Details of the Target
General Information of Target
| Target ID | LDTP07906 | |||||
|---|---|---|---|---|---|---|
| Target Name | Mediator of RNA polymerase II transcription subunit 25 (MED25) | |||||
| Gene Name | MED25 | |||||
| Gene ID | 81857 | |||||
| Synonyms |
ACID1; ARC92; PTOV2; Mediator of RNA polymerase II transcription subunit 25; Activator interaction domain-containing protein 1; Activator-recruited cofactor 92 kDa component; ARC92; Mediator complex subunit 25; p78
|
|||||
| 3D Structure | ||||||
| Sequence |
MVPGSEGPARAGSVVADVVFVIEGTANLGPYFEGLRKHYLLPAIEYFNGGPPAETDFGGD
YGGTQYSLVVFNTVDCAPESYVQCHAPTSSAYEFVTWLDGIKFMGGGGESCSLIAEGLST ALQLFDDFKKMREQIGQTHRVCLLICNSPPYLLPAVESTTYSGCTTENLVQQIGERGIHF SIVSPRKLPALRLLFEKAAPPALLEPLQPPTDVSQDPRHMVLVRGLVLPVGGGSAPGPLQ SKQPVPLPPAAPSGATLSAAPQQPLPPVPPQYQVPGNLSAAQVAAQNAVEAAKNQKAGLG PRFSPITPLQQAAPGVGPPFSQAPAPQLPPGPPGAPKPPPASQPSLVSTVAPGSGLAPTA QPGAPSMAGTVAPGGVSGPSPAQLGAPALGGQQSVSNKLLAWSGVLEWQEKPKPASVDAN TKLTRSLPCQVYVNHGENLKTEQWPQKLIMQLIPQQLLTTLGPLFRNSRMVQFHFTNKDL ESLKGLYRIMGNGFAGCVHFPHTAPCEVRVLMLLYSSKKKIFMGLIPYDQSGFVNGIRQV ITNHKQVQQQKLEQQQRGMGGQQAPPGLGPILEDQARPSQNLLQLRPPQPQPQGTVGASG ATGQPQPQGTAQPPPGAPQGPPGAASGPPPPGPILRPQNPGANPQLRSLLLNPPPPQTGV PPPQASLHHLQPPGAPALLPPPHQGLGQPQLGPPLLHPPPAQSWPAQLPPRAPLPGQMLL SGGPRGPVPQPGLQPSVMEDDILMDLI |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Mediator complex subunit 25 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Component of the Mediator complex, a coactivator involved in the regulated transcription of nearly all RNA polymerase II-dependent genes. Mediator functions as a bridge to convey information from gene-specific regulatory proteins to the basal RNA polymerase II transcription machinery. Mediator is recruited to promoters by direct interactions with regulatory proteins and serves as a scaffold for the assembly of a functional preinitiation complex with RNA polymerase II and the general transcription factors. Required for RARA/RXRA-mediated transcription.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| MELJUSO | SNV: p.P629L | DBIA Probe Info | |||
| MOLT4 | Deletion: p.T64PfsTer150; p.K187SfsTer27 | IA-alkyne Probe Info | |||
| NOMO1 | SNV: p.G237A | . | |||
| RKO | Deletion: p.P566QfsTer31 | DBIA Probe Info | |||
| RPMI8226 | SNV: p.A504V | DBIA Probe Info | |||
| SKMEL28 | SNV: p.L278V | DBIA Probe Info | |||
| TE11 | SNV: p.G722R | DBIA Probe Info | |||
| TOV21G | SNV: p.S258Ter | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
13.89 | LDD0402 | [1] | |
|
STPyne Probe Info |
 |
K545(5.00) | LDD0277 | [2] | |
|
DBIA Probe Info |
 |
C429(2.04) | LDD3312 | [3] | |
|
BTD Probe Info |
 |
C429(1.22) | LDD2094 | [4] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [5] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [5] | |
|
IPM Probe Info |
 |
N.A. | LDD0147 | [6] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [7] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [8] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C008 Probe Info |
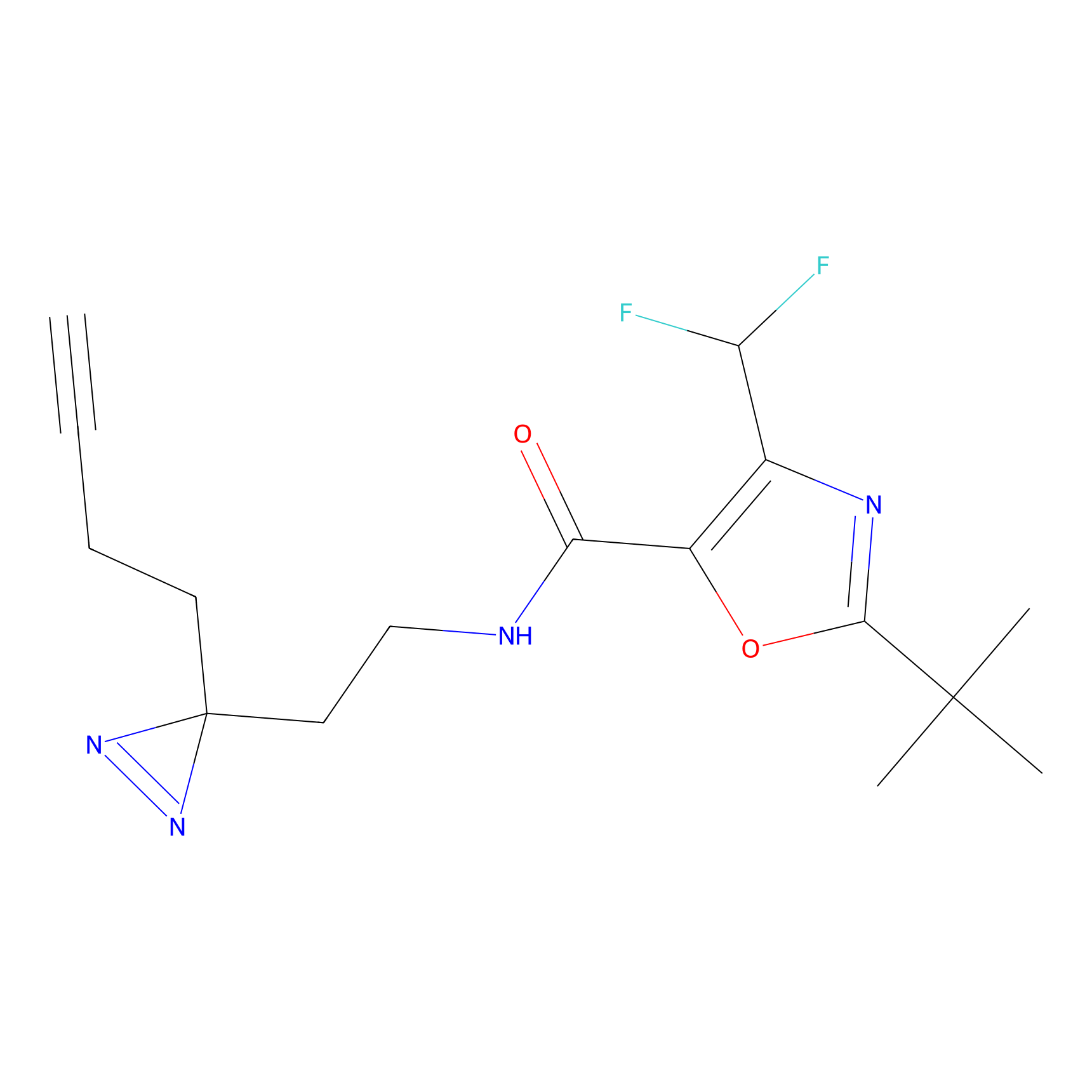 |
5.35 | LDD1717 | [9] | |
|
C010 Probe Info |
 |
5.70 | LDD1719 | [9] | |
|
C059 Probe Info |
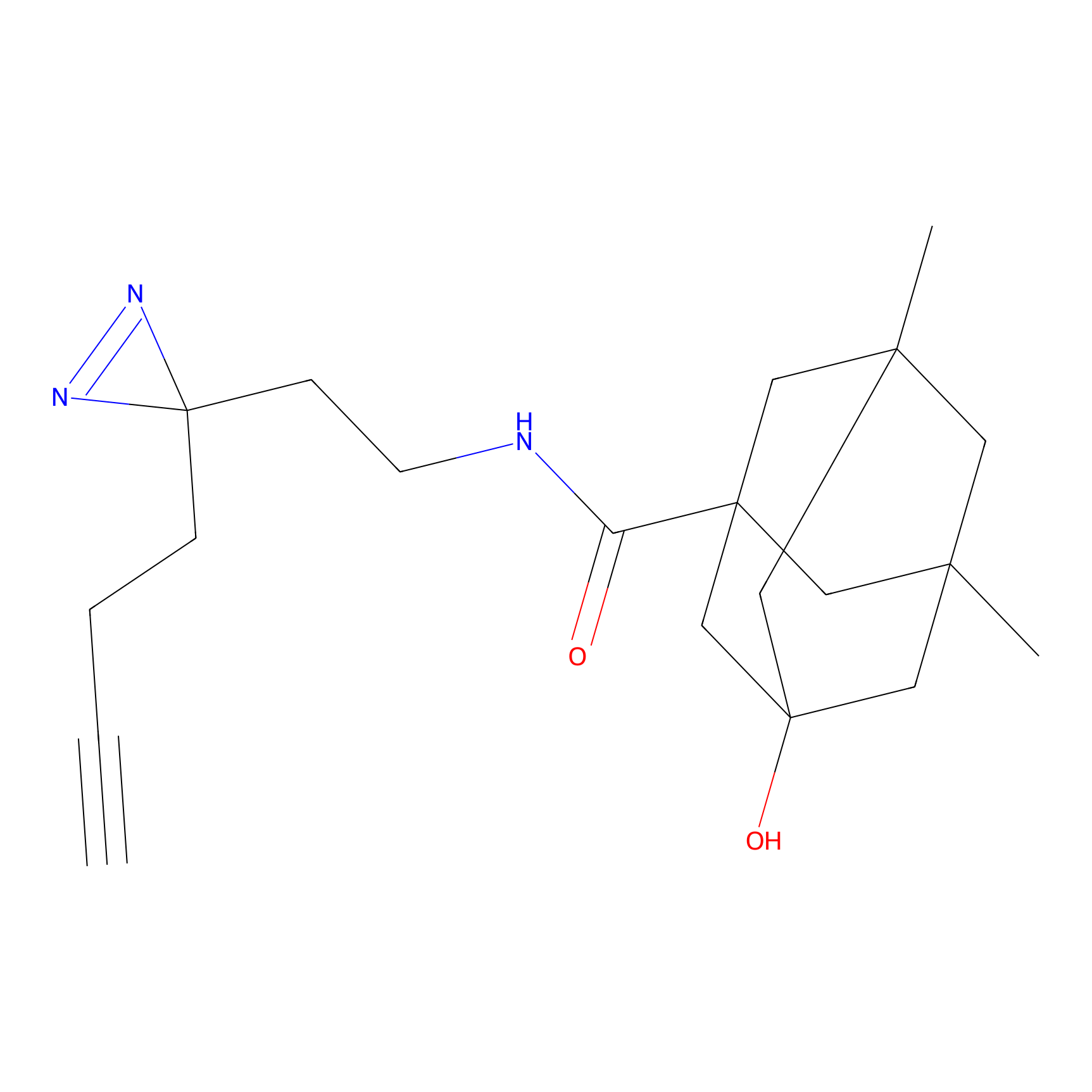 |
36.00 | LDD1756 | [9] | |
|
C064 Probe Info |
 |
5.98 | LDD1761 | [9] | |
|
C067 Probe Info |
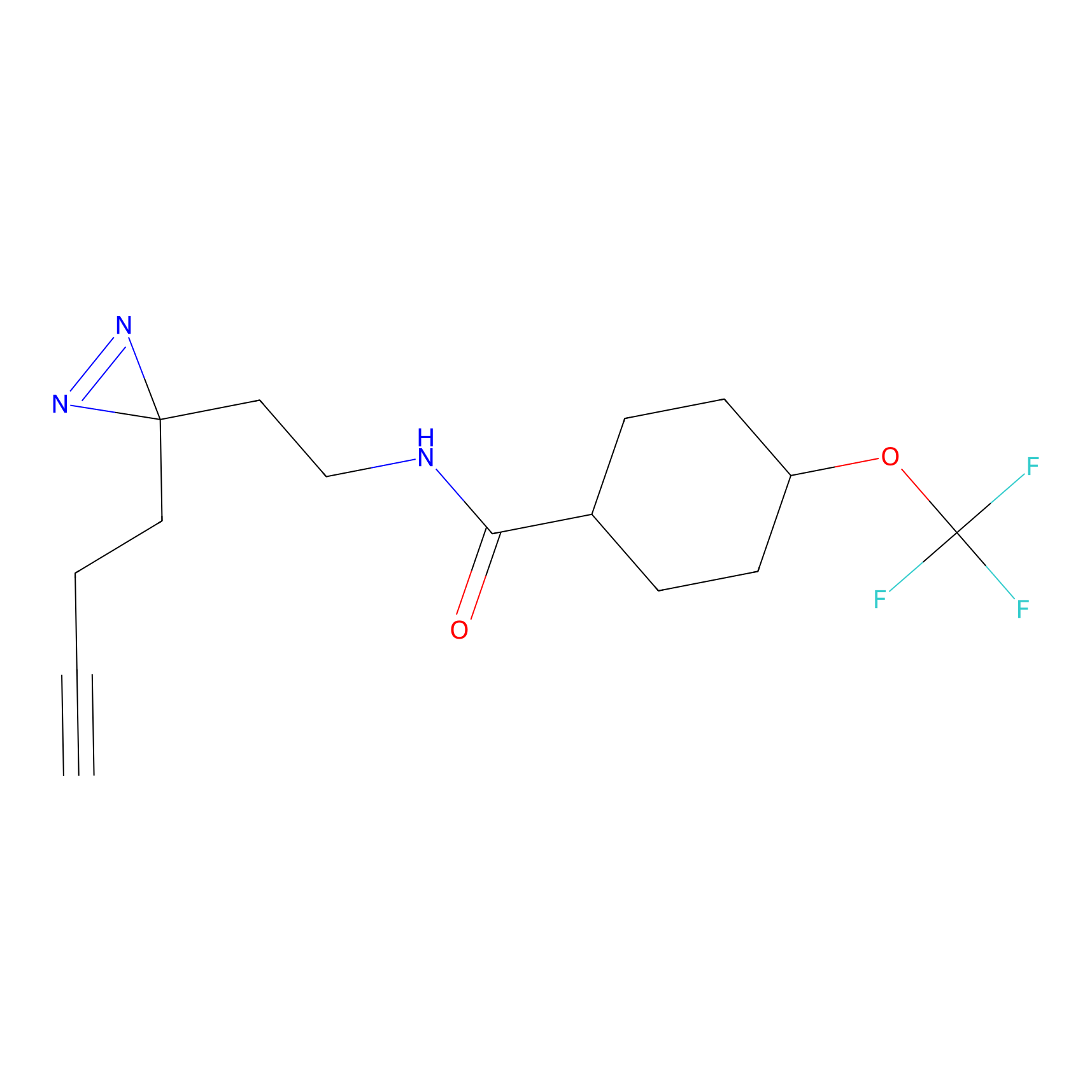 |
5.54 | LDD1763 | [9] | |
|
C070 Probe Info |
 |
13.45 | LDD1766 | [9] | |
|
C082 Probe Info |
 |
6.41 | LDD1774 | [9] | |
|
C083 Probe Info |
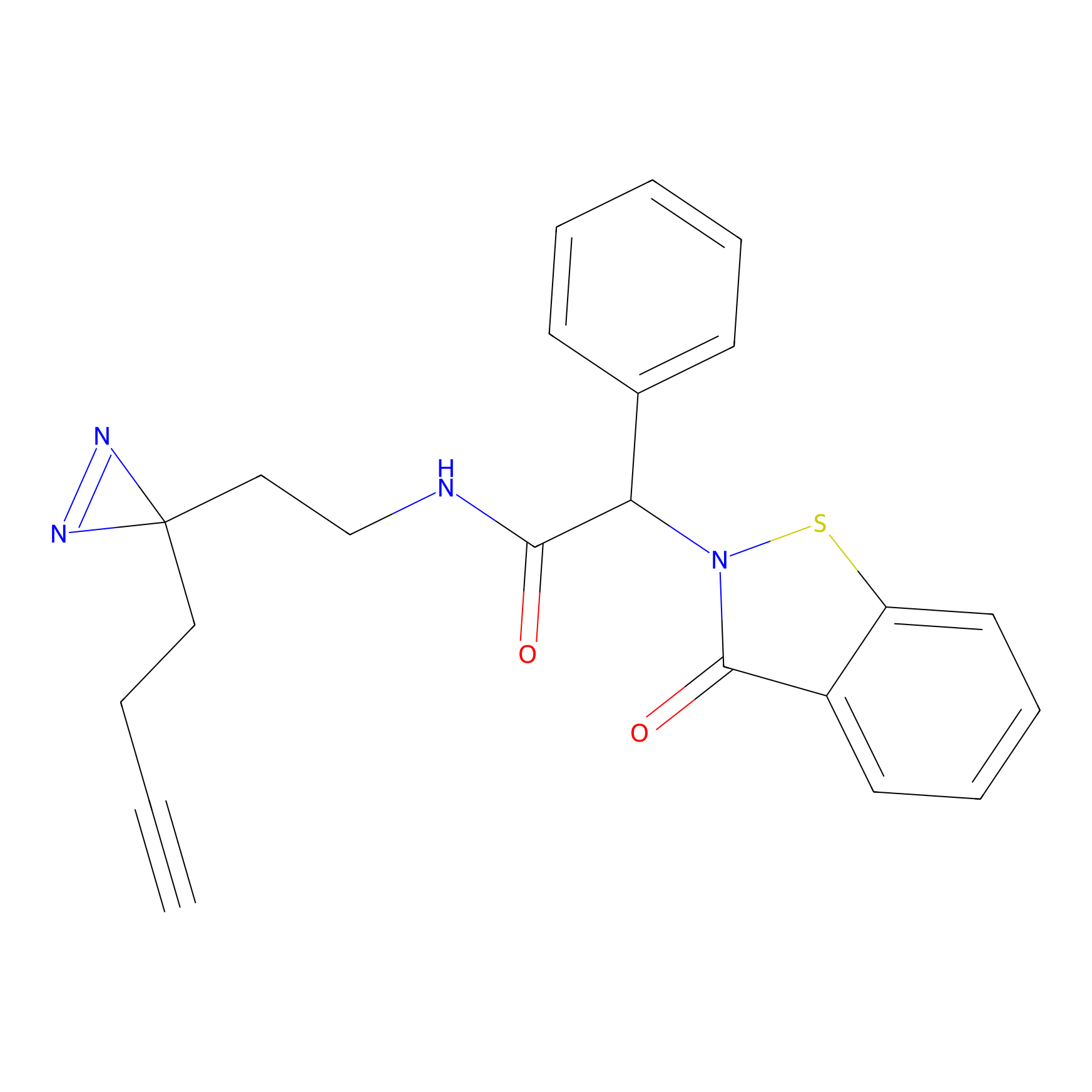 |
5.24 | LDD1775 | [9] | |
|
C085 Probe Info |
 |
9.58 | LDD1777 | [9] | |
|
C107 Probe Info |
 |
10.93 | LDD1794 | [9] | |
|
C108 Probe Info |
 |
8.28 | LDD1795 | [9] | |
|
C130 Probe Info |
 |
6.82 | LDD1812 | [9] | |
|
C160 Probe Info |
 |
6.87 | LDD1840 | [9] | |
|
C178 Probe Info |
 |
16.45 | LDD1857 | [9] | |
|
C198 Probe Info |
 |
14.12 | LDD1874 | [9] | |
|
C219 Probe Info |
 |
10.34 | LDD1893 | [9] | |
|
C220 Probe Info |
 |
12.21 | LDD1894 | [9] | |
|
C277 Probe Info |
 |
19.29 | LDD1947 | [9] | |
|
C363 Probe Info |
 |
41.64 | LDD2024 | [9] | |
|
AEA-DA Probe Info |
 |
5.18 | LDD0333 | [10] | |
|
OEA-DA Probe Info |
 |
20.00 | LDD0046 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C429(1.08) | LDD2103 | [4] |
| LDCM0572 | Fragment10 | Ramos | C429(1.75) | LDD2189 | [11] |
| LDCM0573 | Fragment11 | Ramos | C429(2.99) | LDD2190 | [11] |
| LDCM0575 | Fragment13 | Ramos | C429(0.88) | LDD2192 | [11] |
| LDCM0576 | Fragment14 | Ramos | C429(1.07) | LDD2193 | [11] |
| LDCM0580 | Fragment21 | Ramos | C429(1.07) | LDD2195 | [11] |
| LDCM0582 | Fragment23 | Ramos | C429(0.76) | LDD2196 | [11] |
| LDCM0578 | Fragment27 | Ramos | C429(0.85) | LDD2197 | [11] |
| LDCM0586 | Fragment28 | Ramos | C429(0.65) | LDD2198 | [11] |
| LDCM0588 | Fragment30 | Ramos | C429(1.20) | LDD2199 | [11] |
| LDCM0589 | Fragment31 | Ramos | C429(0.93) | LDD2200 | [11] |
| LDCM0468 | Fragment33 | Ramos | C429(1.13) | LDD2202 | [11] |
| LDCM0596 | Fragment38 | Ramos | C429(0.91) | LDD2203 | [11] |
| LDCM0610 | Fragment52 | Ramos | C429(2.52) | LDD2204 | [11] |
| LDCM0614 | Fragment56 | Ramos | C429(1.13) | LDD2205 | [11] |
| LDCM0571 | Fragment9 | Ramos | C429(1.91) | LDD2188 | [11] |
| LDCM0022 | KB02 | HEK-293T | C429(0.98) | LDD1492 | [12] |
| LDCM0023 | KB03 | HEK-293T | C429(1.00) | LDD1497 | [12] |
| LDCM0024 | KB05 | HMCB | C429(2.04) | LDD3312 | [3] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C429(1.22) | LDD2094 | [4] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C429(0.72) | LDD2098 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C429(0.74) | LDD2109 | [4] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C429(0.74) | LDD2128 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C429(1.00) | LDD2129 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C429(0.91) | LDD2146 | [4] |
| LDCM0559 | Nucleophilic fragment 9b | MDA-MB-231 | C429(2.08) | LDD2153 | [4] |
| LDCM0136 | SR-4559 | HEK-293T | 5.18 | LDD0333 | [10] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| NEDD4-like E3 ubiquitin-protein ligase WWP2 (WWP2) | . | O00308 | |||
Transcription factor
GPCR
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Oxoeicosanoid receptor 1 (OXER1) | G-protein coupled receptor 1 family | Q8TDS5 | |||
Other
References
