Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
HDSF-alk Probe Info |
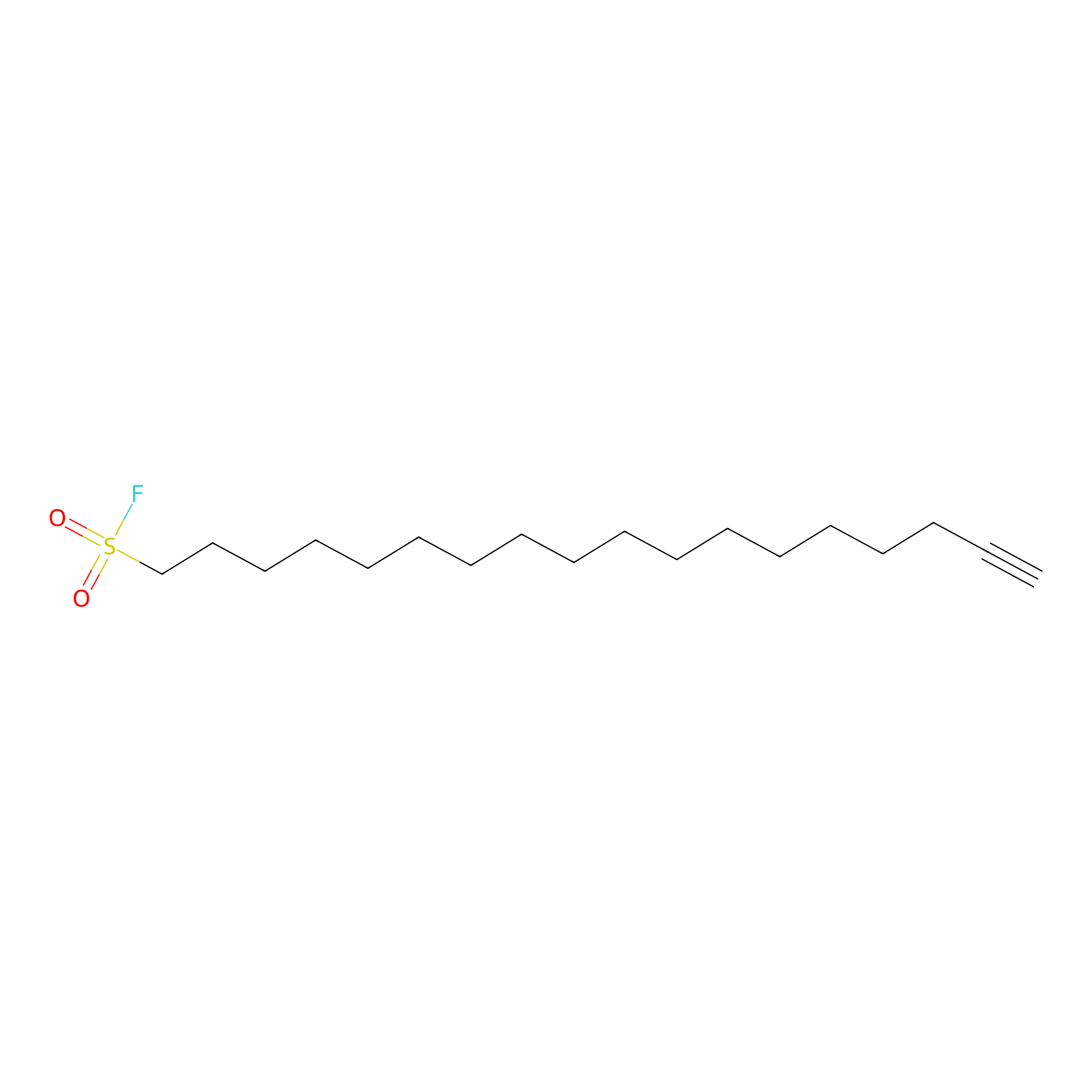 |
1.68 | LDD0197 | [1] | |
|
CY4 Probe Info |
 |
100.00 | LDD0244 | [2] | |
|
FBP2 Probe Info |
 |
3.15 | LDD0317 | [3] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
8.82 | LDD1740 | [4] | |
|
C091 Probe Info |
 |
18.90 | LDD1782 | [4] | |
|
C092 Probe Info |
 |
27.10 | LDD1783 | [4] | |
|
C094 Probe Info |
 |
41.64 | LDD1785 | [4] | |
|
C095 Probe Info |
 |
4.99 | LDD1786 | [4] | |
|
C112 Probe Info |
 |
26.35 | LDD1799 | [4] | |
|
C153 Probe Info |
 |
18.25 | LDD1834 | [4] | |
|
C201 Probe Info |
 |
27.86 | LDD1877 | [4] | |
|
C231 Probe Info |
 |
16.45 | LDD1904 | [4] | |
|
C264 Probe Info |
 |
17.75 | LDD1935 | [4] | |
|
C285 Probe Info |
 |
19.97 | LDD1955 | [4] | |
|
C287 Probe Info |
 |
10.93 | LDD1957 | [4] | |
|
C289 Probe Info |
 |
44.02 | LDD1959 | [4] | |
|
C390 Probe Info |
 |
36.76 | LDD2049 | [4] | |
|
C407 Probe Info |
 |
12.64 | LDD2064 | [4] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [5] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [5] | |
|
FFF probe14 Probe Info |
 |
17.09 | LDD0477 | [5] | |
|
OEA-DA Probe Info |
 |
19.47 | LDD0046 | [6] | |
The Interaction Atlas With This Target
References
