734(0.56); C1732(0.69); C2274(0.79) | LDD0595 | [12] |
| LDCM0279 | AC2 | HCT 116 | C2450(0.78); C2169(0.81); C1155(0.92); C1904(0.93) | LDD0596 | [12] |
| LDCM0280 | AC20 | HCT 116 | C2169(0.17); C1734(0.61); C1732(0.71); C1735(0.73) | LDD0597 | [12] |
| LDCM0281 | AC21 | HCT 116 | C2169(0.18); C1734(0.64); C1732(0.69); C1735(0.78) | LDD0598 | [12] |
| LDCM0282 | AC22 | HCT 116 | C2169(0.16); C1734(0.67); C1732(0.79); C2274(0.84) | LDD0599 | [12] |
| LDCM0283 | AC23 | HCT 116 | C2169(0.14); C1734(0.56); C1732(0.64); C2274(0.79) | LDD0600 | [12] |
| LDCM0284 | AC24 | HCT 116 | C2169(0.12); C1734(0.50); C2274(0.77); C1732(0.80) | LDD0601 | [12] |
| LDCM0285 | AC25 | HCT 116 | C1155(0.65); C880(0.88); C312(0.95); C731(0.97) | LDD0602 | [12] |
| LDCM0286 | AC26 | HCT 116 | C1155(1.00); C312(1.03); C880(1.07); C1865(1.10) | LDD0603 | [12] |
| LDCM0287 | AC27 | HCT 116 | C1155(0.74); C880(0.90); C312(1.08); C731(1.09) | LDD0604 | [12] |
| LDCM0288 | AC28 | HCT 116 | C1155(0.76); C1865(1.12); C312(1.13); C880(1.16) | LDD0605 | [12] |
| LDCM0289 | AC29 | HCT 116 | C880(0.96); C1155(0.99); C731(1.11); C312(1.11) | LDD0606 | [12] |
| LDCM0290 | AC3 | HCT 116 | C2169(0.69); C2450(0.76); C2274(0.84); C1155(0.85) | LDD0607 | [12] |
| LDCM0291 | AC30 | HCT 116 | C1155(0.90); C880(0.95); C1865(1.11); C312(1.13) | LDD0608 | [12] |
| LDCM0292 | AC31 | HCT 116 | C1155(0.82); C880(0.89); C2450(1.07); C731(1.08) | LDD0609 | [12] |
| LDCM0293 | AC32 | HCT 116 | C880(1.05); C312(1.22); C731(1.23); C2450(1.24) | LDD0610 | [12] |
| LDCM0294 | AC33 | HCT 116 | C880(0.92); C1155(1.08); C718(1.11); C730(1.11) | LDD0611 | [12] |
| LDCM0295 | AC34 | HCT 116 | C1155(1.11); C731(1.18); C880(1.19); C718(1.23) | LDD0612 | [12] |
| LDCM0296 | AC35 | HCT 116 | C718(0.88); C731(0.88); C312(0.90); C2450(0.92) | LDD0613 | [12] |
| LDCM0297 | AC36 | HCT 116 | C718(0.76); C731(0.76); C312(0.87); C1865(0.89) | LDD0614 | [12] |
| LDCM0298 | AC37 | HCT 116 | C312(0.83); C880(0.90); C718(0.98); C731(0.98) | LDD0615 | [12] |
| LDCM0299 | AC38 | HCT 116 | C718(0.91); C731(0.91); C730(0.97); C880(0.99) | LDD0616 | [12] |
| LDCM0300 | AC39 | HCT 116 | C718(0.96); C731(0.96); C312(1.05); C1865(1.06) | LDD0617 | [12] |
| LDCM0301 | AC4 | HCT 116 | C2169(0.67); C1155(0.77); C2450(0.77); C2274(0.80) | LDD0618 | [12] |
| LDCM0302 | AC40 | HCT 116 | C718(1.01); C731(1.01); C312(1.09); C2450(1.24) | LDD0619 | [12] |
| LDCM0303 | AC41 | HCT 116 | C718(0.82); C731(0.82); C730(0.89); C312(0.94) | LDD0620 | [12] |
| LDCM0304 | AC42 | HCT 116 | C718(0.92); C731(0.92); C2450(1.07); C730(1.09) | LDD0621 | [12] |
| LDCM0305 | AC43 | HCT 116 | C718(0.92); C731(0.92); C880(1.06); C2450(1.07) | LDD0622 | [12] |
| LDCM0306 | AC44 | HCT 116 | C312(0.99); C718(1.00); C731(1.00); C880(1.00) | LDD0623 | [12] |
| LDCM0307 | AC45 | HCT 116 | C718(1.00); C731(1.00); C2450(1.05); C1865(1.16) | LDD0624 | [12] |
| LDCM0308 | AC46 | HCT 116 | C731(0.88); C730(0.92); C718(0.93); C2450(1.06) | LDD0625 | [12] |
| LDCM0309 | AC47 | HCT 116 | C718(0.91); C731(0.93); C730(0.94); C1735(0.97) | LDD0626 | [12] |
| LDCM0310 | AC48 | HCT 116 | C1155(0.88); C731(0.90); C880(0.98); C730(0.99) | LDD0627 | [12] |
| LDCM0311 | AC49 | HCT 116 | C730(1.10); C731(1.11); C1734(1.18); C1904(1.18) | LDD0628 | [12] |
| LDCM0312 | AC5 | HCT 116 | C2169(0.57); C2450(0.73); C2274(0.75); C1155(0.75) | LDD0629 | [12] |
| LDCM0313 | AC50 | HCT 116 | C1734(0.92); C731(1.05); C730(1.10); C1735(1.14) | LDD0630 | [12] |
| LDCM0314 | AC51 | HCT 116 | C1155(0.71); C880(0.81); C731(0.82); C730(0.82) | LDD0631 | [12] |
| LDCM0315 | AC52 | HCT 116 | C1155(0.78); C731(0.89); C730(0.91); C880(0.96) | LDD0632 | [12] |
| LDCM0316 | AC53 | HCT 116 | C731(1.02); C1904(1.02); C1735(1.04); C2450(1.07) | LDD0633 | [12] |
| LDCM0317 | AC54 | HCT 116 | C880(0.88); C1735(0.95); C1734(1.03); C731(1.05) | LDD0634 | [12] |
| LDCM0318 | AC55 | HCT 116 | C1904(0.85); C880(1.01); C730(1.15); C1735(1.16) | LDD0635 | [12] |
| LDCM0319 | AC56 | HCT 116 | C880(1.20); C1735(1.24); C731(1.25); C730(1.25) | LDD0636 | [12] |
| LDCM0320 | AC57 | HCT 116 | C1865(0.97); C2450(1.06); C1630(1.07); C1650(1.07) | LDD0637 | [12] |
| LDCM0321 | AC58 | HCT 116 | C1865(0.91); C312(0.95); C1732(0.96); C1734(0.96) | LDD0638 | [12] |
| LDCM0322 | AC59 | HCT 116 | C1630(0.70); C1650(0.70); C2450(1.05); C2274(1.11) | LDD0639 | [12] |
| LDCM0323 | AC6 | HCT 116 | C1155(0.78); C1865(0.87); C2274(1.01); C312(1.04) | LDD0640 | [12] |
| LDCM0324 | AC60 | HCT 116 | C2450(1.10); C312(1.16); C731(1.16); C718(1.17) | LDD0641 | [12] |
| LDCM0325 | AC61 | HCT 116 | C1865(0.90); C312(0.92); C1630(0.96); C1650(0.96) | LDD0642 | [12] |
| LDCM0326 | AC62 | HCT 116 | C1630(0.91); C1650(0.91); C1865(1.07); C2450(1.12) | LDD0643 | [12] |
| LDCM0327 | AC63 | HCT 116 | C1630(0.75); C1650(0.75); C312(1.03); C2450(1.07) | LDD0644 | [12] |
| LDCM0328 | AC64 | HCT 116 | C1630(0.66); C1650(0.66); C2450(1.10); C1732(1.13) | LDD0645 | [12] |
| LDCM0329 | AC65 | HCT 116 | C1630(0.85); C1650(0.85); C1155(0.94); C2450(0.96) | LDD0646 | [12] |
| LDCM0330 | AC66 | HCT 116 | C1155(0.87); C1865(0.94); C1630(0.95); C1650(0.95) | LDD0647 | [12] |
| LDCM0331 | AC67 | HCT 116 | C1630(0.80); C1650(0.80); C2450(1.08); C2274(1.21) | LDD0648 | [12] |
| LDCM0332 | AC68 | HCT 116 | C312(0.76); C1732(0.80); C880(0.85); C2450(0.86) | LDD0649 | [12] |
| LDCM0333 | AC69 | HCT 116 | C312(0.73); C2450(0.81); C1732(0.84); C880(0.88) | LDD0650 | [12] |
| LDCM0334 | AC7 | HCT 116 | C1155(0.66); C1865(0.72); C2169(0.90); C2450(0.92) | LDD0651 | [12] |
| LDCM0335 | AC70 | HCT 116 | C312(0.68); C2450(0.76); C2169(0.92); C880(1.00) | LDD0652 | [12] |
| LDCM0336 | AC71 | HCT 116 | C2169(0.72); C880(0.82); C1732(0.87); C2450(0.88) | LDD0653 | [12] |
| LDCM0337 | AC72 | HCT 116 | C2450(0.73); C312(0.85); C718(0.96); C731(0.97) | LDD0654 | [12] |
| LDCM0338 | AC73 | HCT 116 | C2450(0.85); C312(1.02); C880(1.06); C1732(1.26) | LDD0655 | [12] |
| LDCM0339 | AC74 | HCT 116 | C1732(0.95); C880(0.95); C312(1.01); C1155(1.18) | LDD0656 | [12] |
| LDCM0340 | AC75 | HCT 116 | C2450(0.75); C312(0.99); C880(1.00); C1732(1.11) | LDD0657 | [12] |
| LDCM0341 | AC76 | HCT 116 | C2450(0.99); C880(1.00); C312(1.01); C1732(1.09) | LDD0658 | [12] |
| LDCM0342 | AC77 | HCT 116 | C2450(0.76); C1732(0.90); C312(0.91); C718(1.10) | LDD0659 | [12] |
| LDCM0343 | AC78 | HCT 116 | C312(0.79); C718(0.95); C2450(0.95); C731(0.96) | LDD0660 | [12] |
| LDCM0344 | AC79 | HCT 116 | C1732(0.89); C718(0.96); C731(0.98); C2169(1.00) | LDD0661 | [12] |
| LDCM0345 | AC8 | HCT 116 | C1865(0.78); C1155(0.79); C2274(0.95); C731(0.96) | LDD0662 | [12] |
| LDCM0346 | AC80 | HCT 116 | C312(0.70); C880(0.89); C2450(1.05); C730(1.05) | LDD0663 | [12] |
| LDCM0347 | AC81 | HCT 116 | C312(0.34); C2450(0.67); C1732(0.85); C718(0.85) | LDD0664 | [12] |
| LDCM0348 | AC82 | HCT 116 | C2450(0.92); C1732(1.04); C312(1.21); C718(1.44) | LDD0665 | [12] |
| LDCM0349 | AC83 | HCT 116 | C2450(1.26); C1904(1.37); C731(1.41); C730(1.41) | LDD0666 | [12] |
| LDCM0350 | AC84 | HCT 116 | C718(1.13); C731(1.19); C2450(1.20); C730(1.21) | LDD0667 | [12] |
| LDCM0351 | AC85 | HCT 116 | C1692(1.00); C718(1.02); C2169(1.06); C731(1.10) | LDD0668 | [12] |
| LDCM0352 | AC86 | HCT 116 | C718(1.00); C1692(1.04); C731(1.09); C2450(1.20) | LDD0669 | [12] |
| LDCM0353 | AC87 | HCT 116 | C718(0.80); C731(0.99); C312(1.00); C1692(1.00) | LDD0670 | [12] |
| LDCM0354 | AC88 | HCT 116 | C718(0.92); C312(1.02); C1692(1.07); C1904(1.07) | LDD0671 | [12] |
| LDCM0355 | AC89 | HCT 116 | C1692(1.07); C718(1.09); C2450(1.14); C1904(1.15) | LDD0672 | [12] |
| LDCM0357 | AC90 | HCT 116 | C2169(0.78); C312(0.96); C1865(1.04); C880(1.06) | LDD0674 | [12] |
| LDCM0358 | AC91 | HCT 116 | C2450(0.98); C1692(1.15); C730(1.20); C731(1.28) | LDD0675 | [12] |
| LDCM0359 | AC92 | HCT 116 | C1692(1.10); C730(1.14); C2450(1.17); C718(1.17) | LDD0676 | [12] |
| LDCM0360 | AC93 | HCT 116 | C718(0.94); C1692(0.96); C731(1.06); C730(1.10) | LDD0677 | [12] |
| LDCM0361 | AC94 | HCT 116 | C1692(0.81); C718(0.90); C2450(0.90); C731(1.03) | LDD0678 | [12] |
| LDCM0362 | AC95 | HCT 116 | C718(0.83); C731(0.94); C730(1.04); C1692(1.07) | LDD0679 | [12] |
| LDCM0363 | AC96 | HCT 116 | C1692(0.78); C2450(0.86); C718(1.07); C730(1.15) | LDD0680 | [12] |
| LDCM0364 | AC97 | HCT 116 | C2450(1.11); C718(1.13); C731(1.15); C730(1.19) | LDD0681 | [12] |
| LDCM0365 | AC98 | HCT 116 | C312(0.84); C2274(0.87); C1904(1.11); C731(1.19) | LDD0682 | [12] |
| LDCM0366 | AC99 | HCT 116 | C1904(0.78); C2274(0.82); C312(0.86); C2169(0.88) | LDD0683 | [12] |
| LDCM0248 | AKOS034007472 | HCT 116 | C1732(1.09); C1155(0.87); C1630(1.09); C1650(1.09) | LDD0565 | [12] |
| LDCM0356 | AKOS034007680 | HCT 116 | C1155(0.72); C1865(0.73); C731(0.97); C730(0.98) | LDD0673 | [12] |
| LDCM0275 | AKOS034007705 | HCT 116 | C1732(0.85); C1734(0.91); C2450(0.99); C1865(1.03) | LDD0592 | [12] |
| LDCM0156 | Aniline | NCI-H1299 | 11.61 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C880(1.25); C2450(1.10); C1718(1.02); C2169(1.00) | LDD0078 | [12] |
| LDCM0088 | C45 | HEK-293T | 11.91 | LDD0202 | [5] |
| LDCM0108 | Chloroacetamide | HeLa | C1155(0.00); C1718(0.00); C1904(0.00); H2362(0.00) | LDD0222 | [23] |
| LDCM0632 | CL-Sc | Hep-G2 | C1718(20.00); C2450(0.85); C1718(0.52); C312(0.22) | LDD2227 | [16] |
| LDCM0367 | CL1 | HCT 116 | C1630(0.70); C1650(0.70); C312(0.78); C2169(0.86) | LDD0684 | [12] |
| LDCM0368 | CL10 | HCT 116 | C2169(0.69); C2274(0.83); C2450(0.88); C1904(0.97) | LDD0685 | [12] |
| LDCM0369 | CL100 | HCT 116 | C2450(0.76); C2169(0.86); C1904(0.89); C2274(0.91) | LDD0686 | [12] |
| LDCM0370 | CL101 | HCT 116 | C1732(0.68); C1155(0.71); C2450(0.81); C1865(0.82) | LDD0687 | [12] |
| LDCM0371 | CL102 | HCT 116 | C1155(0.77); C1865(0.78); C2450(0.85); C2274(0.86) | LDD0688 | [12] |
| LDCM0372 | CL103 | HCT 116 | C1732(0.72); C1865(0.80); C1734(0.80); C1155(0.82) | LDD0689 | [12] |
| LDCM0373 | CL104 | HCT 116 | C1155(0.72); C1865(0.80); C1734(0.86); C2274(0.88) | LDD0690 | [12] |
| LDCM0374 | CL105 | HCT 116 | C2169(0.17); C1734(0.63); C1732(0.82); C2274(0.88) | LDD0691 | [12] |
| LDCM0375 | CL106 | HCT 116 | C2169(0.15); C1734(0.65); C1732(0.66); C1155(0.75) | LDD0692 | [12] |
| LDCM0376 | CL107 | HCT 116 | C2169(0.17); C1734(0.67); C1732(0.68); C2274(0.82) | LDD0693 | [12] |
| LDCM0377 | CL108 | HCT 116 | C2169(0.13); C1734(0.56); C1732(0.77); C1155(0.78) | LDD0694 | [12] |
| LDCM0378 | CL109 | HCT 116 | C2169(0.17); C1734(0.77); C2274(0.88); C1732(0.96) | LDD0695 | [12] |
| LDCM0379 | CL11 | HCT 116 | C2169(0.66); C1155(1.05); C1904(1.06); C2274(1.06) | LDD0696 | [12] |
| LDCM0380 | CL110 | HCT 116 | C2169(0.10); C1734(0.53); C1732(0.75); C2274(0.79) | LDD0697 | [12] |
| LDCM0381 | CL111 | HCT 116 | C2169(0.13); C1734(0.58); C1732(0.76); C1735(0.79) | LDD0698 | [12] |
| LDCM0382 | CL112 | HCT 116 | C1155(0.79); C312(0.86); C1865(0.96); C718(1.01) | LDD0699 | [12] |
| LDCM0383 | CL113 | HCT 116 | C880(1.00); C1155(1.06); C731(1.09); C312(1.10) | LDD0700 | [12] |
| LDCM0384 | CL114 | HCT 116 | C312(0.94); C731(0.94); C1155(0.97); C1865(0.99) | LDD0701 | [12] |
| LDCM0385 | CL115 | HCT 116 | C880(0.92); C731(1.05); C312(1.06); C718(1.09) | LDD0702 | [12] |
| LDCM0386 | CL116 | HCT 116 | C880(0.92); C1155(1.00); C312(1.07); C731(1.07) | LDD0703 | [12] |
| LDCM0387 | CL117 | HCT 116 | C718(1.22); C731(1.22); C312(1.36); C1155(1.38) | LDD0704 | [12] |
| LDCM0388 | CL118 | HCT 116 | C2450(0.97); C718(0.97); C731(0.97); C880(1.05) | LDD0705 | [12] |
| LDCM0389 | CL119 | HCT 116 | C2450(0.91); C718(0.96); C731(0.96); C312(1.01) | LDD0706 | [12] |
| LDCM0390 | CL12 | HCT 116 | C2169(0.73); C2450(0.95); C2274(0.98); C1904(1.14) | LDD0707 | [12] |
| LDCM0391 | CL120 | HCT 116 | C312(0.95); C2450(0.97); C880(1.04); C718(1.08) | LDD0708 | [12] |
| LDCM0392 | CL121 | HCT 116 | C2450(0.72); C880(0.72); C1865(1.11); C1155(1.19) | LDD0709 | [12] |
| LDCM0393 | CL122 | HCT 116 | C1155(0.85); C731(0.94); C1904(0.98); C1865(1.00) | LDD0710 | [12] |
| LDCM0394 | CL123 | HCT 116 | C1735(0.99); C1904(1.01); C1865(1.02); C731(1.04) | LDD0711 | [12] |
| LDCM0395 | CL124 | HCT 116 | C880(1.07); C731(1.08); C730(1.10); C1904(1.14) | LDD0712 | [12] |
| LDCM0396 | CL125 | HCT 116 | C2450(0.79); C312(0.85); C718(0.89); C730(0.89) | LDD0713 | [12] |
| LDCM0397 | CL126 | HCT 116 | C1630(0.78); C1650(0.78); C1865(0.85); C312(0.87) | LDD0714 | [12] |
| LDCM0398 | CL127 | HCT 116 | C1630(0.53); C1650(0.53); C2450(0.82); C1155(0.89) | LDD0715 | [12] |
| LDCM0399 | CL128 | HCT 116 | C1630(0.74); C1650(0.74); C2450(0.87); C1865(0.90) | LDD0716 | [12] |
| LDCM0400 | CL13 | HCT 116 | C2169(0.82); C2274(0.88); C1630(0.92); C1650(0.92) | LDD0717 | [12] |
| LDCM0401 | CL14 | HCT 116 | C1630(0.84); C1650(0.84); C1904(0.87); C2169(0.94) | LDD0718 | [12] |
| LDCM0402 | CL15 | HCT 116 | C2169(0.73); C1692(0.76); C1155(0.81); C1904(0.87) | LDD0719 | [12] |
| LDCM0403 | CL16 | HCT 116 | C2274(0.92); C1630(0.95); C1650(0.95); C2169(0.96) | LDD0720 | [12] |
| LDCM0404 | CL17 | HCT 116 | C2169(0.85); C718(0.87); C731(0.87); C1692(0.93) | LDD0721 | [12] |
| LDCM0405 | CL18 | HCT 116 | C2450(0.90); C880(0.91); C718(1.00); C731(1.00) | LDD0722 | [12] |
| LDCM0406 | CL19 | HCT 116 | C2169(0.76); C1692(0.79); C1732(0.94); C1155(0.98) | LDD0723 | [12] |
| LDCM0407 | CL2 | HCT 116 | C312(0.68); C1630(0.81); C1650(0.81); C880(0.91) | LDD0724 | [12] |
| LDCM0408 | CL20 | HCT 116 | C2169(0.75); C1692(0.90); C2450(0.94); C718(1.00) | LDD0725 | [12] |
| LDCM0409 | CL21 | HCT 116 | C2169(0.72); C2450(0.89); C1692(0.94); C718(0.98) | LDD0726 | [12] |
| LDCM0410 | CL22 | HCT 116 | C2169(0.65); C1692(0.77); C1155(0.88); C730(0.98) | LDD0727 | [12] |
| LDCM0411 | CL23 | HCT 116 | C2169(0.92); C1630(0.97); C1650(0.97); C880(1.05) | LDD0728 | [12] |
| LDCM0412 | CL24 | HCT 116 | C1732(1.31); C1155(1.13); C1630(1.49); C1650(1.49) | LDD0729 | [12] |
| LDCM0413 | CL25 | HCT 116 | C1732(1.63); C1155(1.03); C1630(1.24); C1650(1.24) | LDD0730 | [12] |
| LDCM0414 | CL26 | HCT 116 | C1732(2.47); C1155(1.11); C1630(1.17); C1650(1.17) | LDD0731 | [12] |
| LDCM0415 | CL27 | HCT 116 | C1732(1.65); C1155(1.07); C1630(1.30); C1650(1.30) | LDD0732 | [12] |
| LDCM0416 | CL28 | HCT 116 | C1732(1.46); C1155(1.22); C1630(1.30); C1650(1.30) | LDD0733 | [12] |
| LDCM0417 | CL29 | HCT 116 | C1732(1.18); C1155(1.21); C1630(1.37); C1650(1.37) | LDD0734 | [12] |
| LDCM0418 | CL3 | HCT 116 | C1155(1.04); C1630(1.00); C1650(1.00); C1692(0.94) | LDD0735 | [12] |
| LDCM0419 | CL30 | HCT 116 | C1732(1.21); C1155(1.04); C1630(1.03); C1650(1.03) | LDD0736 | [12] |
| LDCM0420 | CL31 | HCT 116 | C1732(1.53); C1155(0.87); C1735(0.63); C1865(0.80) | LDD0737 | [12] |
| LDCM0421 | CL32 | HCT 116 | C1732(1.51); C1155(1.05); C1734(1.28); C1735(0.91) | LDD0738 | [12] |
| LDCM0422 | CL33 | HCT 116 | C1732(1.12); C1155(0.99); C1734(1.11); C1735(0.79) | LDD0739 | [12] |
| LDCM0423 | CL34 | HCT 116 | C1732(1.20); C1155(1.04); C1734(0.90); C1735(1.11) | LDD0740 | [12] |
| LDCM0424 | CL35 | HCT 116 | C1732(5.32); C1155(1.13); C1734(1.07); C1735(1.04) | LDD0741 | [12] |
| LDCM0425 | CL36 | HCT 116 | C1732(1.34); C1155(1.10); C1734(0.90); C1735(0.79) | LDD0742 | [12] |
| LDCM0426 | CL37 | HCT 116 | C1732(1.33); C1155(1.08); C1734(0.92); C1735(1.67) | LDD0743 | [12] |
| LDCM0428 | CL39 | HCT 116 | C1732(1.70); C1155(1.01); C1734(1.24); C1735(0.74) | LDD0745 | [12] |
| LDCM0429 | CL4 | HCT 116 | C1155(1.01); C1630(1.22); C1650(1.22); C1692(0.89) | LDD0746 | [12] |
| LDCM0430 | CL40 | HCT 116 | C1732(1.23); C1155(0.97); C1734(1.10); C1735(0.65) | LDD0747 | [12] |
| LDCM0431 | CL41 | HCT 116 | C1732(1.32); C1155(1.19); C1734(1.06); C1735(0.58) | LDD0748 | [12] |
| LDCM0432 | CL42 | HCT 116 | C1732(2.04); C1155(0.91); C1734(0.85); C1735(0.67) | LDD0749 | [12] |
| LDCM0433 | CL43 | HCT 116 | C1732(1.80); C1155(1.03); C1734(0.99); C1735(0.71) | LDD0750 | [12] |
| LDCM0434 | CL44 | HCT 116 | C1732(1.96); C1155(0.88); C1734(0.89); C1735(0.64) | LDD0751 | [12] |
| LDCM0435 | CL45 | HCT 116 | C1732(1.85); C1155(1.05); C1734(0.95); C1735(0.90) | LDD0752 | [12] |
| LDCM0436 | CL46 | HCT 116 | C1732(1.05); C1155(0.96); C1630(0.76); C1650(0.76) | LDD0753 | [12] |
| LDCM0437 | CL47 | HCT 116 | C1732(0.94); C1155(0.84); C1630(0.57); C1650(0.57) | LDD0754 | [12] |
| LDCM0438 | CL48 | HCT 116 | C1732(0.95); C1155(0.79); C1630(0.74); C1650(0.74) | LDD0755 | [12] |
| LDCM0439 | CL49 | HCT 116 | C1732(0.82); C1155(0.82); C1630(0.85); C1650(0.85) | LDD0756 | [12] |
| LDCM0440 | CL5 | HCT 116 | C1155(1.07); C1630(1.01); C1650(1.01); C1692(0.92) | LDD0757 | [12] |
| LDCM0441 | CL50 | HCT 116 | C1732(0.96); C1155(0.85); C1630(0.96); C1650(0.96) | LDD0758 | [12] |
| LDCM0442 | CL51 | HCT 116 | C1732(1.03); C1155(1.15); C1630(1.18); C1650(1.18) | LDD0759 | [12] |
| LDCM0443 | CL52 | HCT 116 | C1732(1.13); C1155(1.11); C1630(1.12); C1650(1.12) | LDD0760 | [12] |
| LDCM0444 | CL53 | HCT 116 | C1732(1.00); C1155(0.95); C1630(0.84); C1650(0.84) | LDD0761 | [12] |
| LDCM0445 | CL54 | HCT 116 | C1732(0.99); C1155(1.00); C1630(0.76); C1650(0.76) | LDD0762 | [12] |
| LDCM0446 | CL55 | HCT 116 | C1732(0.88); C1155(0.95); C1630(0.95); C1650(0.95) | LDD0763 | [12] |
| LDCM0447 | CL56 | HCT 116 | C1732(0.95); C1155(0.85); C1630(0.78); C1650(0.78) | LDD0764 | [12] |
| LDCM0448 | CL57 | HCT 116 | C1732(0.92); C1155(0.75); C1630(0.82); C1650(0.82) | LDD0765 | [12] |
| LDCM0449 | CL58 | HCT 116 | C1732(0.99); C1155(0.85); C1630(0.76); C1650(0.76) | LDD0766 | [12] |
| LDCM0450 | CL59 | HCT 116 | C1732(0.91); C1155(0.84); C1630(0.72); C1650(0.72) | LDD0767 | [12] |
| LDCM0451 | CL6 | HCT 116 | C1155(1.17); C1630(0.89); C1650(0.89); C1692(1.03) | LDD0768 | [12] |
| LDCM0452 | CL60 | HCT 116 | C1732(1.13); C1155(0.89); C1630(0.75); C1650(0.75) | LDD0769 | [12] |
| LDCM0453 | CL61 | HCT 116 | C1732(0.87); C1155(1.21); C1630(1.22); C1650(1.22) | LDD0770 | [12] |
| LDCM0454 | CL62 | HCT 116 | C1732(0.98); C1155(1.32); C1630(1.15); C1650(1.15) | LDD0771 | [12] |
| LDCM0455 | CL63 | HCT 116 | C1732(0.98); C1155(0.97); C1630(1.38); C1650(1.38) | LDD0772 | [12] |
| LDCM0456 | CL64 | HCT 116 | C1732(0.70); C1155(0.95); C1630(1.26); C1650(1.26) | LDD0773 | [12] |
| LDCM0457 | CL65 | HCT 116 | C1732(0.80); C1155(1.03); C1630(1.10); C1650(1.10) | LDD0774 | [12] |
| LDCM0458 | CL66 | HCT 116 | C1732(0.89); C1155(1.02); C1630(1.49); C1650(1.49) | LDD0775 | [12] |
| LDCM0459 | CL67 | HCT 116 | C1732(0.88); C1155(1.03); C1630(1.34); C1650(1.34) | LDD0776 | [12] |
| LDCM0460 | CL68 | HCT 116 | C1732(0.75); C1155(1.10); C1630(1.09); C1650(1.09) | LDD0777 | [12] |
| LDCM0461 | CL69 | HCT 116 | C1732(1.23); C1155(1.10); C1630(0.76); C1650(0.76) | LDD0778 | [12] |
| LDCM0462 | CL7 | HCT 116 | C1155(1.00); C1630(1.05); C1650(1.05); C1692(0.79) | LDD0779 | [12] |
| LDCM0463 | CL70 | HCT 116 | C1732(0.97); C1155(1.10); C1630(1.12); C1650(1.12) | LDD0780 | [12] |
| LDCM0464 | CL71 | HCT 116 | C1732(0.98); C1155(1.07); C1630(1.31); C1650(1.31) | LDD0781 | [12] |
| LDCM0465 | CL72 | HCT 116 | C1732(0.83); C1155(1.30); C1630(1.03); C1650(1.03) | LDD0782 | [12] |
| LDCM0466 | CL73 | HCT 116 | C1732(0.85); C1155(1.37); C1630(1.16); C1650(1.16) | LDD0783 | [12] |
| LDCM0467 | CL74 | HCT 116 | C1732(0.94); C1155(1.11); C1630(1.32); C1650(1.32) | LDD0784 | [12] |
| LDCM0469 | CL76 | HCT 116 | C1732(0.72); C1155(0.75); C1630(0.96); C1650(0.96) | LDD0786 | [12] |
| LDCM0470 | CL77 | HCT 116 | C1732(0.77); C1155(0.73); C1630(0.77); C1650(0.77) | LDD0787 | [12] |
| LDCM0471 | CL78 | HCT 116 | C1732(0.97); C1155(0.84); C1630(1.04); C1650(1.04) | LDD0788 | [12] |
| LDCM0472 | CL79 | HCT 116 | C1732(0.75); C1155(0.71); C1630(1.24); C1650(1.24) | LDD0789 | [12] |
| LDCM0473 | CL8 | HCT 116 | C1155(1.05); C1630(1.30); C1650(1.30); C1692(1.09) | LDD0790 | [12] |
| LDCM0474 | CL80 | HCT 116 | C1732(0.90); C1155(0.78); C1630(0.86); C1650(0.86) | LDD0791 | [12] |
| LDCM0475 | CL81 | HCT 116 | C1732(0.76); C1155(0.68); C1630(0.92); C1650(0.92) | LDD0792 | [12] |
| LDCM0476 | CL82 | HCT 116 | C1732(0.67); C1155(0.66); C1630(1.21); C1650(1.21) | LDD0793 | [12] |
| LDCM0477 | CL83 | HCT 116 | C1732(0.81); C1155(0.73); C1630(1.18); C1650(1.18) | LDD0794 | [12] |
| LDCM0478 | CL84 | HCT 116 | C1732(0.88); C1155(0.83); C1630(1.32); C1650(1.32) | LDD0795 | [12] |
| LDCM0479 | CL85 | HCT 116 | C1732(0.93); C1155(0.75); C1630(1.05); C1650(1.05) | LDD0796 | [12] |
| LDCM0480 | CL86 | HCT 116 | C1732(0.82); C1155(0.73); C1630(0.76); C1650(0.76) | LDD0797 | [12] |
| LDCM0481 | CL87 | HCT 116 | C1732(0.75); C1155(0.72); C1630(0.76); C1650(0.76) | LDD0798 | [12] |
| LDCM0482 | CL88 | HCT 116 | C1732(0.77); C1155(0.69); C1630(1.38); C1650(1.38) | LDD0799 | [12] |
| LDCM0483 | CL89 | HCT 116 | C1732(0.80); C1155(0.75); C1630(1.53); C1650(1.53) | LDD0800 | [12] |
| LDCM0484 | CL9 | HCT 116 | C1155(1.11); C1630(0.95); C1650(0.95); C1692(0.85) | LDD0801 | [12] |
| LDCM0485 | CL90 | HCT 116 | C1732(0.70); C1155(0.60); C1630(0.92); C1650(0.92) | LDD0802 | [12] |
| LDCM0486 | CL91 | HCT 116 | C1732(0.88); C1155(0.86); C1734(0.87); C1735(0.88) | LDD0803 | [12] |
| LDCM0487 | CL92 | HCT 116 | C1732(0.89); C1155(0.91); C1734(0.97); C1735(0.89) | LDD0804 | [12] |
| LDCM0488 | CL93 | HCT 116 | C1732(0.71); C1155(0.81); C1734(0.67); C1735(0.71) | LDD0805 | [12] |
| LDCM0489 | CL94 | HCT 116 | C1732(0.95); C1155(0.90); C1734(0.92); C1735(0.95) | LDD0806 | [12] |
| LDCM0490 | CL95 | HCT 116 | C1732(0.94); C1155(0.89); C1734(0.91); C1735(0.94) | LDD0807 | [12] |
| LDCM0491 | CL96 | HCT 116 | C1732(0.92); C1155(0.80); C1734(0.94); C1735(0.92) | LDD0808 | [12] |
| LDCM0492 | CL97 | HCT 116 | C1732(0.82); C1155(0.87); C1734(0.87); C1735(0.82) | LDD0809 | [12] |
| LDCM0493 | CL98 | HCT 116 | C1732(0.93); C1155(1.02); C1734(0.89); C1735(0.93) | LDD0810 | [12] |
| LDCM0494 | CL99 | HCT 116 | C1732(0.86); C1155(0.85); C1734(0.78); C1735(0.86) | LDD0811 | [12] |
| LDCM0634 | CY-0357 | Hep-G2 | C2169(0.64); C1692(0.24) | LDD2228 | [16] |
| LDCM0028 | Dobutamine | HEK-293T | 4.57 | LDD0180 | [8] |
| LDCM0495 | E2913 | HEK-293T | C1692(1.14); C1155(0.90); C1718(0.98); C880(1.07) | LDD1698 | [27] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C2274(3.55); C2169(1.62); C2450(1.42); C1718(1.07) | LDD1702 | [7] |
| LDCM0625 | F8 | Ramos | C2450(1.05); C2169(1.07) | LDD2187 | [28] |
| LDCM0572 | Fragment10 | MDA-MB-231 | C2298(20.00) | LDD1465 | [29] |
| LDCM0573 | Fragment11 | MDA-MB-231 | C2298(1.34) | LDD1467 | [29] |
| LDCM0574 | Fragment12 | Ramos | C2450(1.26); C1692(0.59) | LDD2191 | [28] |
| LDCM0575 | Fragment13 | Ramos | C2450(1.14); C1692(0.75); C2169(1.00) | LDD2192 | [28] |
| LDCM0576 | Fragment14 | MDA-MB-231 | C2298(20.00) | LDD1471 | [29] |
| LDCM0579 | Fragment20 | Ramos | C2450(2.09); C1692(0.72) | LDD2194 | [28] |
| LDCM0580 | Fragment21 | MDA-MB-231 | C2298(0.89) | LDD1473 | [29] |
| LDCM0582 | Fragment23 | Ramos | C2450(0.69); C1692(0.85) | LDD2196 | [28] |
| LDCM0578 | Fragment27 | Ramos | C2450(0.89); C1692(0.85); C2169(0.77) | LDD2197 | [28] |
| LDCM0586 | Fragment28 | Ramos | C2450(0.83); C1692(0.72); C2169(1.02) | LDD2198 | [28] |
| LDCM0588 | Fragment30 | Ramos | C2450(1.08); C1692(0.93) | LDD2199 | [28] |
| LDCM0589 | Fragment31 | Ramos | C2450(0.97); C1692(0.79) | LDD2200 | [28] |
| LDCM0590 | Fragment32 | Ramos | C2450(1.66); C2169(1.08) | LDD2201 | [28] |
| LDCM0468 | Fragment33 | HCT 116 | C1732(0.89); C1155(1.11); C1630(1.37); C1650(1.37) | LDD0785 | [12] |
| LDCM0596 | Fragment38 | Ramos | C2450(1.23); C1692(0.73); C2169(0.96) | LDD2203 | [28] |
| LDCM0566 | Fragment4 | MDA-MB-231 | C2298(20.00) | LDD1461 | [29] |
| LDCM0599 | Fragment41 | MDA-MB-231 | C2298(1.21) | LDD1481 | [29] |
| LDCM0603 | Fragment45 | MDA-MB-231 | C2298(20.00) | LDD1482 | [29] |
| LDCM0427 | Fragment51 | HCT 116 | C1732(2.19); C1155(1.06); C1734(1.12); C1735(0.84) | LDD0744 | [12] |
| LDCM0610 | Fragment52 | Ramos | C2450(1.04) | LDD2204 | [28] |
| LDCM0614 | Fragment56 | Ramos | C2450(1.19) | LDD2205 | [28] |
| LDCM0569 | Fragment7 | Ramos | C2450(2.44) | LDD2186 | [28] |
| LDCM0571 | Fragment9 | Ramos | C1692(0.75); C2169(0.37) | LDD2188 | [28] |
| LDCM0116 | HHS-0101 | DM93 | Y404(0.34) | LDD0264 | [11] |
| LDCM0119 | HHS-0401 | DM93 | Y404(0.22) | LDD0267 | [11] |
| LDCM0120 | HHS-0701 | DM93 | Y404(0.21) | LDD0268 | [11] |
| LDCM0107 | IAA | HeLa | H2362(0.00); H2229(0.00) | LDD0221 | [23] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [6] |
| LDCM0022 | KB02 | HCT 116 | C730(16.58); C731(16.58); C880(7.67); C2169(2.61) | LDD0080 | [12] |
| LDCM0023 | KB03 | HCT 116 | C730(20.00); C731(20.00); C880(7.95); C2169(1.70) | LDD0081 | [12] |
| LDCM0024 | KB05 | HCT 116 | C730(20.00); C731(20.00); C880(7.19); C2169(2.41) | LDD0082 | [12] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C1865(0.94) | LDD2102 | [7] |
| LDCM0109 | NEM | HeLa | H2229(0.00); H483(0.00); H474(0.00) | LDD0223 | [23] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C2274(0.64) | LDD2089 | [7] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C2274(0.89) | LDD2093 | [7] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C2274(1.07) | LDD2094 | [7] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C2450(0.38) | LDD2096 | [7] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C2274(0.93) | LDD2098 | [7] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C2274(1.12) | LDD2105 | [7] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C2450(1.06) | LDD2107 | [7] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C2274(2.00); C1865(0.93) | LDD2109 | [7] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C1865(1.11) | LDD2111 | [7] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C2450(1.02) | LDD2118 | [7] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C2274(0.88); C2450(1.03) | LDD2123 | [7] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C2274(1.19); C2450(0.71) | LDD2125 | [7] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C2450(1.22) | LDD2126 | [7] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C2274(1.66); C2450(0.97) | LDD2127 | [7] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C2274(0.80) | LDD2128 | [7] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C2274(1.36) | LDD2136 | [7] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C2450(0.86) | LDD2137 | [7] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C2274(1.12); C2450(0.87) | LDD2140 | [7] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C2274(0.87) | LDD2143 | [7] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C2450(1.89) | LDD2144 | [7] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C2450(1.20); C82(0.35) | LDD2206 | [30] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C82(0.63); C2450(0.59) | LDD2207 | [30] |
| LDCM0131 | RA190 | MM1.R | 4.48 | LDD0299 | [10] |
| LDCM0021 | THZ1 | HCT 116 | C880(1.10); C1732(1.16); C1734(1.16); C1735(1.16) | LDD2173 | [12] |
The Interaction Atlas With This Target
References
