Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 22RV1 | SNV: p.W75Ter | . | |||
| A3KAW | Deletion: p.H175QfsTer27 | . | |||
| BICR22 | SNV: p.E43Ter | . | |||
| CAL62 | Substitution: p.K292E | . | |||
| CALU6 | SNV: p.T158I | . | |||
| DEL | Substitution: p.I121R | . | |||
| EGI1 | Substitution: p.I121R | . | |||
| ETK1 | Substitution: p.I121R | . | |||
| HCC44 | SNV: p.A348S | . | |||
| HEC1 | Substitution: p.I121R | . | |||
| HLF | Substitution: p.I121R | . | |||
| HS578T | Substitution: p.I119L | . | |||
| HSC4 | SNV: p.R135L | . | |||
| IM95 | Deletion: p.R45AfsTer32 | . | |||
| JHH4 | SNV: p.E299D; p.S301P | . | |||
| JHH6 | Substitution: p.L180Q | . | |||
| KASUMI1 | Substitution: p.I121R | . | |||
| KMCH1 | Substitution: p.I121R | . | |||
| KMS12BM | SNV: p.L134H | . | |||
| KURAMOCHI | Substitution: p.I121R | . | |||
| KYM1 | SNV: p.Y147Ter | . | |||
| MFE319 | Deletion: p.K210RfsTer4 | . | |||
| MOLM16 | SNV: p.K170E | . | |||
| MOLT4 | Substitution: p.I121R | . | |||
| NCIH1993 | Substitution: p.I121R | . | |||
| OCILY3 | Deletion: p.S156RfsTer18 | . | |||
| OVKATE | Substitution: p.I121R | . | |||
| RBE | Substitution: p.I121R | . | |||
| RT112 | Substitution: p.D357_V358delinsEM | . | |||
| SH4 | SNV: p.W228L Substitution: p.I121R |
DBIA Probe Info | |||
| SNGM | SNV: p.A313T | . | |||
| SNU5 | Substitution: p.K168Q | . | |||
| SSP25 | Substitution: p.I121R | . | |||
| SUPT1 | SNV: p.T305N | . | |||
| SW1573 | Substitution: p.K292E | . | |||
| SW837 | SNV: p.G186D | . | |||
| T98G | SNV: p.P307S | . | |||
| TC71 | Substitution: p.K292E | . | |||
| TE8 | Substitution: p.I121R | . | |||
| U937 | Deletion: p.K210RfsTer4 | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
8RK64 Probe Info |
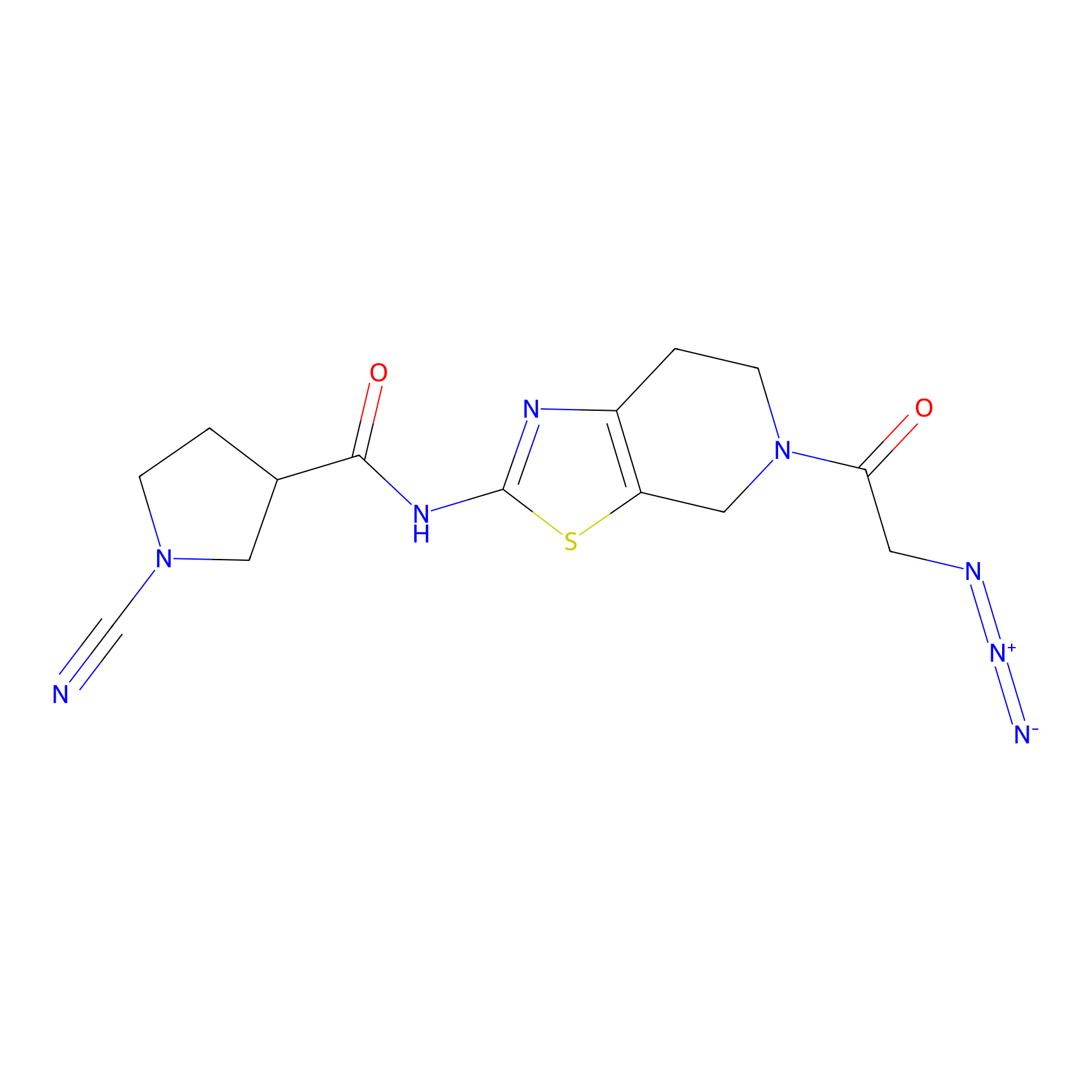 |
N.A. | LDD0039 | [2] | |
|
STPyne Probe Info |
 |
K168(5.37); K210(13.44); K292(20.00) | LDD2217 | [3] | |
|
ONAyne Probe Info |
 |
K340(9.09) | LDD0275 | [4] | |
|
OPA-S-S-alkyne Probe Info |
 |
K92(3.31) | LDD3494 | [5] | |
|
DBIA Probe Info |
 |
C188(17.35) | LDD3327 | [6] | |
|
P12 Probe Info |
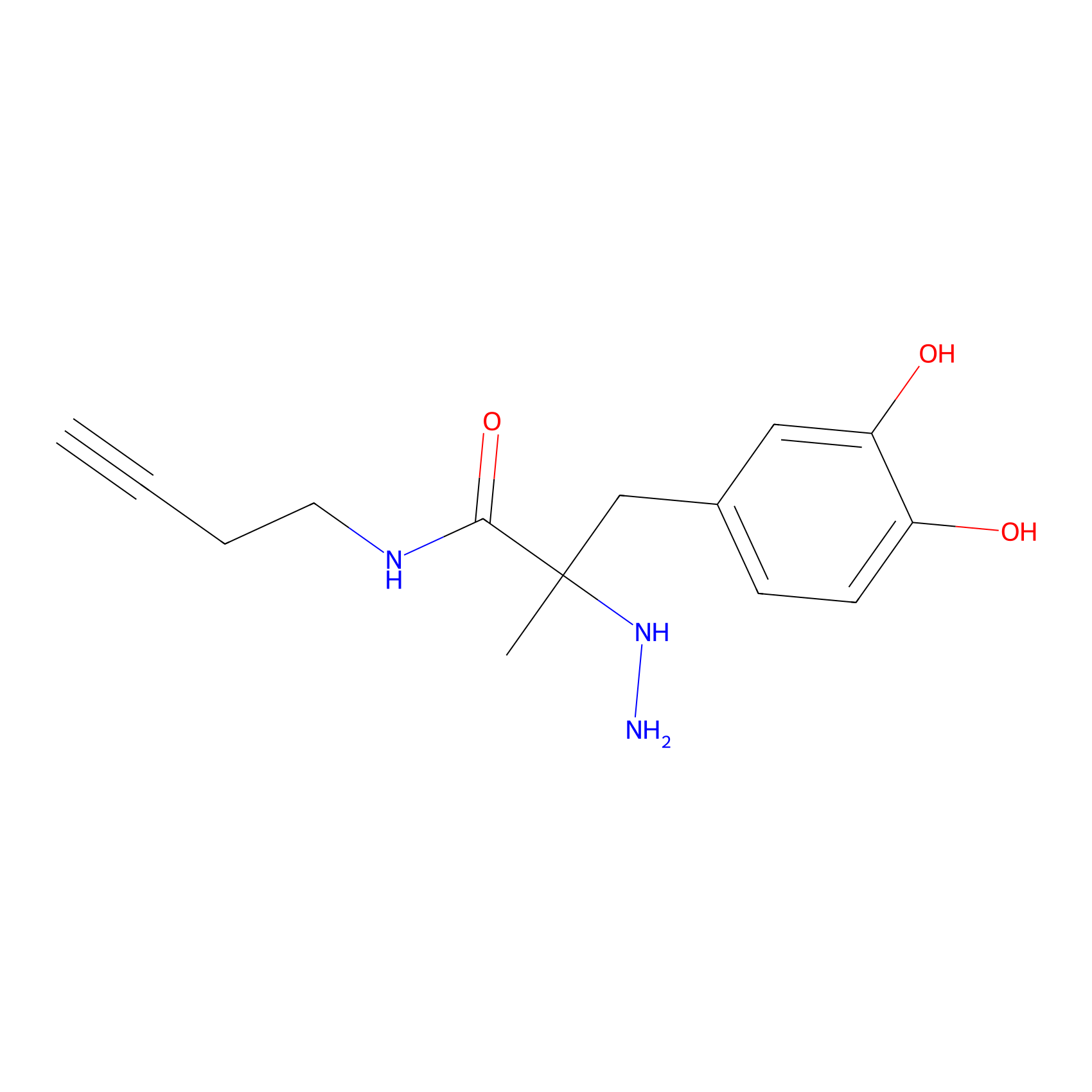 |
12.57 | LDD0202 | [7] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [8] | |
|
BTD Probe Info |
 |
C363(1.83); C283(0.70) | LDD2170 | [9] | |
|
YY4-yne Probe Info |
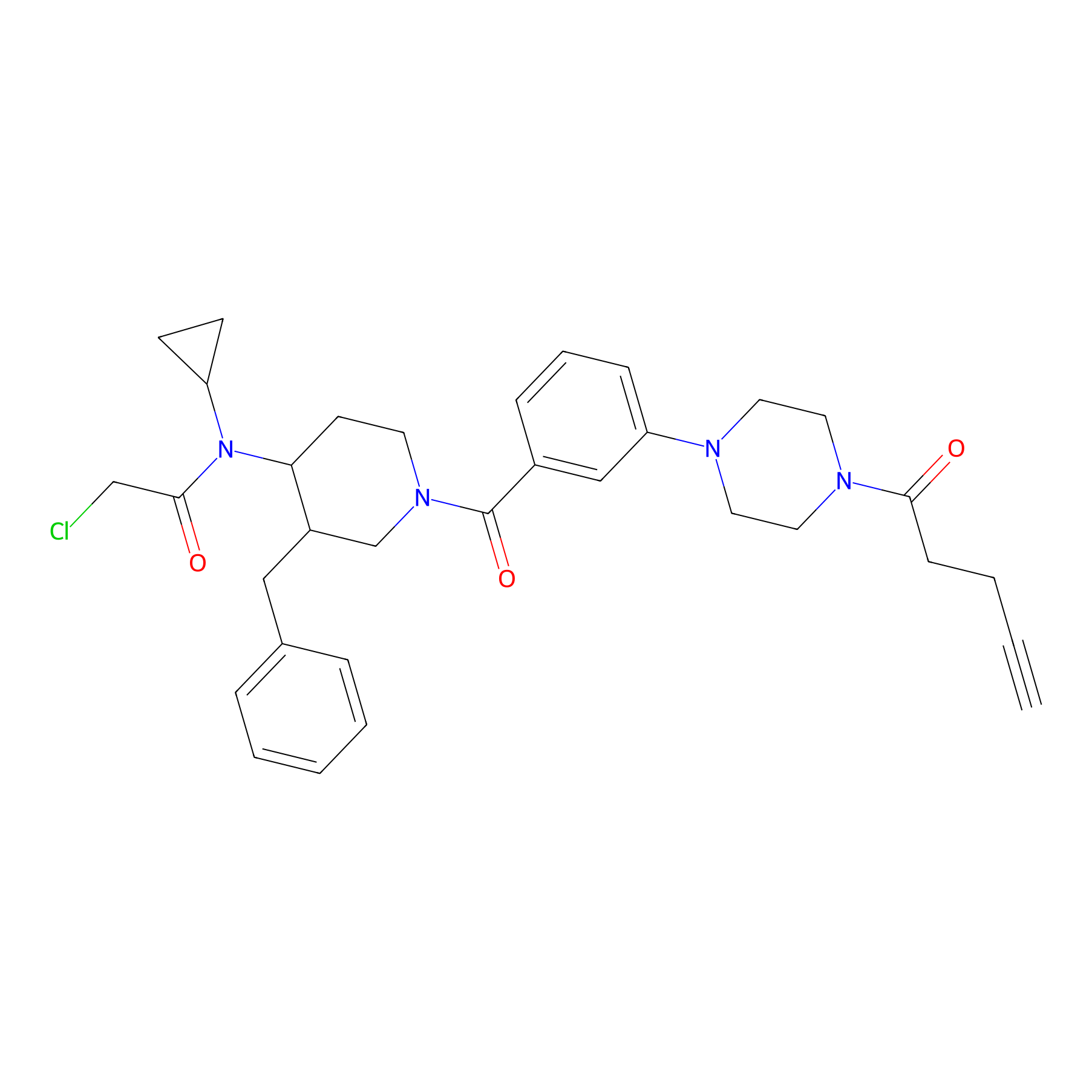 |
2.09 | LDD0400 | [10] | |
|
DA-P3 Probe Info |
 |
11.86 | LDD0179 | [11] | |
|
AHL-Pu-1 Probe Info |
 |
N.A. | LDD0172 | [12] | |
|
IPM Probe Info |
 |
C363(3.50) | LDD1701 | [9] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C188(0.00); C227(0.00); C363(0.00); C283(0.00) | LDD0038 | [13] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [14] | |
|
Lodoacetamide azide Probe Info |
 |
C188(0.00); C227(0.00); C363(0.00); C283(0.00) | LDD0037 | [13] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [15] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [16] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [16] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C004 Probe Info |
 |
5.90 | LDD1714 | [17] | |
|
C022 Probe Info |
 |
7.84 | LDD1728 | [17] | |
|
C106 Probe Info |
 |
17.63 | LDD1793 | [17] | |
|
C158 Probe Info |
 |
10.78 | LDD1838 | [17] | |
|
C170 Probe Info |
 |
24.93 | LDD1850 | [17] | |
|
C186 Probe Info |
 |
11.16 | LDD1864 | [17] | |
|
C223 Probe Info |
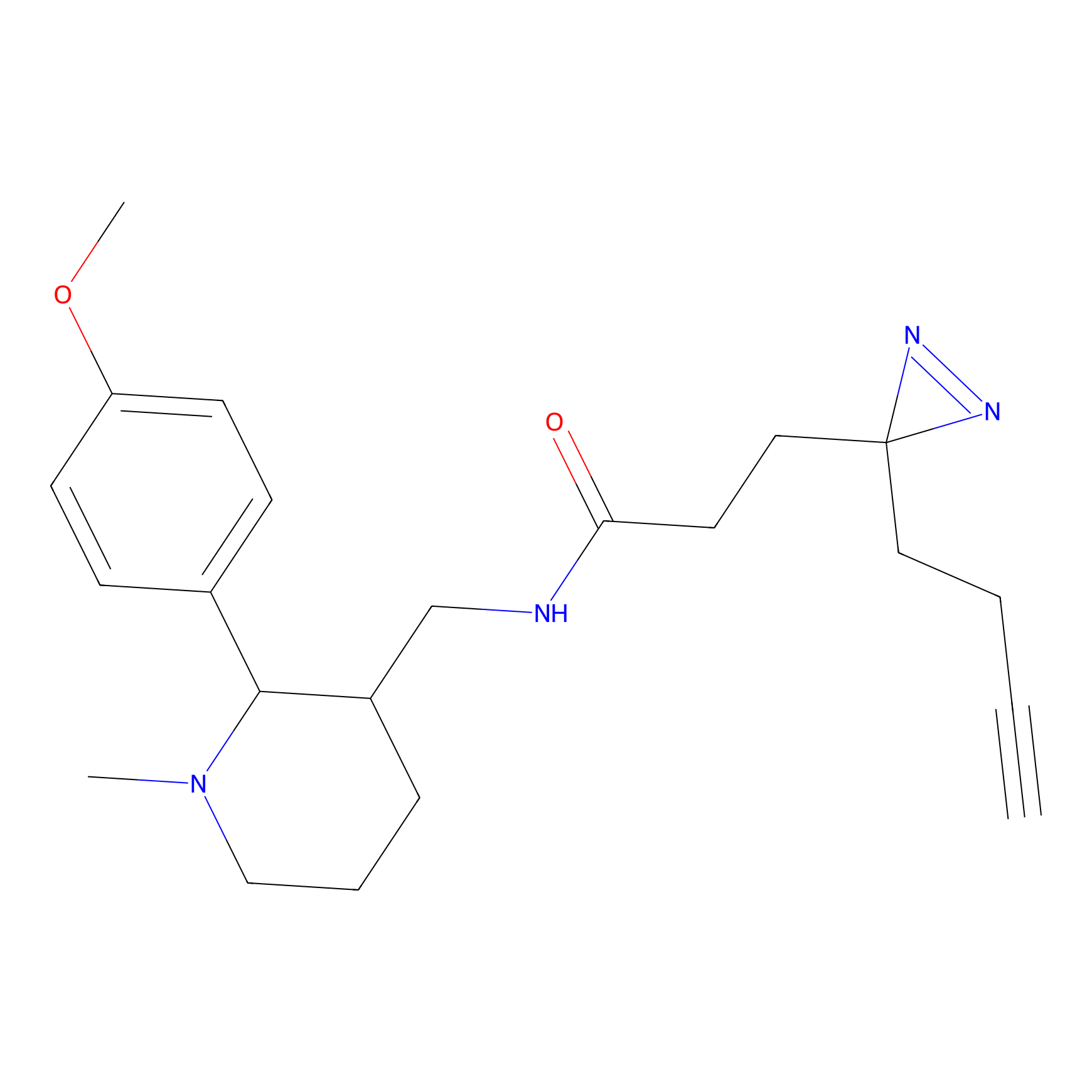 |
5.35 | LDD1897 | [17] | |
|
C302 Probe Info |
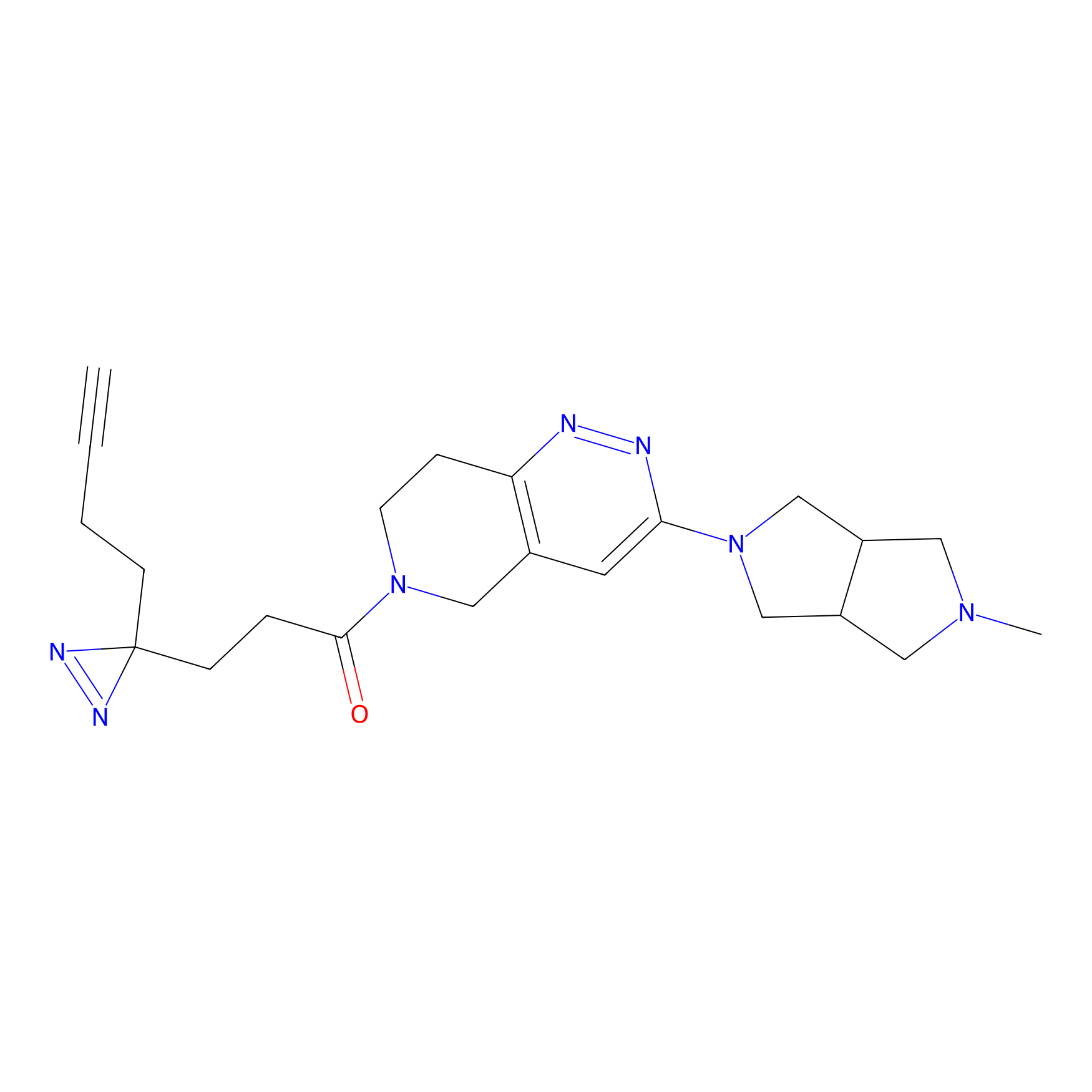 |
5.70 | LDD1971 | [17] | |
|
C310 Probe Info |
 |
19.16 | LDD1977 | [17] | |
|
C313 Probe Info |
 |
12.91 | LDD1980 | [17] | |
|
C350 Probe Info |
 |
25.81 | LDD2011 | [17] | |
|
C354 Probe Info |
 |
7.94 | LDD2015 | [17] | |
|
C425 Probe Info |
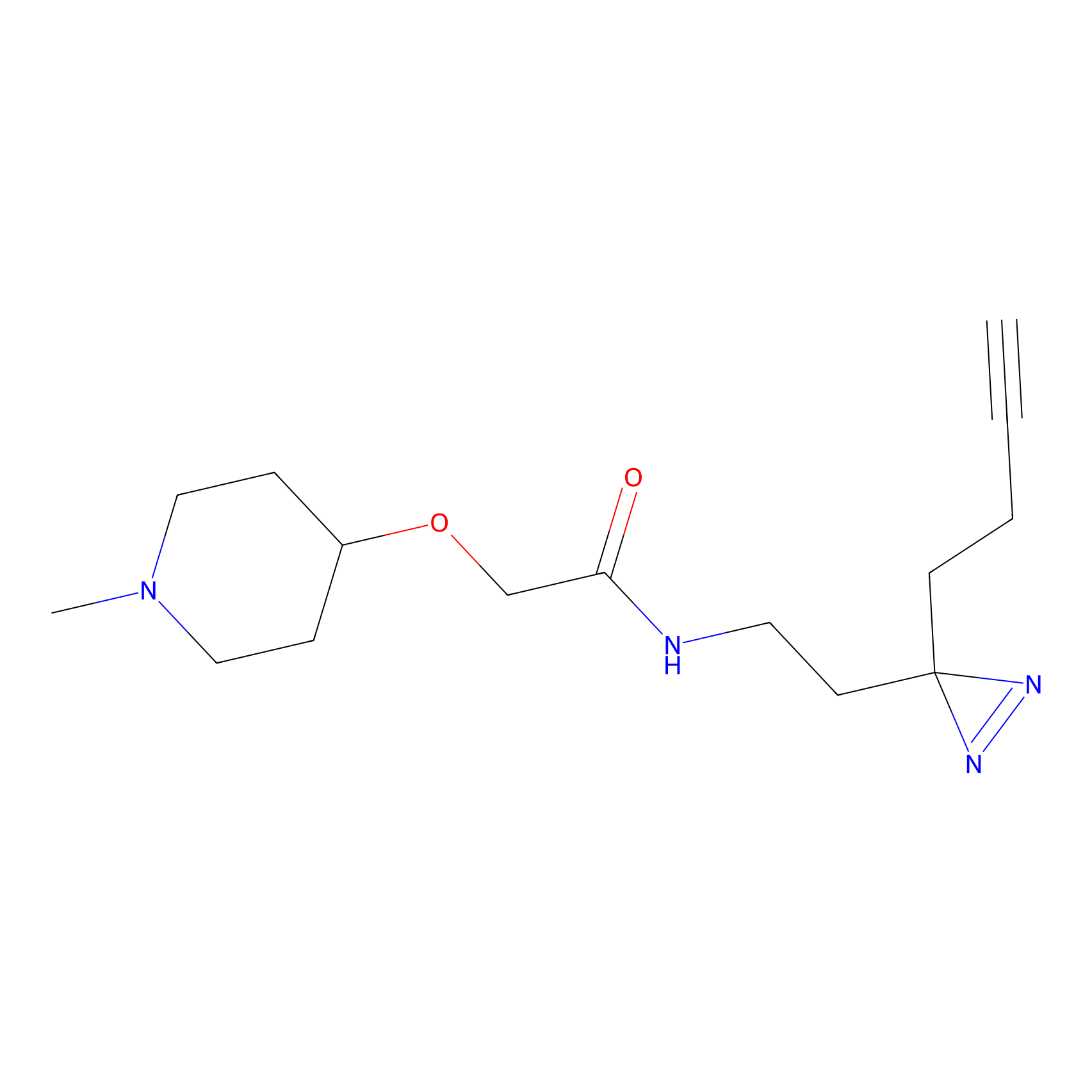 |
9.99 | LDD2080 | [17] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [18] | |
|
FFF probe12 Probe Info |
 |
20.00 | LDD0473 | [18] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [19] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [19] | |
|
Photonaproxen Probe Info |
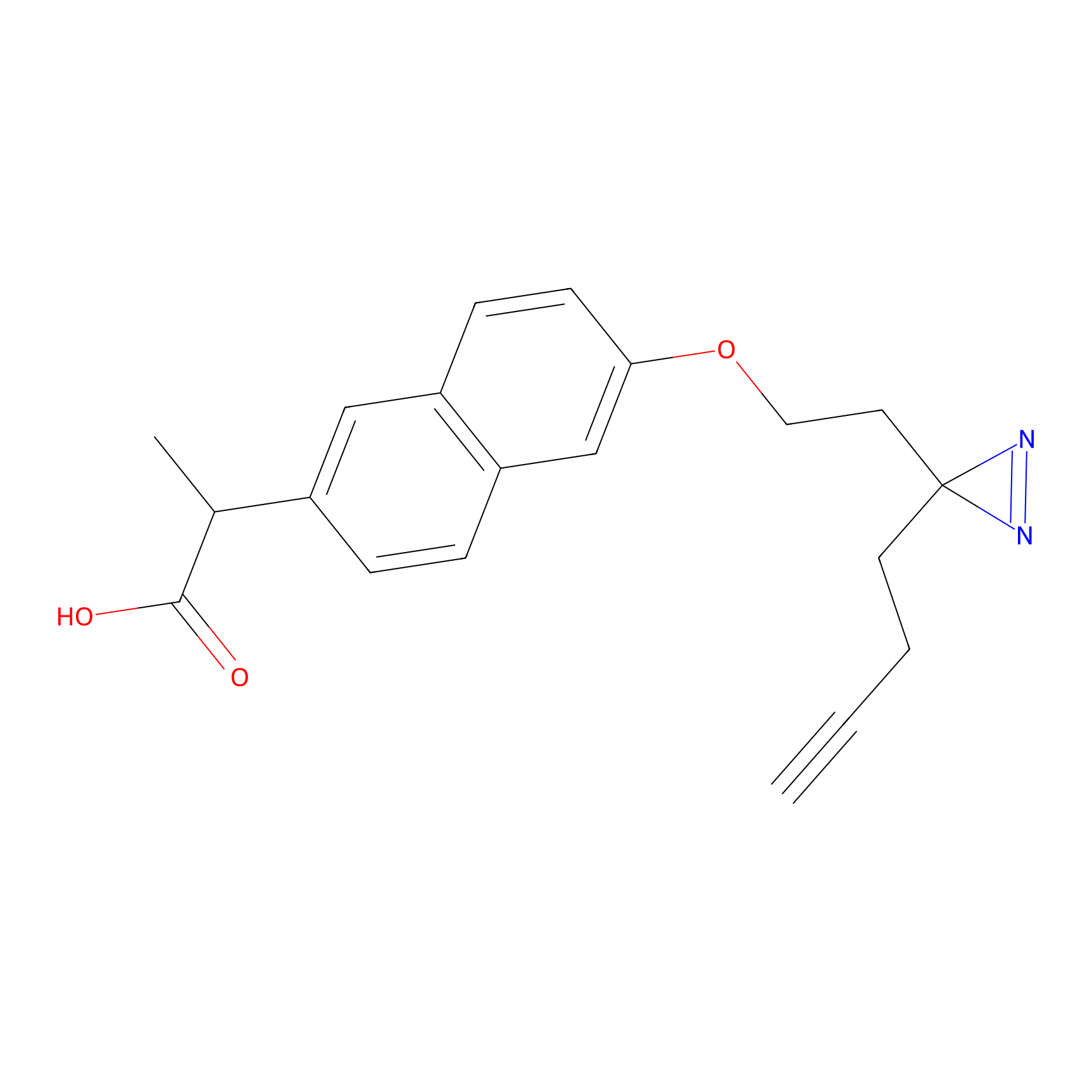 |
N.A. | LDD0157 | [20] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | N.A. | LDD0172 | [12] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | N.A. | LDD0173 | [12] |
| LDCM0545 | Acetamide | MDA-MB-231 | C363(1.83); C283(0.70) | LDD2170 | [9] |
| LDCM0156 | Aniline | NCI-H1299 | 13.42 | LDD0403 | [1] |
| LDCM0088 | C45 | HEK-293T | 12.57 | LDD0202 | [7] |
| LDCM0087 | Capsaicin | HEK-293T | 5.19 | LDD0185 | [11] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [15] |
| LDCM0027 | Dopamine | HEK-293T | 11.86 | LDD0179 | [11] |
| LDCM0625 | F8 | Ramos | C188(0.94); C363(1.17) | LDD2187 | [21] |
| LDCM0572 | Fragment10 | Ramos | C363(1.04) | LDD2189 | [21] |
| LDCM0573 | Fragment11 | Ramos | C188(20.00) | LDD2190 | [21] |
| LDCM0574 | Fragment12 | Ramos | C363(0.86) | LDD2191 | [21] |
| LDCM0575 | Fragment13 | Ramos | C363(1.06) | LDD2192 | [21] |
| LDCM0576 | Fragment14 | Ramos | C188(1.54); C363(0.97) | LDD2193 | [21] |
| LDCM0579 | Fragment20 | Ramos | C363(0.92) | LDD2194 | [21] |
| LDCM0580 | Fragment21 | Ramos | C363(0.80) | LDD2195 | [21] |
| LDCM0582 | Fragment23 | Ramos | C363(0.99) | LDD2196 | [21] |
| LDCM0578 | Fragment27 | Ramos | C363(0.82) | LDD2197 | [21] |
| LDCM0586 | Fragment28 | Ramos | C188(0.61); C363(0.80) | LDD2198 | [21] |
| LDCM0588 | Fragment30 | Ramos | C363(1.09) | LDD2199 | [21] |
| LDCM0589 | Fragment31 | Ramos | C363(0.99) | LDD2200 | [21] |
| LDCM0590 | Fragment32 | Ramos | C363(1.00) | LDD2201 | [21] |
| LDCM0468 | Fragment33 | Ramos | C363(0.95) | LDD2202 | [21] |
| LDCM0596 | Fragment38 | Ramos | C363(1.34) | LDD2203 | [21] |
| LDCM0566 | Fragment4 | Ramos | C363(0.93) | LDD2184 | [21] |
| LDCM0610 | Fragment52 | Ramos | C363(1.17) | LDD2204 | [21] |
| LDCM0614 | Fragment56 | Ramos | C363(1.23) | LDD2205 | [21] |
| LDCM0569 | Fragment7 | Ramos | C363(0.89) | LDD2186 | [21] |
| LDCM0571 | Fragment9 | Ramos | C363(1.08) | LDD2188 | [21] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [8] |
| LDCM0022 | KB02 | Ramos | C363(1.09) | LDD2182 | [21] |
| LDCM0023 | KB03 | MDA-MB-231 | C363(3.50) | LDD1701 | [9] |
| LDCM0024 | KB05 | WM115 | C188(17.35) | LDD3327 | [6] |
| LDCM0154 | YY4 | T cell | 2.09 | LDD0400 | [10] |
The Interaction Atlas With This Target
References
