Details of the Target
General Information of Target
| Target ID | LDTP04703 | |||||
|---|---|---|---|---|---|---|
| Target Name | Tumor protein D52 (TPD52) | |||||
| Gene Name | TPD52 | |||||
| Gene ID | 7163 | |||||
| Synonyms |
Tumor protein D52; Protein N8 |
|||||
| 3D Structure | ||||||
| Sequence |
MDCREMDLYEDYQSPFDFDAGVNKSYLYLSPSGNSSPPGSPTLQKFGLLRTDPVPEEGED
VAATISATETLSEEEQEELRRELAKVEEEIQTLSQVLAAKEKHLAEIKRKLGINSLQELK QNIAKGWQDVTATSAYKKTSETLSQAGQKASAAFSSVGSVITKKLEDVKNSPTFKSFEEK VENLKSKVGGTKPAGGDFGEVLNSAANASATTTEPLPEKTQESL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
TPD52 family
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AZ-9 Probe Info |
 |
10.00 | LDD0393 | [1] | |
|
TH211 Probe Info |
 |
Y136(6.22) | LDD0260 | [2] | |
|
C-Sul Probe Info |
 |
6.38 | LDD0066 | [3] | |
|
TH216 Probe Info |
 |
Y136(16.86) | LDD0259 | [2] | |
|
Probe 1 Probe Info |
 |
Y136(140.46) | LDD3495 | [4] | |
|
HHS-475 Probe Info |
 |
Y136(0.98) | LDD0264 | [5] | |
|
HHS-465 Probe Info |
 |
Y136(10.00) | LDD2237 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [7] | |
|
ATP probe Probe Info |
 |
K163(0.00); K164(0.00); K138(0.00); K185(0.00) | LDD0199 | [8] | |
|
ATP probe Probe Info |
 |
K125(0.00); K85(0.00); K169(0.00); K100(0.00) | LDD0035 | [9] | |
|
NHS Probe Info |
 |
K100(0.00); K120(0.00); K138(0.00) | LDD0010 | [10] | |
|
STPyne Probe Info |
 |
K102(0.00); K149(0.00); K120(0.00); K138(0.00) | LDD0009 | [10] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C003 Probe Info |
 |
15.14 | LDD1713 | [12] | |
|
C055 Probe Info |
 |
15.24 | LDD1752 | [12] | |
|
C056 Probe Info |
 |
25.46 | LDD1753 | [12] | |
|
C112 Probe Info |
 |
20.11 | LDD1799 | [12] | |
|
C134 Probe Info |
 |
24.59 | LDD1816 | [12] | |
|
C143 Probe Info |
 |
14.72 | LDD1825 | [12] | |
|
C201 Probe Info |
 |
34.54 | LDD1877 | [12] | |
|
C228 Probe Info |
 |
15.35 | LDD1901 | [12] | |
|
C264 Probe Info |
 |
17.27 | LDD1935 | [12] | |
|
C326 Probe Info |
 |
6.82 | LDD1990 | [12] | |
|
C350 Probe Info |
 |
22.94 | LDD2011 | [12] | |
|
C362 Probe Info |
 |
55.72 | LDD2023 | [12] | |
|
C388 Probe Info |
 |
37.01 | LDD2047 | [12] | |
|
C390 Probe Info |
 |
22.01 | LDD2049 | [12] | |
|
C426 Probe Info |
 |
15.67 | LDD2081 | [12] | |
|
DFG-out-4 Probe Info |
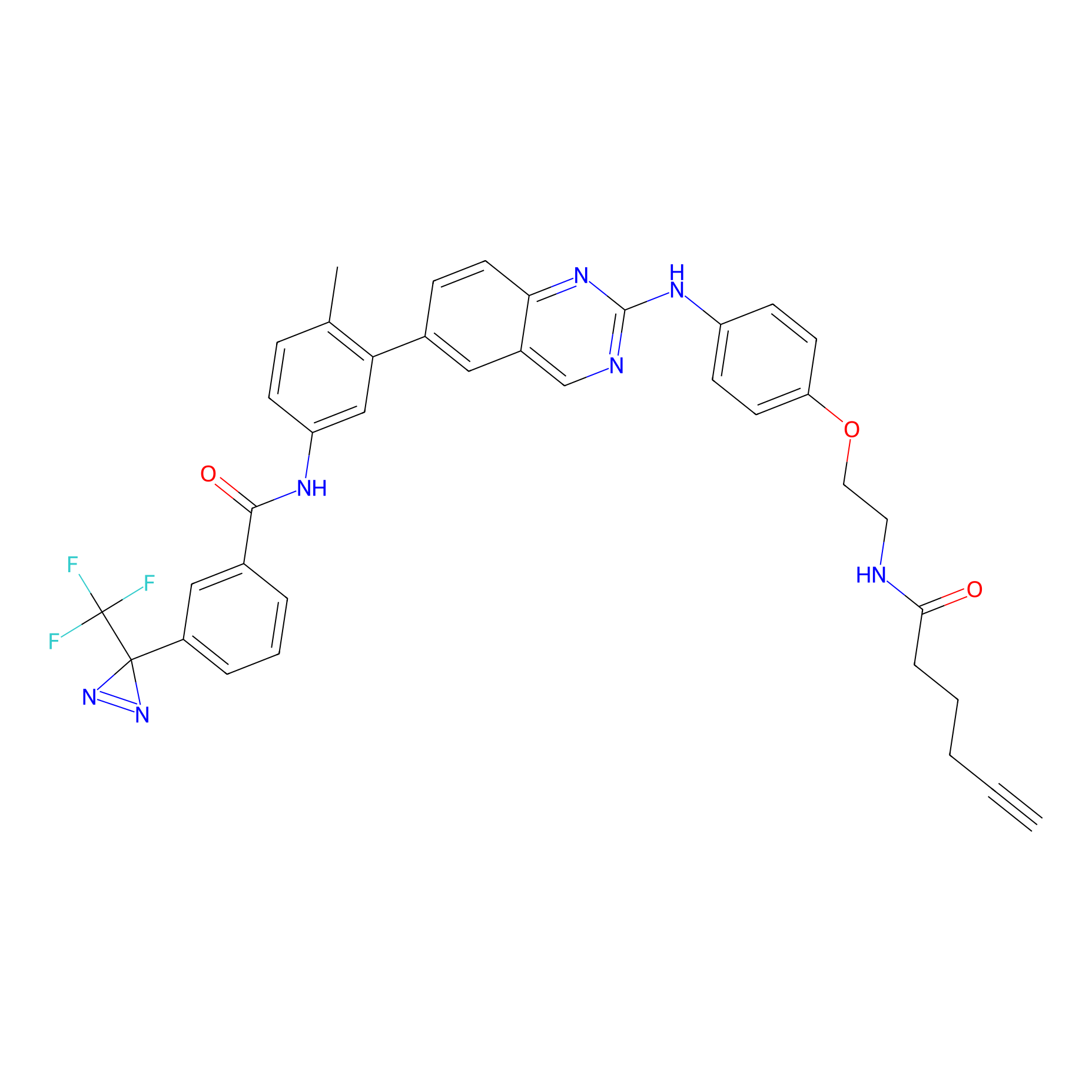 |
6.30 | LDD0075 | [13] | |
|
Staurosporine capture compound Probe Info |
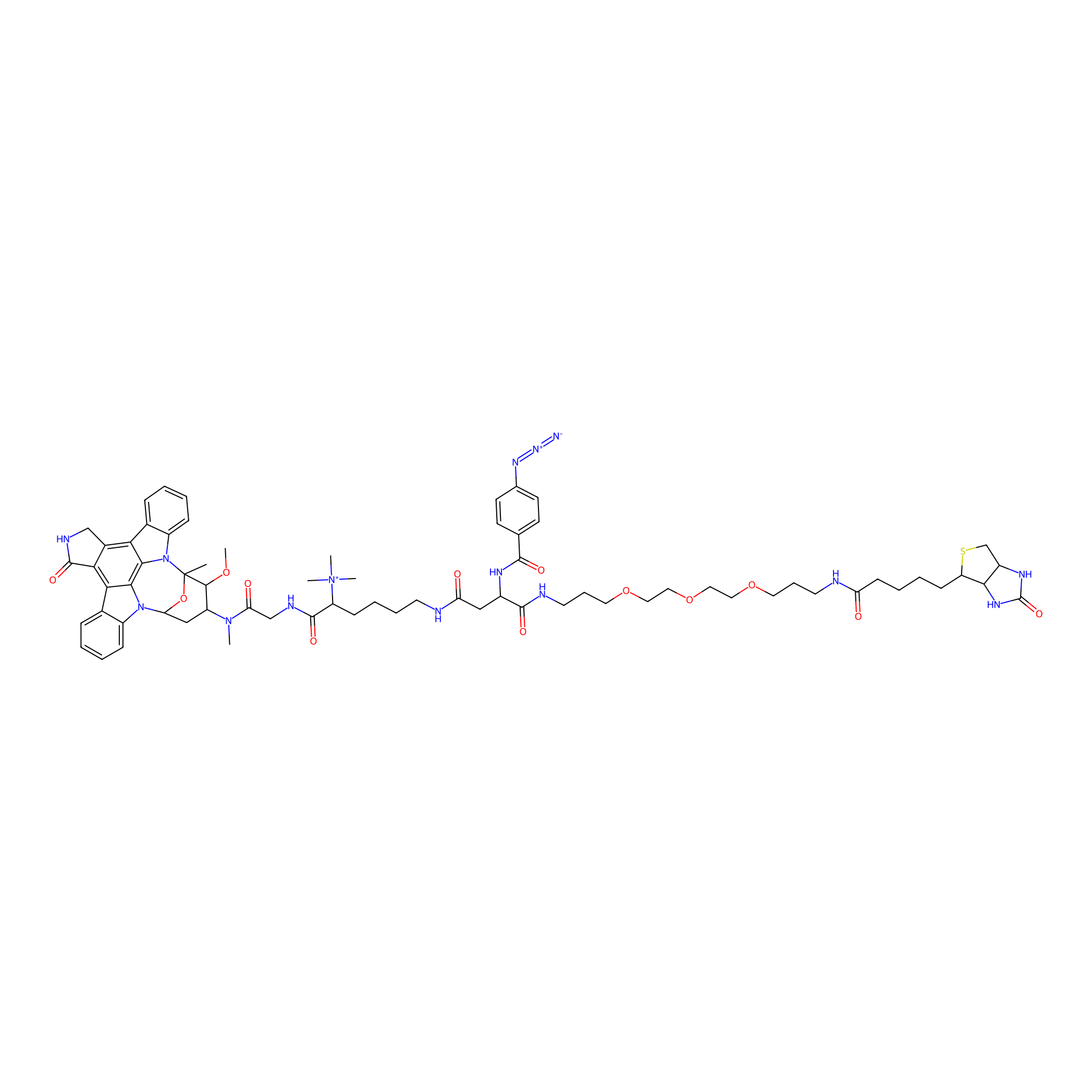 |
N.A. | LDD0083 | [14] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [7] |
| LDCM0017 | DFG-out-2 | A431 | 6.30 | LDD0075 | [13] |
| LDCM0116 | HHS-0101 | DM93 | Y136(0.98) | LDD0264 | [5] |
| LDCM0117 | HHS-0201 | DM93 | Y136(0.84) | LDD0265 | [5] |
| LDCM0118 | HHS-0301 | DM93 | Y136(0.90) | LDD0266 | [5] |
| LDCM0119 | HHS-0401 | DM93 | Y136(0.93) | LDD0267 | [5] |
| LDCM0120 | HHS-0701 | DM93 | Y136(0.65) | LDD0268 | [5] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [7] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [7] |
| LDCM0019 | Staurosporine | Hep-G2 | N.A. | LDD0083 | [14] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Maspardin (SPG21) | AB hydrolase superfamily | Q9NZD8 | |||
| Peroxynitrite isomerase THAP4 (THAP4) | Nitrobindin family | Q8WY91 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Endophilin-B1 (SH3GLB1) | Endophilin family | Q9Y371 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Mitochondrial dynamics protein MIEF1 (MIEF1) | SMCR7 family | Q9NQG6 | |||
| Tumor protein D53 (TPD52L1) | TPD52 family | Q16890 | |||
| Alpha-tocopherol transfer protein (TTPA) | . | P49638 | |||
References
