Details of the Target
General Information of Target
| Target ID | LDTP01532 | |||||
|---|---|---|---|---|---|---|
| Target Name | ZW10 interactor (ZWINT) | |||||
| Gene Name | ZWINT | |||||
| Gene ID | 11130 | |||||
| Synonyms |
ZW10 interactor; ZW10-interacting protein 1; Zwint-1 |
|||||
| 3D Structure | ||||||
| Sequence |
MEAAETEAEAAALEVLAEVAGILEPVGLQEEAELPAKILVEFVVDSQKKDKLLCSQLQVA
DFLQNILAQEDTAKGLDPLASEDTSRQKAIAAKEQWKELKATYREHVEAIKIGLTKALTQ MEEAQRKRTQLREAFEQLQAKKQMAMEKRRAVQNQWQLQQEKHLQHLAEVSAEVRERKTG TQQELDRVFQKLGNLKQQAEQERDKLQRYQTFLQLLYTLQGKLLFPEAEAEAENLPDDKP QQPTRPQEQSTGDTMGRDPGVSFKAVGLQPAGDVNLP |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function | Part of the MIS12 complex, which is required for kinetochore formation and spindle checkpoint activity. Required to target ZW10 to the kinetochore at prometaphase. | |||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| AN3CA | SNV: p.D50G; p.R208M | . | |||
| JHH7 | SNV: p.E5V | Alk-rapa Probe Info | |||
| MEWO | SNV: p.P78S | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STPyne Probe Info |
 |
K111(10.00) | LDD0277 | [1] | |
|
m-APA Probe Info |
 |
15.00 | LDD0403 | [2] | |
|
Curcusone 37 Probe Info |
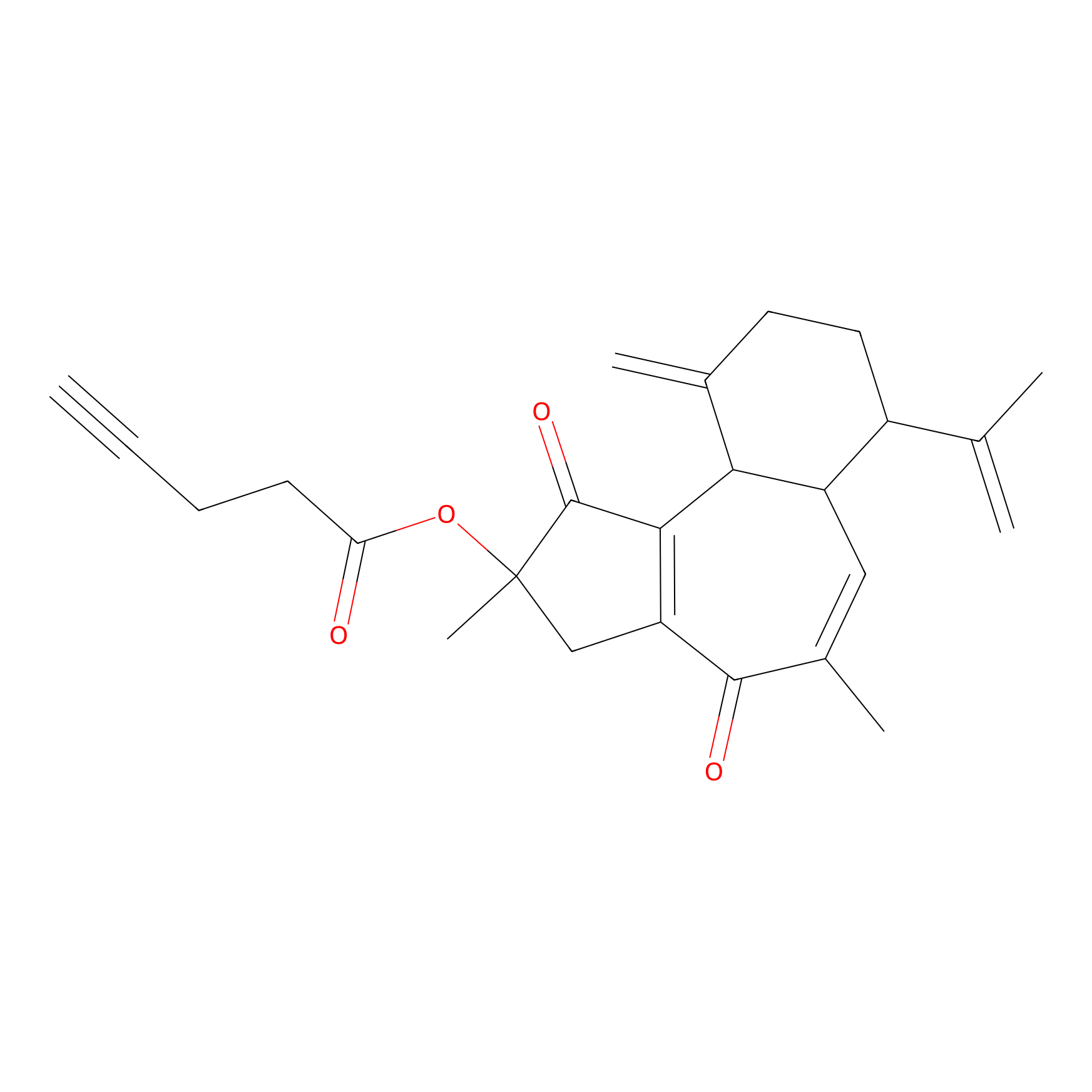 |
2.16 | LDD0188 | [3] | |
|
AHL-Pu-1 Probe Info |
 |
C54(4.70) | LDD0169 | [4] | |
|
DBIA Probe Info |
 |
C54(11.79) | LDD0209 | [5] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [6] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [7] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [6] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [8] | |
|
Compound 11 Probe Info |
 |
N.A. | LDD2213 | [9] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [10] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [11] | |
|
AOyne Probe Info |
 |
14.20 | LDD0443 | [12] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alk-rapa Probe Info |
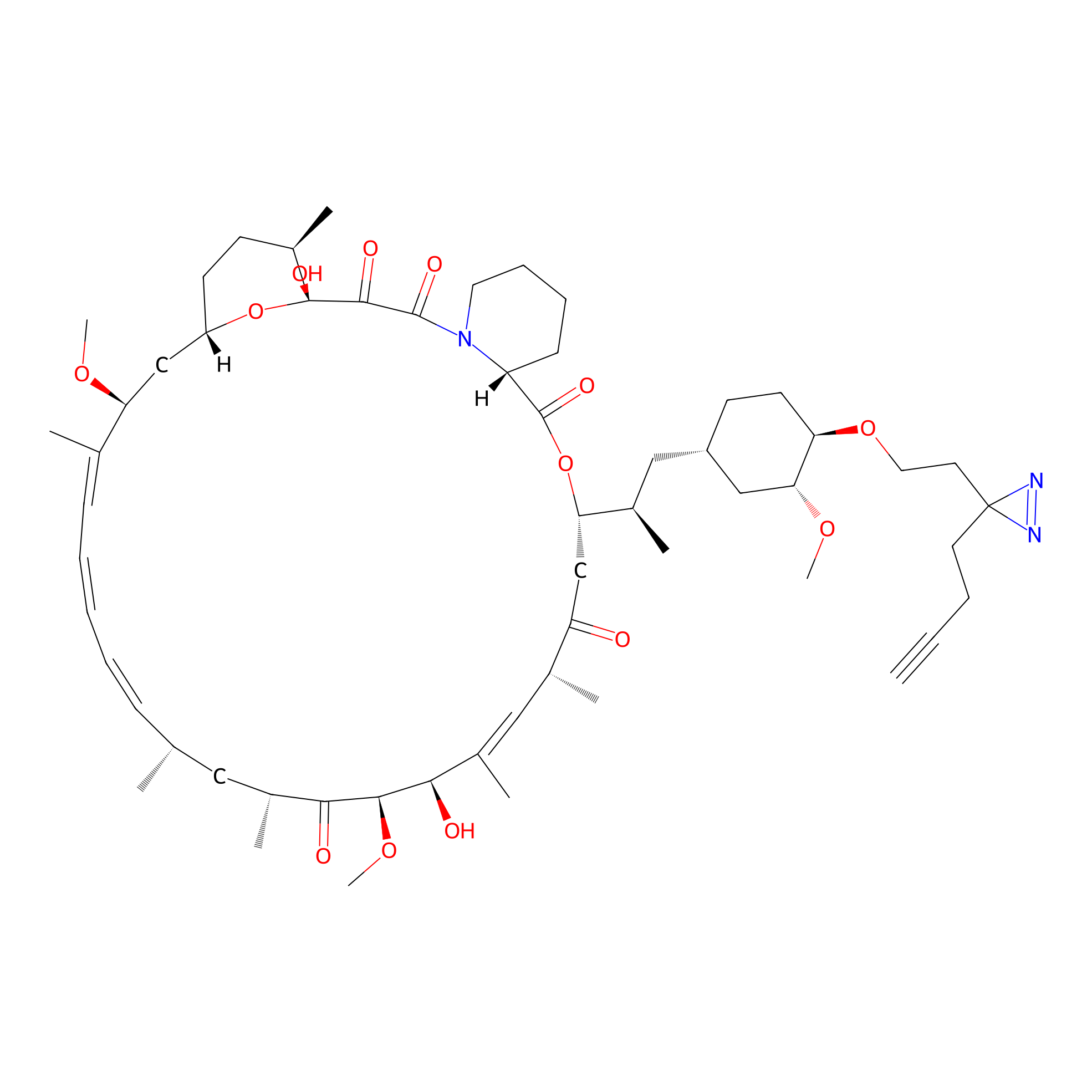 |
3.30 | LDD0213 | [13] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C54(4.70) | LDD0169 | [4] |
| LDCM0214 | AC1 | PaTu 8988t | C54(0.64) | LDD1093 | [14] |
| LDCM0230 | AC113 | PaTu 8988t | C54(1.07) | LDD1109 | [14] |
| LDCM0231 | AC114 | PaTu 8988t | C54(0.79) | LDD1110 | [14] |
| LDCM0232 | AC115 | PaTu 8988t | C54(1.06) | LDD1111 | [14] |
| LDCM0233 | AC116 | PaTu 8988t | C54(0.92) | LDD1112 | [14] |
| LDCM0234 | AC117 | PaTu 8988t | C54(1.29) | LDD1113 | [14] |
| LDCM0235 | AC118 | PaTu 8988t | C54(0.83) | LDD1114 | [14] |
| LDCM0236 | AC119 | PaTu 8988t | C54(0.92) | LDD1115 | [14] |
| LDCM0238 | AC120 | PaTu 8988t | C54(0.86) | LDD1117 | [14] |
| LDCM0239 | AC121 | PaTu 8988t | C54(0.82) | LDD1118 | [14] |
| LDCM0240 | AC122 | PaTu 8988t | C54(1.06) | LDD1119 | [14] |
| LDCM0241 | AC123 | PaTu 8988t | C54(1.05) | LDD1120 | [14] |
| LDCM0242 | AC124 | PaTu 8988t | C54(1.12) | LDD1121 | [14] |
| LDCM0243 | AC125 | PaTu 8988t | C54(0.91) | LDD1122 | [14] |
| LDCM0244 | AC126 | PaTu 8988t | C54(0.69) | LDD1123 | [14] |
| LDCM0245 | AC127 | PaTu 8988t | C54(0.68) | LDD1124 | [14] |
| LDCM0259 | AC14 | HEK-293T | C54(1.03) | LDD1512 | [15] |
| LDCM0279 | AC2 | PaTu 8988t | C54(1.12) | LDD1158 | [14] |
| LDCM0281 | AC21 | HEK-293T | C54(1.07) | LDD1520 | [15] |
| LDCM0282 | AC22 | HEK-293T | C54(1.13) | LDD1521 | [15] |
| LDCM0285 | AC25 | PaTu 8988t | C54(0.90) | LDD1164 | [14] |
| LDCM0286 | AC26 | PaTu 8988t | C54(0.96) | LDD1165 | [14] |
| LDCM0287 | AC27 | PaTu 8988t | C54(1.33) | LDD1166 | [14] |
| LDCM0288 | AC28 | PaTu 8988t | C54(1.42) | LDD1167 | [14] |
| LDCM0289 | AC29 | PaTu 8988t | C54(0.95) | LDD1168 | [14] |
| LDCM0290 | AC3 | PaTu 8988t | C54(1.07) | LDD1169 | [14] |
| LDCM0291 | AC30 | PaTu 8988t | C54(1.05) | LDD1170 | [14] |
| LDCM0292 | AC31 | PaTu 8988t | C54(1.04) | LDD1171 | [14] |
| LDCM0293 | AC32 | PaTu 8988t | C54(0.92) | LDD1172 | [14] |
| LDCM0294 | AC33 | PaTu 8988t | C54(1.27) | LDD1173 | [14] |
| LDCM0295 | AC34 | PaTu 8988t | C54(1.34) | LDD1174 | [14] |
| LDCM0296 | AC35 | PaTu 8988t | C54(1.10) | LDD1175 | [14] |
| LDCM0297 | AC36 | PaTu 8988t | C54(1.17) | LDD1176 | [14] |
| LDCM0298 | AC37 | PaTu 8988t | C54(1.57) | LDD1177 | [14] |
| LDCM0299 | AC38 | PaTu 8988t | C54(1.21) | LDD1178 | [14] |
| LDCM0300 | AC39 | PaTu 8988t | C54(1.41) | LDD1179 | [14] |
| LDCM0301 | AC4 | PaTu 8988t | C54(1.05) | LDD1180 | [14] |
| LDCM0302 | AC40 | PaTu 8988t | C54(1.26) | LDD1181 | [14] |
| LDCM0303 | AC41 | PaTu 8988t | C54(0.96) | LDD1182 | [14] |
| LDCM0304 | AC42 | PaTu 8988t | C54(0.99) | LDD1183 | [14] |
| LDCM0305 | AC43 | PaTu 8988t | C54(1.09) | LDD1184 | [14] |
| LDCM0306 | AC44 | PaTu 8988t | C54(1.12) | LDD1185 | [14] |
| LDCM0307 | AC45 | PaTu 8988t | C54(1.14) | LDD1186 | [14] |
| LDCM0308 | AC46 | PaTu 8988t | C54(1.05) | LDD1187 | [14] |
| LDCM0309 | AC47 | PaTu 8988t | C54(1.15) | LDD1188 | [14] |
| LDCM0310 | AC48 | PaTu 8988t | C54(1.55) | LDD1189 | [14] |
| LDCM0311 | AC49 | PaTu 8988t | C54(1.52) | LDD1190 | [14] |
| LDCM0312 | AC5 | PaTu 8988t | C54(1.32) | LDD1191 | [14] |
| LDCM0313 | AC50 | PaTu 8988t | C54(1.65) | LDD1192 | [14] |
| LDCM0314 | AC51 | PaTu 8988t | C54(0.97) | LDD1193 | [14] |
| LDCM0315 | AC52 | PaTu 8988t | C54(1.44) | LDD1194 | [14] |
| LDCM0316 | AC53 | PaTu 8988t | C54(2.09) | LDD1195 | [14] |
| LDCM0317 | AC54 | PaTu 8988t | C54(1.56) | LDD1196 | [14] |
| LDCM0318 | AC55 | PaTu 8988t | C54(1.56) | LDD1197 | [14] |
| LDCM0319 | AC56 | PaTu 8988t | C54(1.40) | LDD1198 | [14] |
| LDCM0323 | AC6 | HEK-293T | C54(1.01) | LDD1562 | [15] |
| LDCM0325 | AC61 | HEK-293T | C54(1.09) | LDD1564 | [15] |
| LDCM0326 | AC62 | HEK-293T | C54(1.08) | LDD1565 | [15] |
| LDCM0248 | AKOS034007472 | HEK-293T | C54(1.08) | LDD1511 | [15] |
| LDCM0156 | Aniline | NCI-H1299 | 15.00 | LDD0403 | [2] |
| LDCM0630 | CCW28-3 | 231MFP | C54(1.91) | LDD2214 | [16] |
| LDCM0368 | CL10 | HEK-293T | C54(2.41) | LDD1572 | [15] |
| LDCM0369 | CL100 | PaTu 8988t | C54(0.88) | LDD1248 | [14] |
| LDCM0372 | CL103 | HEK-293T | C54(0.99) | LDD1576 | [15] |
| LDCM0376 | CL107 | HEK-293T | C54(0.89) | LDD1580 | [15] |
| LDCM0381 | CL111 | HEK-293T | C54(1.08) | LDD1585 | [15] |
| LDCM0382 | CL112 | PaTu 8988t | C54(1.06) | LDD1261 | [14] |
| LDCM0383 | CL113 | PaTu 8988t | C54(1.05) | LDD1262 | [14] |
| LDCM0384 | CL114 | PaTu 8988t | C54(1.01) | LDD1263 | [14] |
| LDCM0385 | CL115 | PaTu 8988t | C54(1.26) | LDD1264 | [14] |
| LDCM0386 | CL116 | PaTu 8988t | C54(0.93) | LDD1265 | [14] |
| LDCM0387 | CL117 | PaTu 8988t | C54(0.93) | LDD1266 | [14] |
| LDCM0388 | CL118 | PaTu 8988t | C54(0.93) | LDD1267 | [14] |
| LDCM0389 | CL119 | PaTu 8988t | C54(1.00) | LDD1268 | [14] |
| LDCM0391 | CL120 | PaTu 8988t | C54(0.89) | LDD1270 | [14] |
| LDCM0392 | CL121 | PaTu 8988t | C54(1.61) | LDD1271 | [14] |
| LDCM0393 | CL122 | PaTu 8988t | C54(1.51) | LDD1272 | [14] |
| LDCM0394 | CL123 | PaTu 8988t | C54(2.58) | LDD1273 | [14] |
| LDCM0395 | CL124 | PaTu 8988t | C54(1.27) | LDD1274 | [14] |
| LDCM0398 | CL127 | HEK-293T | C54(1.05) | LDD1602 | [15] |
| LDCM0402 | CL15 | HEK-293T | C54(1.51) | LDD1606 | [15] |
| LDCM0409 | CL21 | HEK-293T | C54(1.59) | LDD1613 | [15] |
| LDCM0410 | CL22 | HEK-293T | C54(0.96) | LDD1614 | [15] |
| LDCM0415 | CL27 | HEK-293T | C54(0.96) | LDD1619 | [15] |
| LDCM0418 | CL3 | HEK-293T | C54(1.20) | LDD1622 | [15] |
| LDCM0422 | CL33 | HEK-293T | C54(1.98) | LDD1626 | [15] |
| LDCM0423 | CL34 | HEK-293T | C54(1.10) | LDD1627 | [15] |
| LDCM0428 | CL39 | HEK-293T | C54(1.20) | LDD1632 | [15] |
| LDCM0435 | CL45 | HEK-293T | C54(1.39) | LDD1639 | [15] |
| LDCM0436 | CL46 | HEK-293T | C54(1.07) | LDD1640 | [15] |
| LDCM0448 | CL57 | HEK-293T | C54(1.38) | LDD1651 | [15] |
| LDCM0449 | CL58 | HEK-293T | C54(1.06) | LDD1652 | [15] |
| LDCM0453 | CL61 | PaTu 8988t | C54(1.52) | LDD1332 | [14] |
| LDCM0454 | CL62 | PaTu 8988t | C54(1.86) | LDD1333 | [14] |
| LDCM0455 | CL63 | PaTu 8988t | C54(1.03) | LDD1334 | [14] |
| LDCM0456 | CL64 | PaTu 8988t | C54(0.99) | LDD1335 | [14] |
| LDCM0457 | CL65 | PaTu 8988t | C54(1.07) | LDD1336 | [14] |
| LDCM0458 | CL66 | PaTu 8988t | C54(0.92) | LDD1337 | [14] |
| LDCM0459 | CL67 | PaTu 8988t | C54(0.74) | LDD1338 | [14] |
| LDCM0460 | CL68 | PaTu 8988t | C54(1.07) | LDD1339 | [14] |
| LDCM0461 | CL69 | PaTu 8988t | C54(1.35) | LDD1340 | [14] |
| LDCM0463 | CL70 | PaTu 8988t | C54(0.86) | LDD1342 | [14] |
| LDCM0464 | CL71 | PaTu 8988t | C54(1.03) | LDD1343 | [14] |
| LDCM0465 | CL72 | PaTu 8988t | C54(0.93) | LDD1344 | [14] |
| LDCM0466 | CL73 | PaTu 8988t | C54(1.15) | LDD1345 | [14] |
| LDCM0467 | CL74 | PaTu 8988t | C54(0.95) | LDD1346 | [14] |
| LDCM0469 | CL76 | PaTu 8988t | C54(1.10) | LDD1348 | [14] |
| LDCM0470 | CL77 | PaTu 8988t | C54(1.29) | LDD1349 | [14] |
| LDCM0471 | CL78 | PaTu 8988t | C54(1.28) | LDD1350 | [14] |
| LDCM0472 | CL79 | PaTu 8988t | C54(1.60) | LDD1351 | [14] |
| LDCM0474 | CL80 | PaTu 8988t | C54(1.28) | LDD1353 | [14] |
| LDCM0475 | CL81 | PaTu 8988t | C54(1.25) | LDD1354 | [14] |
| LDCM0476 | CL82 | PaTu 8988t | C54(1.22) | LDD1355 | [14] |
| LDCM0477 | CL83 | PaTu 8988t | C54(1.41) | LDD1356 | [14] |
| LDCM0478 | CL84 | PaTu 8988t | C54(1.29) | LDD1357 | [14] |
| LDCM0479 | CL85 | PaTu 8988t | C54(1.56) | LDD1358 | [14] |
| LDCM0480 | CL86 | PaTu 8988t | C54(1.32) | LDD1359 | [14] |
| LDCM0481 | CL87 | PaTu 8988t | C54(1.54) | LDD1360 | [14] |
| LDCM0482 | CL88 | PaTu 8988t | C54(1.53) | LDD1361 | [14] |
| LDCM0483 | CL89 | PaTu 8988t | C54(1.45) | LDD1362 | [14] |
| LDCM0484 | CL9 | HEK-293T | C54(1.22) | LDD1687 | [15] |
| LDCM0485 | CL90 | PaTu 8988t | C54(1.54) | LDD1364 | [14] |
| LDCM0486 | CL91 | PaTu 8988t | C54(0.84) | LDD1365 | [14] |
| LDCM0487 | CL92 | PaTu 8988t | C54(0.65) | LDD1366 | [14] |
| LDCM0488 | CL93 | PaTu 8988t | C54(0.80) | LDD1367 | [14] |
| LDCM0489 | CL94 | PaTu 8988t | C54(1.10) | LDD1368 | [14] |
| LDCM0490 | CL95 | PaTu 8988t | C54(0.91) | LDD1369 | [14] |
| LDCM0491 | CL96 | PaTu 8988t | C54(0.74) | LDD1370 | [14] |
| LDCM0492 | CL97 | PaTu 8988t | C54(0.73) | LDD1371 | [14] |
| LDCM0493 | CL98 | PaTu 8988t | C54(0.94) | LDD1372 | [14] |
| LDCM0494 | CL99 | PaTu 8988t | C54(0.75) | LDD1373 | [14] |
| LDCM0033 | Curcusone1d | MCF-7 | 2.16 | LDD0188 | [3] |
| LDCM0495 | E2913 | HEK-293T | C54(1.02) | LDD1698 | [15] |
| LDCM0572 | Fragment10 | Ramos | C54(1.03) | LDD1390 | [17] |
| LDCM0573 | Fragment11 | MDA-MB-231 | C54(3.26) | LDD1391 | [17] |
| LDCM0574 | Fragment12 | MDA-MB-231 | C54(20.00) | LDD1393 | [17] |
| LDCM0575 | Fragment13 | Ramos | C54(0.91) | LDD1396 | [17] |
| LDCM0576 | Fragment14 | MDA-MB-231 | C54(2.45) | LDD1397 | [17] |
| LDCM0579 | Fragment20 | Ramos | C54(2.41) | LDD1403 | [17] |
| LDCM0580 | Fragment21 | MDA-MB-231 | C54(1.78) | LDD1404 | [17] |
| LDCM0584 | Fragment25 | MDA-MB-231 | C54(1.10) | LDD1411 | [17] |
| LDCM0585 | Fragment26 | Ramos | C54(0.86) | LDD1412 | [17] |
| LDCM0578 | Fragment27 | MDA-MB-231 | C54(0.76) | LDD1401 | [17] |
| LDCM0586 | Fragment28 | Ramos | C54(1.07) | LDD1416 | [17] |
| LDCM0587 | Fragment29 | MDA-MB-231 | C54(1.05) | LDD1417 | [17] |
| LDCM0588 | Fragment30 | MDA-MB-231 | C54(1.86) | LDD1419 | [17] |
| LDCM0590 | Fragment32 | MDA-MB-231 | C54(4.52) | LDD1423 | [17] |
| LDCM0468 | Fragment33 | PaTu 8988t | C54(1.00) | LDD1347 | [14] |
| LDCM0592 | Fragment34 | MDA-MB-231 | C54(0.76) | LDD1427 | [17] |
| LDCM0593 | Fragment35 | Ramos | C54(1.16) | LDD1430 | [17] |
| LDCM0595 | Fragment37 | Ramos | C54(1.05) | LDD1432 | [17] |
| LDCM0596 | Fragment38 | MDA-MB-231 | C54(1.11) | LDD1433 | [17] |
| LDCM0566 | Fragment4 | MDA-MB-231 | C54(3.35) | LDD1378 | [17] |
| LDCM0599 | Fragment41 | Ramos | C54(1.45) | LDD1439 | [17] |
| LDCM0600 | Fragment42 | Ramos | C54(1.06) | LDD1440 | [17] |
| LDCM0601 | Fragment43 | MDA-MB-231 | C54(2.79) | LDD1441 | [17] |
| LDCM0602 | Fragment44 | MDA-MB-231 | C54(0.96) | LDD1443 | [17] |
| LDCM0607 | Fragment49 | MDA-MB-231 | C54(4.60) | LDD1448 | [17] |
| LDCM0427 | Fragment51 | Ramos | C54(0.64) | LDD1451 | [17] |
| LDCM0610 | Fragment52 | MDA-MB-231 | C54(1.26) | LDD1452 | [17] |
| LDCM0611 | Fragment53 | MDA-MB-231 | C54(0.98) | LDD1454 | [17] |
| LDCM0614 | Fragment56 | MDA-MB-231 | C54(1.05) | LDD1458 | [17] |
| LDCM0569 | Fragment7 | MDA-MB-231 | C54(1.56) | LDD1383 | [17] |
| LDCM0570 | Fragment8 | Ramos | C54(1.25) | LDD1386 | [17] |
| LDCM0571 | Fragment9 | MDA-MB-231 | C54(1.91) | LDD1387 | [17] |
| LDCM0022 | KB02 | MDA-MB-231 | C54(20.00) | LDD1374 | [17] |
| LDCM0023 | KB03 | Jurkat | C54(11.79) | LDD0209 | [5] |
| LDCM0024 | KB05 | NCI-H1155 | C54(4.18) | LDD3342 | [18] |
| LDCM0090 | Rapamycin | JHH-7 | 3.30 | LDD0213 | [13] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Tumor susceptibility gene 101 protein (TSG101) | Ubiquitin-conjugating enzyme family | Q99816 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Beclin-1 (BECN1) | Beclin family | Q14457 | |||
| Nucleoporin p54 (NUP54) | NUP54 family | Q7Z3B4 | |||
| Nucleoporin p58/p45 (NUP58) | NUP58 family | Q9BVL2 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Krueppel-like factor 11 (KLF11) | Sp1 C2H2-type zinc-finger protein family | O14901 | |||
| Mesogenin-1 (MSGN1) | . | A6NI15 | |||
Other
References
