Details of the Target
General Information of Target
| Target ID | LDTP04970 | |||||
|---|---|---|---|---|---|---|
| Target Name | Small ribosomal subunit protein uS11 (RPS14) | |||||
| Gene Name | RPS14 | |||||
| Gene ID | 6208 | |||||
| Synonyms |
Small ribosomal subunit protein uS11; 40S ribosomal protein S14 |
|||||
| 3D Structure | ||||||
| Sequence |
MAPRKGKEKKEEQVISLGPQVAEGENVFGVCHIFASFNDTFVHVTDLSGKETICRVTGGM
KVKADRDESSPYAAMLAAQDVAQRCKELGITALHIKLRATGGNRTKTPGPGAQSALRALA RSGMKIGRIEDVTPIPSDSTRRKGGRRGRRL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Universal ribosomal protein uS11 family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Component of the small ribosomal subunit. The ribosome is a large ribonucleoprotein complex responsible for the synthesis of proteins in the cell. Part of the small subunit (SSU) processome, first precursor of the small eukaryotic ribosomal subunit. During the assembly of the SSU processome in the nucleolus, many ribosome biogenesis factors, an RNA chaperone and ribosomal proteins associate with the nascent pre-rRNA and work in concert to generate RNA folding, modifications, rearrangements and cleavage as well as targeted degradation of pre-ribosomal RNA by the RNA exosome.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
8.89 | LDD0402 | [1] | |
|
P1 Probe Info |
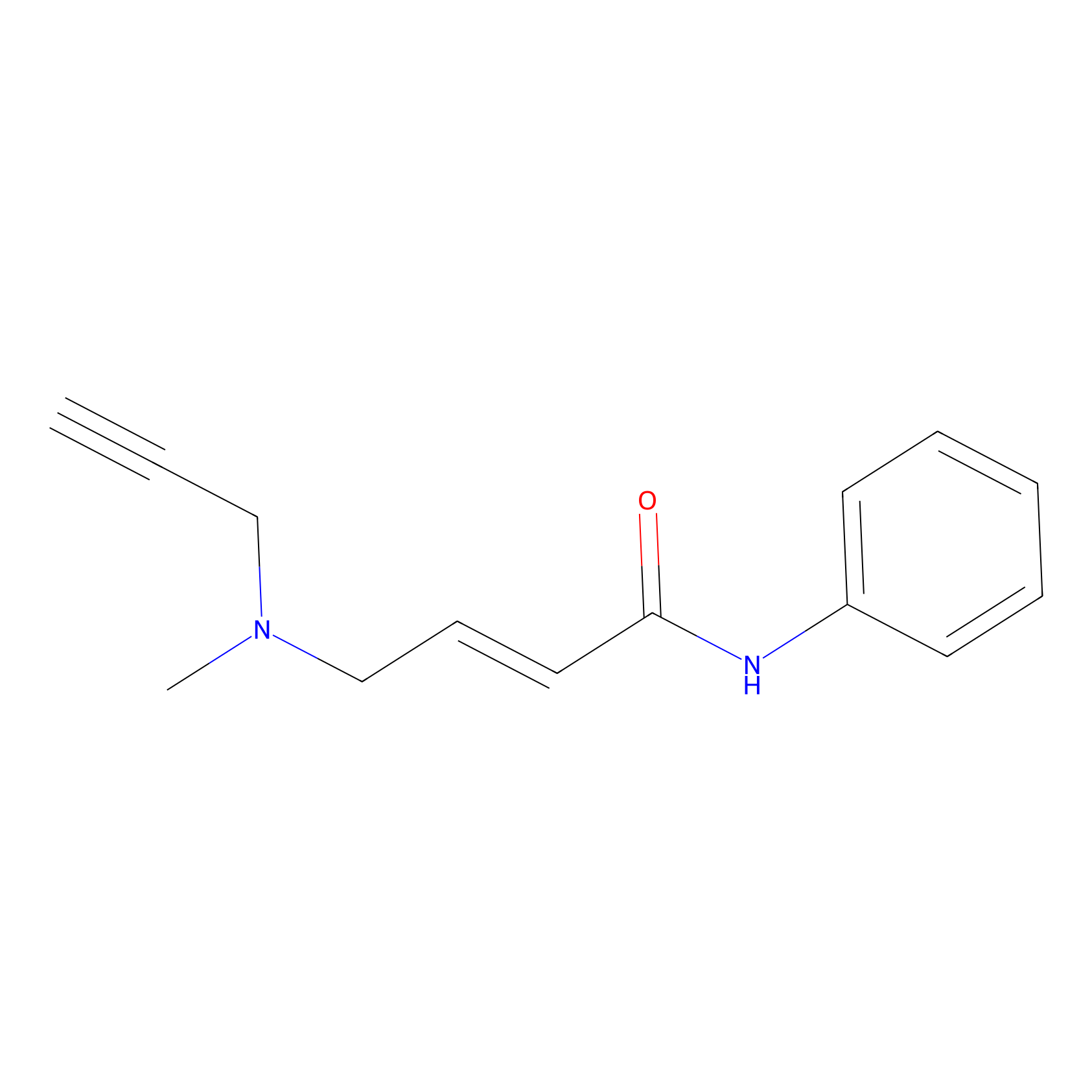 |
10.00 | LDD0448 | [2] | |
|
P2 Probe Info |
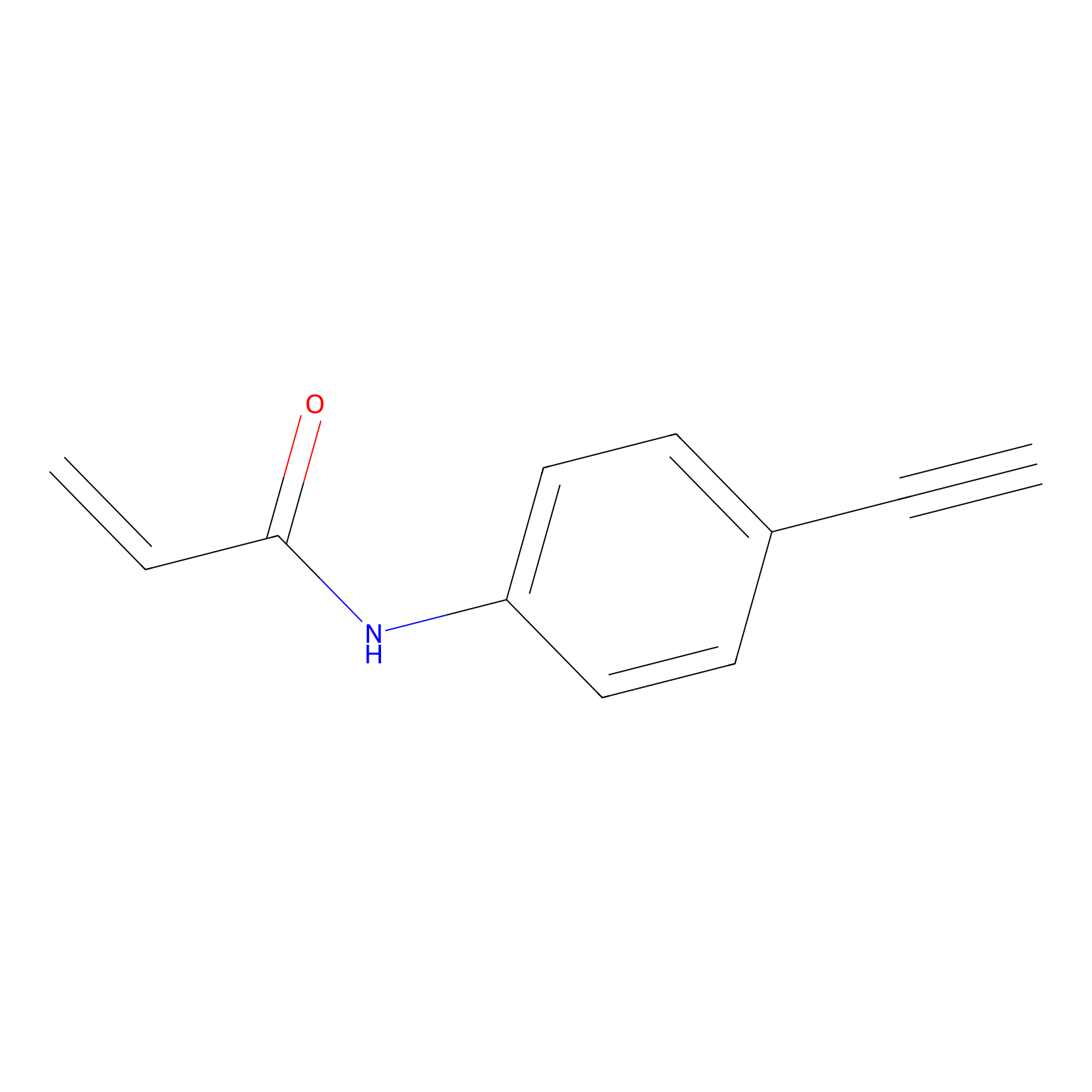 |
10.00 | LDD0449 | [2] | |
|
P3 Probe Info |
 |
10.00 | LDD0450 | [2] | |
|
P8 Probe Info |
 |
5.73 | LDD0451 | [2] | |
|
A-EBA Probe Info |
 |
4.37 | LDD0215 | [3] | |
|
CY4 Probe Info |
 |
5.96 | LDD0244 | [4] | |
|
N1 Probe Info |
 |
5.38 | LDD0242 | [4] | |
|
C-Sul Probe Info |
 |
2.55 | LDD0066 | [5] | |
|
TH211 Probe Info |
 |
Y72(13.68) | LDD0257 | [6] | |
|
TH214 Probe Info |
 |
Y72(20.00) | LDD0258 | [6] | |
|
TH216 Probe Info |
 |
Y72(20.00) | LDD0259 | [6] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [7] | |
|
STPyne Probe Info |
 |
K106(10.00); K125(0.89); K61(8.08); K63(6.34) | LDD0277 | [8] | |
|
AZ-9 Probe Info |
 |
E87(0.83) | LDD2208 | [9] | |
|
ONAyne Probe Info |
 |
K106(0.00); K125(0.00) | LDD0273 | [8] | |
|
OPA-S-S-alkyne Probe Info |
 |
K106(5.29); K125(5.61) | LDD3494 | [10] | |
|
Probe 1 Probe Info |
 |
Y72(10.72) | LDD3495 | [11] | |
|
DBIA Probe Info |
 |
C31(2.88) | LDD3491 | [12] | |
|
BTD Probe Info |
 |
C85(0.09) | LDD2124 | [13] | |
|
HHS-482 Probe Info |
 |
Y72(0.78) | LDD0285 | [14] | |
|
HHS-475 Probe Info |
 |
Y72(4.05) | LDD0264 | [15] | |
|
ATP probe Probe Info |
 |
K63(0.00); K106(0.00); K96(0.00); K86(0.00) | LDD0199 | [16] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C85(0.00); C31(0.00) | LDD0038 | [17] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [18] | |
|
IPIAA_H Probe Info |
 |
N.A. | LDD0030 | [19] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [19] | |
|
Lodoacetamide azide Probe Info |
 |
C85(0.00); C31(0.00) | LDD0037 | [17] | |
|
2PCA Probe Info |
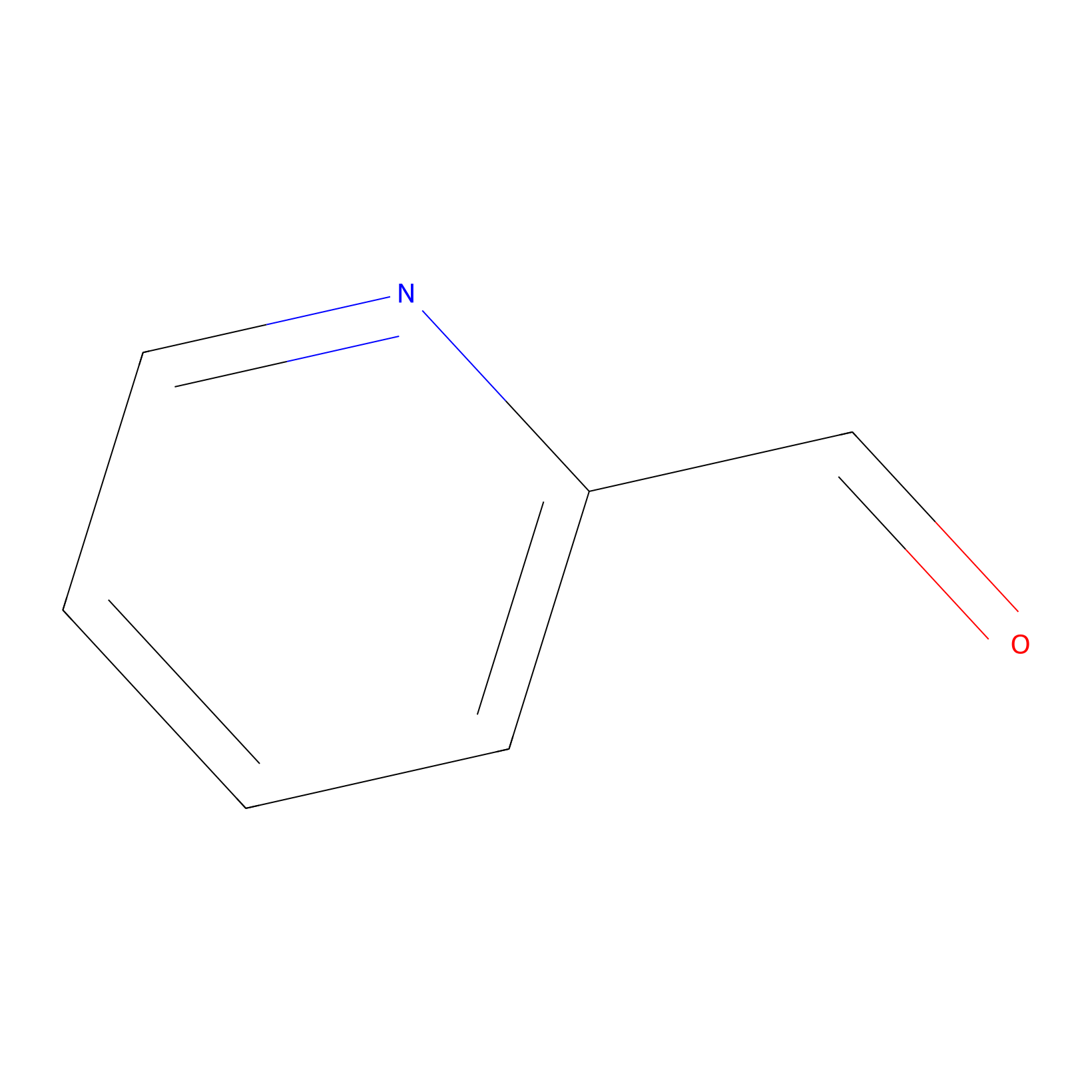 |
N.A. | LDD0034 | [20] | |
|
ATP probe Probe Info |
 |
K106(0.00); K96(0.00) | LDD0035 | [21] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [22] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [22] | |
|
NAIA_4 Probe Info |
 |
C31(0.00); C54(0.00); C85(0.00) | LDD2226 | [23] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [22] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [24] | |
|
SF Probe Info |
 |
Y72(0.00); K106(0.00) | LDD0028 | [25] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [26] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [26] | |
|
Acrolein Probe Info |
 |
H43(0.00); H94(0.00) | LDD0217 | [27] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [27] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [27] | |
|
AOyne Probe Info |
 |
13.40 | LDD0443 | [28] | |
|
NAIA_5 Probe Info |
 |
C85(0.00); C54(0.00); C31(0.00) | LDD2223 | [23] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [29] | |
|
IMP2070 Probe Info |
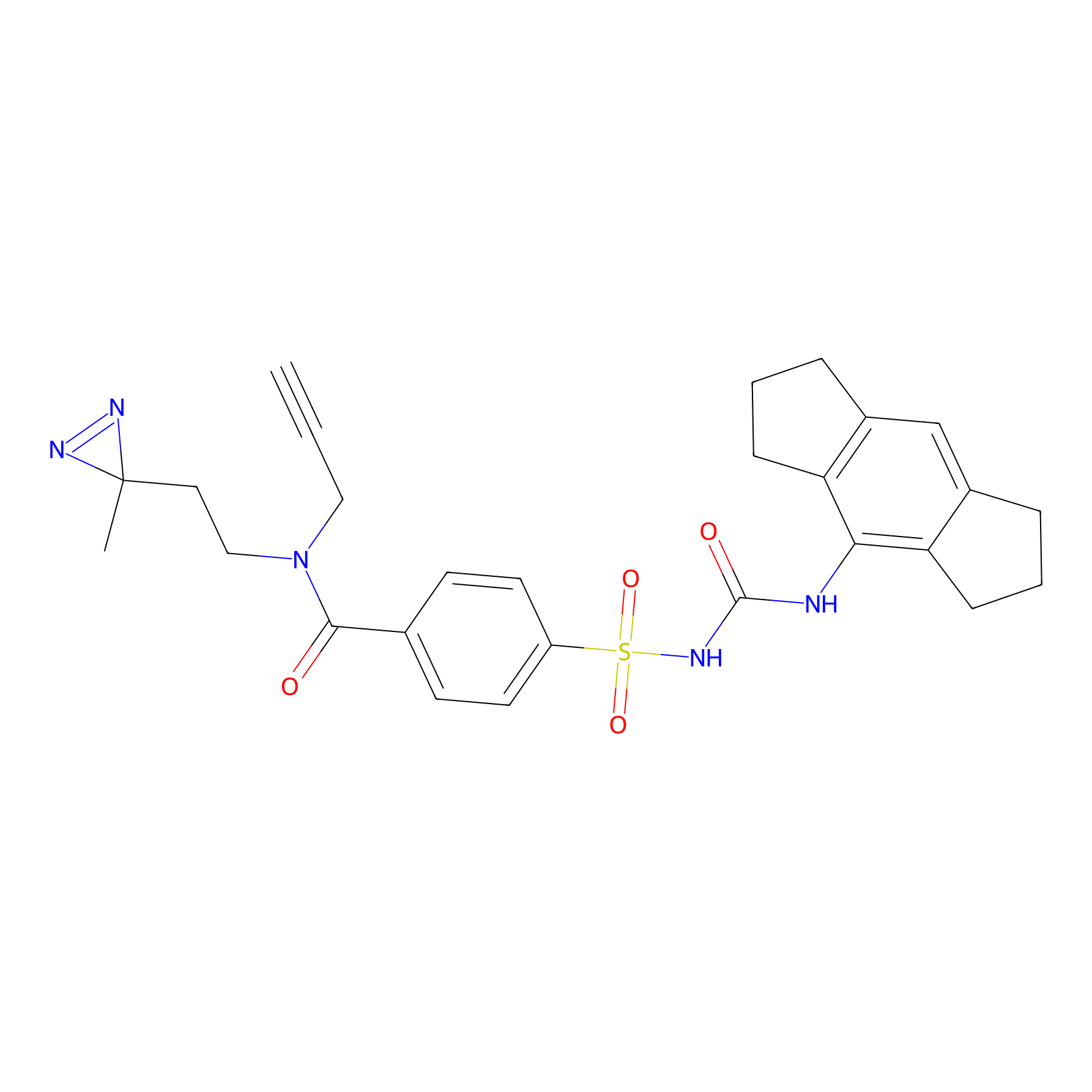 |
1.63 | LDD0368 | [30] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C31(1.08) | LDD1507 | [31] |
| LDCM0215 | AC10 | HEK-293T | C31(1.03) | LDD1508 | [31] |
| LDCM0226 | AC11 | HEK-293T | C31(0.89); C85(1.09) | LDD1509 | [31] |
| LDCM0237 | AC12 | HEK-293T | C31(0.99); C85(1.11) | LDD1510 | [31] |
| LDCM0259 | AC14 | HEK-293T | C31(1.16); C85(1.35) | LDD1512 | [31] |
| LDCM0270 | AC15 | HEK-293T | C31(0.94); C85(1.05) | LDD1513 | [31] |
| LDCM0276 | AC17 | HEK-293T | C31(0.96) | LDD1515 | [31] |
| LDCM0277 | AC18 | HEK-293T | C31(0.91) | LDD1516 | [31] |
| LDCM0278 | AC19 | HEK-293T | C31(0.75); C85(1.18) | LDD1517 | [31] |
| LDCM0279 | AC2 | HEK-293T | C31(1.03) | LDD1518 | [31] |
| LDCM0280 | AC20 | HEK-293T | C31(1.10); C85(1.15) | LDD1519 | [31] |
| LDCM0281 | AC21 | HEK-293T | C31(0.88); C85(1.14) | LDD1520 | [31] |
| LDCM0282 | AC22 | HEK-293T | C31(1.16); C85(1.17) | LDD1521 | [31] |
| LDCM0283 | AC23 | HEK-293T | C31(0.96); C85(1.01) | LDD1522 | [31] |
| LDCM0284 | AC24 | HEK-293T | C31(1.08) | LDD1523 | [31] |
| LDCM0285 | AC25 | HEK-293T | C31(1.10) | LDD1524 | [31] |
| LDCM0286 | AC26 | HEK-293T | C31(0.96) | LDD1525 | [31] |
| LDCM0287 | AC27 | HEK-293T | C31(0.93); C85(1.06) | LDD1526 | [31] |
| LDCM0288 | AC28 | HEK-293T | C31(1.02); C85(1.07) | LDD1527 | [31] |
| LDCM0289 | AC29 | HEK-293T | C31(0.95); C85(1.23) | LDD1528 | [31] |
| LDCM0290 | AC3 | HEK-293T | C31(0.93); C85(0.90) | LDD1529 | [31] |
| LDCM0291 | AC30 | HEK-293T | C31(1.02); C85(1.31) | LDD1530 | [31] |
| LDCM0292 | AC31 | HEK-293T | C31(0.95); C85(0.96) | LDD1531 | [31] |
| LDCM0293 | AC32 | HEK-293T | C31(1.11) | LDD1532 | [31] |
| LDCM0294 | AC33 | HEK-293T | C31(1.01) | LDD1533 | [31] |
| LDCM0295 | AC34 | HEK-293T | C31(0.95) | LDD1534 | [31] |
| LDCM0296 | AC35 | HEK-293T | C31(0.88); C85(1.07) | LDD1535 | [31] |
| LDCM0297 | AC36 | HEK-293T | C31(1.09); C85(1.23) | LDD1536 | [31] |
| LDCM0298 | AC37 | HEK-293T | C31(1.12); C85(1.38) | LDD1537 | [31] |
| LDCM0299 | AC38 | HEK-293T | C31(1.05); C85(1.46) | LDD1538 | [31] |
| LDCM0300 | AC39 | HEK-293T | C31(1.01); C85(1.11) | LDD1539 | [31] |
| LDCM0301 | AC4 | HEK-293T | C31(1.04); C85(1.01) | LDD1540 | [31] |
| LDCM0302 | AC40 | HEK-293T | C31(1.12) | LDD1541 | [31] |
| LDCM0303 | AC41 | HEK-293T | C31(1.00) | LDD1542 | [31] |
| LDCM0304 | AC42 | HEK-293T | C31(0.93) | LDD1543 | [31] |
| LDCM0305 | AC43 | HEK-293T | C31(0.87); C85(1.07) | LDD1544 | [31] |
| LDCM0306 | AC44 | HEK-293T | C31(0.97); C85(1.13) | LDD1545 | [31] |
| LDCM0307 | AC45 | HEK-293T | C31(0.97); C85(1.35) | LDD1546 | [31] |
| LDCM0308 | AC46 | HEK-293T | C31(1.08); C85(1.38) | LDD1547 | [31] |
| LDCM0309 | AC47 | HEK-293T | C31(0.92); C85(1.10) | LDD1548 | [31] |
| LDCM0310 | AC48 | HEK-293T | C31(1.10) | LDD1549 | [31] |
| LDCM0311 | AC49 | HEK-293T | C31(0.95) | LDD1550 | [31] |
| LDCM0312 | AC5 | HEK-293T | C31(0.92); C85(1.08) | LDD1551 | [31] |
| LDCM0313 | AC50 | HEK-293T | C31(0.97) | LDD1552 | [31] |
| LDCM0314 | AC51 | HEK-293T | C31(0.85); C85(1.17) | LDD1553 | [31] |
| LDCM0315 | AC52 | HEK-293T | C31(0.98); C85(1.12) | LDD1554 | [31] |
| LDCM0316 | AC53 | HEK-293T | C31(0.93); C85(1.31) | LDD1555 | [31] |
| LDCM0317 | AC54 | HEK-293T | C31(0.99); C85(1.36) | LDD1556 | [31] |
| LDCM0318 | AC55 | HEK-293T | C31(0.85); C85(1.10) | LDD1557 | [31] |
| LDCM0319 | AC56 | HEK-293T | C31(0.92) | LDD1558 | [31] |
| LDCM0320 | AC57 | HEK-293T | C31(1.01) | LDD1559 | [31] |
| LDCM0321 | AC58 | HEK-293T | C31(0.93) | LDD1560 | [31] |
| LDCM0322 | AC59 | HEK-293T | C31(0.85); C85(0.89) | LDD1561 | [31] |
| LDCM0323 | AC6 | HEK-293T | C31(0.99); C85(1.01) | LDD1562 | [31] |
| LDCM0324 | AC60 | HEK-293T | C31(1.05); C85(0.91) | LDD1563 | [31] |
| LDCM0325 | AC61 | HEK-293T | C31(0.91); C85(0.96) | LDD1564 | [31] |
| LDCM0326 | AC62 | HEK-293T | C31(0.97); C85(1.16) | LDD1565 | [31] |
| LDCM0327 | AC63 | HEK-293T | C31(0.91); C85(0.83) | LDD1566 | [31] |
| LDCM0328 | AC64 | HEK-293T | C31(1.00) | LDD1567 | [31] |
| LDCM0334 | AC7 | HEK-293T | C31(0.92); C85(0.81) | LDD1568 | [31] |
| LDCM0345 | AC8 | HEK-293T | C31(1.06) | LDD1569 | [31] |
| LDCM0248 | AKOS034007472 | HEK-293T | C31(0.86); C85(1.23) | LDD1511 | [31] |
| LDCM0356 | AKOS034007680 | HEK-293T | C31(0.97) | LDD1570 | [31] |
| LDCM0275 | AKOS034007705 | HEK-293T | C31(0.99) | LDD1514 | [31] |
| LDCM0108 | Chloroacetamide | HeLa | H43(0.00); H94(0.00); C85(0.00); H32(0.00) | LDD0222 | [27] |
| LDCM0632 | CL-Sc | Hep-G2 | C85(0.71) | LDD2227 | [23] |
| LDCM0367 | CL1 | HEK-293T | C31(1.06); C85(0.76) | LDD1571 | [31] |
| LDCM0368 | CL10 | HEK-293T | C31(1.19); C85(0.71) | LDD1572 | [31] |
| LDCM0369 | CL100 | HEK-293T | C31(1.04) | LDD1573 | [31] |
| LDCM0370 | CL101 | HEK-293T | C31(0.92); C85(0.94) | LDD1574 | [31] |
| LDCM0371 | CL102 | HEK-293T | C31(0.96); C85(0.98) | LDD1575 | [31] |
| LDCM0372 | CL103 | HEK-293T | C31(1.04); C85(1.26) | LDD1576 | [31] |
| LDCM0373 | CL104 | HEK-293T | C31(0.97) | LDD1577 | [31] |
| LDCM0374 | CL105 | HEK-293T | C31(0.96); C85(0.94) | LDD1578 | [31] |
| LDCM0375 | CL106 | HEK-293T | C31(0.96); C85(1.11) | LDD1579 | [31] |
| LDCM0376 | CL107 | HEK-293T | C31(0.96); C85(1.21) | LDD1580 | [31] |
| LDCM0377 | CL108 | HEK-293T | C31(0.95) | LDD1581 | [31] |
| LDCM0378 | CL109 | HEK-293T | C31(1.03); C85(1.01) | LDD1582 | [31] |
| LDCM0379 | CL11 | HEK-293T | C31(0.97); C85(0.70) | LDD1583 | [31] |
| LDCM0380 | CL110 | HEK-293T | C31(0.93); C85(0.93) | LDD1584 | [31] |
| LDCM0381 | CL111 | HEK-293T | C31(1.00); C85(1.02) | LDD1585 | [31] |
| LDCM0382 | CL112 | HEK-293T | C31(0.99) | LDD1586 | [31] |
| LDCM0383 | CL113 | HEK-293T | C31(1.13); C85(0.99) | LDD1587 | [31] |
| LDCM0384 | CL114 | HEK-293T | C31(1.06); C85(0.90) | LDD1588 | [31] |
| LDCM0385 | CL115 | HEK-293T | C31(1.07); C85(1.20) | LDD1589 | [31] |
| LDCM0386 | CL116 | HEK-293T | C31(1.05) | LDD1590 | [31] |
| LDCM0387 | CL117 | HEK-293T | C31(0.95); C85(1.04) | LDD1591 | [31] |
| LDCM0388 | CL118 | HEK-293T | C31(0.92); C85(1.13) | LDD1592 | [31] |
| LDCM0389 | CL119 | HEK-293T | C31(1.04); C85(1.23) | LDD1593 | [31] |
| LDCM0390 | CL12 | HEK-293T | C31(1.18) | LDD1594 | [31] |
| LDCM0391 | CL120 | HEK-293T | C31(0.99) | LDD1595 | [31] |
| LDCM0392 | CL121 | HEK-293T | C31(0.88); C85(1.17) | LDD1596 | [31] |
| LDCM0393 | CL122 | HEK-293T | C31(0.92); C85(1.02) | LDD1597 | [31] |
| LDCM0394 | CL123 | HEK-293T | C31(0.91); C85(1.01) | LDD1598 | [31] |
| LDCM0395 | CL124 | HEK-293T | C31(0.95) | LDD1599 | [31] |
| LDCM0396 | CL125 | HEK-293T | C31(0.77); C85(1.04) | LDD1600 | [31] |
| LDCM0397 | CL126 | HEK-293T | C31(0.84); C85(0.79) | LDD1601 | [31] |
| LDCM0398 | CL127 | HEK-293T | C31(0.75); C85(0.84) | LDD1602 | [31] |
| LDCM0399 | CL128 | HEK-293T | C31(0.96) | LDD1603 | [31] |
| LDCM0400 | CL13 | HEK-293T | C31(0.92); C85(0.93) | LDD1604 | [31] |
| LDCM0401 | CL14 | HEK-293T | C31(0.95); C85(1.02) | LDD1605 | [31] |
| LDCM0402 | CL15 | HEK-293T | C31(0.91); C85(1.14) | LDD1606 | [31] |
| LDCM0403 | CL16 | HEK-293T | C31(1.01) | LDD1607 | [31] |
| LDCM0404 | CL17 | HEK-293T | C31(0.94) | LDD1608 | [31] |
| LDCM0405 | CL18 | HEK-293T | C31(0.90) | LDD1609 | [31] |
| LDCM0406 | CL19 | HEK-293T | C31(0.80); C85(1.20) | LDD1610 | [31] |
| LDCM0407 | CL2 | HEK-293T | C31(0.99); C85(1.01) | LDD1611 | [31] |
| LDCM0408 | CL20 | HEK-293T | C31(0.92); C85(0.94) | LDD1612 | [31] |
| LDCM0409 | CL21 | HEK-293T | C31(0.83); C85(1.03) | LDD1613 | [31] |
| LDCM0410 | CL22 | HEK-293T | C31(1.03); C85(1.11) | LDD1614 | [31] |
| LDCM0411 | CL23 | HEK-293T | C31(0.93); C85(1.01) | LDD1615 | [31] |
| LDCM0412 | CL24 | HEK-293T | C31(1.01) | LDD1616 | [31] |
| LDCM0413 | CL25 | HEK-293T | C31(0.93); C85(0.82) | LDD1617 | [31] |
| LDCM0414 | CL26 | HEK-293T | C31(1.05); C85(1.17) | LDD1618 | [31] |
| LDCM0415 | CL27 | HEK-293T | C31(0.91); C85(1.22) | LDD1619 | [31] |
| LDCM0416 | CL28 | HEK-293T | C31(0.93) | LDD1620 | [31] |
| LDCM0417 | CL29 | HEK-293T | C31(0.96) | LDD1621 | [31] |
| LDCM0418 | CL3 | HEK-293T | C31(1.00); C85(0.95) | LDD1622 | [31] |
| LDCM0419 | CL30 | HEK-293T | C31(0.89) | LDD1623 | [31] |
| LDCM0420 | CL31 | HEK-293T | C31(0.73); C85(1.07) | LDD1624 | [31] |
| LDCM0421 | CL32 | HEK-293T | C31(0.92); C85(1.09) | LDD1625 | [31] |
| LDCM0422 | CL33 | HEK-293T | C31(0.74); C85(0.85) | LDD1626 | [31] |
| LDCM0423 | CL34 | HEK-293T | C31(0.96); C85(1.07) | LDD1627 | [31] |
| LDCM0424 | CL35 | HEK-293T | C31(0.94); C85(0.91) | LDD1628 | [31] |
| LDCM0425 | CL36 | HEK-293T | C31(1.06) | LDD1629 | [31] |
| LDCM0426 | CL37 | HEK-293T | C31(0.99); C85(0.77) | LDD1630 | [31] |
| LDCM0428 | CL39 | HEK-293T | C31(0.96); C85(1.11) | LDD1632 | [31] |
| LDCM0429 | CL4 | HEK-293T | C31(0.88) | LDD1633 | [31] |
| LDCM0430 | CL40 | HEK-293T | C31(1.07) | LDD1634 | [31] |
| LDCM0431 | CL41 | HEK-293T | C31(1.00) | LDD1635 | [31] |
| LDCM0432 | CL42 | HEK-293T | C31(0.86) | LDD1636 | [31] |
| LDCM0433 | CL43 | HEK-293T | C31(0.90); C85(1.01) | LDD1637 | [31] |
| LDCM0434 | CL44 | HEK-293T | C31(0.92); C85(0.94) | LDD1638 | [31] |
| LDCM0435 | CL45 | HEK-293T | C31(0.83); C85(0.96) | LDD1639 | [31] |
| LDCM0436 | CL46 | HEK-293T | C31(1.02); C85(1.03) | LDD1640 | [31] |
| LDCM0437 | CL47 | HEK-293T | C31(0.91); C85(0.92) | LDD1641 | [31] |
| LDCM0438 | CL48 | HEK-293T | C31(0.98) | LDD1642 | [31] |
| LDCM0439 | CL49 | HEK-293T | C31(0.88); C85(0.88) | LDD1643 | [31] |
| LDCM0440 | CL5 | HEK-293T | C31(0.87) | LDD1644 | [31] |
| LDCM0441 | CL50 | HEK-293T | C31(1.07); C85(1.04) | LDD1645 | [31] |
| LDCM0443 | CL52 | HEK-293T | C31(1.08) | LDD1646 | [31] |
| LDCM0444 | CL53 | HEK-293T | C31(1.00) | LDD1647 | [31] |
| LDCM0445 | CL54 | HEK-293T | C31(0.82) | LDD1648 | [31] |
| LDCM0446 | CL55 | HEK-293T | C31(0.77); C85(1.07) | LDD1649 | [31] |
| LDCM0447 | CL56 | HEK-293T | C31(1.02); C85(0.98) | LDD1650 | [31] |
| LDCM0448 | CL57 | HEK-293T | C31(0.83); C85(1.06) | LDD1651 | [31] |
| LDCM0449 | CL58 | HEK-293T | C31(0.98); C85(1.25) | LDD1652 | [31] |
| LDCM0450 | CL59 | HEK-293T | C31(0.86); C85(1.02) | LDD1653 | [31] |
| LDCM0451 | CL6 | HEK-293T | C31(0.85) | LDD1654 | [31] |
| LDCM0452 | CL60 | HEK-293T | C31(1.01) | LDD1655 | [31] |
| LDCM0453 | CL61 | HEK-293T | C31(1.08); C85(0.71) | LDD1656 | [31] |
| LDCM0454 | CL62 | HEK-293T | C31(1.09); C85(1.02) | LDD1657 | [31] |
| LDCM0455 | CL63 | HEK-293T | C31(0.98); C85(1.15) | LDD1658 | [31] |
| LDCM0456 | CL64 | HEK-293T | C31(1.02) | LDD1659 | [31] |
| LDCM0457 | CL65 | HEK-293T | C31(1.00) | LDD1660 | [31] |
| LDCM0458 | CL66 | HEK-293T | C31(0.85) | LDD1661 | [31] |
| LDCM0459 | CL67 | HEK-293T | C31(0.84); C85(1.00) | LDD1662 | [31] |
| LDCM0460 | CL68 | HEK-293T | C31(0.90); C85(0.92) | LDD1663 | [31] |
| LDCM0461 | CL69 | HEK-293T | C31(0.84); C85(1.08) | LDD1664 | [31] |
| LDCM0462 | CL7 | HEK-293T | C31(0.77); C85(0.82) | LDD1665 | [31] |
| LDCM0463 | CL70 | HEK-293T | C31(1.01); C85(1.07) | LDD1666 | [31] |
| LDCM0464 | CL71 | HEK-293T | C31(0.91); C85(1.03) | LDD1667 | [31] |
| LDCM0465 | CL72 | HEK-293T | C31(1.11) | LDD1668 | [31] |
| LDCM0466 | CL73 | HEK-293T | C31(0.95); C85(0.91) | LDD1669 | [31] |
| LDCM0467 | CL74 | HEK-293T | C31(0.92); C85(1.05) | LDD1670 | [31] |
| LDCM0469 | CL76 | HEK-293T | C31(1.00) | LDD1672 | [31] |
| LDCM0470 | CL77 | HEK-293T | C31(0.95) | LDD1673 | [31] |
| LDCM0471 | CL78 | HEK-293T | C31(0.89) | LDD1674 | [31] |
| LDCM0472 | CL79 | HEK-293T | C31(0.84); C85(1.20) | LDD1675 | [31] |
| LDCM0473 | CL8 | HEK-293T | C31(0.88); C85(0.53) | LDD1676 | [31] |
| LDCM0474 | CL80 | HEK-293T | C31(0.91); C85(1.02) | LDD1677 | [31] |
| LDCM0475 | CL81 | HEK-293T | C31(0.79); C85(1.26) | LDD1678 | [31] |
| LDCM0476 | CL82 | HEK-293T | C31(0.98); C85(1.28) | LDD1679 | [31] |
| LDCM0477 | CL83 | HEK-293T | C31(0.86); C85(0.92) | LDD1680 | [31] |
| LDCM0478 | CL84 | HEK-293T | C31(0.93) | LDD1681 | [31] |
| LDCM0479 | CL85 | HEK-293T | C31(0.96); C85(0.92) | LDD1682 | [31] |
| LDCM0480 | CL86 | HEK-293T | C31(0.92); C85(0.82) | LDD1683 | [31] |
| LDCM0481 | CL87 | HEK-293T | C31(0.80); C85(0.89) | LDD1684 | [31] |
| LDCM0482 | CL88 | HEK-293T | C31(0.85) | LDD1685 | [31] |
| LDCM0483 | CL89 | HEK-293T | C31(0.79) | LDD1686 | [31] |
| LDCM0484 | CL9 | HEK-293T | C31(0.80); C85(0.90) | LDD1687 | [31] |
| LDCM0485 | CL90 | HEK-293T | C31(0.76) | LDD1688 | [31] |
| LDCM0486 | CL91 | HEK-293T | C31(0.72); C85(0.83) | LDD1689 | [31] |
| LDCM0487 | CL92 | HEK-293T | C31(0.84); C85(0.88) | LDD1690 | [31] |
| LDCM0488 | CL93 | HEK-293T | C31(0.81); C85(1.03) | LDD1691 | [31] |
| LDCM0489 | CL94 | HEK-293T | C31(1.00); C85(1.09) | LDD1692 | [31] |
| LDCM0490 | CL95 | HEK-293T | C31(0.78); C85(0.74) | LDD1693 | [31] |
| LDCM0491 | CL96 | HEK-293T | C31(0.93) | LDD1694 | [31] |
| LDCM0492 | CL97 | HEK-293T | C31(0.89); C85(0.79) | LDD1695 | [31] |
| LDCM0493 | CL98 | HEK-293T | C31(1.14); C85(0.85) | LDD1696 | [31] |
| LDCM0494 | CL99 | HEK-293T | C31(0.97); C85(0.85) | LDD1697 | [31] |
| LDCM0495 | E2913 | HEK-293T | C31(1.02); C85(1.19) | LDD1698 | [31] |
| LDCM0625 | F8 | Ramos | C85(2.80) | LDD2187 | [32] |
| LDCM0572 | Fragment10 | Ramos | C85(0.40) | LDD2189 | [32] |
| LDCM0573 | Fragment11 | Ramos | C85(0.00) | LDD2190 | [32] |
| LDCM0574 | Fragment12 | Ramos | C85(0.70) | LDD2191 | [32] |
| LDCM0575 | Fragment13 | Ramos | C85(0.32) | LDD2192 | [32] |
| LDCM0576 | Fragment14 | Ramos | C85(0.49) | LDD2193 | [32] |
| LDCM0579 | Fragment20 | Ramos | C85(1.02) | LDD2194 | [32] |
| LDCM0580 | Fragment21 | Ramos | C85(0.36) | LDD2195 | [32] |
| LDCM0582 | Fragment23 | Ramos | C85(1.13) | LDD2196 | [32] |
| LDCM0578 | Fragment27 | Ramos | C85(2.15) | LDD2197 | [32] |
| LDCM0586 | Fragment28 | Ramos | C85(1.12) | LDD2198 | [32] |
| LDCM0588 | Fragment30 | Ramos | C85(0.51) | LDD2199 | [32] |
| LDCM0589 | Fragment31 | Ramos | C85(0.55) | LDD2200 | [32] |
| LDCM0590 | Fragment32 | Ramos | C85(0.85) | LDD2201 | [32] |
| LDCM0468 | Fragment33 | HEK-293T | C31(0.93); C85(1.20) | LDD1671 | [31] |
| LDCM0596 | Fragment38 | Ramos | C85(0.60) | LDD2203 | [32] |
| LDCM0566 | Fragment4 | Ramos | C85(1.83) | LDD2184 | [32] |
| LDCM0427 | Fragment51 | HEK-293T | C31(1.01); C85(0.95) | LDD1631 | [31] |
| LDCM0610 | Fragment52 | Ramos | C85(0.72) | LDD2204 | [32] |
| LDCM0614 | Fragment56 | Ramos | C85(0.18) | LDD2205 | [32] |
| LDCM0569 | Fragment7 | Ramos | C85(0.45) | LDD2186 | [32] |
| LDCM0571 | Fragment9 | Ramos | C85(0.28) | LDD2188 | [32] |
| LDCM0116 | HHS-0101 | DM93 | Y72(4.05) | LDD0264 | [15] |
| LDCM0117 | HHS-0201 | DM93 | Y72(10.44) | LDD0265 | [15] |
| LDCM0118 | HHS-0301 | DM93 | Y72(10.35) | LDD0266 | [15] |
| LDCM0119 | HHS-0401 | DM93 | Y72(0.93) | LDD0267 | [15] |
| LDCM0120 | HHS-0701 | DM93 | Y72(0.47) | LDD0268 | [15] |
| LDCM0107 | IAA | HeLa | H43(0.00); H94(0.00) | LDD0221 | [27] |
| LDCM0123 | JWB131 | DM93 | Y72(0.78) | LDD0285 | [14] |
| LDCM0124 | JWB142 | DM93 | Y72(0.40) | LDD0286 | [14] |
| LDCM0125 | JWB146 | DM93 | Y72(1.05) | LDD0287 | [14] |
| LDCM0126 | JWB150 | DM93 | Y72(2.75) | LDD0288 | [14] |
| LDCM0127 | JWB152 | DM93 | Y72(1.74) | LDD0289 | [14] |
| LDCM0128 | JWB198 | DM93 | Y72(0.87) | LDD0290 | [14] |
| LDCM0129 | JWB202 | DM93 | Y72(0.32) | LDD0291 | [14] |
| LDCM0130 | JWB211 | DM93 | Y72(0.96) | LDD0292 | [14] |
| LDCM0022 | KB02 | HEK-293T | C31(1.04) | LDD1492 | [31] |
| LDCM0023 | KB03 | HEK-293T | C31(0.87) | LDD1497 | [31] |
| LDCM0024 | KB05 | WM793 | C31(2.88) | LDD3491 | [12] |
| LDCM0145 | MCC950 | THP-1 | 1.63 | LDD0368 | [30] |
| LDCM0109 | NEM | HeLa | H43(0.00); H94(0.00) | LDD0223 | [27] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C85(0.09) | LDD2124 | [13] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C85(0.14) | LDD2126 | [13] |
| LDCM0557 | Nucleophilic fragment 8b | MDA-MB-231 | C85(0.08) | LDD2151 | [13] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C85(1.14) | LDD2207 | [33] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Adenylate kinase isoenzyme 6 (AK6) | Adenylate kinase family | Q9Y3D8 | |||
| 26S proteasome non-ATPase regulatory subunit 9 (PSMD9) | Proteasome subunit p27 family | O00233 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Small ribosomal subunit protein eS1 (RPS3A) | Eukaryotic ribosomal protein eS1 family | P61247 | |||
| KRR1 small subunit processome component homolog (KRR1) | KRR1 family | Q13601 | |||
References
