Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
2.27 | LDD0318 | [1] | |
|
FBP2 Probe Info |
 |
3.95 | LDD0317 | [1] | |
|
Curcusone 37 Probe Info |
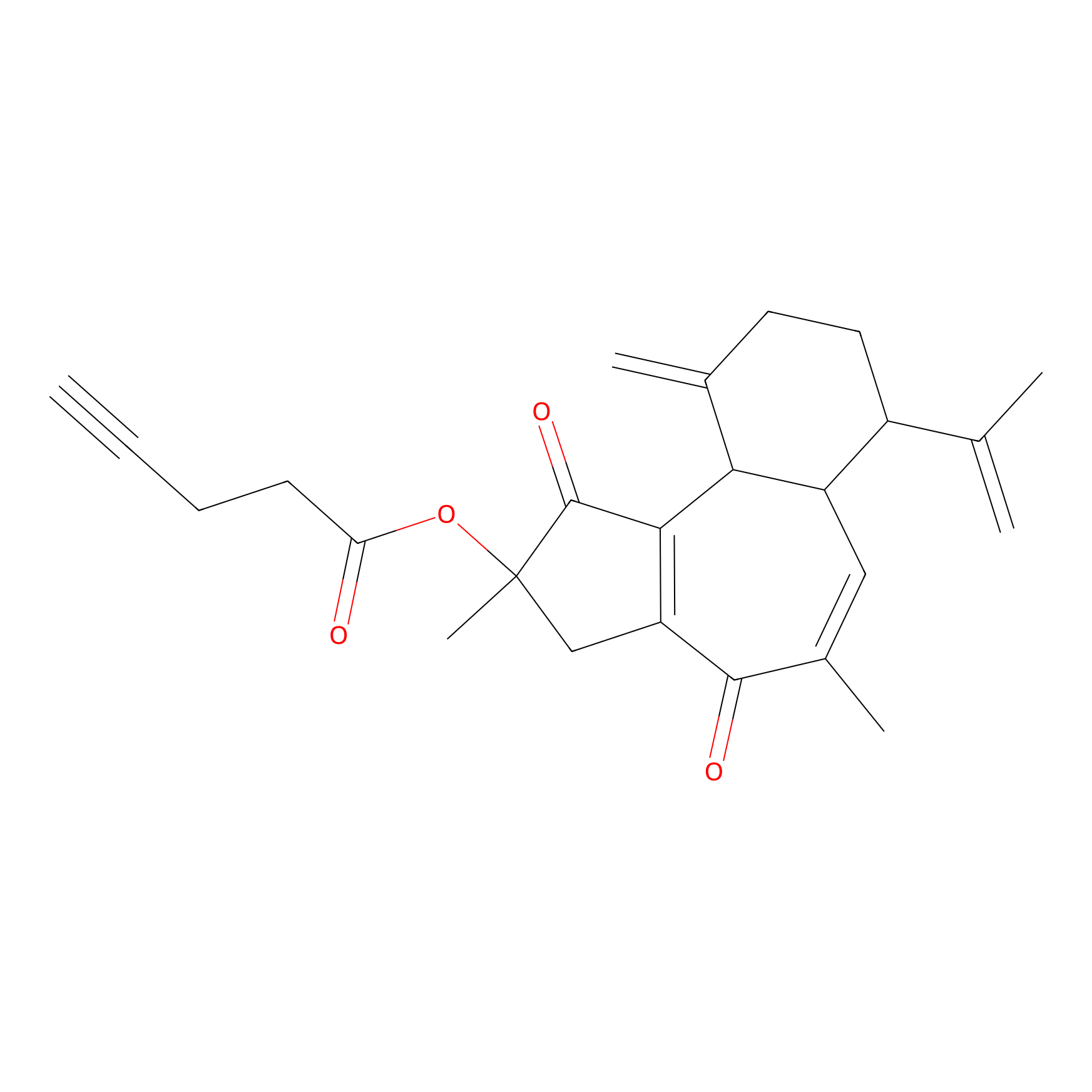 |
4.23 | LDD0188 | [2] | |
|
Alkyne-RA190 Probe Info |
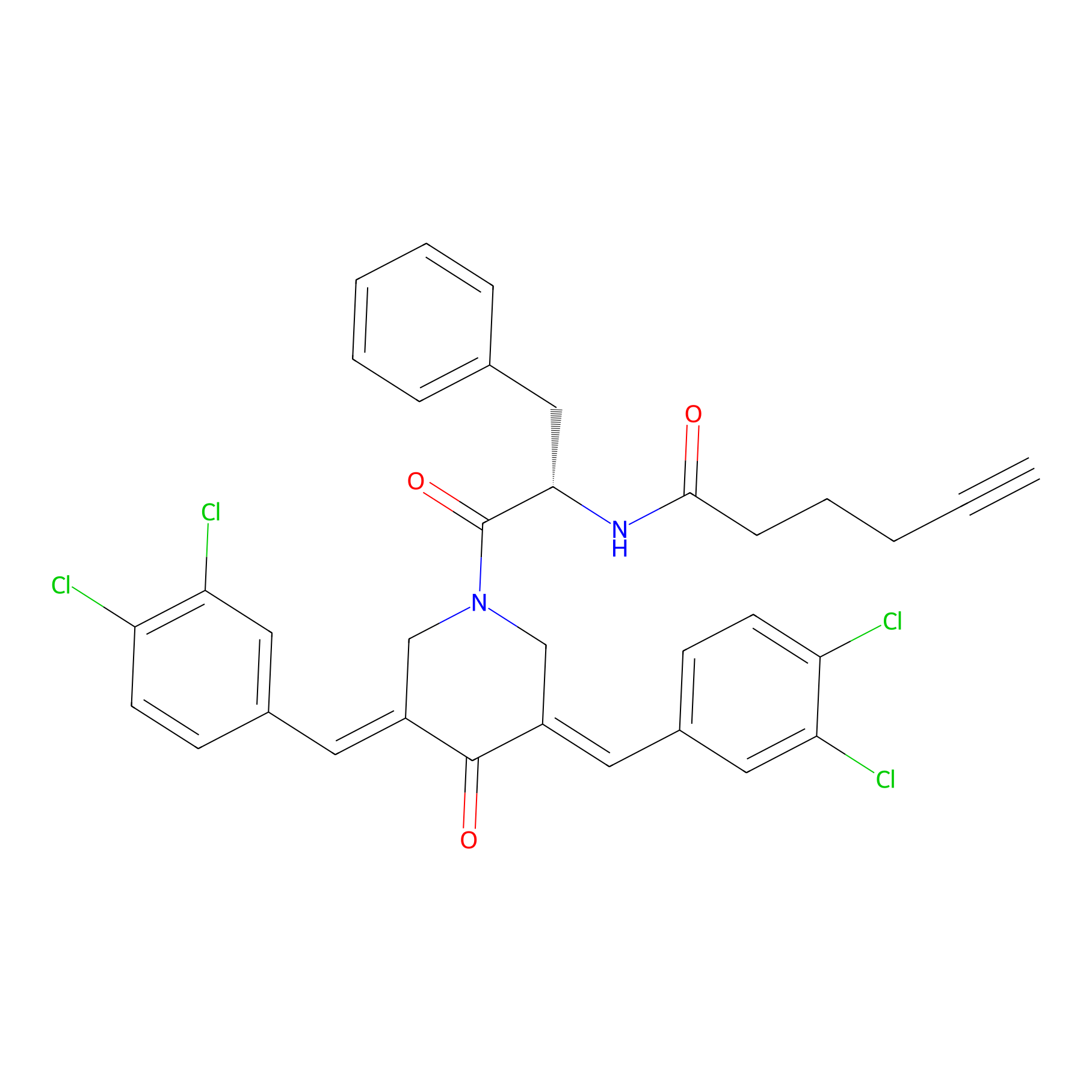 |
2.24 | LDD0302 | [3] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0165 | [4] | |
|
AOyne Probe Info |
 |
12.20 | LDD0443 | [5] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C010 Probe Info |
 |
5.10 | LDD1719 | [6] | |
|
C022 Probe Info |
 |
7.73 | LDD1728 | [6] | |
|
C040 Probe Info |
 |
20.39 | LDD1740 | [6] | |
|
C041 Probe Info |
 |
6.54 | LDD1741 | [6] | |
|
C112 Probe Info |
 |
17.51 | LDD1799 | [6] | |
|
C169 Probe Info |
 |
23.10 | LDD1849 | [6] | |
|
C170 Probe Info |
 |
9.19 | LDD1850 | [6] | |
|
C210 Probe Info |
 |
64.89 | LDD1884 | [6] | |
|
C239 Probe Info |
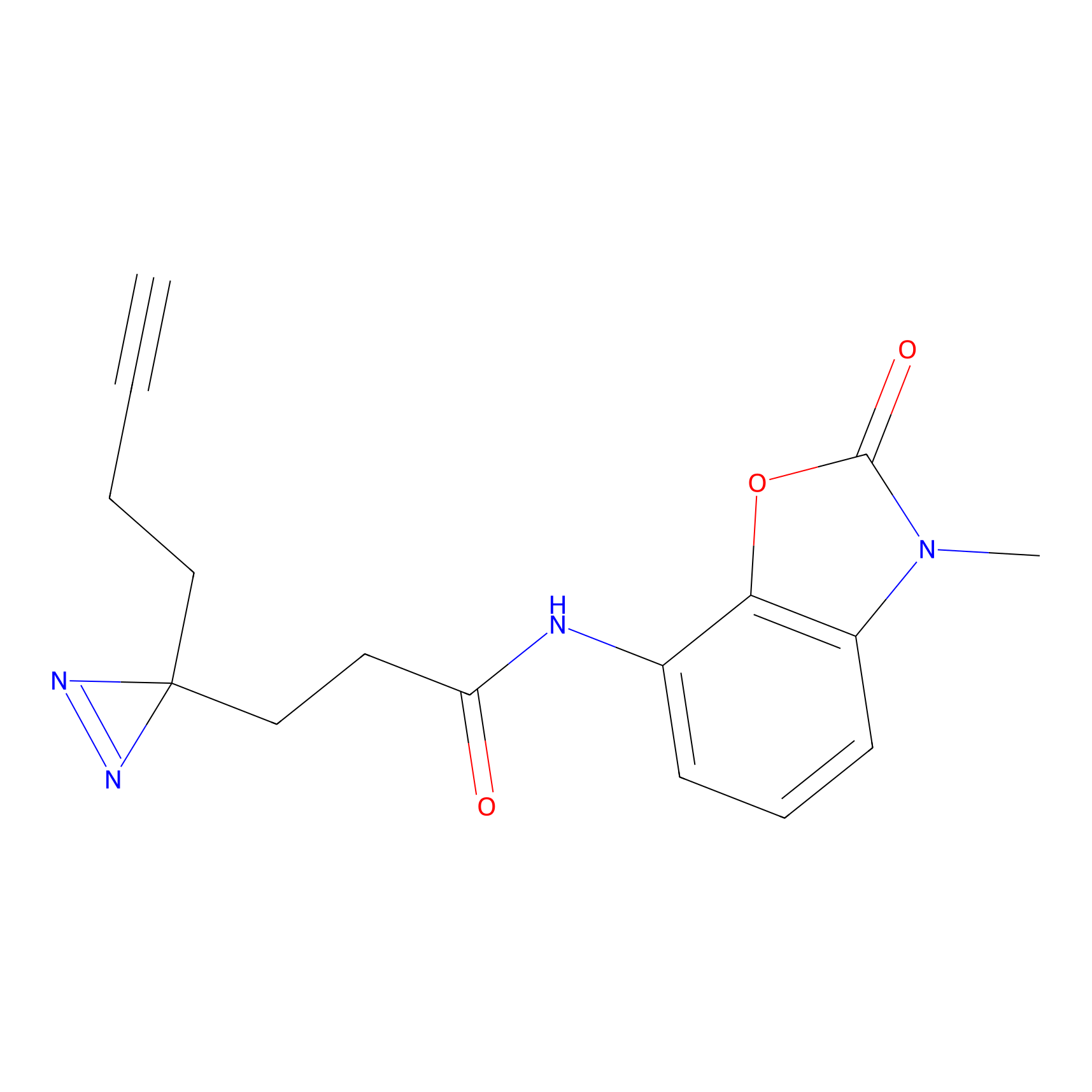 |
15.56 | LDD1912 | [6] | |
|
C278 Probe Info |
 |
53.45 | LDD1948 | [6] | |
|
C310 Probe Info |
 |
11.55 | LDD1977 | [6] | |
|
C355 Probe Info |
 |
30.06 | LDD2016 | [6] | |
|
C356 Probe Info |
 |
8.57 | LDD2017 | [6] | |
|
C362 Probe Info |
 |
34.06 | LDD2023 | [6] | |
|
C388 Probe Info |
 |
45.57 | LDD2047 | [6] | |
|
C403 Probe Info |
 |
18.51 | LDD2061 | [6] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References
