Details of the Target
General Information of Target
| Target ID | LDTP05920 | |||||
|---|---|---|---|---|---|---|
| Target Name | Calcium/calmodulin-dependent protein kinase type II subunit gamma (CAMK2G) | |||||
| Gene Name | CAMK2G | |||||
| Gene ID | 818 | |||||
| Synonyms |
CAMK; CAMK-II; CAMKG; Calcium/calmodulin-dependent protein kinase type II subunit gamma; CaM kinase II subunit gamma; CaMK-II subunit gamma; EC 2.7.11.17 |
|||||
| 3D Structure | ||||||
| Sequence |
MATTATCTRFTDDYQLFEELGKGAFSVVRRCVKKTSTQEYAAKIINTKKLSARDHQKLER
EARICRLLKHPNIVRLHDSISEEGFHYLVFDLVTGGELFEDIVAREYYSEADASHCIHQI LESVNHIHQHDIVHRDLKPENLLLASKCKGAAVKLADFGLAIEVQGEQQAWFGFAGTPGY LSPEVLRKDPYGKPVDIWACGVILYILLVGYPPFWDEDQHKLYQQIKAGAYDFPSPEWDT VTPEAKNLINQMLTINPAKRITADQALKHPWVCQRSTVASMMHRQETVECLRKFNARRKL KGAILTTMLVSRNFSAAKSLLNKKSDGGVKKRKSSSSVHLMPQSNNKNSLVSPAQEPAPL QTAMEPQTTVVHNATDGIKGSTESCNTTTEDEDLKGRVPEGRSSRDRTAPSAGMQPQPSL CSSAMRKQEIIKITEQLIEAINNGDFEAYTKICDPGLTSFEPEALGNLVEGMDFHKFYFE NLLSKNSKPIHTTILNPHVHVIGEDAACIAYIRLTQYIDGQGRPRTSQSEETRVWHRRDG KWLNVHYHCSGAPAAPLQ |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Protein kinase superfamily, CAMK Ser/Thr protein kinase family, CaMK subfamily
|
|||||
| Subcellular location |
Sarcoplasmic reticulum membrane
|
|||||
| Function |
Calcium/calmodulin-dependent protein kinase that functions autonomously after Ca(2+)/calmodulin-binding and autophosphorylation, and is involved in sarcoplasmic reticulum Ca(2+) transport in skeletal muscle and may function in dendritic spine and synapse formation and neuronal plasticity. In slow-twitch muscles, is involved in regulation of sarcoplasmic reticulum (SR) Ca(2+) transport and in fast-twitch muscle participates in the control of Ca(2+) release from the SR through phosphorylation of the ryanodine receptor-coupling factor triadin. In the central nervous system, it is involved in the regulation of neurite formation and arborization. It may participate in the promotion of dendritic spine and synapse formation and maintenance of synaptic plasticity which enables long-term potentiation (LTP) and hippocampus-dependent learning. In response to interferon-gamma (IFN-gamma) stimulation, catalyzes phosphorylation of STAT1, stimulating the JAK-STAT signaling pathway.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
TH211 Probe Info |
 |
Y231(20.00); Y517(11.98) | LDD0257 | [2] | |
|
TH216 Probe Info |
 |
Y517(5.32) | LDD0259 | [2] | |
|
DBIA Probe Info |
 |
C273(4.05) | LDD3310 | [3] | |
|
HPAP Probe Info |
 |
3.00 | LDD0062 | [4] | |
|
ATP probe Probe Info |
 |
K147(0.00); K43(0.00); K138(0.00); K227(0.00) | LDD0199 | [5] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C385(0.00); C273(0.00); C453(0.00); C290(0.00) | LDD0038 | [6] | |
|
IA-alkyne Probe Info |
 |
C273(0.00); C290(0.00); C549(0.00); C148(0.00) | LDD0036 | [6] | |
|
Lodoacetamide azide Probe Info |
 |
C385(0.00); C273(0.00); C453(0.00); C290(0.00) | LDD0037 | [6] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [7] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [8] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [9] | |
|
IPM Probe Info |
 |
C273(0.00); C7(0.00) | LDD0147 | [8] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [9] | |
|
Phosphinate-6 Probe Info |
 |
C273(0.00); C148(0.00) | LDD0018 | [10] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [11] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [11] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [11] | |
|
AOyne Probe Info |
 |
14.50 | LDD0443 | [12] | |
|
NAIA_5 Probe Info |
 |
C273(0.00); C7(0.00) | LDD2223 | [13] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C017 Probe Info |
 |
6.50 | LDD1725 | [14] | |
|
C055 Probe Info |
 |
11.88 | LDD1752 | [14] | |
|
C056 Probe Info |
 |
16.11 | LDD1753 | [14] | |
|
C063 Probe Info |
 |
11.08 | LDD1760 | [14] | |
|
C085 Probe Info |
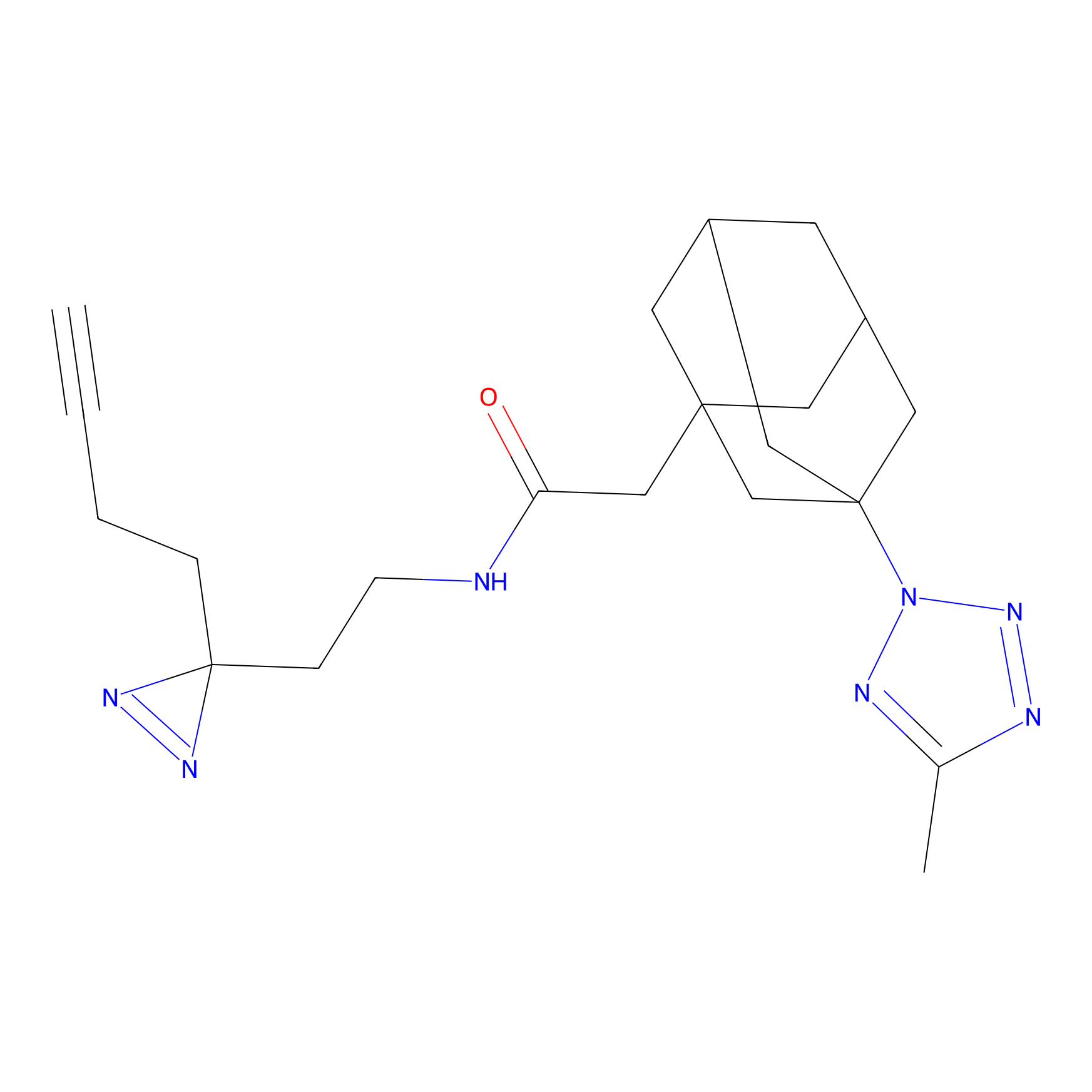 |
6.19 | LDD1777 | [14] | |
|
C159 Probe Info |
 |
12.21 | LDD1839 | [14] | |
|
C212 Probe Info |
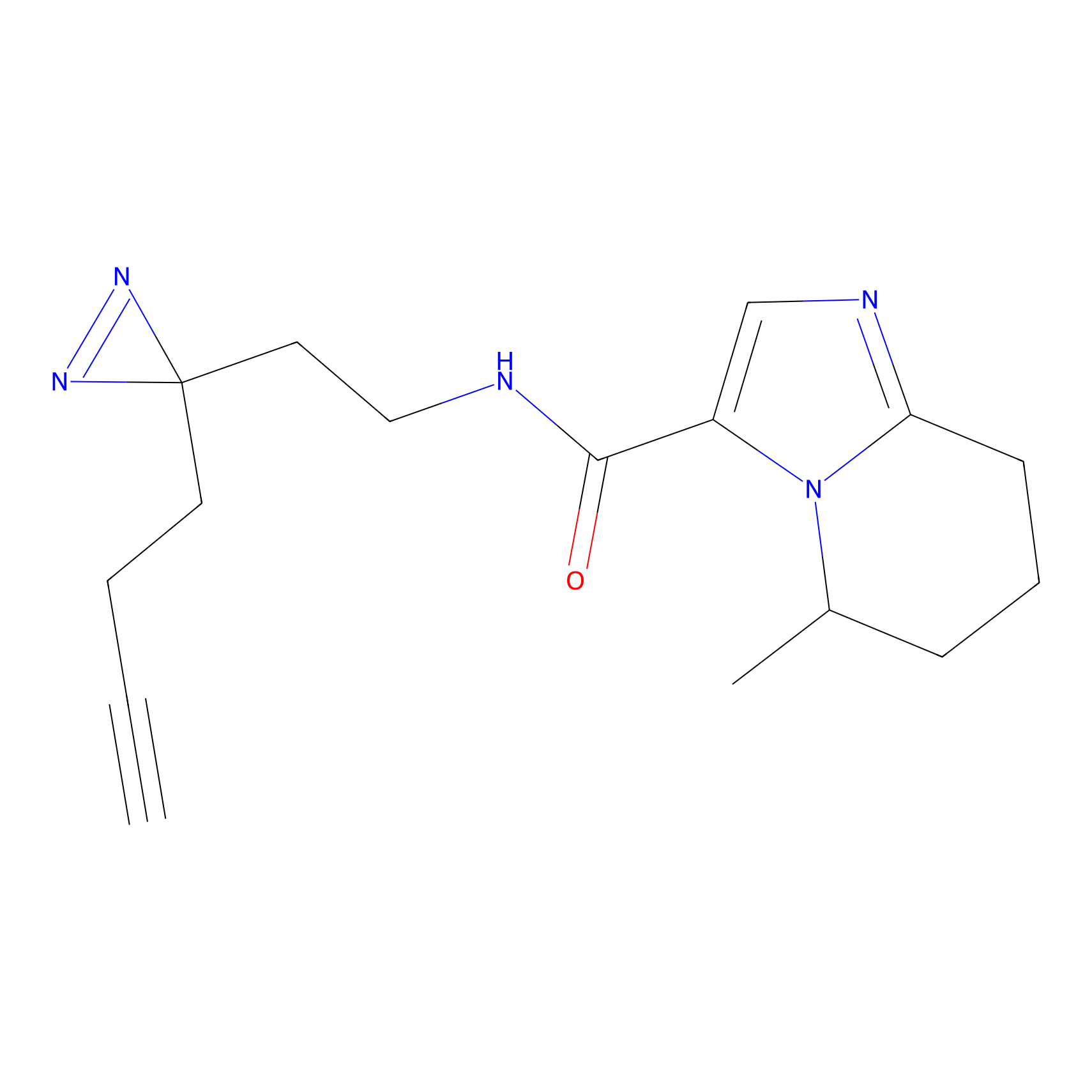 |
8.94 | LDD1886 | [14] | |
|
C216 Probe Info |
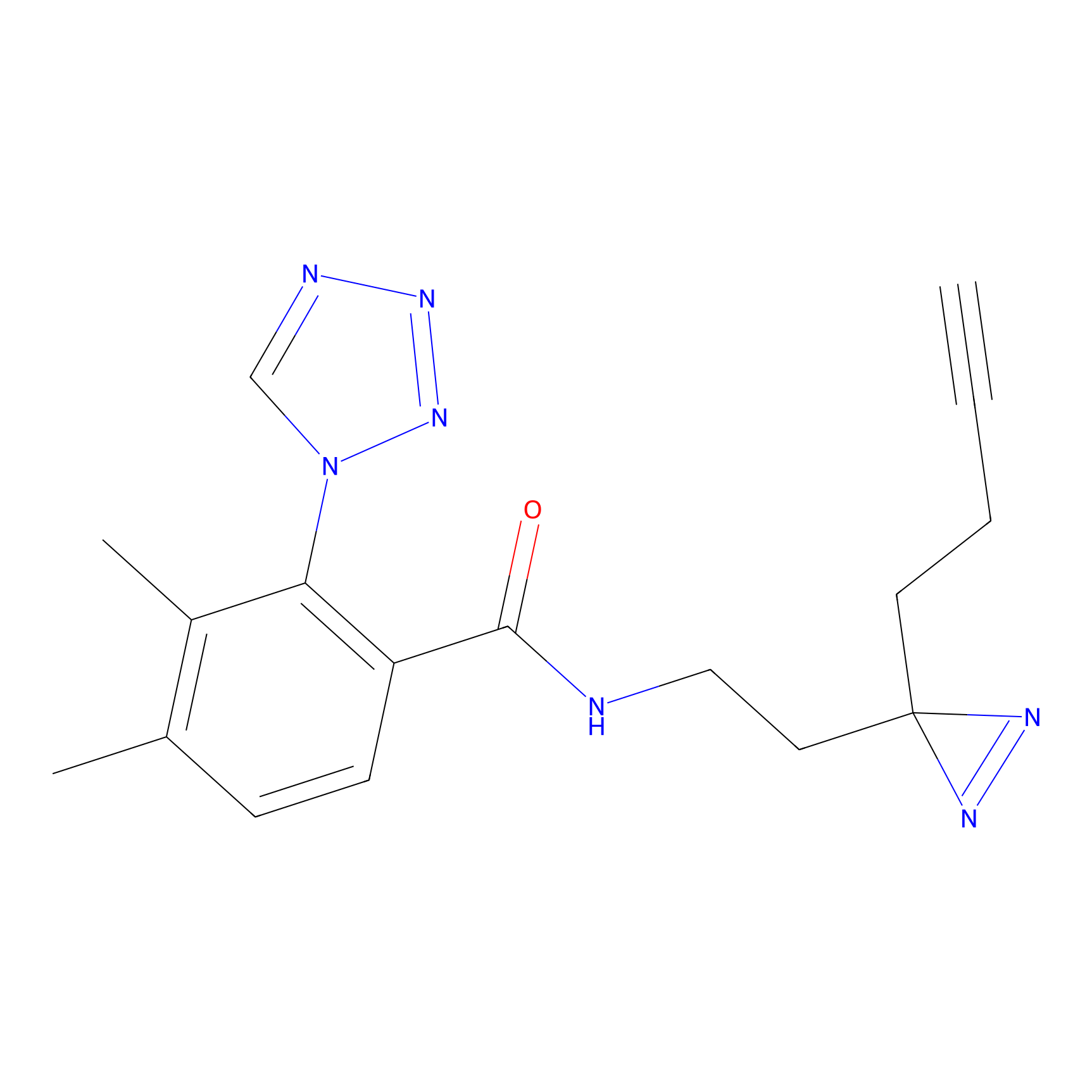 |
6.54 | LDD1890 | [14] | |
|
C218 Probe Info |
 |
14.12 | LDD1892 | [14] | |
|
C280 Probe Info |
 |
15.35 | LDD1950 | [14] | |
|
C282 Probe Info |
 |
21.56 | LDD1952 | [14] | |
|
C284 Probe Info |
 |
55.72 | LDD1954 | [14] | |
|
C338 Probe Info |
 |
18.77 | LDD2001 | [14] | |
|
C403 Probe Info |
 |
16.68 | LDD2061 | [14] | |
|
FFF probe11 Probe Info |
 |
7.26 | LDD0472 | [15] | |
|
FFF probe13 Probe Info |
 |
9.91 | LDD0476 | [15] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [15] | |
|
FFF probe3 Probe Info |
 |
20.00 | LDD0465 | [15] | |
|
FFF probe6 Probe Info |
 |
11.01 | LDD0468 | [15] | |
|
LD-F Probe Info |
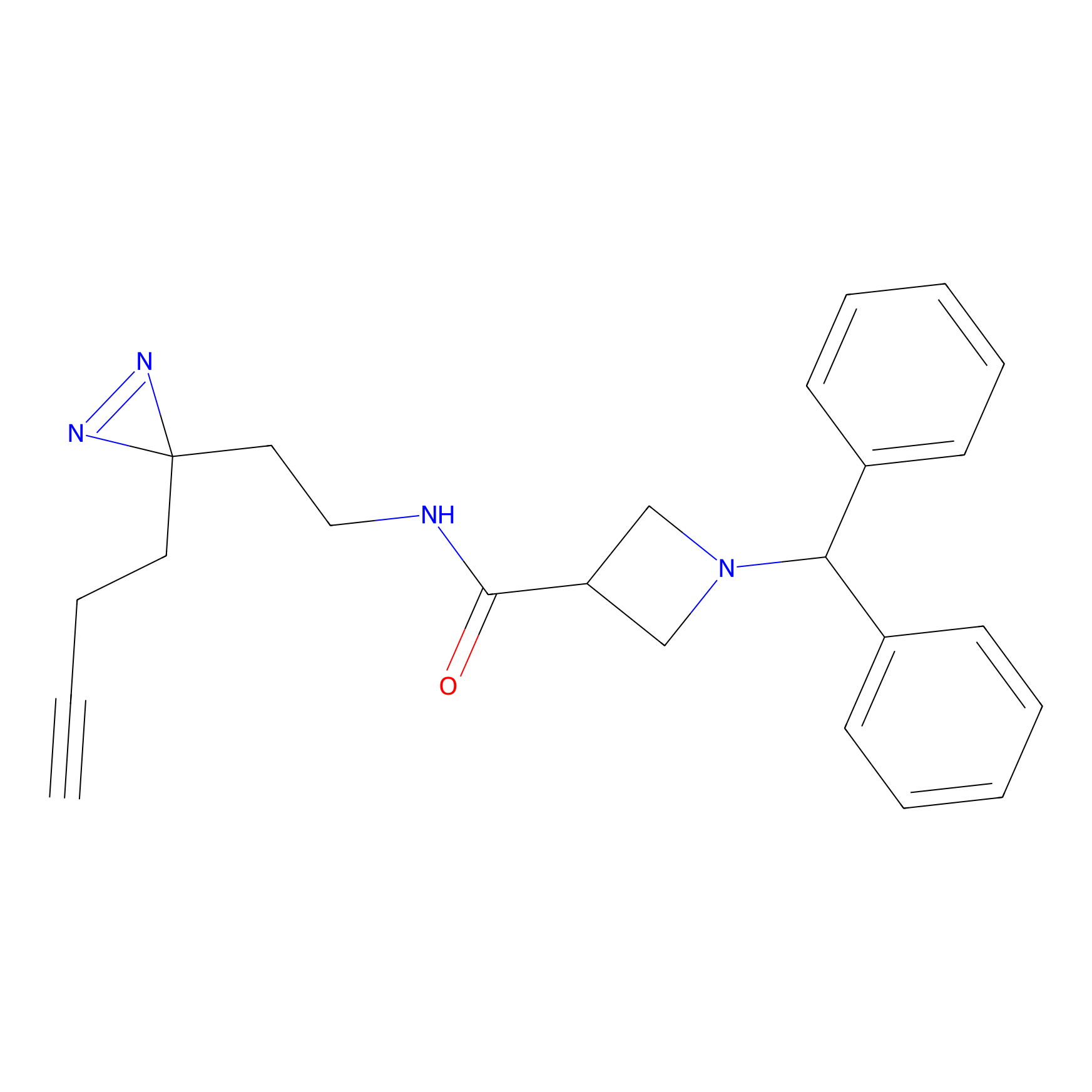 |
L457(0.00); E461(0.00); S459(0.00) | LDD0015 | [16] | |
|
OEA-DA Probe Info |
 |
3.83 | LDD0046 | [17] | |
|
Probe 12 Probe Info |
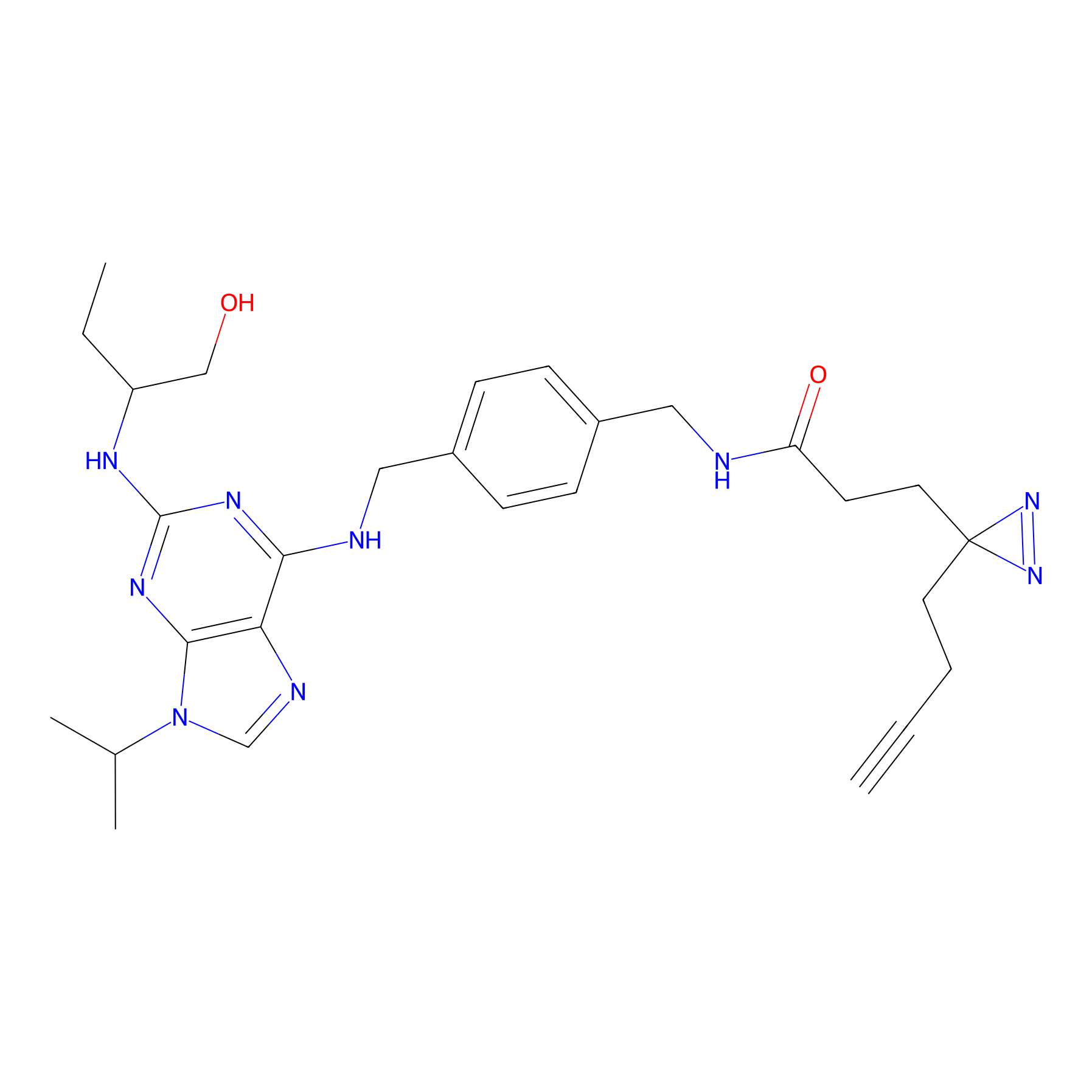 |
N.A. | LDD0420 | [18] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C290(1.10); C453(1.09); C385(0.87) | LDD1507 | [19] |
| LDCM0215 | AC10 | HEK-293T | C290(1.00); C453(0.98); C385(0.98); C421(1.44) | LDD1508 | [19] |
| LDCM0226 | AC11 | HEK-293T | C290(1.00); C453(0.96); C385(1.08) | LDD1509 | [19] |
| LDCM0237 | AC12 | HEK-293T | C290(1.00); C453(1.00); C385(0.96) | LDD1510 | [19] |
| LDCM0259 | AC14 | HEK-293T | C290(0.99); C453(0.96); C385(1.01); C421(0.95) | LDD1512 | [19] |
| LDCM0270 | AC15 | HEK-293T | C290(1.11); C453(1.08); C385(1.08) | LDD1513 | [19] |
| LDCM0276 | AC17 | HEK-293T | C290(1.03); C453(0.97); C385(1.04) | LDD1515 | [19] |
| LDCM0277 | AC18 | HEK-293T | C290(1.01); C453(0.90); C385(0.98); C421(1.10) | LDD1516 | [19] |
| LDCM0278 | AC19 | HEK-293T | C290(1.12); C453(0.85); C385(1.19) | LDD1517 | [19] |
| LDCM0279 | AC2 | HEK-293T | C290(0.98); C453(0.96); C385(0.98); C421(1.22) | LDD1518 | [19] |
| LDCM0280 | AC20 | HEK-293T | C290(1.02); C453(1.01); C385(1.07) | LDD1519 | [19] |
| LDCM0281 | AC21 | HEK-293T | C290(1.01); C453(1.06); C385(0.96) | LDD1520 | [19] |
| LDCM0282 | AC22 | HEK-293T | C290(1.05); C453(1.02); C385(0.99); C421(1.05) | LDD1521 | [19] |
| LDCM0283 | AC23 | HEK-293T | C290(1.07); C453(1.20); C385(1.07) | LDD1522 | [19] |
| LDCM0284 | AC24 | HEK-293T | C290(0.93); C453(1.06); C421(1.09) | LDD1523 | [19] |
| LDCM0285 | AC25 | HEK-293T | C290(1.04); C453(1.07); C385(0.87) | LDD1524 | [19] |
| LDCM0286 | AC26 | HEK-293T | C290(1.02); C453(1.01); C385(1.07); C421(1.18) | LDD1525 | [19] |
| LDCM0287 | AC27 | HEK-293T | C290(1.00); C453(0.94); C385(1.06) | LDD1526 | [19] |
| LDCM0288 | AC28 | HEK-293T | C290(1.01); C453(1.05); C385(1.02) | LDD1527 | [19] |
| LDCM0289 | AC29 | HEK-293T | C290(1.03); C453(0.94); C385(1.00) | LDD1528 | [19] |
| LDCM0290 | AC3 | HEK-293T | C290(0.99); C453(0.86); C385(1.06) | LDD1529 | [19] |
| LDCM0291 | AC30 | HEK-293T | C290(0.96); C453(1.13); C385(1.00); C421(0.91) | LDD1530 | [19] |
| LDCM0292 | AC31 | HEK-293T | C290(1.13); C453(1.07); C385(1.15) | LDD1531 | [19] |
| LDCM0293 | AC32 | HEK-293T | C290(0.99); C453(1.05); C421(0.92) | LDD1532 | [19] |
| LDCM0294 | AC33 | HEK-293T | C290(0.99); C453(1.08); C385(0.99) | LDD1533 | [19] |
| LDCM0295 | AC34 | HEK-293T | C290(0.99); C453(0.94); C385(0.94); C421(1.20) | LDD1534 | [19] |
| LDCM0296 | AC35 | HEK-293T | C290(1.04); C453(0.88); C385(1.07) | LDD1535 | [19] |
| LDCM0297 | AC36 | HEK-293T | C290(1.04); C453(1.17); C385(0.99) | LDD1536 | [19] |
| LDCM0298 | AC37 | HEK-293T | C290(1.07); C453(0.93); C385(1.05) | LDD1537 | [19] |
| LDCM0299 | AC38 | HEK-293T | C290(0.99); C453(0.90); C385(0.97); C421(0.98) | LDD1538 | [19] |
| LDCM0300 | AC39 | HEK-293T | C290(1.02); C453(1.29); C385(1.12) | LDD1539 | [19] |
| LDCM0301 | AC4 | HEK-293T | C290(0.99); C453(1.23); C385(0.92) | LDD1540 | [19] |
| LDCM0302 | AC40 | HEK-293T | C290(1.08); C453(1.04); C421(1.04) | LDD1541 | [19] |
| LDCM0303 | AC41 | HEK-293T | C290(1.15); C453(1.10); C385(1.09) | LDD1542 | [19] |
| LDCM0304 | AC42 | HEK-293T | C290(1.00); C453(1.05); C385(1.07); C421(1.25) | LDD1543 | [19] |
| LDCM0305 | AC43 | HEK-293T | C290(1.01); C453(0.97); C385(1.13) | LDD1544 | [19] |
| LDCM0306 | AC44 | HEK-293T | C290(1.02); C453(0.89); C385(0.99) | LDD1545 | [19] |
| LDCM0307 | AC45 | HEK-293T | C290(1.08); C453(0.97); C385(0.88) | LDD1546 | [19] |
| LDCM0308 | AC46 | HEK-293T | C290(1.02); C453(0.93); C385(1.02); C421(1.01) | LDD1547 | [19] |
| LDCM0309 | AC47 | HEK-293T | C290(1.09); C453(1.27); C385(1.01) | LDD1548 | [19] |
| LDCM0310 | AC48 | HEK-293T | C290(1.08); C453(1.04); C421(1.13) | LDD1549 | [19] |
| LDCM0311 | AC49 | HEK-293T | C290(1.04); C453(1.06); C385(0.98) | LDD1550 | [19] |
| LDCM0312 | AC5 | HEK-293T | C290(1.09); C453(0.90); C385(0.94) | LDD1551 | [19] |
| LDCM0313 | AC50 | HEK-293T | C290(0.98); C453(1.04); C385(0.98); C421(1.19) | LDD1552 | [19] |
| LDCM0314 | AC51 | HEK-293T | C290(0.99); C453(1.03); C385(1.14) | LDD1553 | [19] |
| LDCM0315 | AC52 | HEK-293T | C290(1.07); C453(0.99); C385(0.94) | LDD1554 | [19] |
| LDCM0316 | AC53 | HEK-293T | C290(1.01); C453(1.03); C385(0.93) | LDD1555 | [19] |
| LDCM0317 | AC54 | HEK-293T | C290(0.96); C453(1.06); C385(1.01); C421(0.81) | LDD1556 | [19] |
| LDCM0318 | AC55 | HEK-293T | C290(1.07); C453(1.09); C385(1.01) | LDD1557 | [19] |
| LDCM0319 | AC56 | HEK-293T | C290(1.01); C453(0.98); C421(1.12) | LDD1558 | [19] |
| LDCM0320 | AC57 | HEK-293T | C290(1.09); C453(1.19); C385(1.06) | LDD1559 | [19] |
| LDCM0321 | AC58 | HEK-293T | C290(1.03); C453(1.02); C385(0.97); C421(1.18) | LDD1560 | [19] |
| LDCM0322 | AC59 | HEK-293T | C290(1.11); C453(1.01); C385(1.10) | LDD1561 | [19] |
| LDCM0323 | AC6 | HEK-293T | C290(1.02); C453(1.05); C385(0.95); C421(0.80) | LDD1562 | [19] |
| LDCM0324 | AC60 | HEK-293T | C290(1.04); C453(1.11); C385(0.90) | LDD1563 | [19] |
| LDCM0325 | AC61 | HEK-293T | C290(1.00); C453(0.96); C385(0.92) | LDD1564 | [19] |
| LDCM0326 | AC62 | HEK-293T | C290(0.97); C453(0.99); C385(0.95); C421(0.96) | LDD1565 | [19] |
| LDCM0327 | AC63 | HEK-293T | C290(1.13); C453(1.08); C385(0.95) | LDD1566 | [19] |
| LDCM0328 | AC64 | HEK-293T | C290(0.99); C453(1.01); C421(0.93) | LDD1567 | [19] |
| LDCM0334 | AC7 | HEK-293T | C290(1.08); C453(1.09); C385(1.04) | LDD1568 | [19] |
| LDCM0345 | AC8 | HEK-293T | C290(0.99); C453(0.97); C421(1.14) | LDD1569 | [19] |
| LDCM0248 | AKOS034007472 | HEK-293T | C290(1.07); C453(0.93); C385(0.96) | LDD1511 | [19] |
| LDCM0356 | AKOS034007680 | HEK-293T | C290(1.07); C453(1.18); C385(1.06) | LDD1570 | [19] |
| LDCM0275 | AKOS034007705 | HEK-293T | C290(0.98); C453(0.99); C421(1.48) | LDD1514 | [19] |
| LDCM0156 | Aniline | NCI-H1299 | 11.90 | LDD0403 | [1] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [11] |
| LDCM0632 | CL-Sc | Hep-G2 | C273(0.93); C7(0.90) | LDD2227 | [13] |
| LDCM0367 | CL1 | HEK-293T | C290(1.02); C453(1.10); C385(1.19) | LDD1571 | [19] |
| LDCM0368 | CL10 | HEK-293T | C290(0.85); C453(0.76); C385(0.85); C421(1.04) | LDD1572 | [19] |
| LDCM0369 | CL100 | HEK-293T | C290(1.02); C453(0.89) | LDD1573 | [19] |
| LDCM0370 | CL101 | HEK-293T | C290(0.97); C453(1.11); C385(1.13) | LDD1574 | [19] |
| LDCM0371 | CL102 | HEK-293T | C290(0.97); C453(0.64) | LDD1575 | [19] |
| LDCM0372 | CL103 | HEK-293T | C290(1.00); C453(1.39); C385(1.03) | LDD1576 | [19] |
| LDCM0373 | CL104 | HEK-293T | C290(0.98); C453(1.00) | LDD1577 | [19] |
| LDCM0374 | CL105 | HEK-293T | C290(1.00); C453(1.08); C385(1.05) | LDD1578 | [19] |
| LDCM0375 | CL106 | HEK-293T | C290(0.98); C453(0.77) | LDD1579 | [19] |
| LDCM0376 | CL107 | HEK-293T | C290(1.03); C453(0.99); C385(1.03) | LDD1580 | [19] |
| LDCM0377 | CL108 | HEK-293T | C290(1.04); C453(0.94) | LDD1581 | [19] |
| LDCM0378 | CL109 | HEK-293T | C290(1.11); C453(1.02); C385(1.00) | LDD1582 | [19] |
| LDCM0379 | CL11 | HEK-293T | C290(1.08); C453(0.99); C385(0.99) | LDD1583 | [19] |
| LDCM0380 | CL110 | HEK-293T | C290(0.93); C453(0.75) | LDD1584 | [19] |
| LDCM0381 | CL111 | HEK-293T | C290(0.96); C453(1.08); C385(0.97) | LDD1585 | [19] |
| LDCM0382 | CL112 | HEK-293T | C290(1.04); C453(0.87) | LDD1586 | [19] |
| LDCM0383 | CL113 | HEK-293T | C290(1.01); C453(1.11); C385(1.14) | LDD1587 | [19] |
| LDCM0384 | CL114 | HEK-293T | C290(0.88); C453(0.74) | LDD1588 | [19] |
| LDCM0385 | CL115 | HEK-293T | C290(0.98); C453(1.18); C385(0.96) | LDD1589 | [19] |
| LDCM0386 | CL116 | HEK-293T | C290(0.98); C453(0.92) | LDD1590 | [19] |
| LDCM0387 | CL117 | HEK-293T | C290(1.04); C453(1.18); C385(1.12) | LDD1591 | [19] |
| LDCM0388 | CL118 | HEK-293T | C290(0.99); C453(0.81) | LDD1592 | [19] |
| LDCM0389 | CL119 | HEK-293T | C290(1.00); C453(0.90); C385(1.02) | LDD1593 | [19] |
| LDCM0390 | CL12 | HEK-293T | C290(1.13); C453(0.94); C421(1.26) | LDD1594 | [19] |
| LDCM0391 | CL120 | HEK-293T | C290(1.00); C453(0.95) | LDD1595 | [19] |
| LDCM0392 | CL121 | HEK-293T | C290(1.06); C453(1.09); C385(1.16) | LDD1596 | [19] |
| LDCM0393 | CL122 | HEK-293T | C290(0.99); C453(1.02) | LDD1597 | [19] |
| LDCM0394 | CL123 | HEK-293T | C290(0.95); C453(0.86); C385(0.97) | LDD1598 | [19] |
| LDCM0395 | CL124 | HEK-293T | C290(0.92); C453(0.87) | LDD1599 | [19] |
| LDCM0396 | CL125 | HEK-293T | C290(1.01); C453(1.22); C385(0.94) | LDD1600 | [19] |
| LDCM0397 | CL126 | HEK-293T | C290(0.97); C453(0.76) | LDD1601 | [19] |
| LDCM0398 | CL127 | HEK-293T | C290(1.03); C453(0.94); C385(1.02) | LDD1602 | [19] |
| LDCM0399 | CL128 | HEK-293T | C290(1.02); C453(0.97) | LDD1603 | [19] |
| LDCM0400 | CL13 | HEK-293T | C290(1.00); C453(1.12); C385(1.26) | LDD1604 | [19] |
| LDCM0401 | CL14 | HEK-293T | C290(0.96); C453(0.78) | LDD1605 | [19] |
| LDCM0402 | CL15 | HEK-293T | C290(1.06); C453(1.08); C385(1.04) | LDD1606 | [19] |
| LDCM0403 | CL16 | HEK-293T | C290(1.02); C453(1.04) | LDD1607 | [19] |
| LDCM0404 | CL17 | HEK-293T | C290(1.15); C453(0.98); C385(1.02) | LDD1608 | [19] |
| LDCM0405 | CL18 | HEK-293T | C290(1.07); C453(0.96); C385(1.01); C421(1.17) | LDD1609 | [19] |
| LDCM0406 | CL19 | HEK-293T | C290(1.07); C453(1.06); C385(1.24) | LDD1610 | [19] |
| LDCM0407 | CL2 | HEK-293T | C290(0.94); C453(0.69) | LDD1611 | [19] |
| LDCM0408 | CL20 | HEK-293T | C290(1.08); C453(1.05); C385(1.02) | LDD1612 | [19] |
| LDCM0409 | CL21 | HEK-293T | C290(1.15); C453(0.83); C385(0.78) | LDD1613 | [19] |
| LDCM0410 | CL22 | HEK-293T | C290(0.93); C453(1.02); C385(1.06); C421(0.99) | LDD1614 | [19] |
| LDCM0411 | CL23 | HEK-293T | C290(1.19); C453(1.17); C385(1.04) | LDD1615 | [19] |
| LDCM0412 | CL24 | HEK-293T | C290(1.04); C453(1.07); C421(1.47) | LDD1616 | [19] |
| LDCM0413 | CL25 | HEK-293T | C290(1.08); C453(1.00); C385(1.00) | LDD1617 | [19] |
| LDCM0414 | CL26 | HEK-293T | C290(0.95); C453(0.99) | LDD1618 | [19] |
| LDCM0415 | CL27 | HEK-293T | C290(0.98); C453(0.99); C385(0.91) | LDD1619 | [19] |
| LDCM0416 | CL28 | HEK-293T | C290(0.97); C453(1.00) | LDD1620 | [19] |
| LDCM0417 | CL29 | HEK-293T | C290(1.20); C453(0.97); C385(1.04) | LDD1621 | [19] |
| LDCM0418 | CL3 | HEK-293T | C290(1.03); C453(1.21); C385(1.09) | LDD1622 | [19] |
| LDCM0419 | CL30 | HEK-293T | C290(1.04); C453(1.04); C385(0.99); C421(1.07) | LDD1623 | [19] |
| LDCM0420 | CL31 | HEK-293T | C290(1.02); C453(1.00); C385(1.21) | LDD1624 | [19] |
| LDCM0421 | CL32 | HEK-293T | C290(1.03); C453(0.82); C385(1.00) | LDD1625 | [19] |
| LDCM0422 | CL33 | HEK-293T | C290(1.06); C453(0.90); C385(0.92) | LDD1626 | [19] |
| LDCM0423 | CL34 | HEK-293T | C290(1.00); C453(0.83); C385(0.96); C421(0.82) | LDD1627 | [19] |
| LDCM0424 | CL35 | HEK-293T | C290(1.05); C453(1.13); C385(1.07) | LDD1628 | [19] |
| LDCM0425 | CL36 | HEK-293T | C290(0.99); C453(0.98); C421(0.78) | LDD1629 | [19] |
| LDCM0426 | CL37 | HEK-293T | C290(1.03); C453(0.94); C385(0.93) | LDD1630 | [19] |
| LDCM0428 | CL39 | HEK-293T | C290(0.99); C453(1.00); C385(0.99) | LDD1632 | [19] |
| LDCM0429 | CL4 | HEK-293T | C290(1.04); C453(0.93) | LDD1633 | [19] |
| LDCM0430 | CL40 | HEK-293T | C290(1.05); C453(0.92) | LDD1634 | [19] |
| LDCM0431 | CL41 | HEK-293T | C290(1.02); C453(0.98); C385(0.98) | LDD1635 | [19] |
| LDCM0432 | CL42 | HEK-293T | C290(1.06); C453(1.05); C385(0.97); C421(1.21) | LDD1636 | [19] |
| LDCM0433 | CL43 | HEK-293T | C290(1.03); C453(0.94); C385(1.07) | LDD1637 | [19] |
| LDCM0434 | CL44 | HEK-293T | C290(1.07); C453(0.57); C385(1.00) | LDD1638 | [19] |
| LDCM0435 | CL45 | HEK-293T | C290(1.05); C453(0.84); C385(0.94) | LDD1639 | [19] |
| LDCM0436 | CL46 | HEK-293T | C290(1.02); C453(0.89); C385(0.98); C421(0.86) | LDD1640 | [19] |
| LDCM0437 | CL47 | HEK-293T | C290(1.19); C453(1.16); C385(1.15) | LDD1641 | [19] |
| LDCM0438 | CL48 | HEK-293T | C290(1.04); C453(1.06); C421(0.98) | LDD1642 | [19] |
| LDCM0439 | CL49 | HEK-293T | C290(0.99); C453(1.17); C385(1.13) | LDD1643 | [19] |
| LDCM0440 | CL5 | HEK-293T | C290(1.07); C453(0.99); C385(1.09) | LDD1644 | [19] |
| LDCM0441 | CL50 | HEK-293T | C290(0.94); C453(0.80) | LDD1645 | [19] |
| LDCM0443 | CL52 | HEK-293T | C290(0.98); C453(0.98) | LDD1646 | [19] |
| LDCM0444 | CL53 | HEK-293T | C290(1.10); C453(0.87); C385(0.91) | LDD1647 | [19] |
| LDCM0445 | CL54 | HEK-293T | C290(0.91); C453(0.98); C385(0.97); C421(1.17) | LDD1648 | [19] |
| LDCM0446 | CL55 | HEK-293T | C290(1.07); C453(0.96); C385(1.15) | LDD1649 | [19] |
| LDCM0447 | CL56 | HEK-293T | C290(1.07); C453(0.76); C385(0.93) | LDD1650 | [19] |
| LDCM0448 | CL57 | HEK-293T | C290(1.05); C453(0.90); C385(0.91) | LDD1651 | [19] |
| LDCM0449 | CL58 | HEK-293T | C290(1.10); C453(0.95); C385(1.01); C421(0.95) | LDD1652 | [19] |
| LDCM0450 | CL59 | HEK-293T | C290(1.17); C453(1.23); C385(0.97) | LDD1653 | [19] |
| LDCM0451 | CL6 | HEK-293T | C290(1.02); C453(0.84); C385(0.89); C421(1.24) | LDD1654 | [19] |
| LDCM0452 | CL60 | HEK-293T | C290(1.09); C453(0.93); C421(1.06) | LDD1655 | [19] |
| LDCM0453 | CL61 | HEK-293T | C290(0.98); C453(1.03); C385(1.06) | LDD1656 | [19] |
| LDCM0454 | CL62 | HEK-293T | C290(0.91); C453(0.99) | LDD1657 | [19] |
| LDCM0455 | CL63 | HEK-293T | C290(0.93); C453(1.38); C385(1.04) | LDD1658 | [19] |
| LDCM0456 | CL64 | HEK-293T | C290(1.04); C453(0.94) | LDD1659 | [19] |
| LDCM0457 | CL65 | HEK-293T | C290(1.01); C453(0.99); C385(0.95) | LDD1660 | [19] |
| LDCM0458 | CL66 | HEK-293T | C290(1.01); C453(0.98); C385(1.06); C421(1.36) | LDD1661 | [19] |
| LDCM0459 | CL67 | HEK-293T | C290(1.09); C453(0.99); C385(1.25) | LDD1662 | [19] |
| LDCM0460 | CL68 | HEK-293T | C290(1.07); C453(0.75); C385(0.98) | LDD1663 | [19] |
| LDCM0461 | CL69 | HEK-293T | C290(1.03); C453(1.01); C385(0.94) | LDD1664 | [19] |
| LDCM0462 | CL7 | HEK-293T | C290(1.01); C453(0.92); C385(1.07) | LDD1665 | [19] |
| LDCM0463 | CL70 | HEK-293T | C290(1.06); C453(0.89); C385(1.03); C421(0.88) | LDD1666 | [19] |
| LDCM0464 | CL71 | HEK-293T | C290(1.19); C453(1.22); C385(1.16) | LDD1667 | [19] |
| LDCM0465 | CL72 | HEK-293T | C290(1.07); C453(0.96); C421(0.78) | LDD1668 | [19] |
| LDCM0466 | CL73 | HEK-293T | C290(0.95); C453(1.04); C385(1.03) | LDD1669 | [19] |
| LDCM0467 | CL74 | HEK-293T | C290(0.97); C453(1.07) | LDD1670 | [19] |
| LDCM0469 | CL76 | HEK-293T | C290(1.03); C453(0.95) | LDD1672 | [19] |
| LDCM0470 | CL77 | HEK-293T | C290(1.09); C453(0.81); C385(0.92) | LDD1673 | [19] |
| LDCM0471 | CL78 | HEK-293T | C290(1.04); C453(0.86); C385(0.94); C421(1.34) | LDD1674 | [19] |
| LDCM0472 | CL79 | HEK-293T | C290(1.05); C453(0.90); C385(1.15) | LDD1675 | [19] |
| LDCM0473 | CL8 | HEK-293T | C290(0.91); C453(0.90); C385(0.74) | LDD1676 | [19] |
| LDCM0474 | CL80 | HEK-293T | C290(1.03); C453(0.84); C385(0.99) | LDD1677 | [19] |
| LDCM0475 | CL81 | HEK-293T | C290(1.14); C453(1.01); C385(0.95) | LDD1678 | [19] |
| LDCM0476 | CL82 | HEK-293T | C290(0.97); C453(0.88); C385(1.02); C421(1.07) | LDD1679 | [19] |
| LDCM0477 | CL83 | HEK-293T | C290(1.02); C453(0.89); C385(0.97) | LDD1680 | [19] |
| LDCM0478 | CL84 | HEK-293T | C290(1.12); C453(0.95); C421(0.91) | LDD1681 | [19] |
| LDCM0479 | CL85 | HEK-293T | C290(1.14); C453(1.22); C385(0.95) | LDD1682 | [19] |
| LDCM0480 | CL86 | HEK-293T | C290(1.01); C453(0.82) | LDD1683 | [19] |
| LDCM0481 | CL87 | HEK-293T | C290(0.88); C453(0.97); C385(1.04) | LDD1684 | [19] |
| LDCM0482 | CL88 | HEK-293T | C290(1.02); C453(0.96) | LDD1685 | [19] |
| LDCM0483 | CL89 | HEK-293T | C290(1.14); C453(0.94); C385(0.97) | LDD1686 | [19] |
| LDCM0484 | CL9 | HEK-293T | C290(1.06); C453(0.87); C385(0.81) | LDD1687 | [19] |
| LDCM0485 | CL90 | HEK-293T | C290(0.95); C453(0.80); C385(0.77); C421(1.19) | LDD1688 | [19] |
| LDCM0486 | CL91 | HEK-293T | C290(1.07); C453(0.99); C385(1.06) | LDD1689 | [19] |
| LDCM0487 | CL92 | HEK-293T | C290(0.96); C453(1.01); C385(0.95) | LDD1690 | [19] |
| LDCM0488 | CL93 | HEK-293T | C290(1.05); C453(0.84); C385(0.92) | LDD1691 | [19] |
| LDCM0489 | CL94 | HEK-293T | C290(1.02); C453(0.94); C385(0.97); C421(1.10) | LDD1692 | [19] |
| LDCM0490 | CL95 | HEK-293T | C290(1.03); C453(1.04); C385(0.92) | LDD1693 | [19] |
| LDCM0491 | CL96 | HEK-293T | C290(1.15); C453(0.95); C421(1.17) | LDD1694 | [19] |
| LDCM0492 | CL97 | HEK-293T | C290(1.06); C453(1.10); C385(1.04) | LDD1695 | [19] |
| LDCM0493 | CL98 | HEK-293T | C290(0.99); C453(1.03) | LDD1696 | [19] |
| LDCM0494 | CL99 | HEK-293T | C290(0.98); C453(1.44); C385(0.98) | LDD1697 | [19] |
| LDCM0165 | Compound 9 | HL-60 | N.A. | LDD0420 | [18] |
| LDCM0495 | E2913 | HEK-293T | C290(0.94); C453(0.92); C385(0.98) | LDD1698 | [19] |
| LDCM0625 | F8 | Ramos | C148(0.89); C273(1.66) | LDD2187 | [20] |
| LDCM0572 | Fragment10 | Ramos | C148(0.95) | LDD2189 | [20] |
| LDCM0573 | Fragment11 | Ramos | C148(13.18); C273(15.81) | LDD2190 | [20] |
| LDCM0574 | Fragment12 | Ramos | C148(0.78) | LDD2191 | [20] |
| LDCM0575 | Fragment13 | Ramos | C148(0.93); C273(1.04) | LDD2192 | [20] |
| LDCM0576 | Fragment14 | Ramos | C273(0.76) | LDD2193 | [20] |
| LDCM0579 | Fragment20 | Ramos | C273(0.82) | LDD2194 | [20] |
| LDCM0580 | Fragment21 | Ramos | C273(0.89) | LDD2195 | [20] |
| LDCM0582 | Fragment23 | Ramos | C148(1.03); C273(0.98) | LDD2196 | [20] |
| LDCM0578 | Fragment27 | Ramos | C273(2.02) | LDD2197 | [20] |
| LDCM0586 | Fragment28 | Ramos | C148(0.42); C273(1.74) | LDD2198 | [20] |
| LDCM0588 | Fragment30 | Ramos | C148(1.04); C273(0.87) | LDD2199 | [20] |
| LDCM0589 | Fragment31 | Ramos | C148(0.75); C273(0.81) | LDD2200 | [20] |
| LDCM0590 | Fragment32 | Ramos | C148(0.91) | LDD2201 | [20] |
| LDCM0468 | Fragment33 | HEK-293T | C290(0.95); C453(0.92); C385(1.03) | LDD1671 | [19] |
| LDCM0596 | Fragment38 | Ramos | C148(1.54) | LDD2203 | [20] |
| LDCM0566 | Fragment4 | Ramos | C273(4.20) | LDD2184 | [20] |
| LDCM0427 | Fragment51 | HEK-293T | C290(0.98); C453(1.06) | LDD1631 | [19] |
| LDCM0614 | Fragment56 | Ramos | C148(1.62) | LDD2205 | [20] |
| LDCM0571 | Fragment9 | Ramos | C148(1.06) | LDD2188 | [20] |
| LDCM0022 | KB02 | HEK-293T | C385(0.97); C290(0.91); C421(1.35); C453(1.15) | LDD1492 | [19] |
| LDCM0023 | KB03 | Jurkat | C290(20.00) | LDD0315 | [21] |
| LDCM0024 | KB05 | COLO792 | C273(4.05) | LDD3310 | [3] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C7(0.66) | LDD2206 | [22] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C7(0.76) | LDD2207 | [22] |
| LDCM0014 | Panhematin | HEK-293T | 3.00 | LDD0062 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
References
