Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P1 Probe Info |
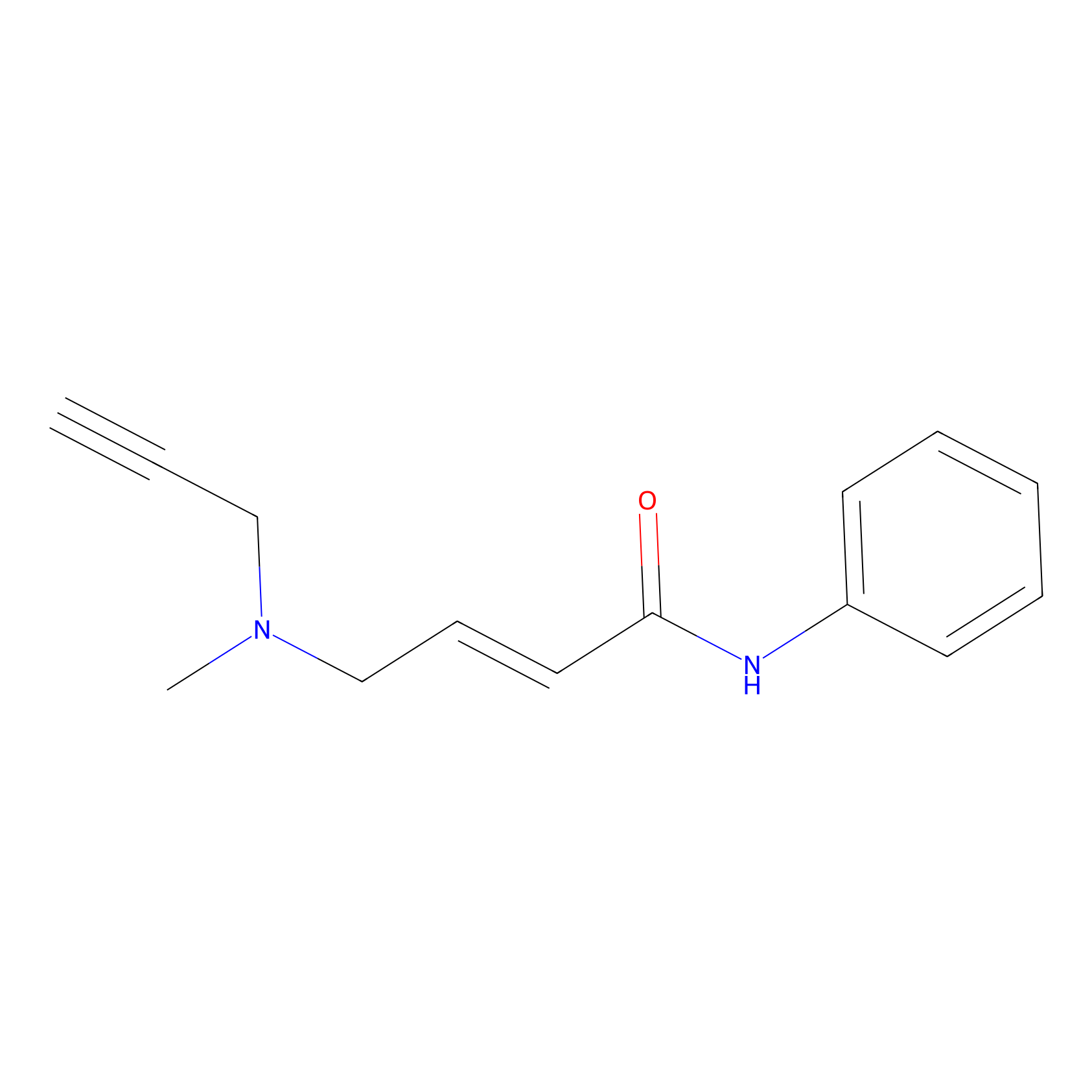 |
1.70 | LDD0452 | [2] | |
|
P2 Probe Info |
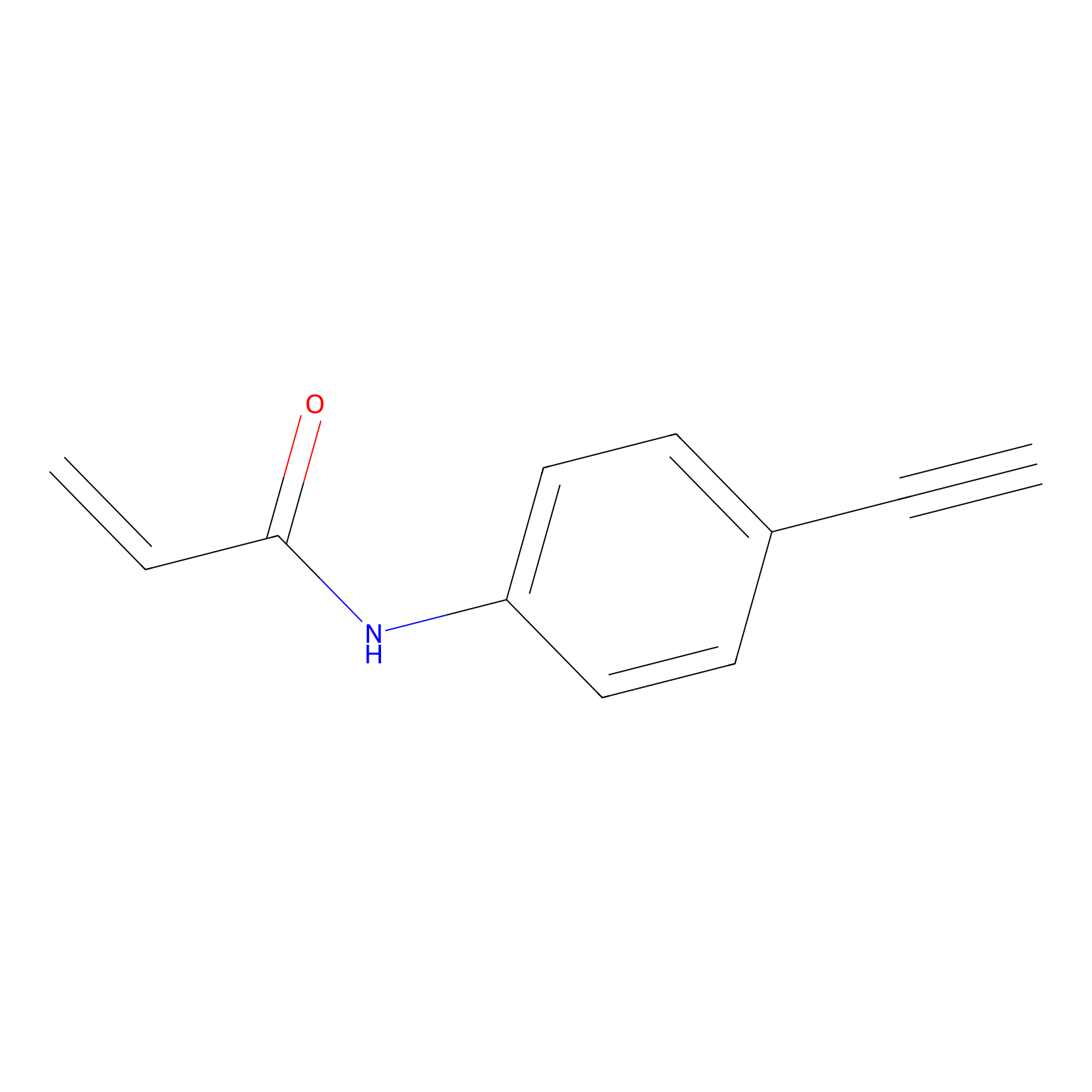 |
2.87 | LDD0453 | [2] | |
|
P3 Probe Info |
 |
1.67 | LDD0454 | [2] | |
|
A-EBA Probe Info |
 |
5.34 | LDD0215 | [3] | |
|
ILS-1 Probe Info |
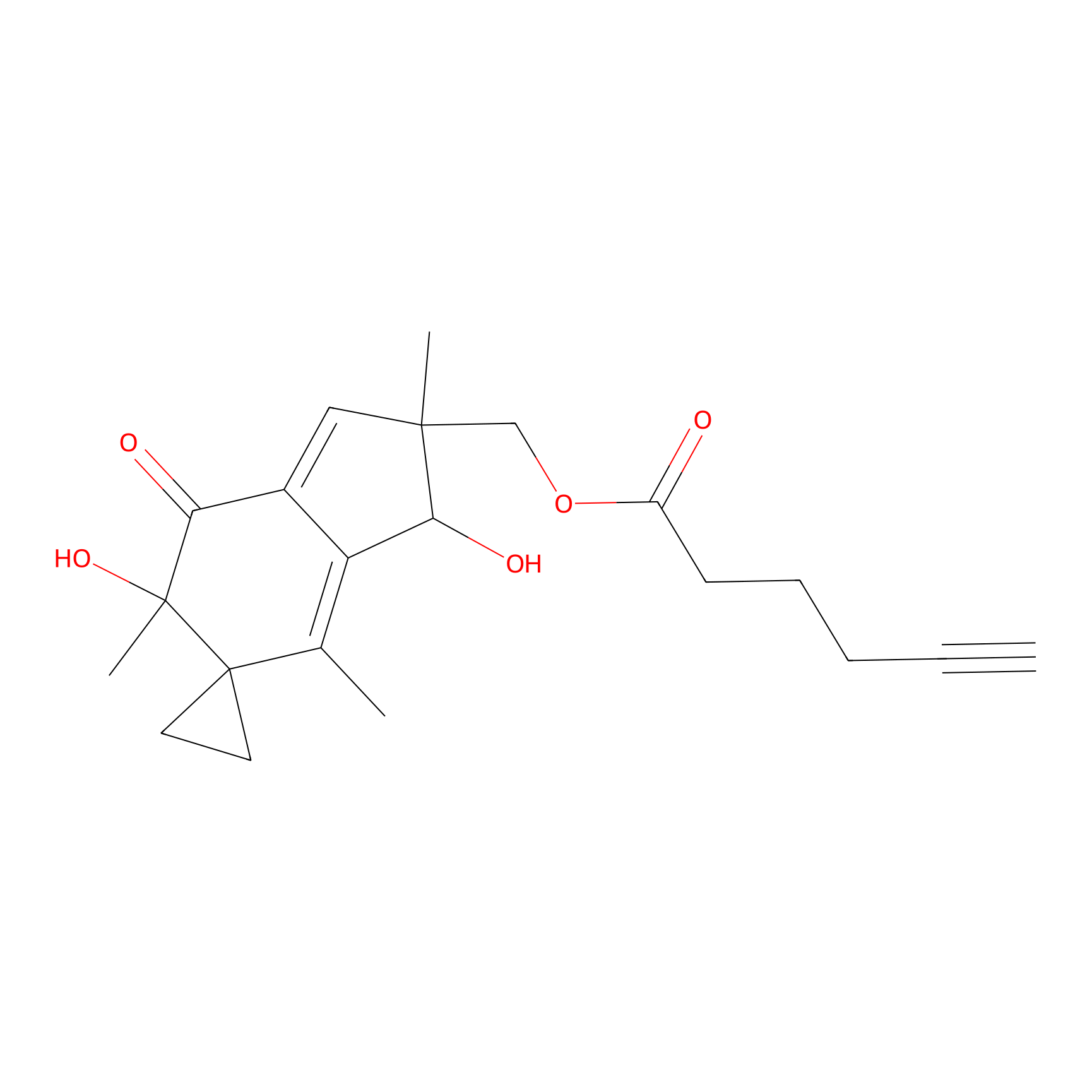 |
3.44 | LDD0415 | [4] | |
|
TH211 Probe Info |
 |
Y89(8.21); Y52(5.05) | LDD0257 | [5] | |
|
TH214 Probe Info |
 |
Y89(20.00); Y73(19.93) | LDD0258 | [5] | |
|
TH216 Probe Info |
 |
Y73(5.07) | LDD0259 | [5] | |
|
YN-1 Probe Info |
 |
2.13 | LDD0444 | [6] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [6] | |
|
AZ-9 Probe Info |
 |
D25(1.15); E54(0.92); E64(1.00); E53(1.00) | LDD2208 | [7] | |
|
ONAyne Probe Info |
 |
K78(0.00); K32(0.00); K80(0.00); K13(0.00) | LDD0273 | [8] | |
|
OPA-S-S-alkyne Probe Info |
 |
K17(2.72); K13(2.72); K92(7.01); K32(7.50) | LDD3494 | [9] | |
|
Probe 1 Probe Info |
 |
Y52(15.82); Y73(67.23); Y89(19.86) | LDD3495 | [10] | |
|
AF-2 Probe Info |
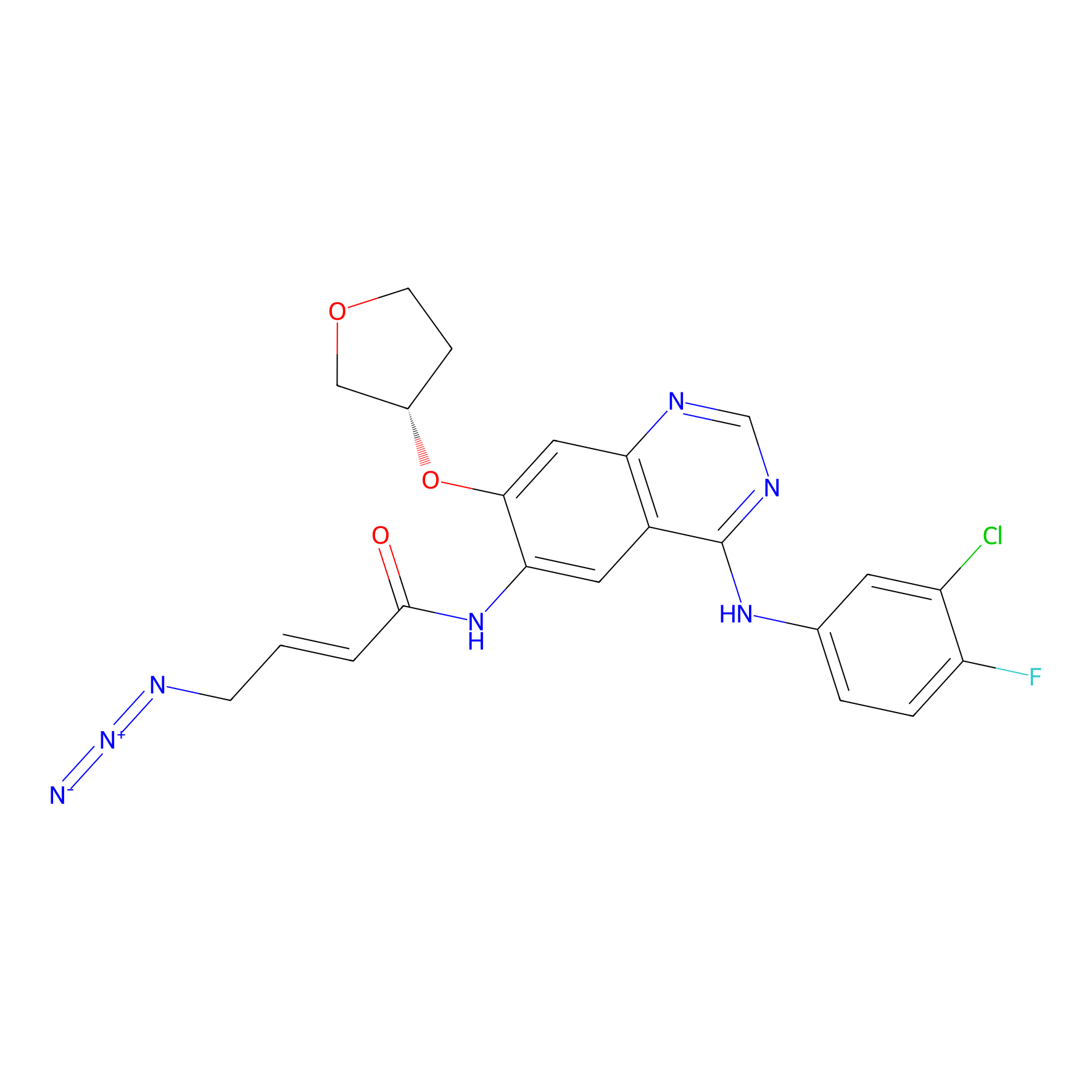 |
1.59 | LDD0422 | [11] | |
|
HHS-482 Probe Info |
 |
Y73(2.36); Y89(1.14) | LDD0285 | [12] | |
|
HHS-475 Probe Info |
 |
Y89(0.70); Y73(1.63) | LDD0264 | [13] | |
|
HHS-465 Probe Info |
 |
Y73(7.14); Y89(2.01) | LDD2237 | [14] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [15] | |
|
ATP probe Probe Info |
 |
K92(0.00); K32(0.00); K80(0.00); K78(0.00) | LDD0199 | [15] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [16] | |
|
CY-1 Probe Info |
 |
D86(0.00); K92(0.00); M85(0.00); Y89(0.00) | LDD0246 | [17] | |
|
aONE Probe Info |
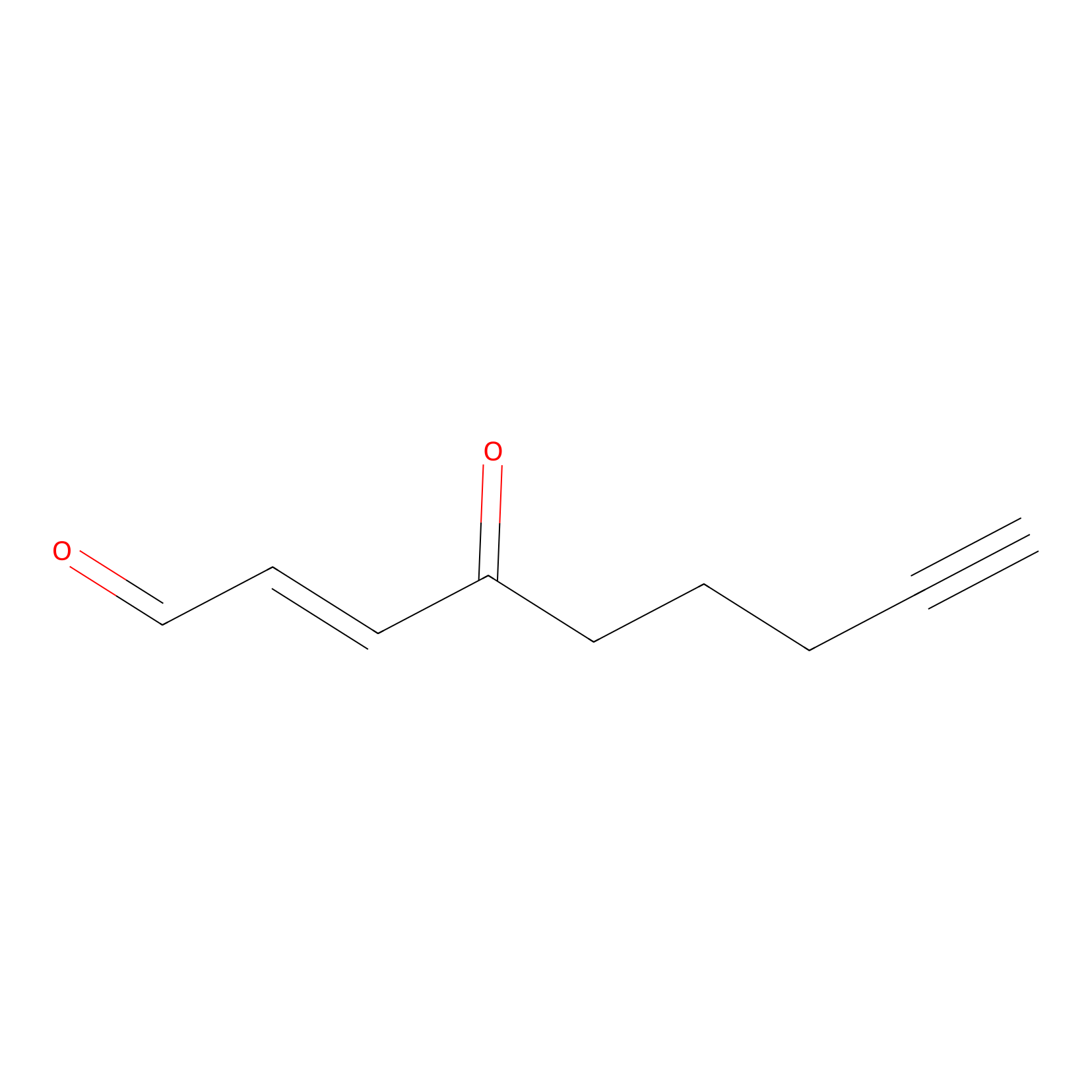 |
K78(0.00); K92(0.00) | LDD0002 | [18] | |
|
1d-yne Probe Info |
 |
K80(0.00); K92(0.00) | LDD0356 | [19] | |
|
NHS Probe Info |
 |
K92(0.00); K80(0.00); K32(0.00); K78(0.00) | LDD0010 | [18] | |
|
OSF Probe Info |
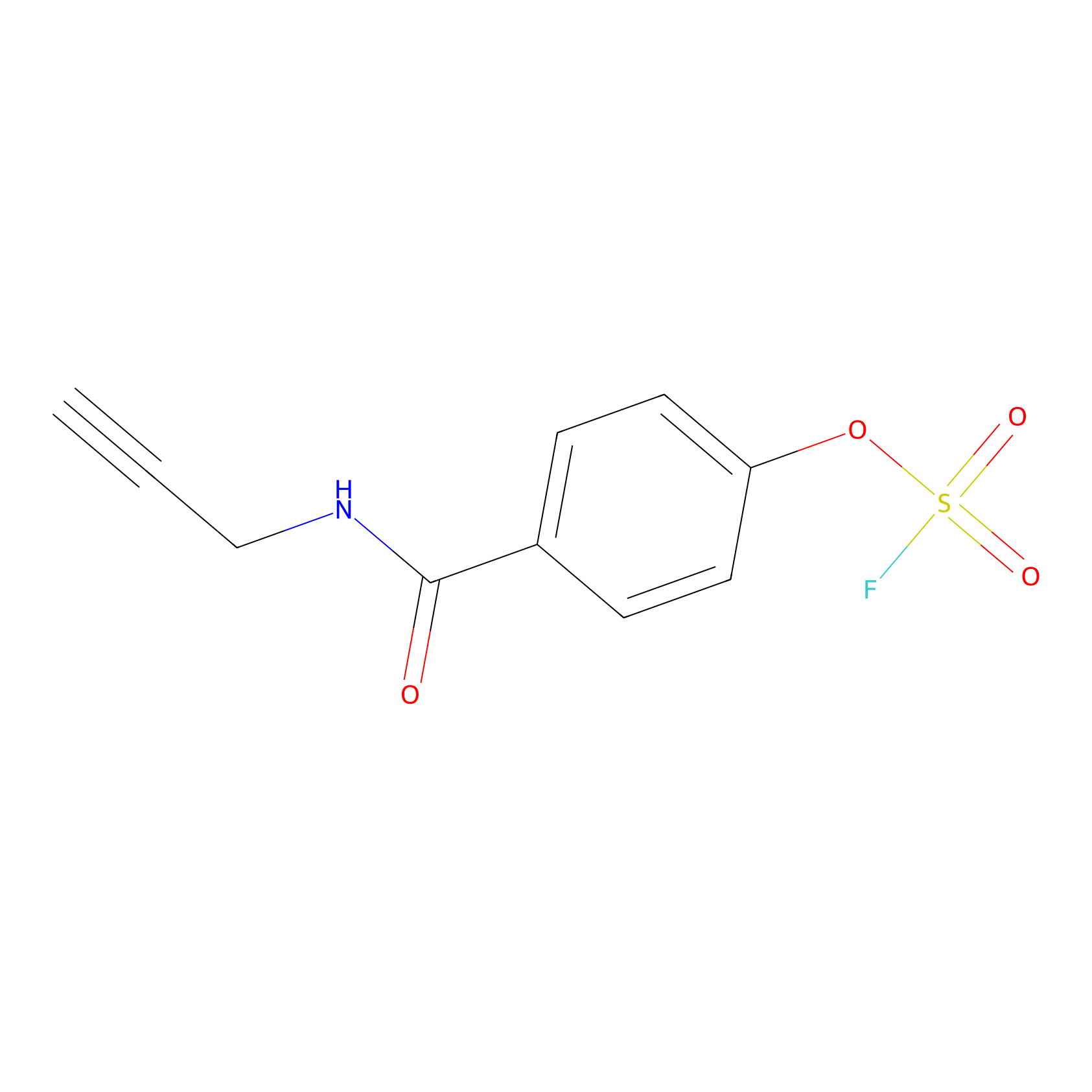 |
N.A. | LDD0029 | [20] | |
|
SF Probe Info |
 |
K92(0.00); Y73(0.00); Y52(0.00); K32(0.00) | LDD0028 | [20] | |
|
STPyne Probe Info |
 |
K92(0.00); K32(0.00); K78(0.00) | LDD0009 | [18] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [19] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [21] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [21] | |
|
Methacrolein Probe Info |
 |
H76(0.00); K32(0.00) | LDD0218 | [21] | |
|
W1 Probe Info |
 |
K32(0.00); K92(0.00); D25(0.00); Y89(0.00) | LDD0236 | [22] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-2 Probe Info |
 |
1.55 | LDD0138 | [23] | |
|
Diazir Probe Info |
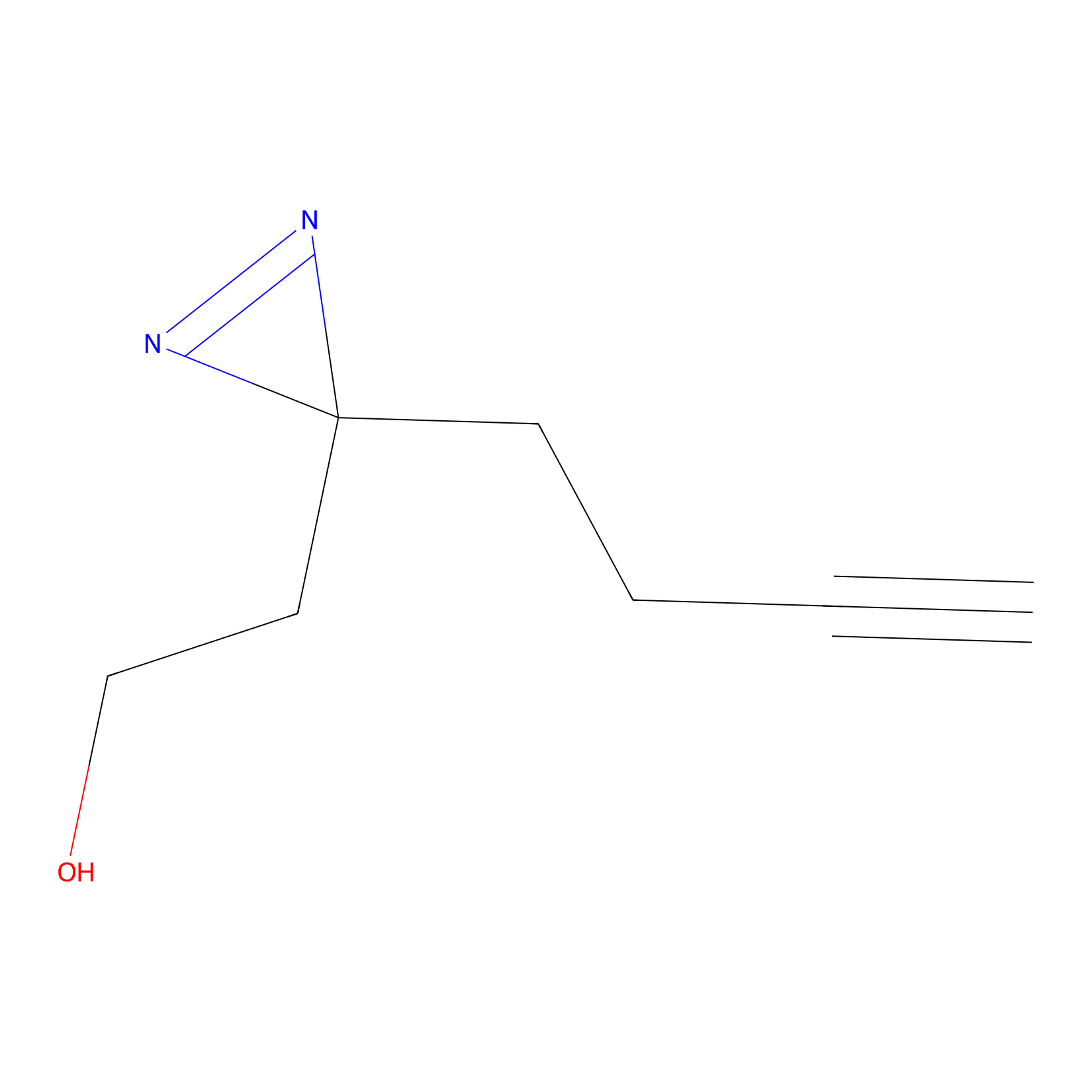 |
N.A. | LDD0011 | [18] | |
|
Photocelecoxib Probe Info |
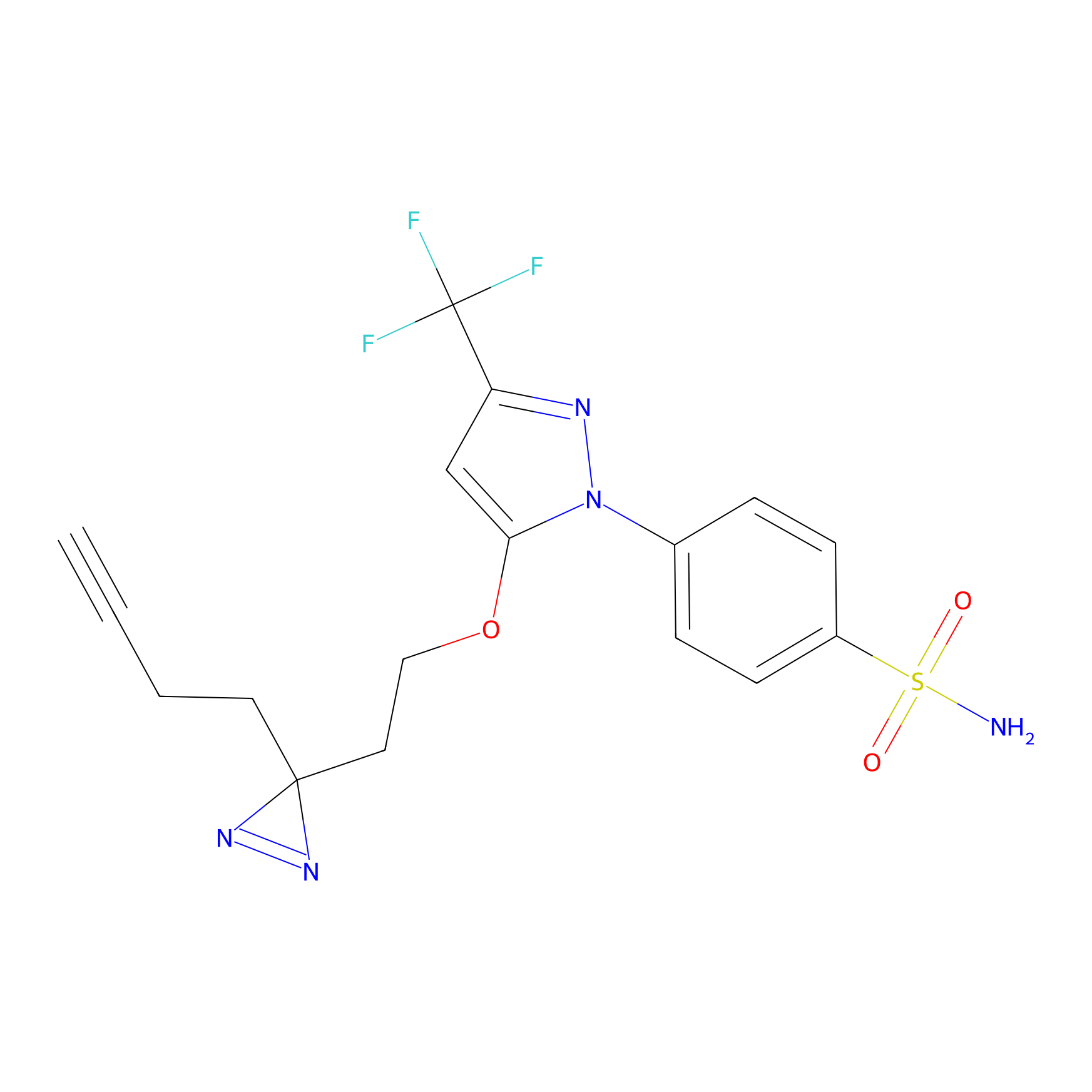 |
Y52(0.00); E54(0.00); E64(0.00); D69(0.00) | LDD0019 | [24] | |
|
Photoindomethacin Probe Info |
 |
N.A. | LDD0155 | [24] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0072 | [25] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0166 | Afatinib | A431 | 1.59 | LDD0422 | [11] |
| LDCM0108 | Chloroacetamide | HeLa | H76(0.00); K32(0.00) | LDD0222 | [21] |
| LDCM0116 | HHS-0101 | DM93 | Y89(0.70); Y73(1.63) | LDD0264 | [13] |
| LDCM0117 | HHS-0201 | DM93 | Y89(0.69); Y73(0.73) | LDD0265 | [13] |
| LDCM0118 | HHS-0301 | DM93 | Y89(0.62); Y73(1.10) | LDD0266 | [13] |
| LDCM0119 | HHS-0401 | DM93 | Y89(0.67); Y73(0.82) | LDD0267 | [13] |
| LDCM0120 | HHS-0701 | DM93 | Y89(0.43); Y73(0.76) | LDD0268 | [13] |
| LDCM0107 | IAA | HeLa | H76(0.00); K32(0.00) | LDD0221 | [21] |
| LDCM0123 | JWB131 | DM93 | Y73(2.36); Y89(1.14) | LDD0285 | [12] |
| LDCM0124 | JWB142 | DM93 | Y73(3.70); Y89(1.53) | LDD0286 | [12] |
| LDCM0125 | JWB146 | DM93 | Y73(1.86); Y89(0.67) | LDD0287 | [12] |
| LDCM0126 | JWB150 | DM93 | Y73(1.25); Y89(3.80) | LDD0288 | [12] |
| LDCM0127 | JWB152 | DM93 | Y73(3.32); Y89(2.09) | LDD0289 | [12] |
| LDCM0128 | JWB198 | DM93 | Y73(1.02); Y89(2.15) | LDD0290 | [12] |
| LDCM0129 | JWB202 | DM93 | Y73(3.04); Y89(0.46) | LDD0291 | [12] |
| LDCM0130 | JWB211 | DM93 | Y73(2.06); Y89(1.07) | LDD0292 | [12] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [21] |
| LDCM0110 | W12 | Hep-G2 | K13(0.50); Q28(0.86); K32(0.87); N26(0.94) | LDD0237 | [22] |
| LDCM0111 | W14 | Hep-G2 | K13(0.65); K78(1.02); H76(1.15); D25(1.19) | LDD0238 | [22] |
| LDCM0112 | W16 | Hep-G2 | R36(0.92); K32(0.93); Q28(0.95); D25(0.99) | LDD0239 | [22] |
| LDCM0113 | W17 | Hep-G2 | K60(0.88); N65(0.88); D25(1.01); N26(1.03) | LDD0240 | [22] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Lethal(3)malignant brain tumor-like protein 1 (L3MBTL1) | . | Q9Y468 | |||
| Peregrin (BRPF1) | . | P55201 | |||
Other
References
