Details of the Target
General Information of Target
| Target ID | LDTP03871 | |||||
|---|---|---|---|---|---|---|
| Target Name | Acidic leucine-rich nuclear phosphoprotein 32 family member A (ANP32A) | |||||
| Gene Name | ANP32A | |||||
| Gene ID | 8125 | |||||
| Synonyms |
C15orf1; LANP; MAPM; PHAP1; Acidic leucine-rich nuclear phosphoprotein 32 family member A; Acidic nuclear phosphoprotein pp32; pp32; Leucine-rich acidic nuclear protein; LANP; Mapmodulin; Potent heat-stable protein phosphatase 2A inhibitor I1PP2A; Putative HLA-DR-associated protein I; PHAPI
|
|||||
| 3D Structure | ||||||
| Sequence |
MEMGRRIHLELRNRTPSDVKELVLDNSRSNEGKLEGLTDEFEELEFLSTINVGLTSIANL
PKLNKLKKLELSDNRVSGGLEVLAEKCPNLTHLNLSGNKIKDLSTIEPLKKLENLKSLDL FNCEVTNLNDYRENVFKLLPQLTYLDGYDRDDKEAPDSDAEGYVEGLDDEEEDEDEEEYD EDAQVVEDEEDEDEEEEGEEEDVSGEEEEDEEGYNDGEVDDEEDEEELGEEERGQKRKRE PEDEGEDDD |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
ANP32 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Multifunctional protein that is involved in the regulation of many processes including tumor suppression, apoptosis, cell cycle progression or transcription. Promotes apoptosis by favouring the activation of caspase-9/CASP9 and allowing apoptosome formation. In addition, plays a role in the modulation of histone acetylation and transcription as part of the INHAT (inhibitor of histone acetyltransferases) complex. Inhibits the histone-acetyltranferase activity of EP300/CREBBP (CREB-binding protein) and EP300/CREBBP-associated factor by histone masking. Preferentially binds to unmodified histone H3 and sterically inhibiting its acetylation and phosphorylation leading to cell growth inhibition. Participates in other biochemical processes such as regulation of mRNA nuclear-to-cytoplasmic translocation and stability by its association with ELAVL1 (Hu-antigen R). Plays a role in E4F1-mediated transcriptional repression as well as inhibition of protein phosphatase 2A.; (Microbial infection) Plays an essential role in influenza A, B and C viral genome replication. Mechanistically, mediates the assembly of the viral replicase asymmetric dimers composed of PB1, PB2 and PA via its N-terminal region. Also plays an essential role in foamy virus mRNA export from the nucleus.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AZ-9 Probe Info |
 |
4.39 | LDD0393 | [1] | |
|
FBPP2 Probe Info |
 |
2.87 | LDD0318 | [2] | |
|
FBP2 Probe Info |
 |
3.09 | LDD0323 | [2] | |
|
TH216 Probe Info |
 |
Y131(10.80) | LDD0259 | [3] | |
|
ONAyne Probe Info |
 |
K68(0.00); K20(0.00) | LDD0273 | [4] | |
|
Probe 1 Probe Info |
 |
Y131(16.37); Y148(19.32) | LDD3495 | [5] | |
|
DBIA Probe Info |
 |
C87(1.17) | LDD3312 | [6] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [7] | |
|
THZ1-DTB Probe Info |
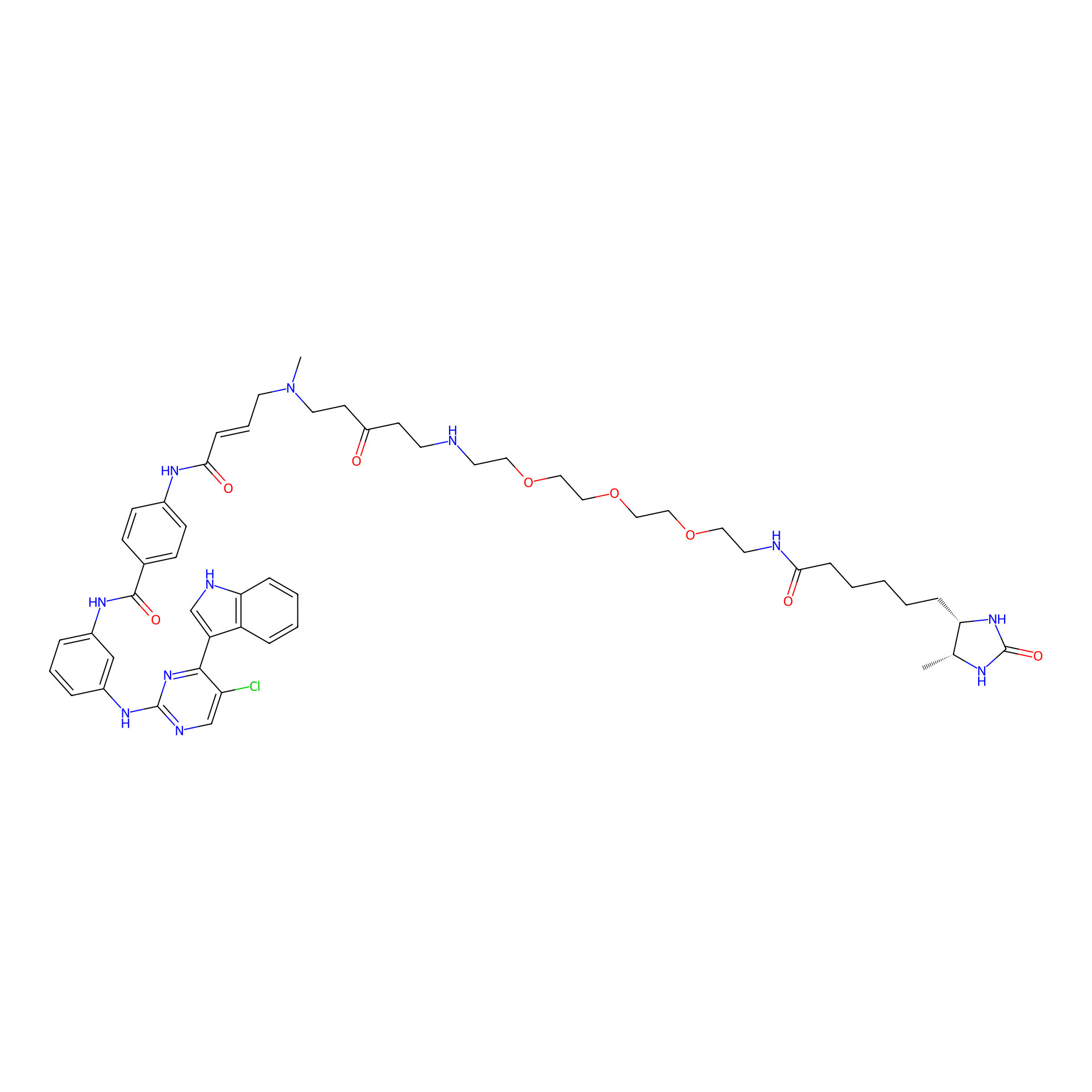 |
C123(1.07) | LDD0460 | [7] | |
|
BTD Probe Info |
 |
C87(1.28) | LDD2090 | [8] | |
|
AHL-Pu-1 Probe Info |
 |
C123(3.35) | LDD0171 | [9] | |
|
HHS-482 Probe Info |
 |
Y131(2.52) | LDD0285 | [10] | |
|
5E-2FA Probe Info |
 |
H92(0.00); H8(0.00) | LDD2235 | [11] | |
|
ATP probe Probe Info |
 |
K99(0.00); K101(0.00); K68(0.00); K20(0.00) | LDD0199 | [12] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [11] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0358 | [13] | |
|
N1 Probe Info |
 |
N.A. | LDD0245 | [14] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C87(0.00); C123(0.00) | LDD0038 | [15] | |
|
IA-alkyne Probe Info |
 |
C87(0.00); C123(0.00) | LDD0032 | [16] | |
|
IPIAA_H Probe Info |
 |
C123(0.00); C87(0.00) | LDD0030 | [17] | |
|
IPIAA_L Probe Info |
 |
C87(0.00); C123(0.00) | LDD0031 | [17] | |
|
Lodoacetamide azide Probe Info |
 |
C87(0.00); C123(0.00) | LDD0037 | [15] | |
|
ATP probe Probe Info |
 |
K99(0.00); K86(0.00) | LDD0035 | [18] | |
|
Compound 10 Probe Info |
 |
N.A. | LDD2216 | [19] | |
|
IPM Probe Info |
 |
C123(0.00); C87(0.00) | LDD0147 | [20] | |
|
OSF Probe Info |
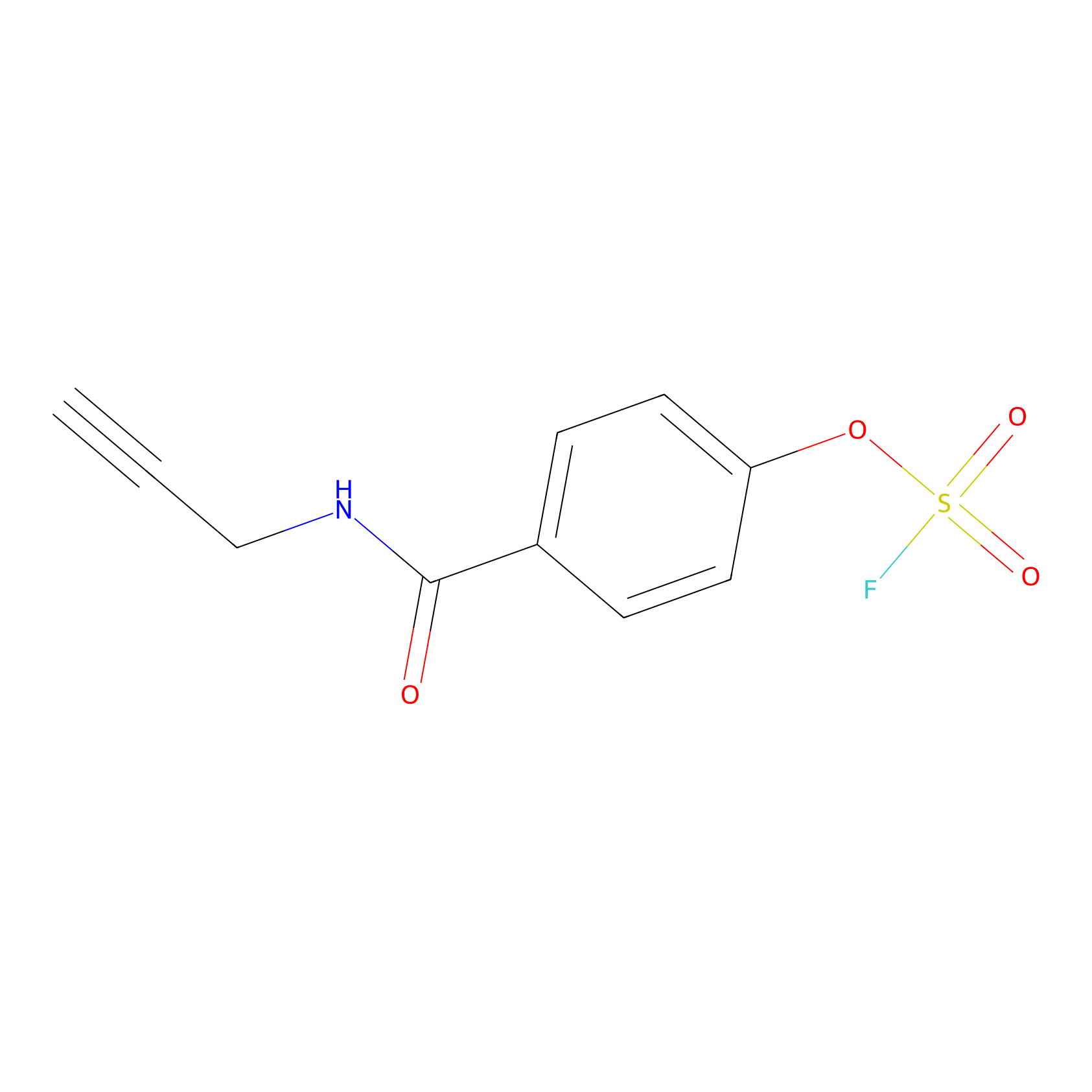 |
N.A. | LDD0029 | [21] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [22] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [20] | |
|
YN-1 Probe Info |
 |
N.A. | LDD0446 | [23] | |
|
1c-yne Probe Info |
 |
K137(0.00); K20(0.00); K116(0.00) | LDD0228 | [13] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [24] | |
|
NAIA_5 Probe Info |
 |
C87(0.00); C123(0.00) | LDD2223 | [25] | |
|
HHS-465 Probe Info |
 |
K101(0.00); K110(0.00); K111(0.00); K99(0.00) | LDD2240 | [26] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C022 Probe Info |
 |
8.63 | LDD1728 | [27] | |
|
FFF probe12 Probe Info |
 |
6.38 | LDD0473 | [28] | |
|
FFF probe13 Probe Info |
 |
6.44 | LDD0475 | [28] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0070 | [29] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C87(0.47) | LDD2142 | [8] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C87(1.11) | LDD2117 | [8] |
| LDCM0510 | 3-(4-(Hydroxydiphenylmethyl)piperidin-1-yl)-3-oxopropanenitrile | MDA-MB-231 | C87(1.07) | LDD2103 | [8] |
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C123(3.35) | LDD0171 | [9] |
| LDCM0214 | AC1 | HEK-293T | C87(0.87); C123(1.10) | LDD1507 | [30] |
| LDCM0215 | AC10 | HEK-293T | C87(0.95); C123(0.90) | LDD1508 | [30] |
| LDCM0226 | AC11 | HEK-293T | C87(0.98); C123(1.01) | LDD1509 | [30] |
| LDCM0237 | AC12 | HEK-293T | C87(1.03); C123(0.97) | LDD1510 | [30] |
| LDCM0259 | AC14 | HEK-293T | C87(1.19); C123(1.00) | LDD1512 | [30] |
| LDCM0270 | AC15 | HEK-293T | C87(0.99); C123(0.79) | LDD1513 | [30] |
| LDCM0276 | AC17 | HEK-293T | C87(0.92); C123(0.89) | LDD1515 | [30] |
| LDCM0277 | AC18 | HEK-293T | C87(1.02); C123(1.01) | LDD1516 | [30] |
| LDCM0278 | AC19 | HEK-293T | C87(0.90); C123(0.89) | LDD1517 | [30] |
| LDCM0279 | AC2 | HEK-293T | C87(0.95); C123(1.02) | LDD1518 | [30] |
| LDCM0280 | AC20 | HEK-293T | C87(1.08); C123(1.00) | LDD1519 | [30] |
| LDCM0281 | AC21 | HEK-293T | C87(1.10); C123(1.06) | LDD1520 | [30] |
| LDCM0282 | AC22 | HEK-293T | C87(1.16); C123(0.99) | LDD1521 | [30] |
| LDCM0283 | AC23 | HEK-293T | C87(1.22); C123(0.95) | LDD1522 | [30] |
| LDCM0284 | AC24 | HEK-293T | C87(1.10); C123(1.09) | LDD1523 | [30] |
| LDCM0285 | AC25 | HEK-293T | C87(0.90); C123(0.96) | LDD1524 | [30] |
| LDCM0286 | AC26 | HEK-293T | C87(0.99); C123(0.98) | LDD1525 | [30] |
| LDCM0287 | AC27 | HEK-293T | C87(1.06); C123(1.05) | LDD1526 | [30] |
| LDCM0288 | AC28 | HEK-293T | C87(1.13); C123(1.09) | LDD1527 | [30] |
| LDCM0289 | AC29 | HEK-293T | C87(1.14); C123(1.08) | LDD1528 | [30] |
| LDCM0290 | AC3 | HEK-293T | C87(0.99); C123(1.12) | LDD1529 | [30] |
| LDCM0291 | AC30 | HEK-293T | C87(1.19); C123(1.04) | LDD1530 | [30] |
| LDCM0292 | AC31 | HEK-293T | C87(1.06); C123(1.04) | LDD1531 | [30] |
| LDCM0293 | AC32 | HEK-293T | C87(1.03); C123(1.06) | LDD1532 | [30] |
| LDCM0294 | AC33 | HEK-293T | C87(0.81); C123(0.87) | LDD1533 | [30] |
| LDCM0295 | AC34 | HEK-293T | C87(0.94); C123(1.00) | LDD1534 | [30] |
| LDCM0296 | AC35 | HEK-293T | C87(0.97); C123(1.04) | LDD1535 | [30] |
| LDCM0297 | AC36 | HEK-293T | C87(1.00); C123(1.09) | LDD1536 | [30] |
| LDCM0298 | AC37 | HEK-293T | C87(1.14); C123(1.18) | LDD1537 | [30] |
| LDCM0299 | AC38 | HEK-293T | C87(1.27); C123(1.00) | LDD1538 | [30] |
| LDCM0300 | AC39 | HEK-293T | C87(0.99); C123(0.91) | LDD1539 | [30] |
| LDCM0301 | AC4 | HEK-293T | C87(1.06); C123(1.21) | LDD1540 | [30] |
| LDCM0302 | AC40 | HEK-293T | C87(1.09); C123(1.04) | LDD1541 | [30] |
| LDCM0303 | AC41 | HEK-293T | C87(0.83); C123(0.83) | LDD1542 | [30] |
| LDCM0304 | AC42 | HEK-293T | C87(0.89); C123(0.93) | LDD1543 | [30] |
| LDCM0305 | AC43 | HEK-293T | C87(0.98); C123(0.93) | LDD1544 | [30] |
| LDCM0306 | AC44 | HEK-293T | C87(0.98); C123(1.09) | LDD1545 | [30] |
| LDCM0307 | AC45 | HEK-293T | C87(1.08); C123(1.16) | LDD1546 | [30] |
| LDCM0308 | AC46 | HEK-293T | C87(1.24); C123(1.07) | LDD1547 | [30] |
| LDCM0309 | AC47 | HEK-293T | C87(0.97); C123(1.11) | LDD1548 | [30] |
| LDCM0310 | AC48 | HEK-293T | C87(1.12); C123(1.11) | LDD1549 | [30] |
| LDCM0311 | AC49 | HEK-293T | C87(0.91); C123(0.90) | LDD1550 | [30] |
| LDCM0312 | AC5 | HEK-293T | C87(1.09); C123(1.27) | LDD1551 | [30] |
| LDCM0313 | AC50 | HEK-293T | C87(1.02); C123(1.00) | LDD1552 | [30] |
| LDCM0314 | AC51 | HEK-293T | C87(1.04); C123(1.06) | LDD1553 | [30] |
| LDCM0315 | AC52 | HEK-293T | C87(0.97); C123(1.00) | LDD1554 | [30] |
| LDCM0316 | AC53 | HEK-293T | C87(1.03); C123(1.12) | LDD1555 | [30] |
| LDCM0317 | AC54 | HEK-293T | C87(1.12); C123(0.92) | LDD1556 | [30] |
| LDCM0318 | AC55 | HEK-293T | C87(1.03); C123(0.94) | LDD1557 | [30] |
| LDCM0319 | AC56 | HEK-293T | C87(1.07); C123(1.08) | LDD1558 | [30] |
| LDCM0320 | AC57 | HEK-293T | C87(0.84); C123(0.87) | LDD1559 | [30] |
| LDCM0321 | AC58 | HEK-293T | C87(0.98); C123(1.11) | LDD1560 | [30] |
| LDCM0322 | AC59 | HEK-293T | C87(1.04); C123(1.11) | LDD1561 | [30] |
| LDCM0323 | AC6 | HEK-293T | C87(1.13); C123(0.97) | LDD1562 | [30] |
| LDCM0324 | AC60 | HEK-293T | C87(0.96); C123(1.08) | LDD1563 | [30] |
| LDCM0325 | AC61 | HEK-293T | C87(1.10); C123(1.28) | LDD1564 | [30] |
| LDCM0326 | AC62 | HEK-293T | C87(1.12); C123(0.99) | LDD1565 | [30] |
| LDCM0327 | AC63 | HEK-293T | C87(0.95); C123(1.10) | LDD1566 | [30] |
| LDCM0328 | AC64 | HEK-293T | C87(1.12); C123(1.35) | LDD1567 | [30] |
| LDCM0334 | AC7 | HEK-293T | C87(1.00); C123(1.02) | LDD1568 | [30] |
| LDCM0345 | AC8 | HEK-293T | C87(1.10); C123(1.19) | LDD1569 | [30] |
| LDCM0248 | AKOS034007472 | HEK-293T | C87(1.09); C123(1.22) | LDD1511 | [30] |
| LDCM0356 | AKOS034007680 | HEK-293T | C87(0.80); C123(0.77) | LDD1570 | [30] |
| LDCM0275 | AKOS034007705 | HEK-293T | C87(1.11); C123(1.00) | LDD1514 | [30] |
| LDCM0630 | CCW28-3 | 231MFP | C123(1.49) | LDD2214 | [31] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [24] |
| LDCM0632 | CL-Sc | Hep-G2 | C123(20.00) | LDD2227 | [25] |
| LDCM0367 | CL1 | HEK-293T | C87(0.90); C123(0.95) | LDD1571 | [30] |
| LDCM0368 | CL10 | HEK-293T | C87(0.82); C123(0.50) | LDD1572 | [30] |
| LDCM0369 | CL100 | HEK-293T | C87(0.92); C123(1.01) | LDD1573 | [30] |
| LDCM0370 | CL101 | HEK-293T | C87(0.91); C123(1.02) | LDD1574 | [30] |
| LDCM0371 | CL102 | HEK-293T | C87(0.92); C123(0.86) | LDD1575 | [30] |
| LDCM0372 | CL103 | HEK-293T | C87(0.99); C123(0.85) | LDD1576 | [30] |
| LDCM0373 | CL104 | HEK-293T | C87(0.88); C123(0.99) | LDD1577 | [30] |
| LDCM0374 | CL105 | HEK-293T | C87(1.01); C123(0.97) | LDD1578 | [30] |
| LDCM0375 | CL106 | HEK-293T | C87(1.02); C123(0.94) | LDD1579 | [30] |
| LDCM0376 | CL107 | HEK-293T | C87(1.09); C123(1.09) | LDD1580 | [30] |
| LDCM0377 | CL108 | HEK-293T | C87(1.07); C123(1.05) | LDD1581 | [30] |
| LDCM0378 | CL109 | HEK-293T | C87(1.02); C123(1.00) | LDD1582 | [30] |
| LDCM0379 | CL11 | HEK-293T | C87(0.89); C123(0.21) | LDD1583 | [30] |
| LDCM0380 | CL110 | HEK-293T | C87(0.95); C123(0.90) | LDD1584 | [30] |
| LDCM0381 | CL111 | HEK-293T | C87(1.03); C123(0.89) | LDD1585 | [30] |
| LDCM0382 | CL112 | HEK-293T | C87(0.96); C123(1.00) | LDD1586 | [30] |
| LDCM0383 | CL113 | HEK-293T | C87(0.92); C123(1.01) | LDD1587 | [30] |
| LDCM0384 | CL114 | HEK-293T | C87(0.92); C123(0.94) | LDD1588 | [30] |
| LDCM0385 | CL115 | HEK-293T | C87(1.03); C123(1.07) | LDD1589 | [30] |
| LDCM0386 | CL116 | HEK-293T | C87(0.95); C123(1.04) | LDD1590 | [30] |
| LDCM0387 | CL117 | HEK-293T | C87(0.97); C123(0.97) | LDD1591 | [30] |
| LDCM0388 | CL118 | HEK-293T | C87(1.00); C123(1.02) | LDD1592 | [30] |
| LDCM0389 | CL119 | HEK-293T | C87(1.06); C123(0.96) | LDD1593 | [30] |
| LDCM0390 | CL12 | HEK-293T | C87(1.03); C123(0.32) | LDD1594 | [30] |
| LDCM0391 | CL120 | HEK-293T | C87(0.92); C123(1.06) | LDD1595 | [30] |
| LDCM0392 | CL121 | HEK-293T | C87(1.00); C123(1.05) | LDD1596 | [30] |
| LDCM0393 | CL122 | HEK-293T | C87(0.99); C123(1.00) | LDD1597 | [30] |
| LDCM0394 | CL123 | HEK-293T | C87(0.93); C123(1.03) | LDD1598 | [30] |
| LDCM0395 | CL124 | HEK-293T | C87(0.95); C123(1.02) | LDD1599 | [30] |
| LDCM0396 | CL125 | HEK-293T | C87(0.98); C123(1.07) | LDD1600 | [30] |
| LDCM0397 | CL126 | HEK-293T | C87(0.94); C123(1.12) | LDD1601 | [30] |
| LDCM0398 | CL127 | HEK-293T | C87(0.96); C123(1.26) | LDD1602 | [30] |
| LDCM0399 | CL128 | HEK-293T | C87(0.98); C123(1.22) | LDD1603 | [30] |
| LDCM0400 | CL13 | HEK-293T | C87(0.78); C123(0.83) | LDD1604 | [30] |
| LDCM0401 | CL14 | HEK-293T | C87(1.01); C123(0.88) | LDD1605 | [30] |
| LDCM0402 | CL15 | HEK-293T | C87(0.75); C123(0.55) | LDD1606 | [30] |
| LDCM0403 | CL16 | HEK-293T | C87(0.91); C123(0.81) | LDD1607 | [30] |
| LDCM0404 | CL17 | HEK-293T | C87(0.58); C123(0.39) | LDD1608 | [30] |
| LDCM0405 | CL18 | HEK-293T | C87(0.92); C123(0.69) | LDD1609 | [30] |
| LDCM0406 | CL19 | HEK-293T | C87(0.93); C123(0.57) | LDD1610 | [30] |
| LDCM0407 | CL2 | HEK-293T | C87(0.93); C123(0.82) | LDD1611 | [30] |
| LDCM0408 | CL20 | HEK-293T | C87(0.96); C123(0.56) | LDD1612 | [30] |
| LDCM0409 | CL21 | HEK-293T | C87(0.93); C123(0.29) | LDD1613 | [30] |
| LDCM0410 | CL22 | HEK-293T | C87(1.12); C123(0.66) | LDD1614 | [30] |
| LDCM0411 | CL23 | HEK-293T | C87(0.99); C123(0.22) | LDD1615 | [30] |
| LDCM0412 | CL24 | HEK-293T | C87(1.09); C123(0.32) | LDD1616 | [30] |
| LDCM0413 | CL25 | HEK-293T | C87(0.86); C123(1.01) | LDD1617 | [30] |
| LDCM0414 | CL26 | HEK-293T | C87(0.99); C123(0.85) | LDD1618 | [30] |
| LDCM0415 | CL27 | HEK-293T | C87(1.03); C123(0.87) | LDD1619 | [30] |
| LDCM0416 | CL28 | HEK-293T | C87(0.96); C123(0.85) | LDD1620 | [30] |
| LDCM0417 | CL29 | HEK-293T | C87(0.93); C123(0.62) | LDD1621 | [30] |
| LDCM0418 | CL3 | HEK-293T | C87(0.92); C123(0.68) | LDD1622 | [30] |
| LDCM0419 | CL30 | HEK-293T | C87(1.06); C123(0.80) | LDD1623 | [30] |
| LDCM0420 | CL31 | HEK-293T | C87(0.96); C123(0.57) | LDD1624 | [30] |
| LDCM0421 | CL32 | HEK-293T | C87(1.03); C123(0.59) | LDD1625 | [30] |
| LDCM0422 | CL33 | HEK-293T | C87(0.91); C123(0.34) | LDD1626 | [30] |
| LDCM0423 | CL34 | HEK-293T | C87(1.13); C123(0.66) | LDD1627 | [30] |
| LDCM0424 | CL35 | HEK-293T | C87(1.10); C123(0.21) | LDD1628 | [30] |
| LDCM0425 | CL36 | HEK-293T | C87(1.14); C123(0.33) | LDD1629 | [30] |
| LDCM0426 | CL37 | HEK-293T | C87(0.94); C123(0.94) | LDD1630 | [30] |
| LDCM0428 | CL39 | HEK-293T | C87(0.95); C123(0.81) | LDD1632 | [30] |
| LDCM0429 | CL4 | HEK-293T | C87(0.76); C123(0.81) | LDD1633 | [30] |
| LDCM0430 | CL40 | HEK-293T | C87(0.95); C123(0.96) | LDD1634 | [30] |
| LDCM0431 | CL41 | HEK-293T | C87(0.85); C123(0.61) | LDD1635 | [30] |
| LDCM0432 | CL42 | HEK-293T | C87(1.01); C123(0.91) | LDD1636 | [30] |
| LDCM0433 | CL43 | HEK-293T | C87(1.18); C123(0.70) | LDD1637 | [30] |
| LDCM0434 | CL44 | HEK-293T | C87(1.07); C123(0.65) | LDD1638 | [30] |
| LDCM0435 | CL45 | HEK-293T | C87(1.01); C123(0.36) | LDD1639 | [30] |
| LDCM0436 | CL46 | HEK-293T | C87(1.22); C123(0.69) | LDD1640 | [30] |
| LDCM0437 | CL47 | HEK-293T | C87(1.06); C123(0.23) | LDD1641 | [30] |
| LDCM0438 | CL48 | HEK-293T | C87(1.10); C123(0.40) | LDD1642 | [30] |
| LDCM0439 | CL49 | HEK-293T | C87(0.96); C123(1.07) | LDD1643 | [30] |
| LDCM0440 | CL5 | HEK-293T | C87(0.73); C123(0.64) | LDD1644 | [30] |
| LDCM0441 | CL50 | HEK-293T | C87(0.92); C123(0.85) | LDD1645 | [30] |
| LDCM0443 | CL52 | HEK-293T | C87(0.92); C123(0.84) | LDD1646 | [30] |
| LDCM0444 | CL53 | HEK-293T | C87(0.76); C123(0.56) | LDD1647 | [30] |
| LDCM0445 | CL54 | HEK-293T | C87(0.88); C123(0.80) | LDD1648 | [30] |
| LDCM0446 | CL55 | HEK-293T | C87(1.03); C123(0.60) | LDD1649 | [30] |
| LDCM0447 | CL56 | HEK-293T | C87(0.94); C123(0.67) | LDD1650 | [30] |
| LDCM0448 | CL57 | HEK-293T | C87(1.00); C123(0.38) | LDD1651 | [30] |
| LDCM0449 | CL58 | HEK-293T | C87(1.20); C123(0.68) | LDD1652 | [30] |
| LDCM0450 | CL59 | HEK-293T | C87(1.00); C123(0.26) | LDD1653 | [30] |
| LDCM0451 | CL6 | HEK-293T | C87(0.73); C123(0.68) | LDD1654 | [30] |
| LDCM0452 | CL60 | HEK-293T | C87(1.14); C123(0.36) | LDD1655 | [30] |
| LDCM0453 | CL61 | HEK-293T | C87(0.93); C123(1.00) | LDD1656 | [30] |
| LDCM0454 | CL62 | HEK-293T | C87(1.00); C123(1.04) | LDD1657 | [30] |
| LDCM0455 | CL63 | HEK-293T | C87(1.02); C123(1.04) | LDD1658 | [30] |
| LDCM0456 | CL64 | HEK-293T | C87(0.83); C123(0.87) | LDD1659 | [30] |
| LDCM0457 | CL65 | HEK-293T | C87(0.81); C123(0.70) | LDD1660 | [30] |
| LDCM0458 | CL66 | HEK-293T | C87(0.88); C123(0.82) | LDD1661 | [30] |
| LDCM0459 | CL67 | HEK-293T | C87(1.02); C123(0.77) | LDD1662 | [30] |
| LDCM0460 | CL68 | HEK-293T | C87(0.99); C123(0.70) | LDD1663 | [30] |
| LDCM0461 | CL69 | HEK-293T | C87(1.08); C123(0.50) | LDD1664 | [30] |
| LDCM0462 | CL7 | HEK-293T | C87(0.90); C123(0.67) | LDD1665 | [30] |
| LDCM0463 | CL70 | HEK-293T | C87(1.20); C123(0.70) | LDD1666 | [30] |
| LDCM0464 | CL71 | HEK-293T | C87(1.03); C123(0.29) | LDD1667 | [30] |
| LDCM0465 | CL72 | HEK-293T | C87(1.02); C123(0.40) | LDD1668 | [30] |
| LDCM0466 | CL73 | HEK-293T | C87(0.93); C123(0.96) | LDD1669 | [30] |
| LDCM0467 | CL74 | HEK-293T | C87(1.03); C123(1.04) | LDD1670 | [30] |
| LDCM0469 | CL76 | HEK-293T | C87(0.91); C123(1.03) | LDD1672 | [30] |
| LDCM0470 | CL77 | HEK-293T | C87(0.76); C123(0.77) | LDD1673 | [30] |
| LDCM0471 | CL78 | HEK-293T | C87(0.97); C123(0.87) | LDD1674 | [30] |
| LDCM0472 | CL79 | HEK-293T | C87(1.16); C123(0.80) | LDD1675 | [30] |
| LDCM0473 | CL8 | HEK-293T | C87(0.71); C123(0.59) | LDD1676 | [30] |
| LDCM0474 | CL80 | HEK-293T | C87(1.23); C123(0.89) | LDD1677 | [30] |
| LDCM0475 | CL81 | HEK-293T | C87(1.12); C123(0.50) | LDD1678 | [30] |
| LDCM0476 | CL82 | HEK-293T | C87(1.26); C123(0.85) | LDD1679 | [30] |
| LDCM0477 | CL83 | HEK-293T | C87(1.10); C123(0.36) | LDD1680 | [30] |
| LDCM0478 | CL84 | HEK-293T | C87(1.10); C123(0.55) | LDD1681 | [30] |
| LDCM0479 | CL85 | HEK-293T | C87(0.91); C123(1.07) | LDD1682 | [30] |
| LDCM0480 | CL86 | HEK-293T | C87(0.89); C123(1.09) | LDD1683 | [30] |
| LDCM0481 | CL87 | HEK-293T | C87(0.91); C123(1.49) | LDD1684 | [30] |
| LDCM0482 | CL88 | HEK-293T | C87(0.89); C123(1.24) | LDD1685 | [30] |
| LDCM0483 | CL89 | HEK-293T | C87(0.81); C123(1.07) | LDD1686 | [30] |
| LDCM0484 | CL9 | HEK-293T | C87(0.92); C123(0.35) | LDD1687 | [30] |
| LDCM0485 | CL90 | HEK-293T | C87(0.69); C123(0.83) | LDD1688 | [30] |
| LDCM0486 | CL91 | HEK-293T | C87(1.02); C123(0.98) | LDD1689 | [30] |
| LDCM0487 | CL92 | HEK-293T | C87(0.95); C123(0.89) | LDD1690 | [30] |
| LDCM0488 | CL93 | HEK-293T | C87(1.02); C123(0.57) | LDD1691 | [30] |
| LDCM0489 | CL94 | HEK-293T | C87(1.09); C123(0.86) | LDD1692 | [30] |
| LDCM0490 | CL95 | HEK-293T | C87(0.70); C123(0.38) | LDD1693 | [30] |
| LDCM0491 | CL96 | HEK-293T | C87(0.99); C123(0.85) | LDD1694 | [30] |
| LDCM0492 | CL97 | HEK-293T | C87(0.92); C123(0.94) | LDD1695 | [30] |
| LDCM0493 | CL98 | HEK-293T | C87(0.99); C123(1.01) | LDD1696 | [30] |
| LDCM0494 | CL99 | HEK-293T | C87(1.10); C123(1.29) | LDD1697 | [30] |
| LDCM0495 | E2913 | HEK-293T | C87(0.86); C123(0.83) | LDD1698 | [30] |
| LDCM0625 | F8 | Ramos | C123(0.54); C87(0.73) | LDD2187 | [32] |
| LDCM0572 | Fragment10 | Ramos | C123(0.68); C87(0.47) | LDD2189 | [32] |
| LDCM0573 | Fragment11 | Ramos | C123(0.30); C87(0.10) | LDD2190 | [32] |
| LDCM0574 | Fragment12 | Ramos | C123(0.68); C87(0.51) | LDD2191 | [32] |
| LDCM0575 | Fragment13 | Ramos | C123(0.84); C87(0.77) | LDD2192 | [32] |
| LDCM0576 | Fragment14 | Ramos | C123(1.08); C87(0.99) | LDD2193 | [32] |
| LDCM0579 | Fragment20 | Ramos | C123(0.74); C87(0.36) | LDD2194 | [32] |
| LDCM0580 | Fragment21 | Ramos | C123(0.93); C87(0.96) | LDD2195 | [32] |
| LDCM0582 | Fragment23 | Ramos | C123(1.18); C87(1.31) | LDD2196 | [32] |
| LDCM0578 | Fragment27 | Ramos | C123(0.98); C87(1.31) | LDD2197 | [32] |
| LDCM0586 | Fragment28 | Ramos | C123(0.42); C87(0.70) | LDD2198 | [32] |
| LDCM0588 | Fragment30 | Ramos | C123(1.12); C87(0.71) | LDD2199 | [32] |
| LDCM0589 | Fragment31 | Ramos | C123(1.36); C87(1.19) | LDD2200 | [32] |
| LDCM0590 | Fragment32 | Ramos | C123(0.74); C87(0.42) | LDD2201 | [32] |
| LDCM0468 | Fragment33 | HEK-293T | C87(1.01); C123(1.08) | LDD1671 | [30] |
| LDCM0596 | Fragment38 | Ramos | C123(0.97); C87(1.75) | LDD2203 | [32] |
| LDCM0566 | Fragment4 | Ramos | C123(0.87); C87(0.57) | LDD2184 | [32] |
| LDCM0427 | Fragment51 | HEK-293T | C87(1.01); C123(0.95) | LDD1631 | [30] |
| LDCM0610 | Fragment52 | Ramos | C123(1.00); C87(0.99) | LDD2204 | [32] |
| LDCM0614 | Fragment56 | Ramos | C123(1.07); C87(0.75) | LDD2205 | [32] |
| LDCM0569 | Fragment7 | Ramos | C123(0.79); C87(0.34) | LDD2186 | [32] |
| LDCM0571 | Fragment9 | Ramos | C123(0.56); C87(0.31) | LDD2188 | [32] |
| LDCM0123 | JWB131 | DM93 | Y131(2.52) | LDD0285 | [10] |
| LDCM0124 | JWB142 | DM93 | Y131(1.17) | LDD0286 | [10] |
| LDCM0125 | JWB146 | DM93 | Y131(0.72) | LDD0287 | [10] |
| LDCM0126 | JWB150 | DM93 | Y131(7.34) | LDD0288 | [10] |
| LDCM0127 | JWB152 | DM93 | Y131(2.51) | LDD0289 | [10] |
| LDCM0128 | JWB198 | DM93 | Y131(0.84) | LDD0290 | [10] |
| LDCM0129 | JWB202 | DM93 | Y131(0.56) | LDD0291 | [10] |
| LDCM0130 | JWB211 | DM93 | Y131(3.02) | LDD0292 | [10] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [7] |
| LDCM0022 | KB02 | HEK-293T | C87(1.00); C123(0.99) | LDD1492 | [30] |
| LDCM0023 | KB03 | HEK-293T | C87(1.00); C123(1.01) | LDD1497 | [30] |
| LDCM0024 | KB05 | HMCB | C87(1.17) | LDD3312 | [6] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C87(0.95) | LDD2102 | [8] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [24] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C87(1.28) | LDD2090 | [8] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C87(1.24) | LDD2092 | [8] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C87(1.02) | LDD2093 | [8] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C87(1.14) | LDD2098 | [8] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C87(1.12) | LDD2099 | [8] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C87(0.33) | LDD2104 | [8] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C87(0.43) | LDD2106 | [8] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C87(1.07) | LDD2107 | [8] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C87(0.68) | LDD2109 | [8] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C87(0.42) | LDD2110 | [8] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C87(1.18) | LDD2111 | [8] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C87(0.99) | LDD2123 | [8] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C87(0.90) | LDD2125 | [8] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C87(0.93) | LDD2127 | [8] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C87(0.98) | LDD2136 | [8] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C87(1.02) | LDD2137 | [8] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C87(0.51) | LDD2141 | [8] |
| LDCM0131 | RA190 | MM1.R | C87(1.69); C123(1.36) | LDD0304 | [33] |
| LDCM0021 | THZ1 | HeLa S3 | C123(1.07) | LDD0460 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Serine/threonine-protein phosphatase 2A catalytic subunit alpha isoform (PPP2CA) | PPP phosphatase family | P67775 | |||
| Transcription factor E4F1 (E4F1) | . | Q66K89 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Ataxin-1 (ATXN1) | ATXN1 family | P54253 | |||
| Axin-1 (AXIN1) | . | O15169 | |||
References
