Details of the Target
General Information of Target
| Target ID | LDTP11328 | |||||
|---|---|---|---|---|---|---|
| Target Name | Splicing factor 3B subunit 5 (SF3B5) | |||||
| Gene Name | SF3B5 | |||||
| Gene ID | 83443 | |||||
| Synonyms |
SF3B10; Splicing factor 3B subunit 5; SF3b5; Pre-mRNA-splicing factor SF3b 10 kDa subunit |
|||||
| 3D Structure | ||||||
| Sequence |
MLSRPKPGESEVDLLHFQSQFLAAGAAPAVQLVKKGNRGGGDANSDRPPLQDHRDVVMLD
NLPDLPPALVPSPPKRARPSPGHCLPEDEDPEERLRRHDQHITAVLTKIIERDTSSVAVN LPVPSGVAFPAVFLRSRDTQGKSATSGKRSIFAQEIAARRIAEAKGPSVGEVVPNVGPPE GAVTCETPTPRNQGCQLPGSSHSFQGPNLVTGKGLRDQEAEQEAQTIHEENIARLQAMAP EEILQEQQRLLAQLDPSLVAFLRSHSHTQEQTGETASEEQRPGGPSANVTKEEPLMSAFA SEPRKRDKLEPEAPALALPVTPQKEWLHMDTVELEKLHWTQDLPPVRRQQTQERMQARFS LQGELLAPDVDLPTHLGLHHHGEEAERAGYSLQELFHLTRSQVSQQRALALHVLAQVISR AQAGEFGDRLAGSVLSLLLDAGFLFLLRFSLDDRVDGVIATAIRALRALLVAPGDEELLD STFSWYHGALTFPLMPSQEDKEDEDEDEECPAGKAKRKSPEEESRPPPDLARHDVIKGLL ATSLLPRLRYVLEVTYPGPAVVLDILAVLIRLARHSLESATRVLECPRLIETIVREFLPT SWSPVGAGPTPSLYKVPCATAMKLLRVLASAGRNIAARLLSSFDLRSRLCRIIAEAPQEL ALPPEEAEMLSTEALRLWAVAASYGQGGYLYRELYPVLMRALQVVPRELSTHPPQPLSMQ RIASLLTLLTQLTLAAGSTPAETISDSAEASLSATPSLVTWTQVSGLQPLVEPCLRQTLK LLSRPEMWRAVGPVPVACLLFLGAYYQAWSQQPSSCPEDWLQDMQRLSEELLLPLLSQPT LGSLWDSLRHCSLLCNPLSCVPALEAPPSLVSLGCSGGCPRLSLAGSASPFPFLTALLSL LNTLAQIHKGLCGQLAAILAAPGLQNYFLQCVAPGAAPHLTPFSAWALRHEYHLQYLALA LAQKAAALQPLPATHAALYHGMALALLSRLLPGSEYLTHELLLSCVFRLEFLPERTSGGP EAADFSDQLSLGSSRVPRCGQGTLLAQACQDLPSIRNCYLTHCSPARASLLASQALHRGE LQRVPTLLLPMPTEPLLPTDWPFLPLIRLYHRASDTPSGLSPTDTMGTAMRVLQWVLVLE SWRPQALWAVPPAARLARLMCVFLVDSELFRESPVQHLVAALLAQLCQPQVLPNLNLDCR LPGLTSFPDLYANFLDHFEAVSFGDHLFGALVLLPLQRRFSVTLRLALFGEHVGALRALS LPLTQLPVSLECYTVPPEDNLALLQLYFRTLVTGALRPRWCPVLYAVAVAHVNSFIFSQD PQSSDEVKAARRSMLQKTWLLADEGLRQHLLHYKLPNSTLPEGFELYSQLPPLRQHYLQR LTSTVLQNGVSET |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
SF3B5 family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Component of the 17S U2 SnRNP complex of the spliceosome, a large ribonucleoprotein complex that removes introns from transcribed pre-mRNAs. The 17S U2 SnRNP complex (1) directly participates in early spliceosome assembly and (2) mediates recognition of the intron branch site during pre-mRNA splicing by promoting the selection of the pre-mRNA branch-site adenosine, the nucleophile for the first step of splicing. Within the 17S U2 SnRNP complex, SF3B4 is part of the SF3B subcomplex, which is required for 'A' complex assembly formed by the stable binding of U2 snRNP to the branchpoint sequence in pre-mRNA. Sequence independent binding of SF3A and SF3B subcomplexes upstream of the branch site is essential, it may anchor U2 snRNP to the pre-mRNA. Also acts as a component of the minor spliceosome, which is involved in the splicing of U12-type introns in pre-mRNAs.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y43(9.85); Y5(9.53); Y18(6.13) | LDD0257 | [1] | |
|
TH216 Probe Info |
 |
Y52(20.00); Y18(9.99) | LDD0259 | [1] | |
|
IPM Probe Info |
 |
N.A. | LDD0241 | [2] | |
|
Probe 1 Probe Info |
 |
Y5(12.63); Y18(19.20) | LDD3495 | [3] | |
|
m-APA Probe Info |
 |
12.07 | LDD0403 | [4] | |
|
BTD Probe Info |
 |
C76(0.96) | LDD2101 | [5] | |
|
HHS-475 Probe Info |
 |
Y18(1.76); Y43(1.84) | LDD0264 | [6] | |
|
DBIA Probe Info |
 |
C76(1.82) | LDD0531 | [7] | |
|
5E-2FA Probe Info |
 |
H23(0.00); H13(0.00); H8(0.00) | LDD2235 | [8] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [9] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [10] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0163 | [11] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [12] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [10] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [13] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [13] | |
|
PF-06672131 Probe Info |
 |
N.A. | LDD0152 | [14] | |
|
SF Probe Info |
 |
Y18(0.00); Y5(0.00) | LDD0028 | [15] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [16] | |
|
Phosphinate-6 Probe Info |
 |
C76(0.00); C41(0.00) | LDD0018 | [17] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [18] | |
|
Acrolein Probe Info |
 |
H8(0.00); H13(0.00); C76(0.00) | LDD0217 | [19] | |
|
Crotonaldehyde Probe Info |
 |
C76(0.00); H8(0.00); H23(0.00) | LDD0219 | [19] | |
|
Methacrolein Probe Info |
 |
C76(0.00); H8(0.00) | LDD0218 | [19] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [20] | |
|
NAIA_5 Probe Info |
 |
C41(0.00); C76(0.00) | LDD2223 | [21] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [22] | |
|
HHS-482 Probe Info |
 |
Y18(0.56) | LDD2239 | [23] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C179 Probe Info |
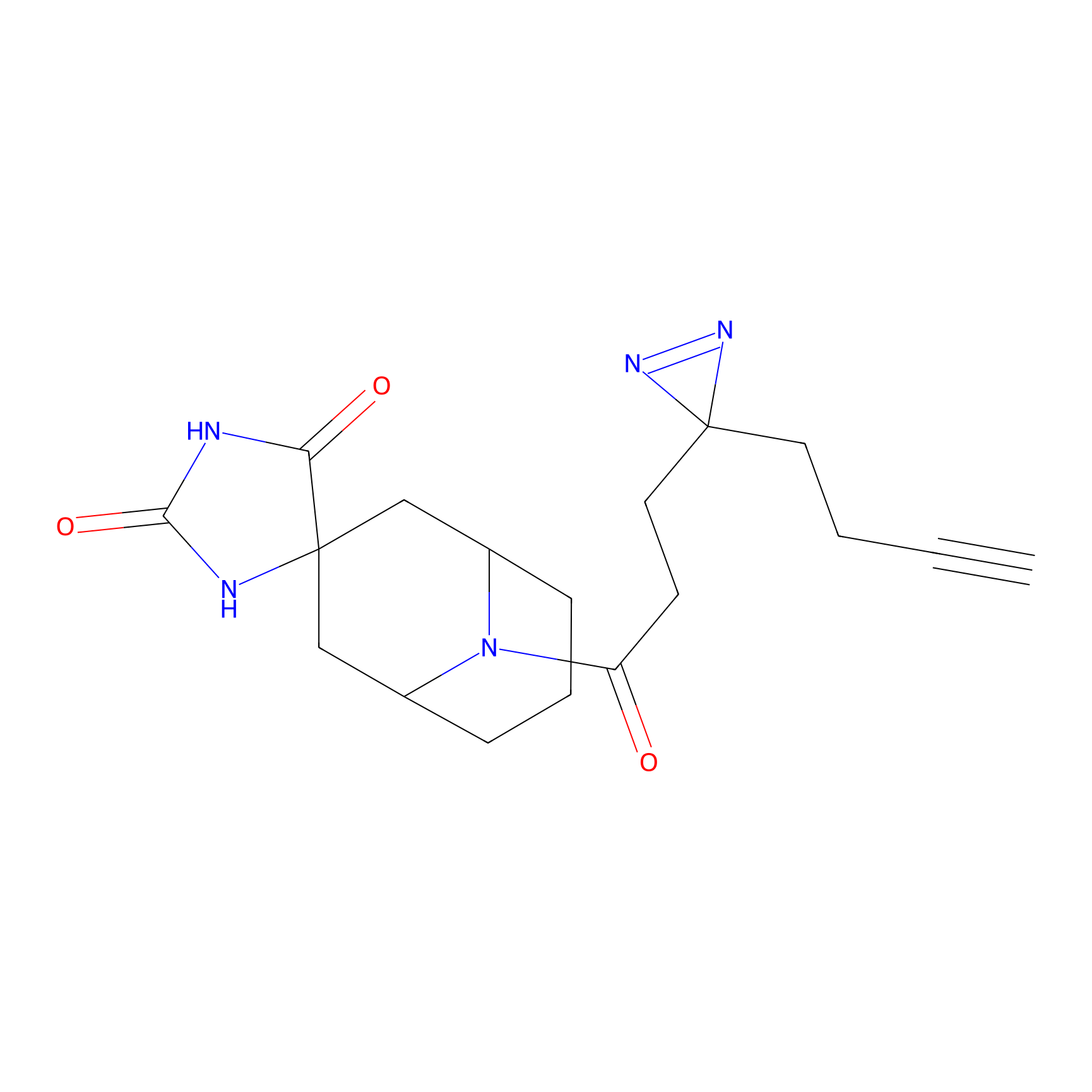 |
7.31 | LDD1858 | [24] | |
|
C277 Probe Info |
 |
9.58 | LDD1947 | [24] | |
|
C343 Probe Info |
 |
19.84 | LDD2005 | [24] | |
|
C344 Probe Info |
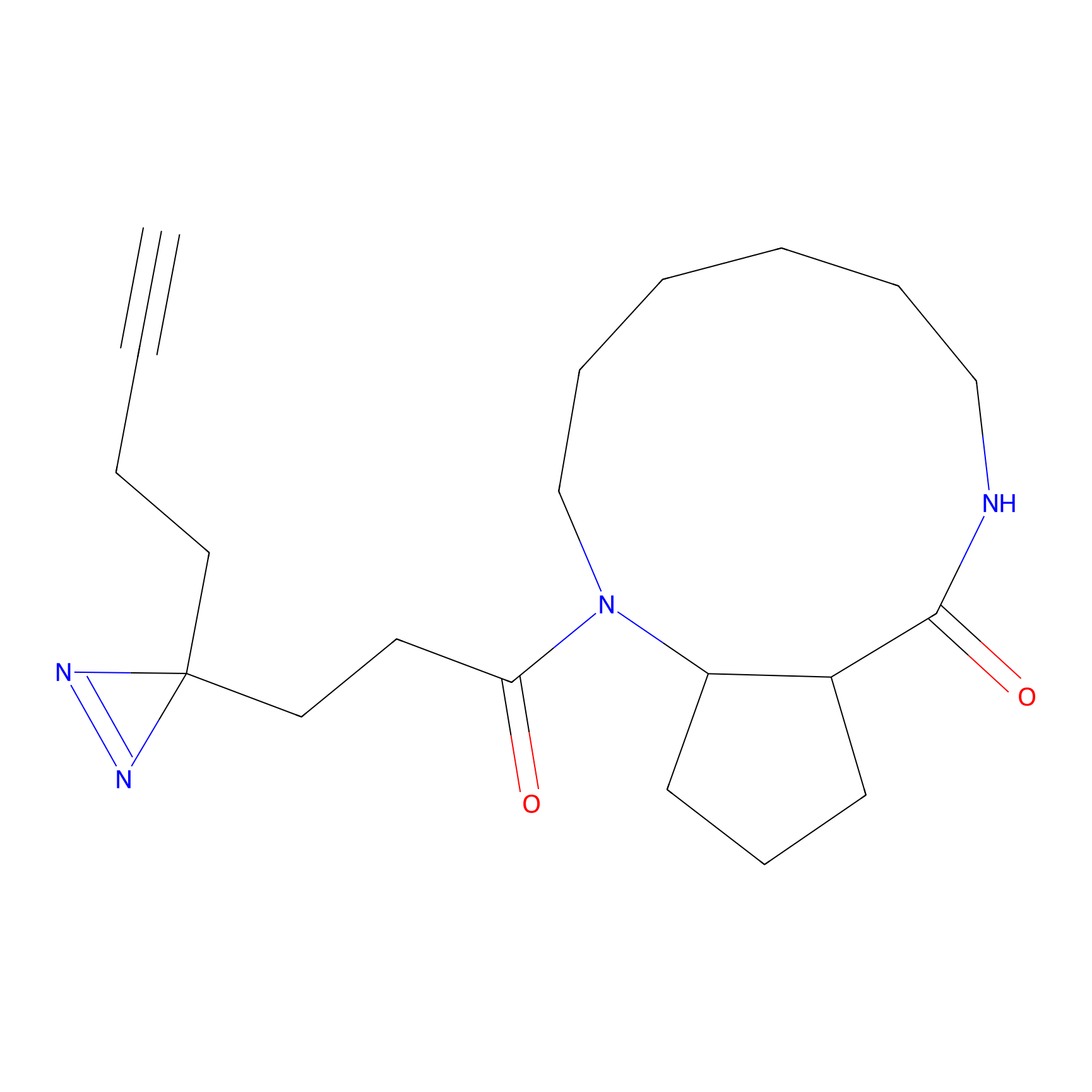 |
6.50 | LDD2006 | [24] | |
|
C346 Probe Info |
 |
12.91 | LDD2007 | [24] | |
|
C347 Probe Info |
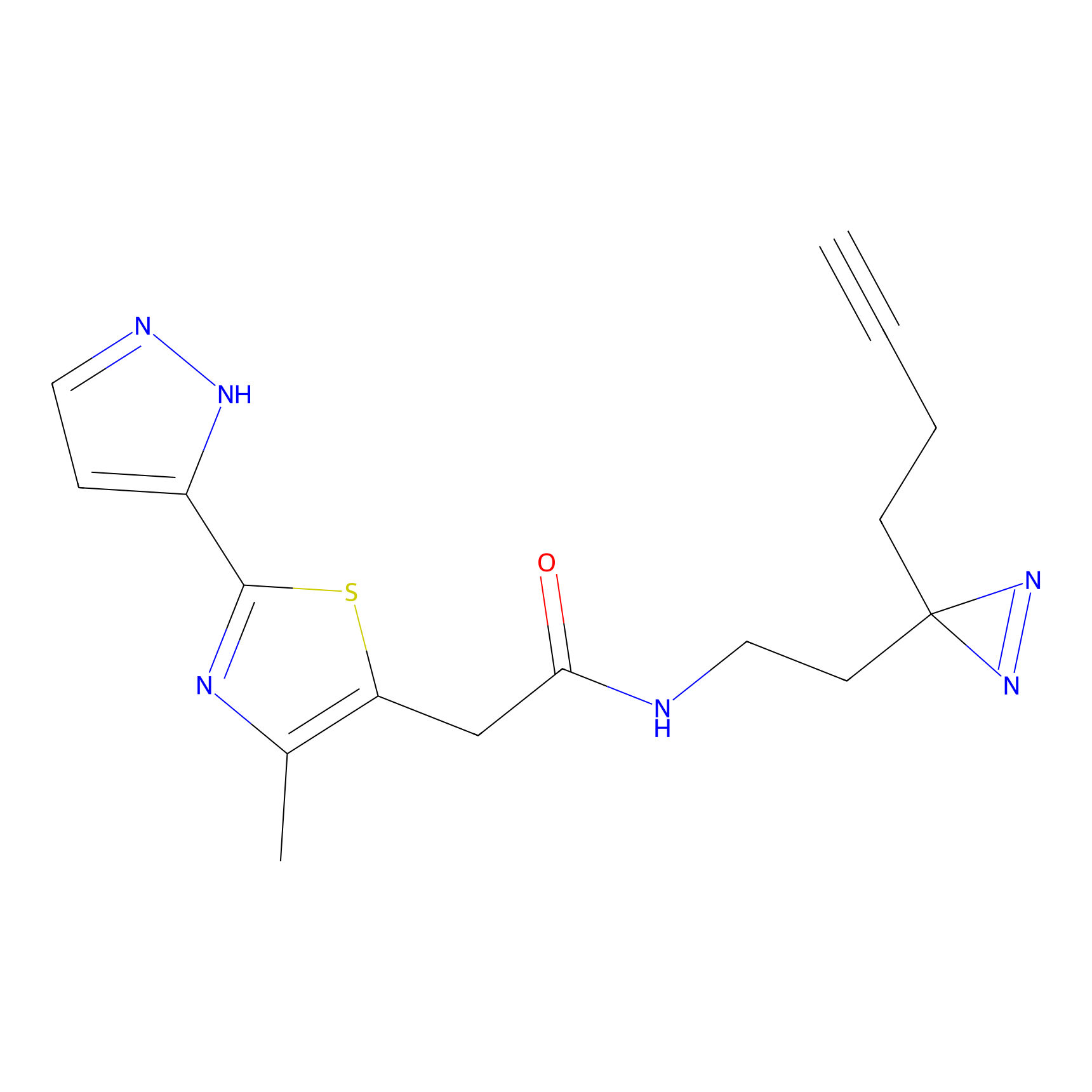 |
9.25 | LDD2008 | [24] | |
|
C399 Probe Info |
 |
6.73 | LDD2058 | [24] | |
|
FFF probe11 Probe Info |
 |
5.37 | LDD0471 | [25] | |
|
FFF probe2 Probe Info |
 |
11.32 | LDD0463 | [25] | |
|
OEA-DA Probe Info |
 |
3.80 | LDD0046 | [26] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C76(0.49) | LDD2112 | [5] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C76(0.96) | LDD2117 | [5] |
| LDCM0214 | AC1 | HCT 116 | C76(1.82) | LDD0531 | [7] |
| LDCM0215 | AC10 | HCT 116 | C76(1.34) | LDD0532 | [7] |
| LDCM0216 | AC100 | HCT 116 | C76(0.72) | LDD0533 | [7] |
| LDCM0217 | AC101 | HCT 116 | C76(0.68) | LDD0534 | [7] |
| LDCM0218 | AC102 | HCT 116 | C76(0.77) | LDD0535 | [7] |
| LDCM0219 | AC103 | HCT 116 | C76(0.62) | LDD0536 | [7] |
| LDCM0220 | AC104 | HCT 116 | C76(0.70) | LDD0537 | [7] |
| LDCM0221 | AC105 | HCT 116 | C76(0.76) | LDD0538 | [7] |
| LDCM0222 | AC106 | HCT 116 | C76(0.65) | LDD0539 | [7] |
| LDCM0223 | AC107 | HCT 116 | C76(0.70) | LDD0540 | [7] |
| LDCM0224 | AC108 | HCT 116 | C76(0.72) | LDD0541 | [7] |
| LDCM0225 | AC109 | HCT 116 | C76(0.77) | LDD0542 | [7] |
| LDCM0226 | AC11 | HCT 116 | C76(1.58) | LDD0543 | [7] |
| LDCM0227 | AC110 | HCT 116 | C76(0.81) | LDD0544 | [7] |
| LDCM0228 | AC111 | HCT 116 | C76(0.65) | LDD0545 | [7] |
| LDCM0229 | AC112 | HCT 116 | C76(0.61) | LDD0546 | [7] |
| LDCM0230 | AC113 | HCT 116 | C76(0.98) | LDD0547 | [7] |
| LDCM0231 | AC114 | HCT 116 | C76(0.91) | LDD0548 | [7] |
| LDCM0232 | AC115 | HCT 116 | C76(0.95) | LDD0549 | [7] |
| LDCM0233 | AC116 | HCT 116 | C76(1.03) | LDD0550 | [7] |
| LDCM0234 | AC117 | HCT 116 | C76(0.96) | LDD0551 | [7] |
| LDCM0235 | AC118 | HCT 116 | C76(1.04) | LDD0552 | [7] |
| LDCM0236 | AC119 | HCT 116 | C76(0.98) | LDD0553 | [7] |
| LDCM0237 | AC12 | HCT 116 | C76(1.04) | LDD0554 | [7] |
| LDCM0238 | AC120 | HCT 116 | C76(0.99) | LDD0555 | [7] |
| LDCM0239 | AC121 | HCT 116 | C76(0.96) | LDD0556 | [7] |
| LDCM0240 | AC122 | HCT 116 | C76(0.93) | LDD0557 | [7] |
| LDCM0241 | AC123 | HCT 116 | C76(0.91) | LDD0558 | [7] |
| LDCM0242 | AC124 | HCT 116 | C76(0.86) | LDD0559 | [7] |
| LDCM0243 | AC125 | HCT 116 | C76(0.86) | LDD0560 | [7] |
| LDCM0244 | AC126 | HCT 116 | C76(0.89) | LDD0561 | [7] |
| LDCM0245 | AC127 | HCT 116 | C76(0.83) | LDD0562 | [7] |
| LDCM0246 | AC128 | HCT 116 | C76(1.51) | LDD0563 | [7] |
| LDCM0247 | AC129 | HCT 116 | C76(1.26) | LDD0564 | [7] |
| LDCM0249 | AC130 | HCT 116 | C76(1.12) | LDD0566 | [7] |
| LDCM0250 | AC131 | HCT 116 | C76(1.00) | LDD0567 | [7] |
| LDCM0251 | AC132 | HCT 116 | C76(0.90) | LDD0568 | [7] |
| LDCM0252 | AC133 | HCT 116 | C76(1.09) | LDD0569 | [7] |
| LDCM0253 | AC134 | HCT 116 | C76(0.74) | LDD0570 | [7] |
| LDCM0254 | AC135 | HCT 116 | C76(0.90) | LDD0571 | [7] |
| LDCM0255 | AC136 | HCT 116 | C76(0.81) | LDD0572 | [7] |
| LDCM0256 | AC137 | HCT 116 | C76(0.81) | LDD0573 | [7] |
| LDCM0257 | AC138 | HCT 116 | C76(0.72) | LDD0574 | [7] |
| LDCM0258 | AC139 | HCT 116 | C76(0.93) | LDD0575 | [7] |
| LDCM0259 | AC14 | HCT 116 | C76(1.57) | LDD0576 | [7] |
| LDCM0260 | AC140 | HCT 116 | C76(0.83) | LDD0577 | [7] |
| LDCM0261 | AC141 | HCT 116 | C76(0.86) | LDD0578 | [7] |
| LDCM0262 | AC142 | HCT 116 | C76(0.94) | LDD0579 | [7] |
| LDCM0263 | AC143 | HCT 116 | C76(0.70) | LDD0580 | [7] |
| LDCM0264 | AC144 | HCT 116 | C76(0.82) | LDD0581 | [7] |
| LDCM0265 | AC145 | HCT 116 | C76(0.91) | LDD0582 | [7] |
| LDCM0266 | AC146 | HCT 116 | C76(0.76) | LDD0583 | [7] |
| LDCM0267 | AC147 | HCT 116 | C76(0.95) | LDD0584 | [7] |
| LDCM0268 | AC148 | HCT 116 | C76(0.87) | LDD0585 | [7] |
| LDCM0269 | AC149 | HCT 116 | C76(0.79) | LDD0586 | [7] |
| LDCM0270 | AC15 | HCT 116 | C76(1.04) | LDD0587 | [7] |
| LDCM0271 | AC150 | HCT 116 | C76(0.79) | LDD0588 | [7] |
| LDCM0272 | AC151 | HCT 116 | C76(0.92) | LDD0589 | [7] |
| LDCM0273 | AC152 | HCT 116 | C76(0.75) | LDD0590 | [7] |
| LDCM0274 | AC153 | HCT 116 | C76(0.78) | LDD0591 | [7] |
| LDCM0621 | AC154 | HCT 116 | C76(0.75) | LDD2158 | [7] |
| LDCM0622 | AC155 | HCT 116 | C76(0.83) | LDD2159 | [7] |
| LDCM0623 | AC156 | HCT 116 | C76(0.89) | LDD2160 | [7] |
| LDCM0624 | AC157 | HCT 116 | C76(0.73) | LDD2161 | [7] |
| LDCM0276 | AC17 | HCT 116 | C76(1.10) | LDD0593 | [7] |
| LDCM0277 | AC18 | HCT 116 | C76(1.04) | LDD0594 | [7] |
| LDCM0278 | AC19 | HCT 116 | C76(1.10) | LDD0595 | [7] |
| LDCM0279 | AC2 | HCT 116 | C76(1.36) | LDD0596 | [7] |
| LDCM0280 | AC20 | HCT 116 | C76(0.99) | LDD0597 | [7] |
| LDCM0281 | AC21 | HCT 116 | C76(1.05) | LDD0598 | [7] |
| LDCM0282 | AC22 | HCT 116 | C76(1.02) | LDD0599 | [7] |
| LDCM0283 | AC23 | HCT 116 | C76(0.99) | LDD0600 | [7] |
| LDCM0284 | AC24 | HCT 116 | C76(1.10) | LDD0601 | [7] |
| LDCM0285 | AC25 | HCT 116 | C76(0.94) | LDD0602 | [7] |
| LDCM0286 | AC26 | HCT 116 | C76(0.76) | LDD0603 | [7] |
| LDCM0287 | AC27 | HCT 116 | C76(0.79) | LDD0604 | [7] |
| LDCM0288 | AC28 | HCT 116 | C76(0.77) | LDD0605 | [7] |
| LDCM0289 | AC29 | HCT 116 | C76(0.73) | LDD0606 | [7] |
| LDCM0290 | AC3 | HCT 116 | C76(1.79) | LDD0607 | [7] |
| LDCM0291 | AC30 | HCT 116 | C76(0.76) | LDD0608 | [7] |
| LDCM0292 | AC31 | HCT 116 | C76(0.79) | LDD0609 | [7] |
| LDCM0293 | AC32 | HCT 116 | C76(0.79) | LDD0610 | [7] |
| LDCM0294 | AC33 | HCT 116 | C76(0.78) | LDD0611 | [7] |
| LDCM0295 | AC34 | HCT 116 | C76(0.66) | LDD0612 | [7] |
| LDCM0296 | AC35 | HCT 116 | C76(1.01) | LDD0613 | [7] |
| LDCM0297 | AC36 | HCT 116 | C76(1.03) | LDD0614 | [7] |
| LDCM0298 | AC37 | HCT 116 | C76(1.16) | LDD0615 | [7] |
| LDCM0299 | AC38 | HCT 116 | C76(1.01) | LDD0616 | [7] |
| LDCM0300 | AC39 | HCT 116 | C76(1.03) | LDD0617 | [7] |
| LDCM0301 | AC4 | HCT 116 | C76(2.20) | LDD0618 | [7] |
| LDCM0302 | AC40 | HCT 116 | C76(1.04) | LDD0619 | [7] |
| LDCM0303 | AC41 | HCT 116 | C76(0.98) | LDD0620 | [7] |
| LDCM0304 | AC42 | HCT 116 | C76(1.04) | LDD0621 | [7] |
| LDCM0305 | AC43 | HCT 116 | C76(1.02) | LDD0622 | [7] |
| LDCM0306 | AC44 | HCT 116 | C76(1.15) | LDD0623 | [7] |
| LDCM0307 | AC45 | HCT 116 | C76(1.28) | LDD0624 | [7] |
| LDCM0308 | AC46 | HCT 116 | C76(0.95) | LDD0625 | [7] |
| LDCM0309 | AC47 | HCT 116 | C76(1.00) | LDD0626 | [7] |
| LDCM0310 | AC48 | HCT 116 | C76(1.08) | LDD0627 | [7] |
| LDCM0311 | AC49 | HCT 116 | C76(1.00) | LDD0628 | [7] |
| LDCM0312 | AC5 | HCT 116 | C76(2.49) | LDD0629 | [7] |
| LDCM0313 | AC50 | HCT 116 | C76(0.94) | LDD0630 | [7] |
| LDCM0314 | AC51 | HCT 116 | C76(0.83) | LDD0631 | [7] |
| LDCM0315 | AC52 | HCT 116 | C76(0.95) | LDD0632 | [7] |
| LDCM0316 | AC53 | HCT 116 | C76(0.88) | LDD0633 | [7] |
| LDCM0317 | AC54 | HCT 116 | C76(1.08) | LDD0634 | [7] |
| LDCM0318 | AC55 | HCT 116 | C76(1.00) | LDD0635 | [7] |
| LDCM0319 | AC56 | HCT 116 | C76(1.61) | LDD0636 | [7] |
| LDCM0320 | AC57 | HCT 116 | C76(1.01) | LDD0637 | [7] |
| LDCM0321 | AC58 | HCT 116 | C76(1.02) | LDD0638 | [7] |
| LDCM0322 | AC59 | HCT 116 | C76(0.97) | LDD0639 | [7] |
| LDCM0323 | AC6 | HCT 116 | C76(0.88) | LDD0640 | [7] |
| LDCM0324 | AC60 | HCT 116 | C76(1.01) | LDD0641 | [7] |
| LDCM0325 | AC61 | HCT 116 | C76(1.29) | LDD0642 | [7] |
| LDCM0326 | AC62 | HCT 116 | C76(1.00) | LDD0643 | [7] |
| LDCM0327 | AC63 | HCT 116 | C76(0.97) | LDD0644 | [7] |
| LDCM0328 | AC64 | HCT 116 | C76(0.98) | LDD0645 | [7] |
| LDCM0329 | AC65 | HCT 116 | C76(1.26) | LDD0646 | [7] |
| LDCM0330 | AC66 | HCT 116 | C76(1.17) | LDD0647 | [7] |
| LDCM0331 | AC67 | HCT 116 | C76(0.95) | LDD0648 | [7] |
| LDCM0332 | AC68 | HCT 116 | C76(0.99) | LDD0649 | [7] |
| LDCM0333 | AC69 | HCT 116 | C76(0.95) | LDD0650 | [7] |
| LDCM0334 | AC7 | HCT 116 | C76(0.83) | LDD0651 | [7] |
| LDCM0335 | AC70 | HCT 116 | C76(0.72) | LDD0652 | [7] |
| LDCM0336 | AC71 | HCT 116 | C76(1.04) | LDD0653 | [7] |
| LDCM0337 | AC72 | HCT 116 | C76(0.91) | LDD0654 | [7] |
| LDCM0338 | AC73 | HCT 116 | C76(0.80) | LDD0655 | [7] |
| LDCM0339 | AC74 | HCT 116 | C76(0.90) | LDD0656 | [7] |
| LDCM0340 | AC75 | HCT 116 | C76(1.10) | LDD0657 | [7] |
| LDCM0341 | AC76 | HCT 116 | C76(0.89) | LDD0658 | [7] |
| LDCM0342 | AC77 | HCT 116 | C76(0.82) | LDD0659 | [7] |
| LDCM0343 | AC78 | HCT 116 | C76(0.92) | LDD0660 | [7] |
| LDCM0344 | AC79 | HCT 116 | C76(0.85) | LDD0661 | [7] |
| LDCM0345 | AC8 | HCT 116 | C76(0.95) | LDD0662 | [7] |
| LDCM0346 | AC80 | HCT 116 | C76(0.98) | LDD0663 | [7] |
| LDCM0347 | AC81 | HCT 116 | C76(0.90) | LDD0664 | [7] |
| LDCM0348 | AC82 | HCT 116 | C76(1.02) | LDD0665 | [7] |
| LDCM0349 | AC83 | HCT 116 | C76(0.77) | LDD0666 | [7] |
| LDCM0350 | AC84 | HCT 116 | C76(0.72) | LDD0667 | [7] |
| LDCM0351 | AC85 | HCT 116 | C76(0.68) | LDD0668 | [7] |
| LDCM0352 | AC86 | HCT 116 | C76(0.75) | LDD0669 | [7] |
| LDCM0353 | AC87 | HCT 116 | C76(0.93) | LDD0670 | [7] |
| LDCM0354 | AC88 | HCT 116 | C76(0.82) | LDD0671 | [7] |
| LDCM0355 | AC89 | HCT 116 | C76(0.89) | LDD0672 | [7] |
| LDCM0357 | AC90 | HCT 116 | C76(0.98) | LDD0674 | [7] |
| LDCM0358 | AC91 | HCT 116 | C76(1.14) | LDD0675 | [7] |
| LDCM0359 | AC92 | HCT 116 | C76(0.95) | LDD0676 | [7] |
| LDCM0360 | AC93 | HCT 116 | C76(0.99) | LDD0677 | [7] |
| LDCM0361 | AC94 | HCT 116 | C76(1.01) | LDD0678 | [7] |
| LDCM0362 | AC95 | HCT 116 | C76(0.91) | LDD0679 | [7] |
| LDCM0363 | AC96 | HCT 116 | C76(1.03) | LDD0680 | [7] |
| LDCM0364 | AC97 | HCT 116 | C76(0.77) | LDD0681 | [7] |
| LDCM0365 | AC98 | HCT 116 | C76(0.85) | LDD0682 | [7] |
| LDCM0366 | AC99 | HCT 116 | C76(0.71) | LDD0683 | [7] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C76(0.63) | LDD2113 | [5] |
| LDCM0248 | AKOS034007472 | HCT 116 | C76(1.21) | LDD0565 | [7] |
| LDCM0356 | AKOS034007680 | HCT 116 | C76(1.49) | LDD0673 | [7] |
| LDCM0275 | AKOS034007705 | HCT 116 | C76(0.99) | LDD0592 | [7] |
| LDCM0156 | Aniline | NCI-H1299 | 12.07 | LDD0403 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | H8(0.00); H13(0.00); C76(0.00); H36(0.00) | LDD0222 | [19] |
| LDCM0367 | CL1 | HCT 116 | C76(0.80) | LDD0684 | [7] |
| LDCM0368 | CL10 | HCT 116 | C76(0.99) | LDD0685 | [7] |
| LDCM0369 | CL100 | HCT 116 | C76(1.78) | LDD0686 | [7] |
| LDCM0370 | CL101 | HCT 116 | C76(0.89) | LDD0687 | [7] |
| LDCM0371 | CL102 | HCT 116 | C76(1.20) | LDD0688 | [7] |
| LDCM0372 | CL103 | HCT 116 | C76(1.17) | LDD0689 | [7] |
| LDCM0373 | CL104 | HCT 116 | C76(1.11) | LDD0690 | [7] |
| LDCM0374 | CL105 | HCT 116 | C76(1.15) | LDD0691 | [7] |
| LDCM0375 | CL106 | HCT 116 | C76(1.14) | LDD0692 | [7] |
| LDCM0376 | CL107 | HCT 116 | C76(1.12) | LDD0693 | [7] |
| LDCM0377 | CL108 | HCT 116 | C76(1.13) | LDD0694 | [7] |
| LDCM0378 | CL109 | HCT 116 | C76(0.99) | LDD0695 | [7] |
| LDCM0379 | CL11 | HCT 116 | C76(0.98) | LDD0696 | [7] |
| LDCM0380 | CL110 | HCT 116 | C76(0.83) | LDD0697 | [7] |
| LDCM0381 | CL111 | HCT 116 | C76(0.93) | LDD0698 | [7] |
| LDCM0382 | CL112 | HCT 116 | C76(0.70) | LDD0699 | [7] |
| LDCM0383 | CL113 | HCT 116 | C76(0.65) | LDD0700 | [7] |
| LDCM0384 | CL114 | HCT 116 | C76(0.57) | LDD0701 | [7] |
| LDCM0385 | CL115 | HCT 116 | C76(0.70) | LDD0702 | [7] |
| LDCM0386 | CL116 | HCT 116 | C76(0.82) | LDD0703 | [7] |
| LDCM0387 | CL117 | HCT 116 | C76(1.34) | LDD0704 | [7] |
| LDCM0388 | CL118 | HCT 116 | C76(1.05) | LDD0705 | [7] |
| LDCM0389 | CL119 | HCT 116 | C76(1.10) | LDD0706 | [7] |
| LDCM0390 | CL12 | HCT 116 | C76(1.19) | LDD0707 | [7] |
| LDCM0391 | CL120 | HCT 116 | C76(1.14) | LDD0708 | [7] |
| LDCM0392 | CL121 | HCT 116 | C76(1.52) | LDD0709 | [7] |
| LDCM0393 | CL122 | HCT 116 | C76(0.91) | LDD0710 | [7] |
| LDCM0394 | CL123 | HCT 116 | C76(0.81) | LDD0711 | [7] |
| LDCM0395 | CL124 | HCT 116 | C76(0.92) | LDD0712 | [7] |
| LDCM0396 | CL125 | HCT 116 | C76(1.06) | LDD0713 | [7] |
| LDCM0397 | CL126 | HCT 116 | C76(1.22) | LDD0714 | [7] |
| LDCM0398 | CL127 | HCT 116 | C76(1.00) | LDD0715 | [7] |
| LDCM0399 | CL128 | HCT 116 | C76(1.05) | LDD0716 | [7] |
| LDCM0400 | CL13 | HCT 116 | C76(1.23) | LDD0717 | [7] |
| LDCM0401 | CL14 | HCT 116 | C76(0.85) | LDD0718 | [7] |
| LDCM0402 | CL15 | HCT 116 | C76(0.93) | LDD0719 | [7] |
| LDCM0403 | CL16 | HCT 116 | C76(1.19) | LDD0720 | [7] |
| LDCM0404 | CL17 | HCT 116 | C76(1.07) | LDD0721 | [7] |
| LDCM0405 | CL18 | HCT 116 | C76(1.09) | LDD0722 | [7] |
| LDCM0406 | CL19 | HCT 116 | C76(1.15) | LDD0723 | [7] |
| LDCM0407 | CL2 | HCT 116 | C76(0.82) | LDD0724 | [7] |
| LDCM0408 | CL20 | HCT 116 | C76(1.19) | LDD0725 | [7] |
| LDCM0409 | CL21 | HCT 116 | C76(1.07) | LDD0726 | [7] |
| LDCM0410 | CL22 | HCT 116 | C76(1.80) | LDD0727 | [7] |
| LDCM0411 | CL23 | HCT 116 | C76(1.12) | LDD0728 | [7] |
| LDCM0412 | CL24 | HCT 116 | C76(1.23) | LDD0729 | [7] |
| LDCM0413 | CL25 | HCT 116 | C76(1.10) | LDD0730 | [7] |
| LDCM0414 | CL26 | HCT 116 | C76(0.98) | LDD0731 | [7] |
| LDCM0415 | CL27 | HCT 116 | C76(1.31) | LDD0732 | [7] |
| LDCM0416 | CL28 | HCT 116 | C76(1.07) | LDD0733 | [7] |
| LDCM0417 | CL29 | HCT 116 | C76(1.25) | LDD0734 | [7] |
| LDCM0418 | CL3 | HCT 116 | C76(0.84) | LDD0735 | [7] |
| LDCM0419 | CL30 | HCT 116 | C76(1.10) | LDD0736 | [7] |
| LDCM0420 | CL31 | HCT 116 | C76(0.83) | LDD0737 | [7] |
| LDCM0421 | CL32 | HCT 116 | C76(0.86) | LDD0738 | [7] |
| LDCM0422 | CL33 | HCT 116 | C76(0.75) | LDD0739 | [7] |
| LDCM0423 | CL34 | HCT 116 | C76(0.77) | LDD0740 | [7] |
| LDCM0424 | CL35 | HCT 116 | C76(0.82) | LDD0741 | [7] |
| LDCM0425 | CL36 | HCT 116 | C76(0.68) | LDD0742 | [7] |
| LDCM0426 | CL37 | HCT 116 | C76(0.74) | LDD0743 | [7] |
| LDCM0428 | CL39 | HCT 116 | C76(0.78) | LDD0745 | [7] |
| LDCM0429 | CL4 | HCT 116 | C76(1.02) | LDD0746 | [7] |
| LDCM0430 | CL40 | HCT 116 | C76(0.67) | LDD0747 | [7] |
| LDCM0431 | CL41 | HCT 116 | C76(0.82) | LDD0748 | [7] |
| LDCM0432 | CL42 | HCT 116 | C76(0.97) | LDD0749 | [7] |
| LDCM0433 | CL43 | HCT 116 | C76(0.81) | LDD0750 | [7] |
| LDCM0434 | CL44 | HCT 116 | C76(0.80) | LDD0751 | [7] |
| LDCM0435 | CL45 | HCT 116 | C76(0.73) | LDD0752 | [7] |
| LDCM0436 | CL46 | HCT 116 | C76(1.19) | LDD0753 | [7] |
| LDCM0437 | CL47 | HCT 116 | C76(1.10) | LDD0754 | [7] |
| LDCM0438 | CL48 | HCT 116 | C76(1.10) | LDD0755 | [7] |
| LDCM0439 | CL49 | HCT 116 | C76(0.99) | LDD0756 | [7] |
| LDCM0440 | CL5 | HCT 116 | C76(0.99) | LDD0757 | [7] |
| LDCM0441 | CL50 | HCT 116 | C76(1.07) | LDD0758 | [7] |
| LDCM0442 | CL51 | HCT 116 | C76(1.10) | LDD0759 | [7] |
| LDCM0443 | CL52 | HCT 116 | C76(1.09) | LDD0760 | [7] |
| LDCM0444 | CL53 | HCT 116 | C76(1.03) | LDD0761 | [7] |
| LDCM0445 | CL54 | HCT 116 | C76(1.12) | LDD0762 | [7] |
| LDCM0446 | CL55 | HCT 116 | C76(1.07) | LDD0763 | [7] |
| LDCM0447 | CL56 | HCT 116 | C76(1.02) | LDD0764 | [7] |
| LDCM0448 | CL57 | HCT 116 | C76(1.13) | LDD0765 | [7] |
| LDCM0449 | CL58 | HCT 116 | C76(1.11) | LDD0766 | [7] |
| LDCM0450 | CL59 | HCT 116 | C76(0.97) | LDD0767 | [7] |
| LDCM0451 | CL6 | HCT 116 | C76(0.83) | LDD0768 | [7] |
| LDCM0452 | CL60 | HCT 116 | C76(0.86) | LDD0769 | [7] |
| LDCM0453 | CL61 | HCT 116 | C76(0.91) | LDD0770 | [7] |
| LDCM0454 | CL62 | HCT 116 | C76(0.99) | LDD0771 | [7] |
| LDCM0455 | CL63 | HCT 116 | C76(0.87) | LDD0772 | [7] |
| LDCM0456 | CL64 | HCT 116 | C76(0.91) | LDD0773 | [7] |
| LDCM0457 | CL65 | HCT 116 | C76(0.97) | LDD0774 | [7] |
| LDCM0458 | CL66 | HCT 116 | C76(1.08) | LDD0775 | [7] |
| LDCM0459 | CL67 | HCT 116 | C76(1.00) | LDD0776 | [7] |
| LDCM0460 | CL68 | HCT 116 | C76(0.81) | LDD0777 | [7] |
| LDCM0461 | CL69 | HCT 116 | C76(0.88) | LDD0778 | [7] |
| LDCM0462 | CL7 | HCT 116 | C76(1.25) | LDD0779 | [7] |
| LDCM0463 | CL70 | HCT 116 | C76(0.87) | LDD0780 | [7] |
| LDCM0464 | CL71 | HCT 116 | C76(0.76) | LDD0781 | [7] |
| LDCM0465 | CL72 | HCT 116 | C76(0.85) | LDD0782 | [7] |
| LDCM0466 | CL73 | HCT 116 | C76(0.72) | LDD0783 | [7] |
| LDCM0467 | CL74 | HCT 116 | C76(0.75) | LDD0784 | [7] |
| LDCM0469 | CL76 | HCT 116 | C76(1.10) | LDD0786 | [7] |
| LDCM0470 | CL77 | HCT 116 | C76(0.89) | LDD0787 | [7] |
| LDCM0471 | CL78 | HCT 116 | C76(0.98) | LDD0788 | [7] |
| LDCM0472 | CL79 | HCT 116 | C76(1.15) | LDD0789 | [7] |
| LDCM0473 | CL8 | HCT 116 | C76(0.77) | LDD0790 | [7] |
| LDCM0474 | CL80 | HCT 116 | C76(1.14) | LDD0791 | [7] |
| LDCM0475 | CL81 | HCT 116 | C76(1.24) | LDD0792 | [7] |
| LDCM0476 | CL82 | HCT 116 | C76(1.34) | LDD0793 | [7] |
| LDCM0477 | CL83 | HCT 116 | C76(1.17) | LDD0794 | [7] |
| LDCM0478 | CL84 | HCT 116 | C76(1.17) | LDD0795 | [7] |
| LDCM0479 | CL85 | HCT 116 | C76(1.03) | LDD0796 | [7] |
| LDCM0480 | CL86 | HCT 116 | C76(1.01) | LDD0797 | [7] |
| LDCM0481 | CL87 | HCT 116 | C76(1.04) | LDD0798 | [7] |
| LDCM0482 | CL88 | HCT 116 | C76(1.11) | LDD0799 | [7] |
| LDCM0483 | CL89 | HCT 116 | C76(1.75) | LDD0800 | [7] |
| LDCM0484 | CL9 | HCT 116 | C76(0.93) | LDD0801 | [7] |
| LDCM0485 | CL90 | HCT 116 | C76(1.10) | LDD0802 | [7] |
| LDCM0486 | CL91 | HCT 116 | C76(1.30) | LDD0803 | [7] |
| LDCM0487 | CL92 | HCT 116 | C76(1.02) | LDD0804 | [7] |
| LDCM0488 | CL93 | HCT 116 | C76(1.09) | LDD0805 | [7] |
| LDCM0489 | CL94 | HCT 116 | C76(1.38) | LDD0806 | [7] |
| LDCM0490 | CL95 | HCT 116 | C76(1.08) | LDD0807 | [7] |
| LDCM0491 | CL96 | HCT 116 | C76(1.25) | LDD0808 | [7] |
| LDCM0492 | CL97 | HCT 116 | C76(1.20) | LDD0809 | [7] |
| LDCM0493 | CL98 | HCT 116 | C76(1.52) | LDD0810 | [7] |
| LDCM0494 | CL99 | HCT 116 | C76(1.76) | LDD0811 | [7] |
| LDCM0634 | CY-0357 | Hep-G2 | C76(1.49) | LDD2228 | [21] |
| LDCM0495 | E2913 | HEK-293T | C76(1.08); C41(0.98) | LDD1698 | [27] |
| LDCM0625 | F8 | Ramos | C76(0.97) | LDD2187 | [28] |
| LDCM0572 | Fragment10 | Ramos | C76(0.51) | LDD2189 | [28] |
| LDCM0573 | Fragment11 | Ramos | C76(0.14) | LDD2190 | [28] |
| LDCM0574 | Fragment12 | Ramos | C76(0.53) | LDD2191 | [28] |
| LDCM0575 | Fragment13 | Ramos | C76(0.84) | LDD2192 | [28] |
| LDCM0576 | Fragment14 | Ramos | C76(0.67) | LDD2193 | [28] |
| LDCM0579 | Fragment20 | Ramos | C76(0.49) | LDD2194 | [28] |
| LDCM0580 | Fragment21 | Ramos | C76(0.73) | LDD2195 | [28] |
| LDCM0582 | Fragment23 | Ramos | C76(0.61) | LDD2196 | [28] |
| LDCM0578 | Fragment27 | Ramos | C76(0.89) | LDD2197 | [28] |
| LDCM0586 | Fragment28 | Ramos | C76(0.29) | LDD2198 | [28] |
| LDCM0588 | Fragment30 | Ramos | C76(1.06) | LDD2199 | [28] |
| LDCM0589 | Fragment31 | Ramos | C76(0.83) | LDD2200 | [28] |
| LDCM0590 | Fragment32 | Ramos | C76(0.51) | LDD2201 | [28] |
| LDCM0468 | Fragment33 | HCT 116 | C76(0.93) | LDD0785 | [7] |
| LDCM0596 | Fragment38 | Ramos | C76(1.01) | LDD2203 | [28] |
| LDCM0566 | Fragment4 | Ramos | C76(0.81) | LDD2184 | [28] |
| LDCM0427 | Fragment51 | HCT 116 | C76(0.61) | LDD0744 | [7] |
| LDCM0610 | Fragment52 | Ramos | C76(1.09) | LDD2204 | [28] |
| LDCM0614 | Fragment56 | Ramos | C76(1.27) | LDD2205 | [28] |
| LDCM0569 | Fragment7 | Ramos | C76(0.72) | LDD2186 | [28] |
| LDCM0571 | Fragment9 | Ramos | C76(0.51) | LDD2188 | [28] |
| LDCM0116 | HHS-0101 | DM93 | Y18(1.76); Y43(1.84) | LDD0264 | [6] |
| LDCM0117 | HHS-0201 | DM93 | Y18(0.70); Y43(4.17) | LDD0265 | [6] |
| LDCM0118 | HHS-0301 | DM93 | Y18(0.67); Y43(2.38) | LDD0266 | [6] |
| LDCM0119 | HHS-0401 | DM93 | Y18(0.62) | LDD0267 | [6] |
| LDCM0120 | HHS-0701 | DM93 | Y18(0.59) | LDD0268 | [6] |
| LDCM0107 | IAA | HeLa | H13(0.00); H8(0.00); C76(0.00); H36(0.00) | LDD0221 | [19] |
| LDCM0022 | KB02 | HEK-293T | C76(1.00) | LDD1492 | [27] |
| LDCM0023 | KB03 | HEK-293T | C76(1.08) | LDD1497 | [27] |
| LDCM0024 | KB05 | G361 | C76(0.87) | LDD3311 | [29] |
| LDCM0109 | NEM | HeLa | H8(0.00); H13(0.00) | LDD0223 | [19] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C76(0.96) | LDD2101 | [5] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C76(1.01) | LDD2107 | [5] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C76(0.47) | LDD2108 | [5] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C76(0.94) | LDD2125 | [5] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C76(1.56) | LDD2136 | [5] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C76(0.81) | LDD2140 | [5] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C76(0.73) | LDD2141 | [5] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C76(1.31) | LDD2206 | [30] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C76(1.15) | LDD2207 | [30] |
| LDCM0131 | RA190 | MM1.R | C76(1.11) | LDD0304 | [31] |
The Interaction Atlas With This Target
References
