Details of the Target
General Information of Target
| Target ID | LDTP09261 | |||||
|---|---|---|---|---|---|---|
| Target Name | Major facilitator superfamily domain-containing protein 8 (MFSD8) | |||||
| Gene Name | MFSD8 | |||||
| Gene ID | 256471 | |||||
| Synonyms |
CLN7; Major facilitator superfamily domain-containing protein 8; Ceroid-lipofuscinosis neuronal protein 7 |
|||||
| 3D Structure | ||||||
| Sequence |
MAGLRNESEQEPLLGDTPGSREWDILETEEHYKSRWRSIRILYLTMFLSSVGFSVVMMSI
WPYLQKIDPTADTSFLGWVIASYSLGQMVASPIFGLWSNYRPRKEPLIVSILISVAANCL YAYLHIPASHNKYYMLVARGLLGIGAGNVAVVRSYTAGATSLQERTSSMANISMCQALGF ILGPVFQTCFTFLGEKGVTWDVIKLQINMYTTPVLLSAFLGILNIILILAILREHRVDDS GRQCKSINFEEASTDEAQVPQGNIDQVAVVAINVLFFVTLFIFALFETIITPLTMDMYAW TQEQAVLYNGIILAALGVEAVVIFLGVKLLSKKIGERAILLGGLIVVWVGFFILLPWGNQ FPKIQWEDLHNNSIPNTTFGEIIIGLWKSPMEDDNERPTGCSIEQAWCLYTPVIHLAQFL TSAVLIGLGYPVCNLMSYTLYSKILGPKPQGVYMGWLTASGSGARILGPMFISQVYAHWG PRWAFSLVCGIIVLTITLLGVVYKRLIALSVRYGRIQE |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Major facilitator superfamily
|
|||||
| Subcellular location |
Lysosome membrane
|
|||||
| Function |
Outward-rectifying chloride channel involved in endolysosomal chloride homeostasis, membrane fusion and function. Conducts chloride currents up to hundreds of picoamperes. Regulates lysosomal calcium content by reducing the lysosomal membrane potential, thereby activating TRPML1 channel and further release of lysosomal calcium ions. Regulates the pH in endolysosomal compartments and may contribute to progressive acidification from endosome to lysosome. Permeable to other halides such as iodide and fluoride ions.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AOyne Probe Info |
 |
11.60 | LDD0443 | [1] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C145 Probe Info |
 |
11.08 | LDD1827 | [2] | |
|
C170 Probe Info |
 |
15.03 | LDD1850 | [2] | |
|
C176 Probe Info |
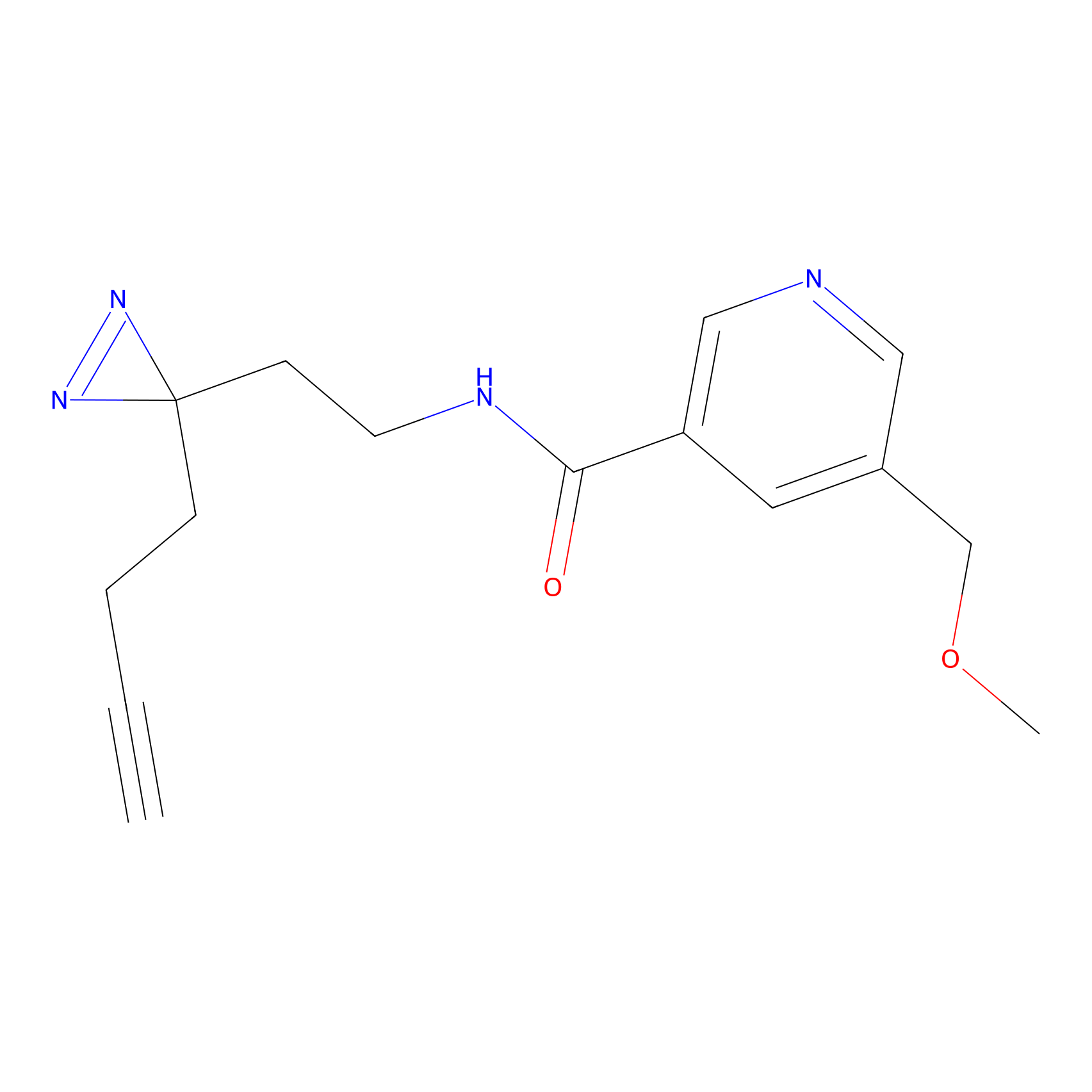 |
5.13 | LDD1855 | [2] | |
|
C177 Probe Info |
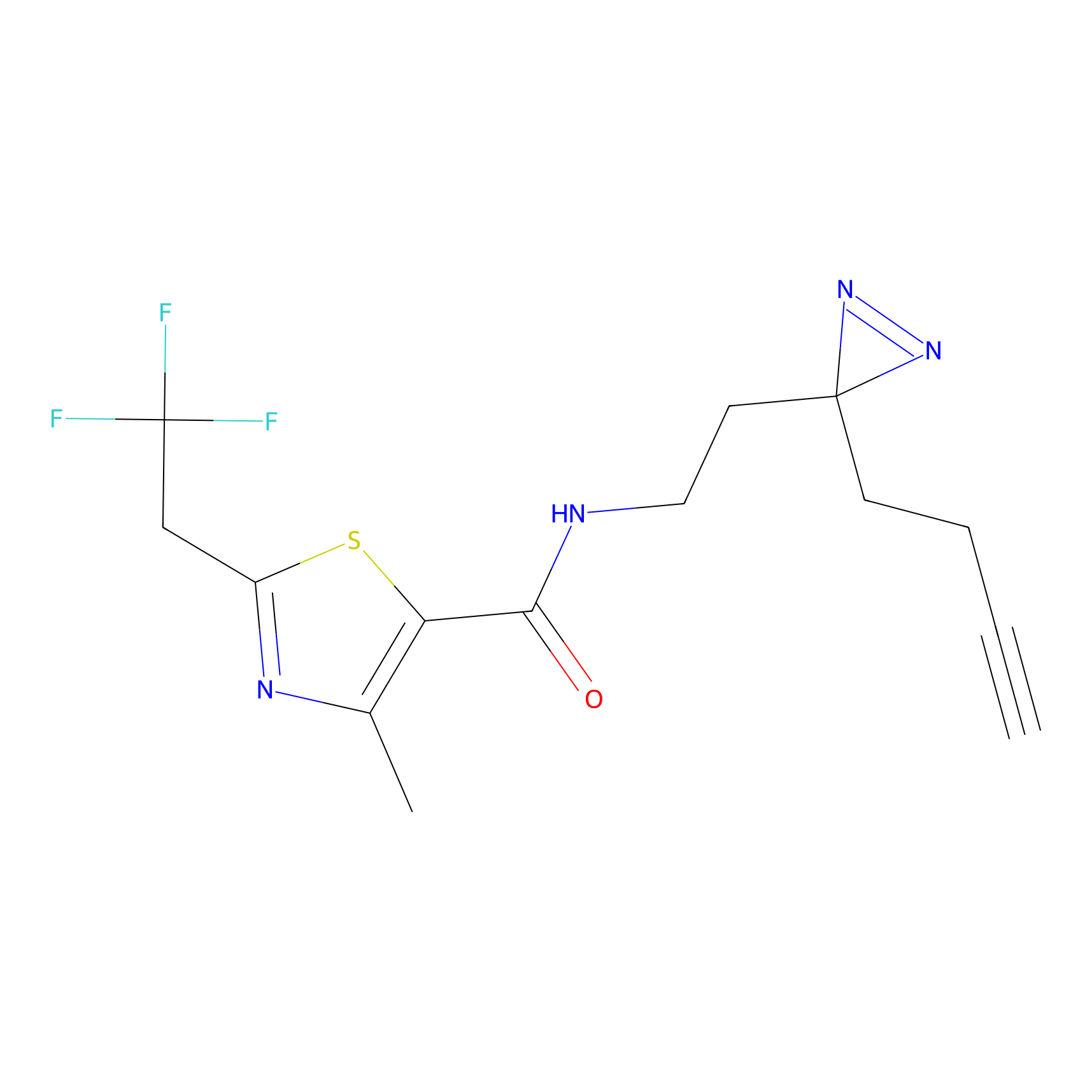 |
14.12 | LDD1856 | [2] | |
|
C178 Probe Info |
 |
21.56 | LDD1857 | [2] | |
|
C229 Probe Info |
 |
20.11 | LDD1902 | [2] | |
|
C231 Probe Info |
 |
20.82 | LDD1904 | [2] | |
|
C232 Probe Info |
 |
43.41 | LDD1905 | [2] | |
|
C233 Probe Info |
 |
21.71 | LDD1906 | [2] | |
|
C234 Probe Info |
 |
7.89 | LDD1907 | [2] | |
|
C235 Probe Info |
 |
29.65 | LDD1908 | [2] | |
|
C265 Probe Info |
 |
12.82 | LDD1936 | [2] | |
|
C270 Probe Info |
 |
8.51 | LDD1940 | [2] | |
|
C273 Probe Info |
 |
5.21 | LDD1943 | [2] | |
|
C277 Probe Info |
 |
7.73 | LDD1947 | [2] | |
|
C303 Probe Info |
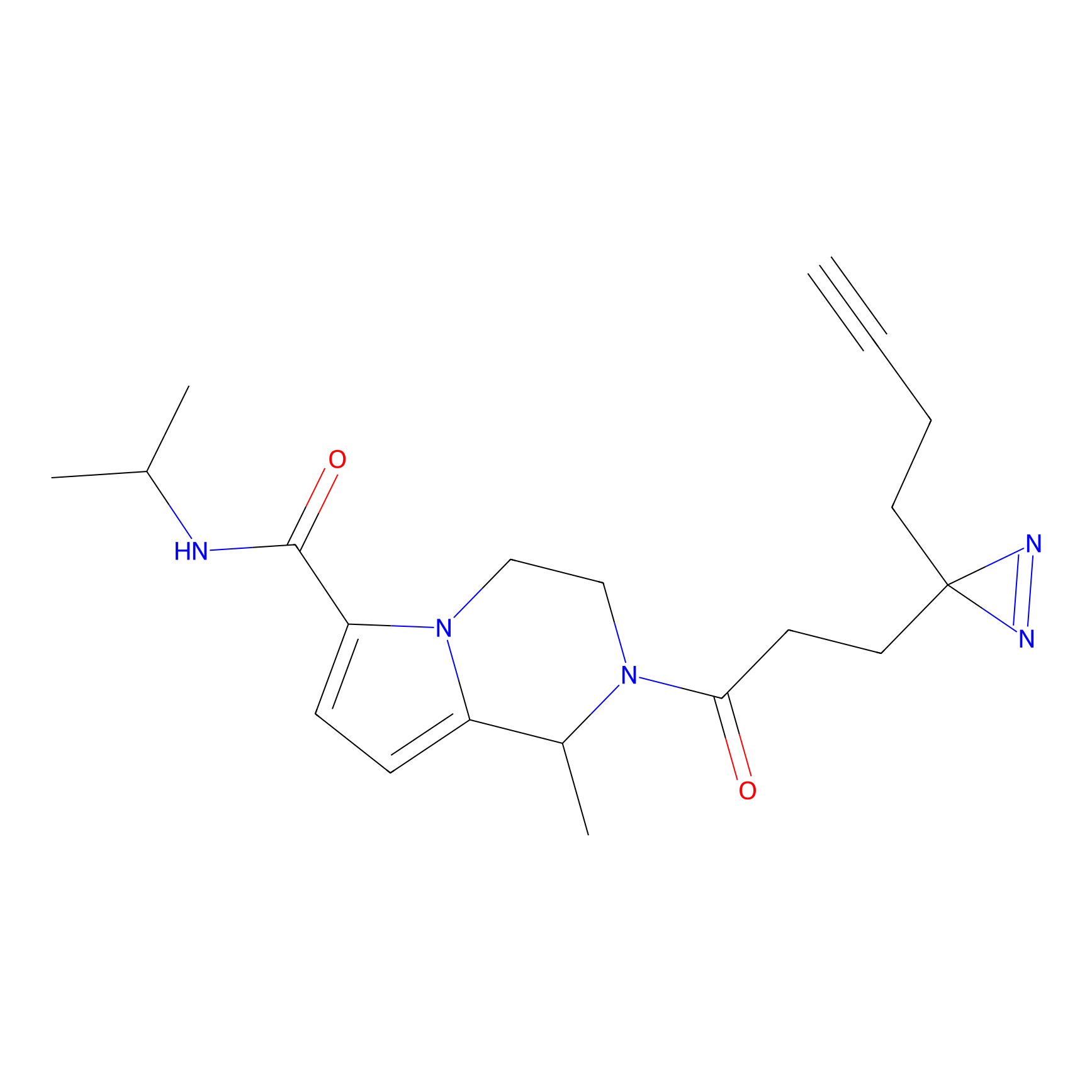 |
4.92 | LDD1972 | [2] | |
|
C307 Probe Info |
 |
8.69 | LDD1975 | [2] | |
|
C310 Probe Info |
 |
9.58 | LDD1977 | [2] | |
|
C424 Probe Info |
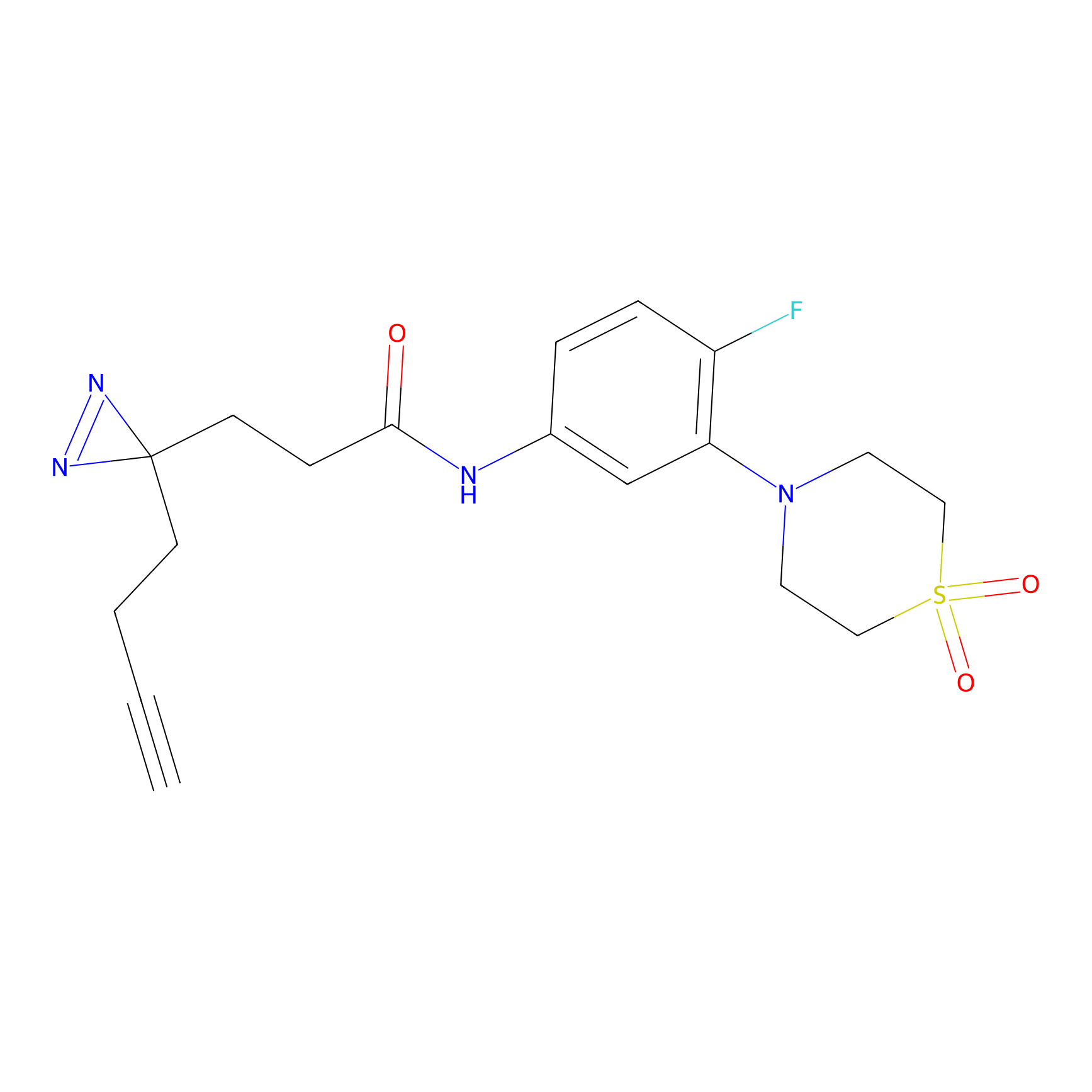 |
9.51 | LDD2079 | [2] | |
|
C426 Probe Info |
 |
31.78 | LDD2081 | [2] | |
|
C428 Probe Info |
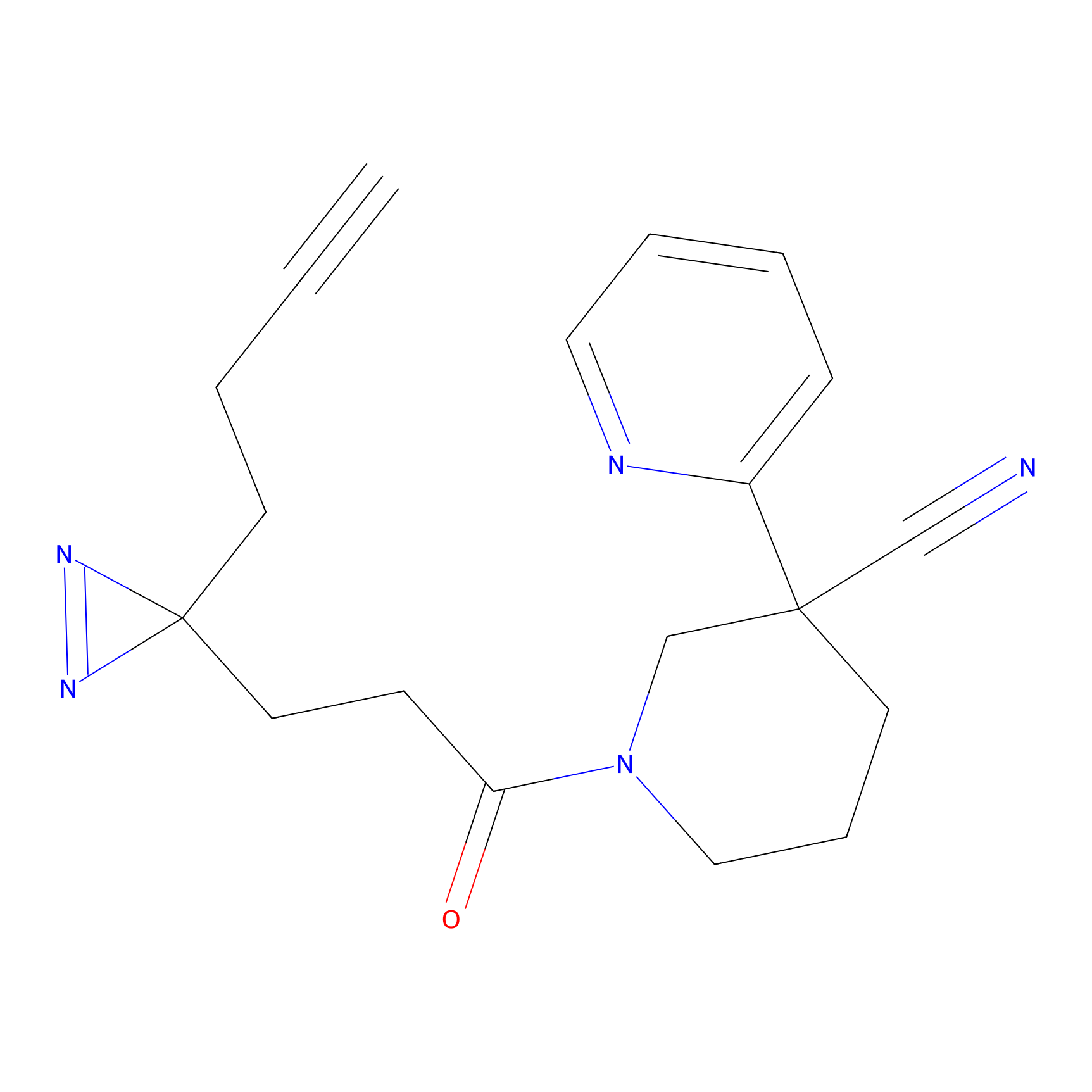 |
8.40 | LDD2083 | [2] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [3] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [3] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [4] | |
References
