Details of the Target
General Information of Target
| Target ID | LDTP05037 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ras-related C3 botulinum toxin substrate 1 (RAC1) | |||||
| Gene Name | RAC1 | |||||
| Gene ID | 5879 | |||||
| Synonyms |
TC25; Ras-related C3 botulinum toxin substrate 1; EC 3.6.5.2; Cell migration-inducing gene 5 protein; Ras-like protein TC25; p21-Rac1 |
|||||
| 3D Structure | ||||||
| Sequence |
MQAIKCVVVGDGAVGKTCLLISYTTNAFPGEYIPTVFDNYSANVMVDGKPVNLGLWDTAG
QEDYDRLRPLSYPQTDVFLICFSLVSPASFENVRAKWYPEVRHHCPNTPIILVGTKLDLR DDKDTIEKLKEKKLTPITYPQGLAMAKEIGAVKYLECSALTQRGLKTVFDEAIRAVLCPP PVKKRKRKCLLL |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Small GTPase superfamily, Rho family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Plasma membrane-associated small GTPase which cycles between active GTP-bound and inactive GDP-bound states. In its active state, binds to a variety of effector proteins to regulate cellular responses such as secretory processes, phagocytosis of apoptotic cells, epithelial cell polarization, neurons adhesion, migration and differentiation, and growth-factor induced formation of membrane ruffles. Rac1 p21/rho GDI heterodimer is the active component of the cytosolic factor sigma 1, which is involved in stimulation of the NADPH oxidase activity in macrophages. Essential for the SPATA13-mediated regulation of cell migration and adhesion assembly and disassembly. Stimulates PKN2 kinase activity. In concert with RAB7A, plays a role in regulating the formation of RBs (ruffled borders) in osteoclasts. In podocytes, promotes nuclear shuttling of NR3C2; this modulation is required for a proper kidney functioning. Required for atypical chemokine receptor ACKR2-induced LIMK1-PAK1-dependent phosphorylation of cofilin (CFL1) and for up-regulation of ACKR2 from endosomal compartment to cell membrane, increasing its efficiency in chemokine uptake and degradation. In neurons, is involved in dendritic spine formation and synaptic plasticity. In hippocampal neurons, involved in spine morphogenesis and synapse formation, through local activation at synapses by guanine nucleotide exchange factors (GEFs), such as ARHGEF6/ARHGEF7/PIX. In synapses, seems to mediate the regulation of F-actin cluster formation performed by SHANK3. In neurons, plays a crucial role in regulating GABA(A) receptor synaptic stability and hence GABAergic inhibitory synaptic transmission through its role in PAK1 activation and eventually F-actin stabilization.; [Isoform B]: Isoform B has an accelerated GEF-independent GDP/GTP exchange and an impaired GTP hydrolysis, which is restored partially by GTPase-activating proteins. It is able to bind to the GTPase-binding domain of PAK but not full-length PAK in a GTP-dependent manner, suggesting that the insertion does not completely abolish effector interaction.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CAL51 | SNV: p.N39S | DBIA Probe Info | |||
| CMK | SNV: p.I21M | DBIA Probe Info | |||
| DEL | SNV: p.V85M; p.A88E | DBIA Probe Info | |||
| HT1080 | SNV: p.N92I | DBIA Probe Info | |||
| IGR1 | SNV: p.P29S | DBIA Probe Info | |||
| JURKAT | SNV: p.C18Y | Compound 10 Probe Info | |||
| KYSE150 | SNV: p.P29S | DBIA Probe Info | |||
| MDAMB157 | SNV: p.P29S | DBIA Probe Info | |||
| MDST8 | SNV: p.D11E | DBIA Probe Info | |||
| SHP77 | SNV: p.Y32C | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
14.93 | LDD0402 | [1] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [2] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [2] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
TH211 Probe Info |
 |
Y154(5.50); Y139(20.00); Y98(10.03) | LDD0260 | [3] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [4] | |
|
ONAyne Probe Info |
 |
K166(1.82) | LDD0274 | [5] | |
|
STPyne Probe Info |
 |
K133(4.93); K147(9.73); K183(7.52); K96(8.42) | LDD0277 | [5] | |
|
BTD Probe Info |
 |
C178(1.35); C157(3.79) | LDD1699 | [6] | |
|
OPA-S-S-alkyne Probe Info |
 |
K153(2.29) | LDD3494 | [7] | |
|
Probe 1 Probe Info |
 |
Y139(14.22) | LDD3495 | [8] | |
|
DBIA Probe Info |
 |
C6(1.61); C124(1.13) | LDD3310 | [9] | |
|
JZ128-DTB Probe Info |
 |
C178(0.00); C157(0.00) | LDD0462 | [10] | |
|
AHL-Pu-1 Probe Info |
 |
C178(5.75) | LDD0171 | [11] | |
|
HHS-482 Probe Info |
 |
Y139(0.50) | LDD0285 | [12] | |
|
HHS-475 Probe Info |
 |
Y98(0.87); Y139(0.98) | LDD0264 | [13] | |
|
HHS-465 Probe Info |
 |
Y139(10.00); Y98(6.31) | LDD2237 | [14] | |
|
ATP probe Probe Info |
 |
K183(0.00); K184(0.00); K166(0.00) | LDD0199 | [15] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C178(0.00); C105(0.00); C81(0.00); C157(0.00) | LDD0038 | [16] | |
|
IA-alkyne Probe Info |
 |
C157(0.00); C178(0.00); C6(0.00) | LDD0032 | [17] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [18] | |
|
IPIAA_L Probe Info |
 |
C6(0.00); C157(0.00); C81(0.00) | LDD0031 | [18] | |
|
Lodoacetamide azide Probe Info |
 |
C178(0.00); C105(0.00); C81(0.00); C157(0.00) | LDD0037 | [16] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [19] | |
|
IPM Probe Info |
 |
C157(0.00); C105(0.00); C178(0.00) | LDD0025 | [20] | |
|
JW-RF-010 Probe Info |
 |
C157(0.00); C178(0.00); C6(0.00); C105(0.00) | LDD0026 | [20] | |
|
TFBX Probe Info |
 |
C157(0.00); C178(0.00); C6(0.00) | LDD0027 | [20] | |
|
WYneN Probe Info |
 |
C105(0.00); C157(0.00); C178(0.00); C6(0.00) | LDD0021 | [21] | |
|
aHNE Probe Info |
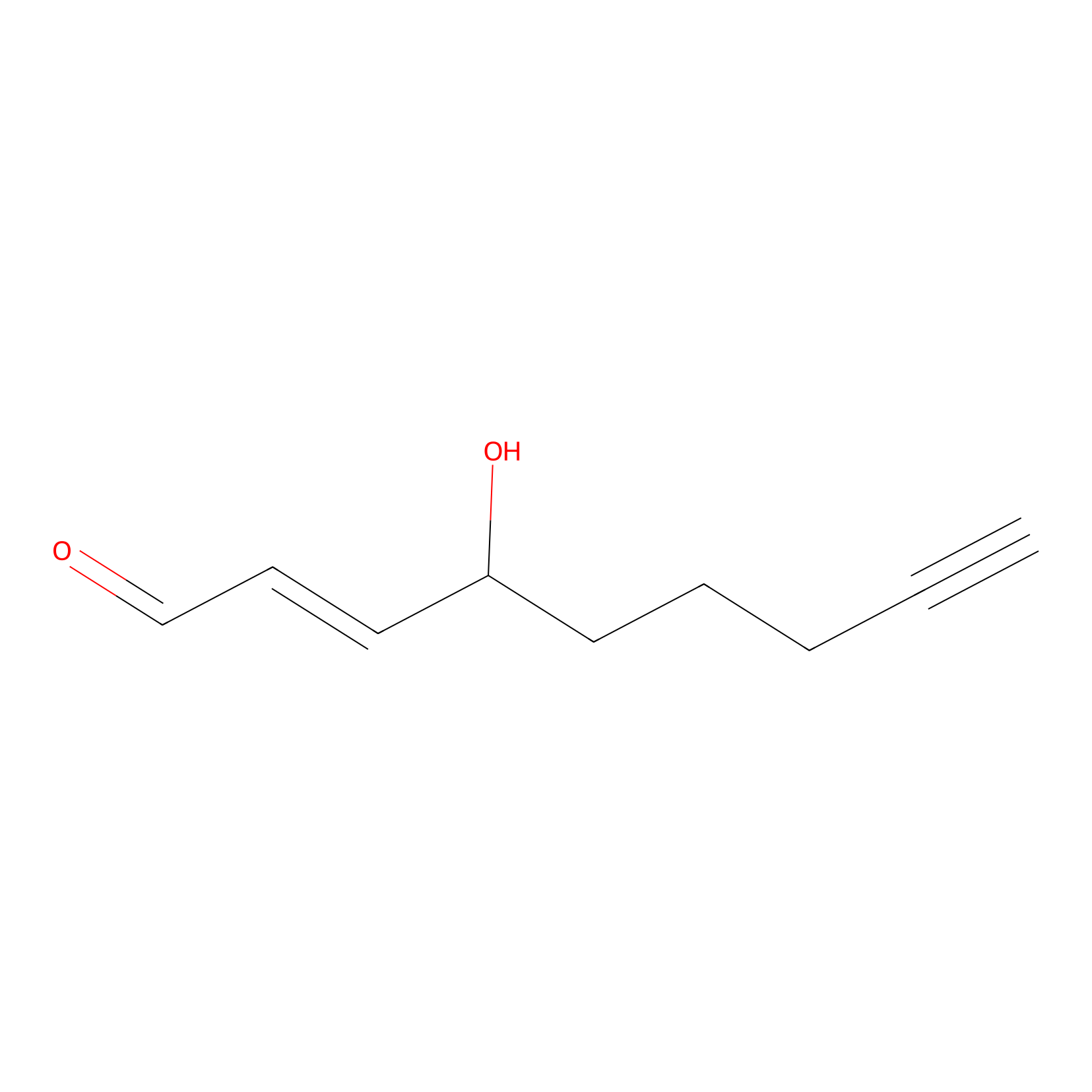 |
N.A. | LDD0001 | [21] | |
|
Compound 10 Probe Info |
 |
C157(0.00); C178(0.00) | LDD2216 | [22] | |
|
Compound 11 Probe Info |
 |
C157(0.00); C178(0.00) | LDD2213 | [22] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0017 | [23] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [24] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [25] | |
|
Acrolein Probe Info |
 |
C105(0.00); C178(0.00); H104(0.00) | LDD0217 | [26] | |
|
Cinnamaldehyde Probe Info |
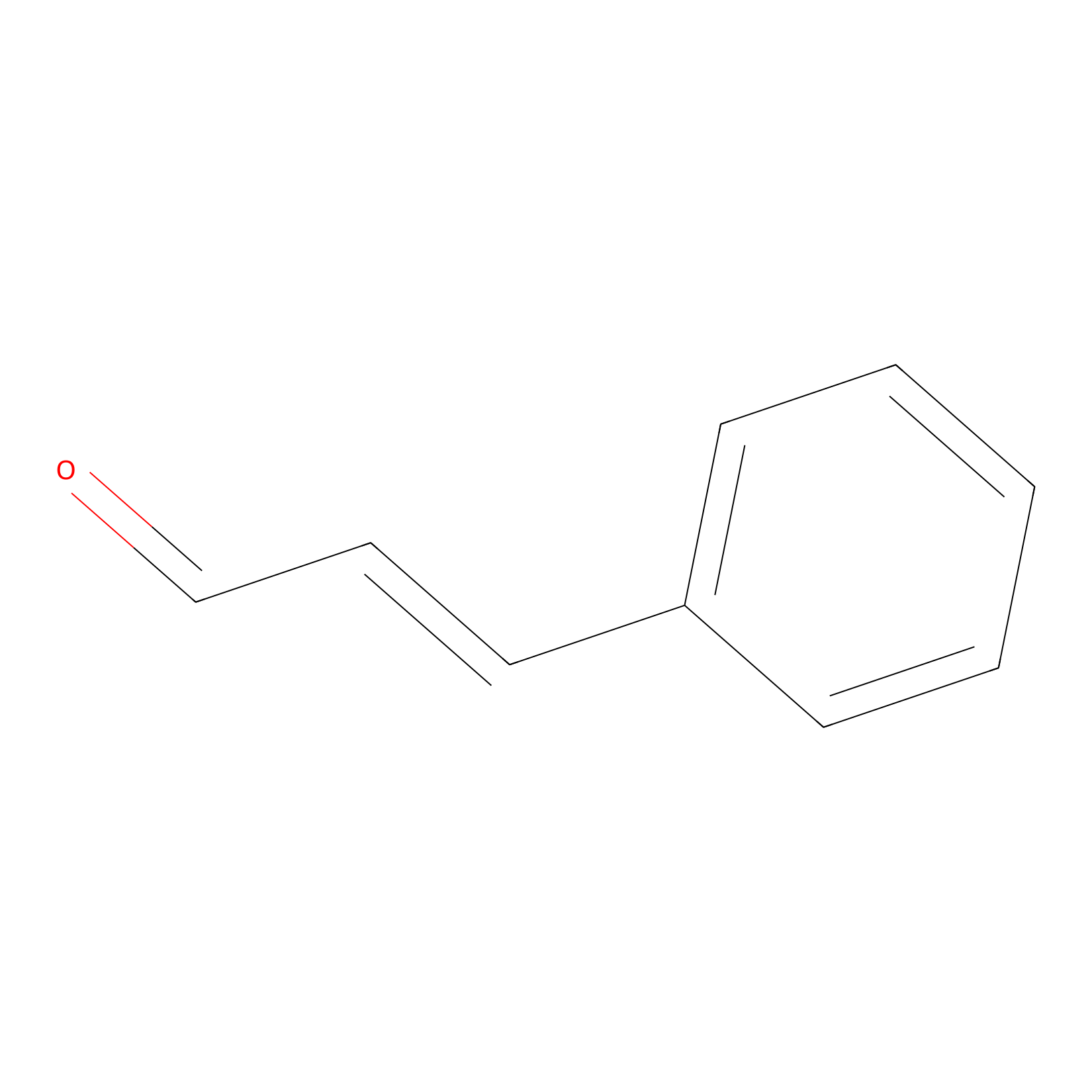 |
N.A. | LDD0220 | [26] | |
|
Crotonaldehyde Probe Info |
 |
C105(0.00); C178(0.00) | LDD0219 | [26] | |
|
Methacrolein Probe Info |
 |
C105(0.00); C178(0.00) | LDD0218 | [26] | |
|
W1 Probe Info |
 |
C105(0.00); C178(0.00) | LDD0236 | [27] | |
|
AOyne Probe Info |
 |
11.30 | LDD0443 | [28] | |
|
MPP-AC Probe Info |
 |
N.A. | LDD0428 | [29] | |
|
NAIA_5 Probe Info |
 |
C178(0.00); C157(0.00); C105(0.00) | LDD2223 | [30] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [29] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C178(0.59); C105(0.39); C157(0.29) | LDD2142 | [6] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C178(0.78) | LDD2112 | [6] |
| LDCM0502 | 1-(Cyanoacetyl)piperidine | MDA-MB-231 | C178(0.62); C105(0.59) | LDD2095 | [6] |
| LDCM0537 | 2-Cyano-N,N-dimethylacetamide | MDA-MB-231 | C178(0.87); C105(0.89) | LDD2130 | [6] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C178(0.80); C105(1.38) | LDD2117 | [6] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C178(1.08); C105(1.61) | LDD2152 | [6] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C178(1.04); C157(1.35) | LDD2103 | [6] |
| LDCM0539 | 3-(4-Isopropylpiperazin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C178(0.58); C105(0.59) | LDD2132 | [6] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C178(0.69); C105(0.59) | LDD2131 | [6] |
| LDCM0025 | 4SU-RNA | HEK-293T | C157(2.70) | LDD0172 | [11] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C178(5.75) | LDD0171 | [11] |
| LDCM0214 | AC1 | HEK-293T | C105(0.85); C157(1.01); C178(0.94); C81(0.91) | LDD1507 | [31] |
| LDCM0215 | AC10 | HEK-293T | C105(0.92); C157(0.95); C178(0.96); C81(0.85) | LDD1508 | [31] |
| LDCM0226 | AC11 | HEK-293T | C105(0.90); C157(1.03); C178(0.92); C18(0.96) | LDD1509 | [31] |
| LDCM0237 | AC12 | HEK-293T | C105(0.94); C157(0.96); C178(0.97); C81(0.97) | LDD1510 | [31] |
| LDCM0259 | AC14 | HEK-293T | C105(0.93); C157(1.10); C178(0.96); C81(0.91) | LDD1512 | [31] |
| LDCM0270 | AC15 | HEK-293T | C105(0.92); C157(0.99); C178(0.92); C81(1.08) | LDD1513 | [31] |
| LDCM0276 | AC17 | HEK-293T | C105(0.92); C157(0.93); C178(0.97); C81(1.02) | LDD1515 | [31] |
| LDCM0277 | AC18 | HEK-293T | C105(1.00); C157(1.18); C178(0.93); C81(0.72) | LDD1516 | [31] |
| LDCM0278 | AC19 | HEK-293T | C105(0.81); C157(1.05); C178(0.97); C18(0.87) | LDD1517 | [31] |
| LDCM0279 | AC2 | HEK-293T | C105(1.00); C157(0.94); C178(1.01); C81(0.95) | LDD1518 | [31] |
| LDCM0280 | AC20 | HEK-293T | C105(0.96); C157(1.02); C178(0.94); C81(1.09) | LDD1519 | [31] |
| LDCM0281 | AC21 | HEK-293T | C105(0.98); C157(0.97); C178(0.95); C81(0.92) | LDD1520 | [31] |
| LDCM0282 | AC22 | HEK-293T | C105(0.93); C157(0.93); C178(0.94); C81(0.93) | LDD1521 | [31] |
| LDCM0283 | AC23 | HEK-293T | C105(0.96); C157(0.98); C178(0.96); C81(0.90) | LDD1522 | [31] |
| LDCM0284 | AC24 | HEK-293T | C105(0.91); C157(1.05); C178(0.99); C81(0.89) | LDD1523 | [31] |
| LDCM0285 | AC25 | HEK-293T | C105(0.95); C157(1.12); C178(0.96); C81(1.02) | LDD1524 | [31] |
| LDCM0286 | AC26 | HEK-293T | C105(0.91); C157(1.13); C178(0.98); C81(0.99) | LDD1525 | [31] |
| LDCM0287 | AC27 | HEK-293T | C105(0.99); C157(0.99); C178(0.96); C18(0.90) | LDD1526 | [31] |
| LDCM0288 | AC28 | HEK-293T | C105(0.91); C157(1.09); C178(0.95); C81(1.03) | LDD1527 | [31] |
| LDCM0289 | AC29 | HEK-293T | C105(0.99); C157(1.04); C178(0.96); C81(0.95) | LDD1528 | [31] |
| LDCM0290 | AC3 | HEK-293T | C105(0.96); C157(0.97); C178(0.94); C18(0.86) | LDD1529 | [31] |
| LDCM0291 | AC30 | HEK-293T | C105(0.98); C157(1.09); C178(0.99); C81(0.97) | LDD1530 | [31] |
| LDCM0292 | AC31 | HEK-293T | C105(0.95); C157(1.17); C178(0.92); C81(0.95) | LDD1531 | [31] |
| LDCM0293 | AC32 | HEK-293T | C105(0.97); C157(0.93); C178(0.97); C81(0.95) | LDD1532 | [31] |
| LDCM0294 | AC33 | HEK-293T | C105(0.89); C157(0.98); C178(0.98); C81(1.13) | LDD1533 | [31] |
| LDCM0295 | AC34 | HEK-293T | C105(0.85); C157(1.29); C178(0.98); C81(0.89) | LDD1534 | [31] |
| LDCM0296 | AC35 | HEK-293T | C105(0.99); C157(1.04); C178(0.96); C18(1.04) | LDD1535 | [31] |
| LDCM0297 | AC36 | HEK-293T | C105(0.90); C157(1.03); C178(0.97); C81(1.06) | LDD1536 | [31] |
| LDCM0298 | AC37 | HEK-293T | C105(0.96); C157(1.02); C178(0.98); C81(1.06) | LDD1537 | [31] |
| LDCM0299 | AC38 | HEK-293T | C105(0.94); C157(0.95); C178(0.99); C81(1.06) | LDD1538 | [31] |
| LDCM0300 | AC39 | HEK-293T | C105(0.92); C157(0.97); C178(0.98); C81(0.87) | LDD1539 | [31] |
| LDCM0301 | AC4 | HEK-293T | C105(1.01); C157(1.03); C178(0.93); C81(1.07) | LDD1540 | [31] |
| LDCM0302 | AC40 | HEK-293T | C105(0.95); C157(1.06); C178(1.01); C81(0.91) | LDD1541 | [31] |
| LDCM0303 | AC41 | HEK-293T | C105(0.92); C157(0.98); C178(0.98); C81(0.99) | LDD1542 | [31] |
| LDCM0304 | AC42 | HEK-293T | C105(0.90); C157(1.18); C178(0.97); C81(0.96) | LDD1543 | [31] |
| LDCM0305 | AC43 | HEK-293T | C105(0.92); C157(1.03); C178(0.99); C18(0.95) | LDD1544 | [31] |
| LDCM0306 | AC44 | HEK-293T | C105(0.95); C157(0.99); C178(0.99); C81(1.05) | LDD1545 | [31] |
| LDCM0307 | AC45 | HEK-293T | C105(0.86); C157(1.05); C178(0.95); C81(0.97) | LDD1546 | [31] |
| LDCM0308 | AC46 | HEK-293T | C105(0.87); C157(0.99); C178(1.00); C81(0.93) | LDD1547 | [31] |
| LDCM0309 | AC47 | HEK-293T | C105(0.86); C157(0.99); C178(0.98); C81(1.02) | LDD1548 | [31] |
| LDCM0310 | AC48 | HEK-293T | C105(0.89); C157(1.25); C178(1.00); C81(1.10) | LDD1549 | [31] |
| LDCM0311 | AC49 | HEK-293T | C105(0.85); C157(1.01); C178(0.96); C81(1.13) | LDD1550 | [31] |
| LDCM0312 | AC5 | HEK-293T | C105(0.97); C157(1.02); C178(1.00); C81(1.00) | LDD1551 | [31] |
| LDCM0313 | AC50 | HEK-293T | C105(0.94); C157(1.32); C178(0.98); C81(0.94) | LDD1552 | [31] |
| LDCM0314 | AC51 | HEK-293T | C105(0.91); C157(1.01); C178(0.97); C18(0.93) | LDD1553 | [31] |
| LDCM0315 | AC52 | HEK-293T | C105(1.00); C157(1.02); C178(0.97); C81(1.15) | LDD1554 | [31] |
| LDCM0316 | AC53 | HEK-293T | C105(0.95); C157(0.99); C178(0.99); C81(1.13) | LDD1555 | [31] |
| LDCM0317 | AC54 | HEK-293T | C105(0.89); C157(0.99); C178(0.99); C81(1.07) | LDD1556 | [31] |
| LDCM0318 | AC55 | HEK-293T | C105(0.93); C157(1.02); C178(0.92); C81(0.93) | LDD1557 | [31] |
| LDCM0319 | AC56 | HEK-293T | C105(0.92); C157(1.16); C178(1.00); C81(1.19) | LDD1558 | [31] |
| LDCM0320 | AC57 | HEK-293T | C105(0.80); C157(1.02); C178(1.01); C81(1.15) | LDD1559 | [31] |
| LDCM0321 | AC58 | HEK-293T | C105(0.87); C157(0.92); C178(0.98); C81(0.70) | LDD1560 | [31] |
| LDCM0322 | AC59 | HEK-293T | C105(0.93); C157(1.00); C178(0.98); C18(1.15) | LDD1561 | [31] |
| LDCM0323 | AC6 | HEK-293T | C105(0.94); C157(0.97); C178(0.99); C81(0.95) | LDD1562 | [31] |
| LDCM0324 | AC60 | HEK-293T | C105(0.82); C157(0.92); C178(0.99); C81(1.18) | LDD1563 | [31] |
| LDCM0325 | AC61 | HEK-293T | C105(0.82); C157(1.00); C178(0.98); C81(0.90) | LDD1564 | [31] |
| LDCM0326 | AC62 | HEK-293T | C105(0.81); C157(1.05); C178(1.01); C81(0.98) | LDD1565 | [31] |
| LDCM0327 | AC63 | HEK-293T | C105(0.84); C157(0.91); C178(0.99); C81(0.89) | LDD1566 | [31] |
| LDCM0328 | AC64 | HEK-293T | C105(0.89); C157(1.12); C178(1.00); C81(1.10) | LDD1567 | [31] |
| LDCM0334 | AC7 | HEK-293T | C105(1.05); C157(0.92); C178(0.98); C81(0.87) | LDD1568 | [31] |
| LDCM0345 | AC8 | HEK-293T | C105(0.97); C157(1.31); C178(0.98); C81(1.05) | LDD1569 | [31] |
| LDCM0545 | Acetamide | MDA-MB-231 | C178(0.35); C105(0.53) | LDD2138 | [6] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C178(0.60); C105(0.76) | LDD2113 | [6] |
| LDCM0248 | AKOS034007472 | HEK-293T | C105(0.90); C157(1.02); C178(0.97); C81(1.01) | LDD1511 | [31] |
| LDCM0356 | AKOS034007680 | HEK-293T | C105(0.91); C157(1.02); C178(0.98); C81(1.06) | LDD1570 | [31] |
| LDCM0275 | AKOS034007705 | HEK-293T | C105(0.97); C157(1.23); C178(1.00); C81(0.95) | LDD1514 | [31] |
| LDCM0156 | Aniline | NCI-H1299 | 11.92 | LDD0403 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C178(0.40); C105(0.80) | LDD2091 | [6] |
| LDCM0108 | Chloroacetamide | HeLa | C105(0.00); C178(0.00) | LDD0222 | [26] |
| LDCM0632 | CL-Sc | Hep-G2 | C178(1.08); C105(0.73); C178(0.70); C157(0.52) | LDD2227 | [30] |
| LDCM0367 | CL1 | HEK-293T | C105(0.78); C157(0.94); C178(0.90); C81(1.20) | LDD1571 | [31] |
| LDCM0368 | CL10 | HEK-293T | C105(0.96); C157(1.20); C178(0.61); C81(0.95) | LDD1572 | [31] |
| LDCM0369 | CL100 | HEK-293T | C105(0.97); C157(1.03); C178(0.91); C81(1.05) | LDD1573 | [31] |
| LDCM0370 | CL101 | HEK-293T | C105(0.88); C157(0.83); C178(0.96); C81(1.16) | LDD1574 | [31] |
| LDCM0371 | CL102 | HEK-293T | C105(0.85); C157(0.86); C178(0.92); C81(1.03) | LDD1575 | [31] |
| LDCM0372 | CL103 | HEK-293T | C105(0.90); C157(0.92); C178(1.00); C81(1.01) | LDD1576 | [31] |
| LDCM0373 | CL104 | HEK-293T | C105(0.98); C157(0.98); C178(0.91); C81(1.27) | LDD1577 | [31] |
| LDCM0374 | CL105 | HEK-293T | C105(0.89); C157(0.97); C178(0.94); C81(0.88) | LDD1578 | [31] |
| LDCM0375 | CL106 | HEK-293T | C105(0.98); C157(0.90); C178(0.92); C81(1.00) | LDD1579 | [31] |
| LDCM0376 | CL107 | HEK-293T | C105(0.99); C157(0.91); C178(0.94); C81(1.06) | LDD1580 | [31] |
| LDCM0377 | CL108 | HEK-293T | C105(0.97); C157(1.05); C178(0.97); C81(1.02) | LDD1581 | [31] |
| LDCM0378 | CL109 | HEK-293T | C105(0.90); C157(0.83); C178(0.98); C81(0.99) | LDD1582 | [31] |
| LDCM0379 | CL11 | HEK-293T | C105(0.68); C157(1.21); C178(0.44); C81(0.91) | LDD1583 | [31] |
| LDCM0380 | CL110 | HEK-293T | C105(0.91); C157(0.89); C178(0.91); C81(0.73) | LDD1584 | [31] |
| LDCM0381 | CL111 | HEK-293T | C105(0.94); C157(1.00); C178(0.92); C81(1.01) | LDD1585 | [31] |
| LDCM0382 | CL112 | HEK-293T | C105(0.94); C157(1.01); C178(0.96); C81(1.02) | LDD1586 | [31] |
| LDCM0383 | CL113 | HEK-293T | C105(0.88); C157(0.94); C178(1.00); C81(1.06) | LDD1587 | [31] |
| LDCM0384 | CL114 | HEK-293T | C105(0.83); C157(0.87); C178(0.90); C81(0.93) | LDD1588 | [31] |
| LDCM0385 | CL115 | HEK-293T | C105(0.99); C157(1.00); C178(0.96); C81(1.01) | LDD1589 | [31] |
| LDCM0386 | CL116 | HEK-293T | C105(0.99); C157(0.96); C178(0.99); C81(1.10) | LDD1590 | [31] |
| LDCM0387 | CL117 | HEK-293T | C105(0.86); C157(0.94); C178(1.01); C81(1.14) | LDD1591 | [31] |
| LDCM0388 | CL118 | HEK-293T | C105(0.91); C157(0.86); C178(1.00); C81(1.13) | LDD1592 | [31] |
| LDCM0389 | CL119 | HEK-293T | C105(0.93); C157(0.98); C178(1.01); C81(0.86) | LDD1593 | [31] |
| LDCM0390 | CL12 | HEK-293T | C105(0.96); C157(1.07); C178(0.83); C81(1.23) | LDD1594 | [31] |
| LDCM0391 | CL120 | HEK-293T | C105(0.96); C157(0.96); C178(1.00); C81(0.97) | LDD1595 | [31] |
| LDCM0392 | CL121 | HEK-293T | C105(0.87); C157(1.04); C178(0.98); C81(1.02) | LDD1596 | [31] |
| LDCM0393 | CL122 | HEK-293T | C105(0.94); C157(0.89); C178(0.92); C81(0.88) | LDD1597 | [31] |
| LDCM0394 | CL123 | HEK-293T | C105(0.87); C157(0.99); C178(0.92); C81(0.92) | LDD1598 | [31] |
| LDCM0395 | CL124 | HEK-293T | C105(0.93); C157(1.12); C178(0.91); C81(1.10) | LDD1599 | [31] |
| LDCM0396 | CL125 | HEK-293T | C105(0.88); C157(0.97); C178(0.96); C81(1.09) | LDD1600 | [31] |
| LDCM0397 | CL126 | HEK-293T | C105(0.83); C157(0.84); C178(0.99); C81(0.86) | LDD1601 | [31] |
| LDCM0398 | CL127 | HEK-293T | C105(0.93); C157(0.98); C178(1.00); C81(1.02) | LDD1602 | [31] |
| LDCM0399 | CL128 | HEK-293T | C105(0.92); C157(0.94); C178(0.99); C81(1.11) | LDD1603 | [31] |
| LDCM0400 | CL13 | HEK-293T | C105(0.84); C157(1.06); C178(0.90); C81(1.02) | LDD1604 | [31] |
| LDCM0401 | CL14 | HEK-293T | C105(0.92); C157(1.04); C178(0.95); C81(0.97) | LDD1605 | [31] |
| LDCM0402 | CL15 | HEK-293T | C105(0.88); C157(1.01); C178(0.89); C81(1.02) | LDD1606 | [31] |
| LDCM0403 | CL16 | HEK-293T | C105(1.04); C157(0.99); C178(0.97); C81(1.04) | LDD1607 | [31] |
| LDCM0404 | CL17 | HEK-293T | C105(0.88); C157(0.97); C178(0.88); C81(1.10) | LDD1608 | [31] |
| LDCM0405 | CL18 | HEK-293T | C105(0.95); C157(1.11); C178(0.98); C81(1.09) | LDD1609 | [31] |
| LDCM0406 | CL19 | HEK-293T | C105(0.99); C157(1.18); C178(1.00); C18(0.79) | LDD1610 | [31] |
| LDCM0407 | CL2 | HEK-293T | C105(1.04); C157(1.08); C178(0.94); C81(1.24) | LDD1611 | [31] |
| LDCM0408 | CL20 | HEK-293T | C105(1.04); C157(1.04); C178(0.86); C81(0.87) | LDD1612 | [31] |
| LDCM0409 | CL21 | HEK-293T | C105(0.90); C157(1.33); C178(0.53); C81(1.00) | LDD1613 | [31] |
| LDCM0410 | CL22 | HEK-293T | C105(1.07); C157(1.11); C178(0.65); C81(1.01) | LDD1614 | [31] |
| LDCM0411 | CL23 | HEK-293T | C105(1.00); C157(1.20); C178(0.41); C81(0.88) | LDD1615 | [31] |
| LDCM0412 | CL24 | HEK-293T | C105(1.02); C157(1.00); C178(0.82); C81(1.14) | LDD1616 | [31] |
| LDCM0413 | CL25 | HEK-293T | C105(0.91); C157(0.99); C178(0.93); C81(0.88) | LDD1617 | [31] |
| LDCM0414 | CL26 | HEK-293T | C105(0.98); C157(1.05); C178(0.93); C81(1.05) | LDD1618 | [31] |
| LDCM0415 | CL27 | HEK-293T | C105(0.98); C157(0.87); C178(0.95); C81(0.94) | LDD1619 | [31] |
| LDCM0416 | CL28 | HEK-293T | C105(1.01); C157(1.02); C178(0.92); C81(1.21) | LDD1620 | [31] |
| LDCM0417 | CL29 | HEK-293T | C105(0.92); C157(1.13); C178(0.99); C81(0.95) | LDD1621 | [31] |
| LDCM0418 | CL3 | HEK-293T | C105(1.10); C157(1.05); C178(0.99); C81(1.11) | LDD1622 | [31] |
| LDCM0419 | CL30 | HEK-293T | C105(0.95); C157(1.01); C178(0.99); C81(0.90) | LDD1623 | [31] |
| LDCM0420 | CL31 | HEK-293T | C105(0.94); C157(1.05); C178(0.91); C18(0.73) | LDD1624 | [31] |
| LDCM0421 | CL32 | HEK-293T | C105(0.98); C157(1.02); C178(0.83); C81(1.02) | LDD1625 | [31] |
| LDCM0422 | CL33 | HEK-293T | C105(0.88); C157(1.14); C178(0.51); C81(0.89) | LDD1626 | [31] |
| LDCM0423 | CL34 | HEK-293T | C105(1.04); C157(1.10); C178(0.63); C81(0.93) | LDD1627 | [31] |
| LDCM0424 | CL35 | HEK-293T | C105(1.10); C157(1.10); C178(0.34); C81(1.00) | LDD1628 | [31] |
| LDCM0425 | CL36 | HEK-293T | C105(1.04); C157(1.40); C178(0.76); C81(1.01) | LDD1629 | [31] |
| LDCM0426 | CL37 | HEK-293T | C105(0.89); C157(0.87); C178(0.97); C81(1.11) | LDD1630 | [31] |
| LDCM0428 | CL39 | HEK-293T | C105(0.94); C157(0.93); C178(0.95); C81(0.79) | LDD1632 | [31] |
| LDCM0429 | CL4 | HEK-293T | C105(1.11); C157(1.14); C178(0.93); C81(1.13) | LDD1633 | [31] |
| LDCM0430 | CL40 | HEK-293T | C105(0.98); C157(0.93); C178(1.02); C81(0.99) | LDD1634 | [31] |
| LDCM0431 | CL41 | HEK-293T | C105(0.84); C157(1.03); C178(0.95); C81(0.87) | LDD1635 | [31] |
| LDCM0432 | CL42 | HEK-293T | C105(0.95); C157(1.26); C178(0.95); C81(0.66) | LDD1636 | [31] |
| LDCM0433 | CL43 | HEK-293T | C105(0.98); C157(1.05); C178(1.01); C18(0.98) | LDD1637 | [31] |
| LDCM0434 | CL44 | HEK-293T | C105(0.92); C157(1.05); C178(0.87); C81(0.99) | LDD1638 | [31] |
| LDCM0435 | CL45 | HEK-293T | C105(0.90); C157(0.99); C178(0.59); C81(0.75) | LDD1639 | [31] |
| LDCM0436 | CL46 | HEK-293T | C105(0.97); C157(1.12); C178(0.64); C81(0.93) | LDD1640 | [31] |
| LDCM0437 | CL47 | HEK-293T | C105(1.02); C157(1.16); C178(0.38); C81(0.97) | LDD1641 | [31] |
| LDCM0438 | CL48 | HEK-293T | C105(0.96); C157(1.18); C178(0.76); C81(1.18) | LDD1642 | [31] |
| LDCM0439 | CL49 | HEK-293T | C105(0.87); C157(1.05); C178(0.94); C81(1.10) | LDD1643 | [31] |
| LDCM0440 | CL5 | HEK-293T | C105(0.98); C157(1.02); C178(0.96); C81(1.21) | LDD1644 | [31] |
| LDCM0441 | CL50 | HEK-293T | C105(0.93); C157(0.88); C178(0.94); C81(1.22) | LDD1645 | [31] |
| LDCM0443 | CL52 | HEK-293T | C105(1.10); C157(0.98); C178(0.92); C81(1.06) | LDD1646 | [31] |
| LDCM0444 | CL53 | HEK-293T | C105(0.93); C157(1.05); C178(0.92); C81(1.01) | LDD1647 | [31] |
| LDCM0445 | CL54 | HEK-293T | C105(0.91); C157(1.25); C178(0.89); C81(0.80) | LDD1648 | [31] |
| LDCM0446 | CL55 | HEK-293T | C105(1.02); C157(1.10); C178(1.00); C18(1.27) | LDD1649 | [31] |
| LDCM0447 | CL56 | HEK-293T | C105(1.09); C157(0.93); C178(0.84); C81(0.83) | LDD1650 | [31] |
| LDCM0448 | CL57 | HEK-293T | C105(0.91); C157(1.06); C178(0.60); C81(0.85) | LDD1651 | [31] |
| LDCM0449 | CL58 | HEK-293T | C105(0.94); C157(1.04); C178(0.64); C81(0.83) | LDD1652 | [31] |
| LDCM0450 | CL59 | HEK-293T | C105(1.01); C157(1.12); C178(0.43); C81(0.85) | LDD1653 | [31] |
| LDCM0451 | CL6 | HEK-293T | C105(0.94); C157(1.16); C178(0.90); C81(0.82) | LDD1654 | [31] |
| LDCM0452 | CL60 | HEK-293T | C105(1.00); C157(1.16); C178(0.78); C81(1.07) | LDD1655 | [31] |
| LDCM0453 | CL61 | HEK-293T | C105(0.90); C157(0.87); C178(0.96); C81(1.01) | LDD1656 | [31] |
| LDCM0454 | CL62 | HEK-293T | C105(0.96); C157(1.02); C178(0.97); C81(1.19) | LDD1657 | [31] |
| LDCM0455 | CL63 | HEK-293T | C105(0.91); C157(0.95); C178(0.92); C81(1.03) | LDD1658 | [31] |
| LDCM0456 | CL64 | HEK-293T | C105(0.91); C157(1.10); C178(0.90); C81(1.08) | LDD1659 | [31] |
| LDCM0457 | CL65 | HEK-293T | C105(0.94); C157(1.03); C178(0.99); C81(1.01) | LDD1660 | [31] |
| LDCM0458 | CL66 | HEK-293T | C105(0.91); C157(1.32); C178(0.97); C81(0.98) | LDD1661 | [31] |
| LDCM0459 | CL67 | HEK-293T | C105(0.94); C157(1.05); C178(0.95); C18(0.91) | LDD1662 | [31] |
| LDCM0460 | CL68 | HEK-293T | C105(0.93); C157(0.95); C178(0.83); C81(0.81) | LDD1663 | [31] |
| LDCM0461 | CL69 | HEK-293T | C105(0.95); C157(1.05); C178(0.67); C81(1.12) | LDD1664 | [31] |
| LDCM0462 | CL7 | HEK-293T | C105(1.03); C157(1.17); C178(0.99); C18(0.62) | LDD1665 | [31] |
| LDCM0463 | CL70 | HEK-293T | C105(0.97); C157(0.97); C178(0.73); C81(0.87) | LDD1666 | [31] |
| LDCM0464 | CL71 | HEK-293T | C105(0.96); C157(1.00); C178(0.47); C81(1.03) | LDD1667 | [31] |
| LDCM0465 | CL72 | HEK-293T | C105(1.03); C157(1.28); C178(0.86); C81(1.00) | LDD1668 | [31] |
| LDCM0466 | CL73 | HEK-293T | C105(0.88); C157(1.08); C178(0.95); C81(1.08) | LDD1669 | [31] |
| LDCM0467 | CL74 | HEK-293T | C105(1.00); C157(0.96); C178(0.90); C81(0.97) | LDD1670 | [31] |
| LDCM0469 | CL76 | HEK-293T | C105(0.96); C157(0.96); C178(0.98); C81(1.06) | LDD1672 | [31] |
| LDCM0470 | CL77 | HEK-293T | C105(0.75); C157(1.06); C178(0.92); C81(0.94) | LDD1673 | [31] |
| LDCM0471 | CL78 | HEK-293T | C105(1.01); C157(1.04); C178(0.97); C81(0.88) | LDD1674 | [31] |
| LDCM0472 | CL79 | HEK-293T | C105(0.97); C157(1.11); C178(0.96); C18(1.07) | LDD1675 | [31] |
| LDCM0473 | CL8 | HEK-293T | C105(0.69); C157(0.97); C178(0.70); C81(0.86) | LDD1676 | [31] |
| LDCM0474 | CL80 | HEK-293T | C105(1.09); C157(1.07); C178(0.91); C81(0.92) | LDD1677 | [31] |
| LDCM0475 | CL81 | HEK-293T | C105(1.00); C157(1.07); C178(0.70); C81(1.13) | LDD1678 | [31] |
| LDCM0476 | CL82 | HEK-293T | C105(1.03); C157(1.01); C178(0.78); C81(0.85) | LDD1679 | [31] |
| LDCM0477 | CL83 | HEK-293T | C105(0.96); C157(1.11); C178(0.58); C81(0.90) | LDD1680 | [31] |
| LDCM0478 | CL84 | HEK-293T | C105(0.94); C157(1.15); C178(0.90); C81(1.28) | LDD1681 | [31] |
| LDCM0479 | CL85 | HEK-293T | C105(0.92); C157(1.01); C178(0.94); C81(1.11) | LDD1682 | [31] |
| LDCM0480 | CL86 | HEK-293T | C105(0.90); C157(0.87); C178(0.98); C81(1.04) | LDD1683 | [31] |
| LDCM0481 | CL87 | HEK-293T | C105(0.87); C157(1.03); C178(0.93); C81(0.93) | LDD1684 | [31] |
| LDCM0482 | CL88 | HEK-293T | C105(1.00); C157(0.96); C178(0.95); C81(1.15) | LDD1685 | [31] |
| LDCM0483 | CL89 | HEK-293T | C105(0.92); C157(1.04); C178(0.99); C81(1.02) | LDD1686 | [31] |
| LDCM0484 | CL9 | HEK-293T | C105(1.05); C157(1.10); C178(0.65); C81(1.02) | LDD1687 | [31] |
| LDCM0485 | CL90 | HEK-293T | C105(0.79); C157(1.04); C178(0.86); C81(0.62) | LDD1688 | [31] |
| LDCM0486 | CL91 | HEK-293T | C105(1.05); C157(1.04); C178(0.97); C18(1.28) | LDD1689 | [31] |
| LDCM0487 | CL92 | HEK-293T | C105(0.84); C157(1.01); C178(0.93); C81(0.92) | LDD1690 | [31] |
| LDCM0488 | CL93 | HEK-293T | C105(0.93); C157(1.08); C178(0.80); C81(1.14) | LDD1691 | [31] |
| LDCM0489 | CL94 | HEK-293T | C105(0.92); C157(0.93); C178(0.90); C81(0.93) | LDD1692 | [31] |
| LDCM0490 | CL95 | HEK-293T | C105(0.81); C157(1.27); C178(0.67); C81(0.89) | LDD1693 | [31] |
| LDCM0491 | CL96 | HEK-293T | C105(0.93); C157(1.29); C178(0.95); C81(1.10) | LDD1694 | [31] |
| LDCM0492 | CL97 | HEK-293T | C105(0.86); C157(0.90); C178(0.93); C81(0.97) | LDD1695 | [31] |
| LDCM0493 | CL98 | HEK-293T | C105(0.95); C157(0.95); C178(0.98); C81(0.93) | LDD1696 | [31] |
| LDCM0494 | CL99 | HEK-293T | C105(0.96); C157(0.89); C178(0.94); C81(0.96) | LDD1697 | [31] |
| LDCM0634 | CY-0357 | Hep-G2 | C178(50.00) | LDD2228 | [30] |
| LDCM0495 | E2913 | HEK-293T | C105(1.01); C157(1.08); C178(0.99); C81(1.11) | LDD1698 | [31] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C105(1.37); C178(0.85) | LDD1702 | [6] |
| LDCM0625 | F8 | Ramos | C178(1.20); C157(0.73); C6(0.50); C105(8.96) | LDD2187 | [32] |
| LDCM0572 | Fragment10 | Ramos | C178(0.61); C157(0.61); C6(0.40); C105(0.37) | LDD2189 | [32] |
| LDCM0573 | Fragment11 | Ramos | C178(0.05); C157(0.00) | LDD2190 | [32] |
| LDCM0574 | Fragment12 | Ramos | C178(0.70); C157(0.87); C6(0.61); C105(0.29) | LDD2191 | [32] |
| LDCM0575 | Fragment13 | Ramos | C178(1.05); C157(0.68); C6(1.04); C105(0.41) | LDD2192 | [32] |
| LDCM0576 | Fragment14 | Ramos | C178(0.67); C157(0.46); C6(0.64); C105(0.87) | LDD2193 | [32] |
| LDCM0579 | Fragment20 | Ramos | C178(0.49); C157(0.76); C6(0.51); C105(0.42) | LDD2194 | [32] |
| LDCM0580 | Fragment21 | Ramos | C178(1.01); C157(0.83); C6(0.77); C105(0.36) | LDD2195 | [32] |
| LDCM0582 | Fragment23 | Ramos | C178(1.24); C157(0.76); C6(0.62) | LDD2196 | [32] |
| LDCM0578 | Fragment27 | Ramos | C178(1.00); C157(0.78); C6(0.96) | LDD2197 | [32] |
| LDCM0586 | Fragment28 | Ramos | C178(0.67); C157(0.44); C6(0.62); C105(0.68) | LDD2198 | [32] |
| LDCM0588 | Fragment30 | Ramos | C178(1.20); C157(1.05); C6(1.18); C105(0.48) | LDD2199 | [32] |
| LDCM0589 | Fragment31 | Ramos | C178(0.97); C157(0.90); C6(0.84); C105(1.56) | LDD2200 | [32] |
| LDCM0590 | Fragment32 | Ramos | C178(0.63); C157(0.61); C6(0.41); C105(0.42) | LDD2201 | [32] |
| LDCM0468 | Fragment33 | HEK-293T | C105(0.99); C157(0.96); C178(0.92); C81(1.00) | LDD1671 | [31] |
| LDCM0596 | Fragment38 | Ramos | C178(0.92); C157(0.79); C6(0.90); C105(0.55) | LDD2203 | [32] |
| LDCM0566 | Fragment4 | Ramos | C178(0.83); C157(0.91); C6(0.59); C105(0.72) | LDD2184 | [32] |
| LDCM0427 | Fragment51 | HEK-293T | C105(0.91); C157(0.89); C178(0.95); C81(0.98) | LDD1631 | [31] |
| LDCM0610 | Fragment52 | Ramos | C178(1.12); C157(0.84); C6(1.67); C105(0.75) | LDD2204 | [32] |
| LDCM0614 | Fragment56 | Ramos | C178(1.54); C157(0.89); C6(1.57); C105(0.19) | LDD2205 | [32] |
| LDCM0569 | Fragment7 | Ramos | C178(0.62); C157(0.81); C6(0.46); C105(0.53) | LDD2186 | [32] |
| LDCM0571 | Fragment9 | Ramos | C178(0.58); C157(0.79); C6(0.56); C105(0.09) | LDD2188 | [32] |
| LDCM0116 | HHS-0101 | DM93 | Y98(0.87); Y139(0.98) | LDD0264 | [13] |
| LDCM0117 | HHS-0201 | DM93 | Y98(0.87); Y139(0.98) | LDD0265 | [13] |
| LDCM0118 | HHS-0301 | DM93 | Y98(0.85); Y139(0.87) | LDD0266 | [13] |
| LDCM0119 | HHS-0401 | DM93 | Y98(0.97); Y139(0.98) | LDD0267 | [13] |
| LDCM0120 | HHS-0701 | DM93 | Y139(0.86); Y98(0.98) | LDD0268 | [13] |
| LDCM0107 | IAA | HeLa | C105(0.00); C178(0.00); H104(0.00); H103(0.00) | LDD0221 | [26] |
| LDCM0123 | JWB131 | DM93 | Y139(0.50) | LDD0285 | [12] |
| LDCM0124 | JWB142 | DM93 | Y139(0.53) | LDD0286 | [12] |
| LDCM0125 | JWB146 | DM93 | Y139(0.90) | LDD0287 | [12] |
| LDCM0126 | JWB150 | DM93 | Y139(3.07) | LDD0288 | [12] |
| LDCM0127 | JWB152 | DM93 | Y139(1.56); Y154(1.16) | LDD0289 | [12] |
| LDCM0128 | JWB198 | DM93 | Y139(0.31); Y154(0.15) | LDD0290 | [12] |
| LDCM0129 | JWB202 | DM93 | Y139(0.64); Y154(0.47) | LDD0291 | [12] |
| LDCM0130 | JWB211 | DM93 | Y139(0.54) | LDD0292 | [12] |
| LDCM0179 | JZ128 | PC-3 | C178(0.00); C157(0.00) | LDD0462 | [10] |
| LDCM0022 | KB02 | Ramos | C178(0.62); C157(0.95); C6(0.44); C105(0.46) | LDD2182 | [32] |
| LDCM0023 | KB03 | MDA-MB-231 | C178(0.97) | LDD1701 | [6] |
| LDCM0024 | KB05 | COLO792 | C6(1.61); C124(1.13) | LDD3310 | [9] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C178(1.91) | LDD2102 | [6] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C178(0.43); C105(0.88) | LDD2121 | [6] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [26] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C178(0.53); C105(0.87) | LDD2089 | [6] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C178(1.46) | LDD2090 | [6] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C178(1.12); C105(0.92) | LDD2092 | [6] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C178(1.01); C105(1.38) | LDD2093 | [6] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C178(1.94); C105(0.79) | LDD2094 | [6] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C178(0.61); C105(0.54) | LDD2096 | [6] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C178(0.76) | LDD2097 | [6] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C178(0.80); C105(1.07) | LDD2098 | [6] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C178(0.80); C105(1.36) | LDD2099 | [6] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C178(0.59); C105(0.44); C157(0.45) | LDD2100 | [6] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C178(0.64) | LDD2101 | [6] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C178(0.77); C105(0.34) | LDD2104 | [6] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C178(1.39); C105(1.44) | LDD2105 | [6] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C178(0.95); C157(0.26) | LDD2106 | [6] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C178(0.81); C105(0.95) | LDD2107 | [6] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C178(0.37); C157(0.47) | LDD2108 | [6] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C178(0.60); C105(1.10) | LDD2109 | [6] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C178(3.68); C157(0.36) | LDD2110 | [6] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C178(0.87); C105(1.06) | LDD2111 | [6] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C178(1.08); C105(0.49) | LDD2114 | [6] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C178(0.39); C105(0.62) | LDD2115 | [6] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C178(0.66); C105(0.72) | LDD2116 | [6] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C178(0.75); C105(0.74) | LDD2118 | [6] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C178(2.93); C105(2.07) | LDD2119 | [6] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C178(0.83) | LDD2120 | [6] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C178(0.87); C105(0.70) | LDD2122 | [6] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C178(0.75); C105(1.20) | LDD2123 | [6] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C178(0.62); C105(1.05) | LDD2124 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C178(0.67); C105(1.14) | LDD2125 | [6] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C178(0.60) | LDD2126 | [6] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C178(0.76); C105(1.87) | LDD2127 | [6] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C105(0.63) | LDD2128 | [6] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C178(0.82); C105(1.16) | LDD2129 | [6] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C178(0.50) | LDD2133 | [6] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C178(0.37); C105(0.56) | LDD2134 | [6] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C178(0.93); C105(1.11) | LDD2135 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C178(0.99); C105(1.51) | LDD2136 | [6] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C178(0.82); C105(1.37) | LDD2137 | [6] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C178(2.81); C105(1.65) | LDD1700 | [6] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C178(0.65); C105(1.06) | LDD2140 | [6] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C178(0.59); C105(0.64) | LDD2141 | [6] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C178(0.97); C105(0.84) | LDD2143 | [6] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C178(3.59); C105(1.58) | LDD2144 | [6] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C178(4.79); C157(4.81) | LDD2145 | [6] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C178(0.77); C105(1.07) | LDD2146 | [6] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C178(3.47) | LDD2147 | [6] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C178(0.35); C105(0.55) | LDD2148 | [6] |
| LDCM0555 | Nucleophilic fragment 7b | MDA-MB-231 | C178(0.76); C105(0.90) | LDD2149 | [6] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C178(0.41); C105(0.54) | LDD2150 | [6] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C178(0.92); C105(0.81) | LDD2151 | [6] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C178(1.62) | LDD2153 | [6] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C178(0.89) | LDD2206 | [33] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C178(0.74) | LDD2207 | [33] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transcription factor Jun (JUN) | BZIP family | P05412 | |||
| THAP domain-containing protein 3 (THAP3) | . | Q8WTV1 | |||
Other
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Azathioprine | Small molecular drug | DB00993 | |||
| Dextromethorphan | Small molecular drug | DB00514 | |||
Investigative
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Guanosine-5'-diphosphate | Small molecular drug | DB04315 | |||
Discontinued
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Eht-1864 | Small molecular drug | D0M9ER | |||
References
