Details of the Target
General Information of Target
| Target ID | LDTP04112 | |||||
|---|---|---|---|---|---|---|
| Target Name | Microtubule-associated protein 1B (MAP1B) | |||||
| Gene Name | MAP1B | |||||
| Gene ID | 4131 | |||||
| Synonyms |
Microtubule-associated protein 1B; MAP-1B) [Cleaved into: MAP1B heavy chain; MAP1 light chain LC1] |
|||||
| 3D Structure | ||||||
| Sequence |
MATVVVEATEPEPSGSIANPAASTSPSLSHRFLDSKFYLLVVVGEIVTEEHLRRAIGNIE
LGIRSWDTNLIECNLDQELKLFVSRHSARFSPEVPGQKILHHRSDVLETVVLINPSDEAV STEVRLMITDAARHKLLVLTGQCFENTGELILQSGSFSFQNFIEIFTDQEIGELLSTTHP ANKASLTLFCPEEGDWKNSNLDRHNLQDFINIKLNSASILPEMEGLSEFTEYLSESVEVP SPFDILEPPTSGGFLKLSKPCCYIFPGGRGDSALFAVNGFNMLINGGSERKSCFWKLIRH LDRVDSILLTHIGDDNLPGINSMLQRKIAELEEEQSQGSTTNSDWMKNLISPDLGVVFLN VPENLKNPEPNIKMKRSIEEACFTLQYLNKLSMKPEPLFRSVGNTIDPVILFQKMGVGKL EMYVLNPVKSSKEMQYFMQQWTGTNKDKAEFILPNGQEVDLPISYLTSVSSLIVWHPANP AEKIIRVLFPGNSTQYNILEGLEKLKHLDFLKQPLATQKDLTGQVPTPVVKQTKLKQRAD SRESLKPAAKPLPSKSVRKESKEETPEVTKVNHVEKPPKVESKEKVMVKKDKPIKTETKP SVTEKEVPSKEEPSPVKAEVAEKQATDVKPKAAKEKTVKKETKVKPEDKKEEKEKPKKEV AKKEDKTPIKKEEKPKKEEVKKEVKKEIKKEEKKEPKKEVKKETPPKEVKKEVKKEEKKE VKKEEKEPKKEIKKLPKDAKKSSTPLSEAKKPAALKPKVPKKEESVKKDSVAAGKPKEKG KIKVIKKEGKAAEAVAAAVGTGATTAAVMAAAGIAAIGPAKELEAERSLMSSPEDLTKDF EELKAEEVDVTKDIKPQLELIEDEEKLKETEPVEAYVIQKEREVTKGPAESPDEGITTTE GEGECEQTPEELEPVEKQGVDDIEKFEDEGAGFEESSETGDYEEKAETEEAEEPEEDGEE HVCVSASKHSPTEDEESAKAEADAYIREKRESVASGDDRAEEDMDEAIEKGEAEQSEEEA DEEDKAEDAREEEYEPEKMEAEDYVMAVVDKAAEAGGAEEQYGFLTTPTKQLGAQSPGRE PASSIHDETLPGGSESEATASDEENREDQPEEFTATSGYTQSTIEISSEPTPMDEMSTPR DVMSDETNNEETESPSQEFVNITKYESSLYSQEYSKPADVTPLNGFSEGSKTDATDGKDY NASASTISPPSSMEEDKFSRSALRDAYCSEVKASTTLDIKDSISAVSSEKVSPSKSPSLS PSPPSPLEKTPLGERSVNFSLTPNEIKVSAEAEVAPVSPEVTQEVVEEHCASPEDKTLEV VSPSQSVTGSAGHTPYYQSPTDEKSSHLPTEVIEKPPAVPVSFEFSDAKDENERASVSPM DEPVPDSESPIEKVLSPLRSPPLIGSESAYESFLSADDKASGRGAESPFEEKSGKQGSPD QVSPVSEMTSTSLYQDKQEGKSTDFAPIKEDFGQEKKTDDVEAMSSQPALALDERKLGDV SPTQIDVSQFGSFKEDTKMSISEGTVSDKSATPVDEGVAEDTYSHMEGVASVSTASVATS SFPEPTTDDVSPSLHAEVGSPHSTEVDDSLSVSVVQTPTTFQETEMSPSKEECPRPMSIS PPDFSPKTAKSRTPVQDHRSEQSSMSIEFGQESPEQSLAMDFSRQSPDHPTVGAGVLHIT ENGPTEVDYSPSDMQDSSLSHKIPPMEEPSYTQDNDLSELISVSQVEASPSTSSAHTPSQ IASPLQEDTLSDVAPPRDMSLYASLTSEKVQSLEGEKLSPKSDISPLTPRESSPLYSPTF SDSTSAVKEKTATCHSSSSPPIDAASAEPYGFRASVLFDTMQHHLALNRDLSTPGLEKDS GGKTPGDFSYAYQKPEETTRSPDEEDYDYESYEKTTRTSDVGGYYYEKIERTTKSPSDSG YSYETIGKTTKTPEDGDYSYEIIEKTTRTPEEGGYSYDISEKTTSPPEVSGYSYEKTERS RRLLDDISNGYDDSEDGGHTLGDPSYSYETTEKITSFPESEGYSYETSTKTTRTPDTSTY CYETAEKITRTPQASTYSYETSDLCYTAEKKSPSEARQDVDLCLVSSCEYKHPKTELSPS FINPNPLEWFASEEPTEESEKPLTQSGGAPPPPGGKQQGRQCDETPPTSVSESAPSQTDS DVPPETEECPSITADANIDSEDESETIPTDKTVTYKHMDPPPAPVQDRSPSPRHPDVSMV DPEALAIEQNLGKALKKDLKEKTKTKKPGTKTKSSSPVKKSDGKSKPLAASPKPAGLKES SDKVSRVASPKKKESVEKAAKPTTTPEVKAARGEEKDKETKNAANASASKSAKTATAGPG TTKTTKSSAVPPGLPVYLDLCYIPNHSNSKNVDVEFFKRVRSSYYVVSGNDPAAEEPSRA VLDALLEGKAQWGSNMQVTLIPTHDSEVMREWYQETHEKQQDLNIMVLASSSTVVMQDES FPACKIEL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
MAP1 family
|
|||||
| Subcellular location |
Cytoplasm; Cytoplasm, cytoskeleton
|
|||||
| Function |
Facilitates tyrosination of alpha-tubulin in neuronal microtubules. Phosphorylated MAP1B is required for proper microtubule dynamics and plays a role in the cytoskeletal changes that accompany neuronal differentiation and neurite extension. Possibly MAP1B binds to at least two tubulin subunits in the polymer, and this bridging of subunits might be involved in nucleating microtubule polymerization and in stabilizing microtubules. Acts as a positive cofactor in DAPK1-mediated autophagic vesicle formation and membrane blebbing.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
DAyne Probe Info |
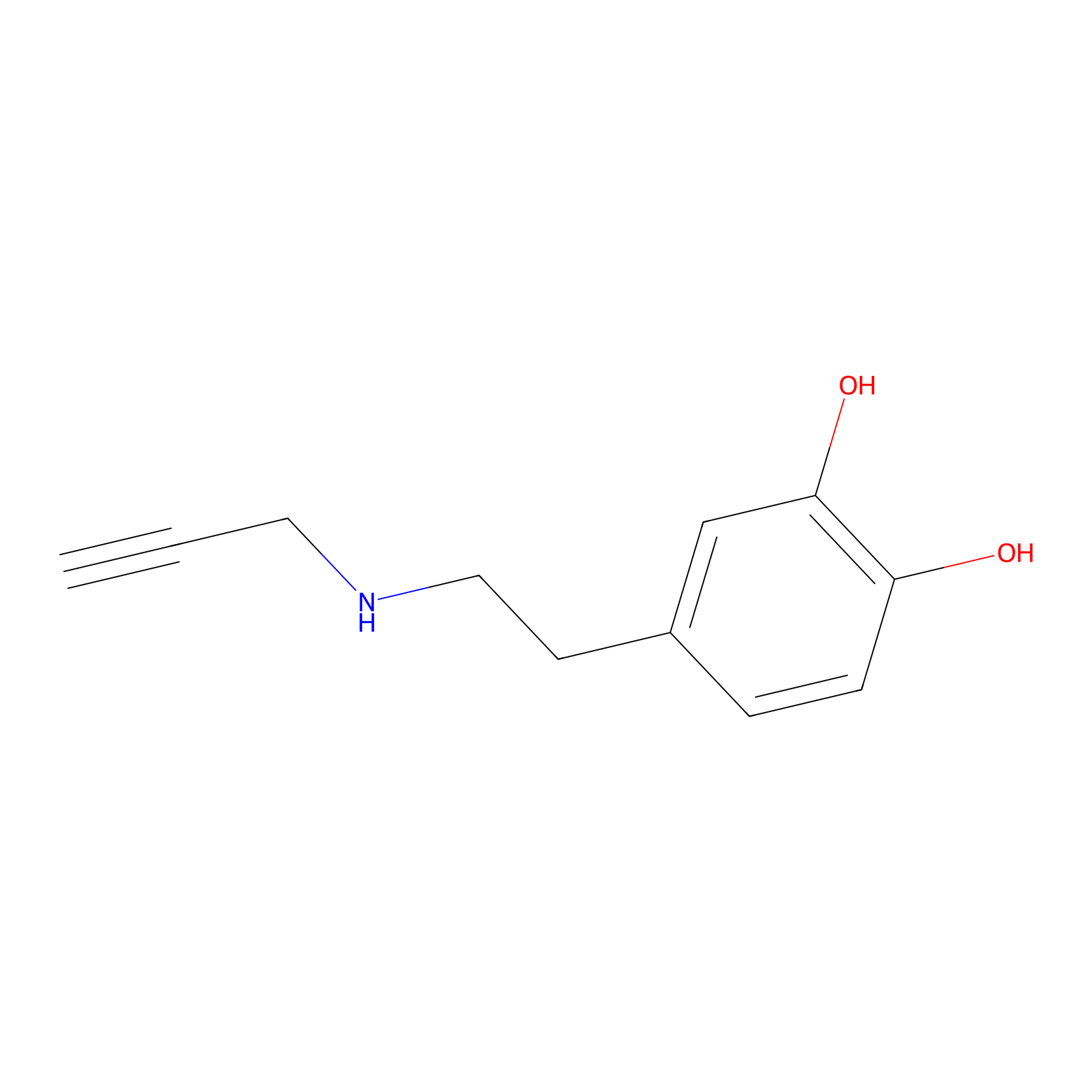 |
4.20 | LDD0261 | [2] | |
|
CY-1 Probe Info |
 |
100.00 | LDD0243 | [3] | |
|
TH211 Probe Info |
 |
Y263(12.11) | LDD0257 | [4] | |
|
STPyne Probe Info |
 |
K1355(6.00); K1976(10.00); K2237(10.00); K2273(9.92) | LDD0277 | [5] | |
|
ONAyne Probe Info |
 |
K2091(10.00); K506(10.00) | LDD0275 | [5] | |
|
BTD Probe Info |
 |
C1228(0.70) | LDD2089 | [6] | |
|
AHL-Pu-1 Probe Info |
 |
C382(4.98) | LDD0168 | [7] | |
|
HHS-475 Probe Info |
 |
Y1923(0.06); Y1336(0.36); Y2023(0.62); Y1957(0.62) | LDD0264 | [8] | |
|
HHS-465 Probe Info |
 |
Y1062(5.07); Y1227(2.18); Y1337(10.00); Y1830(10.00) | LDD2237 | [9] | |
|
DBIA Probe Info |
 |
C2041(1.18); C2065(1.06); C963(1.01); C382(0.99) | LDD0078 | [10] | |
|
IA-alkyne Probe Info |
 |
C2361(0.00); C2041(0.00) | LDD0032 | [11] | |
|
NAIA_4 Probe Info |
 |
C382(0.00); C905(0.00); C963(0.00); C1228(0.00) | LDD2226 | [12] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [13] | |
|
WYneN Probe Info |
 |
C1814(0.00); C2041(0.00) | LDD0021 | [14] | |
|
WYneO Probe Info |
 |
C1814(0.00); C1228(0.00); C2041(0.00) | LDD0022 | [14] | |
|
DA-P3 Probe Info |
 |
N.A. | LDD0178 | [15] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0356 | [16] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [14] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [14] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [17] | |
|
VSF Probe Info |
 |
C1814(0.00); C1228(0.00) | LDD0007 | [14] | |
|
Acrolein Probe Info |
 |
C963(0.00); C73(0.00); C190(0.00); C2361(0.00) | LDD0217 | [18] | |
|
Methacrolein Probe Info |
 |
C2041(0.00); C190(0.00); C963(0.00) | LDD0218 | [18] | |
|
W1 Probe Info |
 |
C2041(0.00); C190(0.00); C73(0.00) | LDD0236 | [19] | |
|
AOyne Probe Info |
 |
10.90 | LDD0443 | [20] | |
|
HHS-482 Probe Info |
 |
Y1062(1.80); Y1336(2.51); Y1872(1.45); Y1906(2.13) | LDD2239 | [9] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FFF probe12 Probe Info |
 |
20.00 | LDD0473 | [21] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [21] | |
|
FFF probe3 Probe Info |
 |
18.77 | LDD0465 | [21] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C261(1.16) | LDD2117 | [6] |
| LDCM0025 | 4SU-RNA | HEK-293T | C382(4.98) | LDD0168 | [7] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C382(8.22); C1814(9.43); C1310(3.13); C1228(2.22) | LDD0169 | [7] |
| LDCM0214 | AC1 | HCT 116 | C2041(1.09) | LDD0531 | [10] |
| LDCM0215 | AC10 | HCT 116 | C2041(1.14); C2065(1.18) | LDD0532 | [10] |
| LDCM0216 | AC100 | HCT 116 | C2041(0.89) | LDD0533 | [10] |
| LDCM0217 | AC101 | HCT 116 | C2041(0.91) | LDD0534 | [10] |
| LDCM0218 | AC102 | HCT 116 | C2041(0.86) | LDD0535 | [10] |
| LDCM0219 | AC103 | HCT 116 | C2041(1.27) | LDD0536 | [10] |
| LDCM0220 | AC104 | HCT 116 | C2041(1.32) | LDD0537 | [10] |
| LDCM0221 | AC105 | HCT 116 | C2041(1.27) | LDD0538 | [10] |
| LDCM0222 | AC106 | HCT 116 | C2041(1.14) | LDD0539 | [10] |
| LDCM0223 | AC107 | HCT 116 | C2041(1.85) | LDD0540 | [10] |
| LDCM0224 | AC108 | HCT 116 | C2041(1.21) | LDD0541 | [10] |
| LDCM0225 | AC109 | HCT 116 | C2041(1.66) | LDD0542 | [10] |
| LDCM0226 | AC11 | HCT 116 | C2041(1.12); C2065(1.24) | LDD0543 | [10] |
| LDCM0227 | AC110 | HCT 116 | C2041(0.94) | LDD0544 | [10] |
| LDCM0228 | AC111 | HCT 116 | C2041(1.68) | LDD0545 | [10] |
| LDCM0229 | AC112 | HCT 116 | C2041(1.21) | LDD0546 | [10] |
| LDCM0230 | AC113 | HCT 116 | C2041(0.97) | LDD0547 | [10] |
| LDCM0231 | AC114 | HCT 116 | C2041(1.05) | LDD0548 | [10] |
| LDCM0232 | AC115 | HCT 116 | C2041(1.16) | LDD0549 | [10] |
| LDCM0233 | AC116 | HCT 116 | C2041(1.28) | LDD0550 | [10] |
| LDCM0234 | AC117 | HCT 116 | C2041(0.97) | LDD0551 | [10] |
| LDCM0235 | AC118 | HCT 116 | C2041(1.02) | LDD0552 | [10] |
| LDCM0236 | AC119 | HCT 116 | C2041(1.10) | LDD0553 | [10] |
| LDCM0237 | AC12 | HCT 116 | C2041(0.88); C2065(0.97) | LDD0554 | [10] |
| LDCM0238 | AC120 | HCT 116 | C2041(1.23) | LDD0555 | [10] |
| LDCM0239 | AC121 | HCT 116 | C2041(0.92) | LDD0556 | [10] |
| LDCM0240 | AC122 | HCT 116 | C2041(1.00) | LDD0557 | [10] |
| LDCM0241 | AC123 | HCT 116 | C2041(0.93) | LDD0558 | [10] |
| LDCM0242 | AC124 | HCT 116 | C2041(0.97) | LDD0559 | [10] |
| LDCM0243 | AC125 | HCT 116 | C2041(1.00) | LDD0560 | [10] |
| LDCM0244 | AC126 | HCT 116 | C2041(1.17) | LDD0561 | [10] |
| LDCM0245 | AC127 | HCT 116 | C2041(1.19) | LDD0562 | [10] |
| LDCM0246 | AC128 | HCT 116 | C2041(0.94); C2083(0.93); C2088(0.93) | LDD0563 | [10] |
| LDCM0247 | AC129 | HCT 116 | C2041(0.75); C2083(1.05); C2088(1.05) | LDD0564 | [10] |
| LDCM0249 | AC130 | HCT 116 | C2041(0.95); C2083(1.03); C2088(1.03) | LDD0566 | [10] |
| LDCM0250 | AC131 | HCT 116 | C2041(0.88); C2083(0.74); C2088(0.74) | LDD0567 | [10] |
| LDCM0251 | AC132 | HCT 116 | C2041(0.74); C2083(0.61); C2088(0.61) | LDD0568 | [10] |
| LDCM0252 | AC133 | HCT 116 | C2041(0.89); C2083(0.82); C2088(0.82) | LDD0569 | [10] |
| LDCM0253 | AC134 | HCT 116 | C2041(0.91); C2083(0.92); C2088(0.92) | LDD0570 | [10] |
| LDCM0254 | AC135 | HCT 116 | C2041(0.93); C2083(1.07); C2088(1.07) | LDD0571 | [10] |
| LDCM0255 | AC136 | HCT 116 | C2041(0.88); C2083(0.56); C2088(0.56) | LDD0572 | [10] |
| LDCM0256 | AC137 | HCT 116 | C2041(0.79); C2083(0.69); C2088(0.69) | LDD0573 | [10] |
| LDCM0257 | AC138 | HCT 116 | C2041(0.89); C2083(0.85); C2088(0.85) | LDD0574 | [10] |
| LDCM0258 | AC139 | HCT 116 | C2041(0.86); C2083(0.89); C2088(0.89) | LDD0575 | [10] |
| LDCM0259 | AC14 | HCT 116 | C2041(1.08); C2065(1.22) | LDD0576 | [10] |
| LDCM0260 | AC140 | HCT 116 | C2041(1.02); C2083(0.74); C2088(0.74) | LDD0577 | [10] |
| LDCM0261 | AC141 | HCT 116 | C2041(0.88); C2083(0.91); C2088(0.91) | LDD0578 | [10] |
| LDCM0262 | AC142 | HCT 116 | C2041(0.80); C2083(0.60); C2088(0.60) | LDD0579 | [10] |
| LDCM0263 | AC143 | HCT 116 | C2041(1.18) | LDD0580 | [10] |
| LDCM0264 | AC144 | HCT 116 | C2041(1.32) | LDD0581 | [10] |
| LDCM0265 | AC145 | HCT 116 | C2041(1.10) | LDD0582 | [10] |
| LDCM0266 | AC146 | HCT 116 | C2041(1.36) | LDD0583 | [10] |
| LDCM0267 | AC147 | HCT 116 | C2041(1.26) | LDD0584 | [10] |
| LDCM0268 | AC148 | HCT 116 | C2041(2.53) | LDD0585 | [10] |
| LDCM0269 | AC149 | HCT 116 | C2041(1.93) | LDD0586 | [10] |
| LDCM0270 | AC15 | HCT 116 | C2041(1.22); C2065(1.41) | LDD0587 | [10] |
| LDCM0271 | AC150 | HCT 116 | C2041(1.20) | LDD0588 | [10] |
| LDCM0272 | AC151 | HCT 116 | C2041(1.06) | LDD0589 | [10] |
| LDCM0273 | AC152 | HCT 116 | C2041(1.36) | LDD0590 | [10] |
| LDCM0274 | AC153 | HCT 116 | C2041(2.12) | LDD0591 | [10] |
| LDCM0621 | AC154 | HCT 116 | C2041(1.39) | LDD2158 | [10] |
| LDCM0622 | AC155 | HCT 116 | C2041(1.46) | LDD2159 | [10] |
| LDCM0623 | AC156 | HCT 116 | C2041(1.20) | LDD2160 | [10] |
| LDCM0624 | AC157 | HCT 116 | C2041(1.25) | LDD2161 | [10] |
| LDCM0276 | AC17 | HCT 116 | C2041(0.78); C2065(1.19) | LDD0593 | [10] |
| LDCM0277 | AC18 | HCT 116 | C2041(1.07); C2065(1.40) | LDD0594 | [10] |
| LDCM0278 | AC19 | HCT 116 | C2041(0.85); C2065(1.57) | LDD0595 | [10] |
| LDCM0279 | AC2 | HCT 116 | C2041(1.07) | LDD0596 | [10] |
| LDCM0280 | AC20 | HCT 116 | C2041(0.78); C2065(1.36) | LDD0597 | [10] |
| LDCM0281 | AC21 | HCT 116 | C2041(0.85); C2065(1.91) | LDD0598 | [10] |
| LDCM0282 | AC22 | HCT 116 | C2041(0.86); C2065(1.53) | LDD0599 | [10] |
| LDCM0283 | AC23 | HCT 116 | C2041(0.80); C2065(1.30) | LDD0600 | [10] |
| LDCM0284 | AC24 | HCT 116 | C2041(0.74); C2065(1.14) | LDD0601 | [10] |
| LDCM0285 | AC25 | HCT 116 | C2041(0.95); C2065(1.49) | LDD0602 | [10] |
| LDCM0286 | AC26 | HCT 116 | C2041(1.02); C2065(1.26) | LDD0603 | [10] |
| LDCM0287 | AC27 | HCT 116 | C2041(1.11); C2065(1.45) | LDD0604 | [10] |
| LDCM0288 | AC28 | HCT 116 | C2041(1.35); C2065(1.71) | LDD0605 | [10] |
| LDCM0289 | AC29 | HCT 116 | C2041(1.16); C2065(1.53) | LDD0606 | [10] |
| LDCM0290 | AC3 | HCT 116 | C2041(1.11) | LDD0607 | [10] |
| LDCM0291 | AC30 | HCT 116 | C2041(1.08); C2065(1.37) | LDD0608 | [10] |
| LDCM0292 | AC31 | HCT 116 | C2041(1.08); C2065(2.18) | LDD0609 | [10] |
| LDCM0293 | AC32 | HCT 116 | C2041(1.41); C2065(1.67) | LDD0610 | [10] |
| LDCM0294 | AC33 | HCT 116 | C2041(1.32); C2065(1.70) | LDD0611 | [10] |
| LDCM0295 | AC34 | HCT 116 | C2041(1.50); C2065(2.20) | LDD0612 | [10] |
| LDCM0296 | AC35 | HCT 116 | C2083(1.00); C2088(1.00) | LDD0613 | [10] |
| LDCM0297 | AC36 | HCT 116 | C2083(1.05); C2088(1.05) | LDD0614 | [10] |
| LDCM0298 | AC37 | HCT 116 | C2083(0.96); C2088(0.96) | LDD0615 | [10] |
| LDCM0299 | AC38 | HCT 116 | C2083(1.08); C2088(1.08) | LDD0616 | [10] |
| LDCM0300 | AC39 | HCT 116 | C2083(0.89); C2088(0.89) | LDD0617 | [10] |
| LDCM0301 | AC4 | HCT 116 | C2041(1.17) | LDD0618 | [10] |
| LDCM0302 | AC40 | HCT 116 | C2083(0.99); C2088(0.99) | LDD0619 | [10] |
| LDCM0303 | AC41 | HCT 116 | C2083(1.40); C2088(1.40) | LDD0620 | [10] |
| LDCM0304 | AC42 | HCT 116 | C2083(1.06); C2088(1.06) | LDD0621 | [10] |
| LDCM0305 | AC43 | HCT 116 | C2083(1.50); C2088(1.50) | LDD0622 | [10] |
| LDCM0306 | AC44 | HCT 116 | C2083(0.89); C2088(0.89) | LDD0623 | [10] |
| LDCM0307 | AC45 | HCT 116 | C2083(1.00); C2088(1.00) | LDD0624 | [10] |
| LDCM0308 | AC46 | HCT 116 | C2065(0.96); C2041(1.11) | LDD0625 | [10] |
| LDCM0309 | AC47 | HCT 116 | C2041(1.05); C2065(1.11) | LDD0626 | [10] |
| LDCM0310 | AC48 | HCT 116 | C2065(1.01); C2041(1.02) | LDD0627 | [10] |
| LDCM0311 | AC49 | HCT 116 | C2041(1.33); C2065(2.03) | LDD0628 | [10] |
| LDCM0312 | AC5 | HCT 116 | C2041(1.00) | LDD0629 | [10] |
| LDCM0313 | AC50 | HCT 116 | C2041(1.13); C2065(1.45) | LDD0630 | [10] |
| LDCM0314 | AC51 | HCT 116 | C2065(0.86); C2041(0.98) | LDD0631 | [10] |
| LDCM0315 | AC52 | HCT 116 | C2065(0.87); C2041(1.12) | LDD0632 | [10] |
| LDCM0316 | AC53 | HCT 116 | C2065(1.18); C2041(1.35) | LDD0633 | [10] |
| LDCM0317 | AC54 | HCT 116 | C2041(1.07); C2065(1.35) | LDD0634 | [10] |
| LDCM0318 | AC55 | HCT 116 | C2041(1.39); C2065(1.68) | LDD0635 | [10] |
| LDCM0319 | AC56 | HCT 116 | C2041(1.20); C2065(2.42) | LDD0636 | [10] |
| LDCM0320 | AC57 | HCT 116 | C2041(1.34) | LDD0637 | [10] |
| LDCM0321 | AC58 | HCT 116 | C2041(1.65) | LDD0638 | [10] |
| LDCM0322 | AC59 | HCT 116 | C2041(1.38) | LDD0639 | [10] |
| LDCM0323 | AC6 | HCT 116 | C2065(1.01); C2041(1.11) | LDD0640 | [10] |
| LDCM0324 | AC60 | HCT 116 | C2041(1.62) | LDD0641 | [10] |
| LDCM0325 | AC61 | HCT 116 | C2041(1.10) | LDD0642 | [10] |
| LDCM0326 | AC62 | HCT 116 | C2041(1.36) | LDD0643 | [10] |
| LDCM0327 | AC63 | HCT 116 | C2041(1.32) | LDD0644 | [10] |
| LDCM0328 | AC64 | HCT 116 | C2041(1.31) | LDD0645 | [10] |
| LDCM0329 | AC65 | HCT 116 | C2041(1.00) | LDD0646 | [10] |
| LDCM0330 | AC66 | HCT 116 | C2041(0.93) | LDD0647 | [10] |
| LDCM0331 | AC67 | HCT 116 | C2041(1.28) | LDD0648 | [10] |
| LDCM0332 | AC68 | HCT 116 | C2041(0.82) | LDD0649 | [10] |
| LDCM0333 | AC69 | HCT 116 | C2041(0.65) | LDD0650 | [10] |
| LDCM0334 | AC7 | HCT 116 | C2065(0.88); C2041(0.89) | LDD0651 | [10] |
| LDCM0335 | AC70 | HCT 116 | C2041(0.75) | LDD0652 | [10] |
| LDCM0336 | AC71 | HCT 116 | C2041(0.78) | LDD0653 | [10] |
| LDCM0337 | AC72 | HCT 116 | C2041(1.02) | LDD0654 | [10] |
| LDCM0338 | AC73 | HCT 116 | C2041(1.08) | LDD0655 | [10] |
| LDCM0339 | AC74 | HCT 116 | C2041(0.91) | LDD0656 | [10] |
| LDCM0340 | AC75 | HCT 116 | C2041(0.84) | LDD0657 | [10] |
| LDCM0341 | AC76 | HCT 116 | C2041(0.84) | LDD0658 | [10] |
| LDCM0342 | AC77 | HCT 116 | C2041(0.91) | LDD0659 | [10] |
| LDCM0343 | AC78 | HCT 116 | C2041(0.81) | LDD0660 | [10] |
| LDCM0344 | AC79 | HCT 116 | C2041(0.87) | LDD0661 | [10] |
| LDCM0345 | AC8 | HCT 116 | C2041(1.06); C2065(1.29) | LDD0662 | [10] |
| LDCM0346 | AC80 | HCT 116 | C2041(0.95) | LDD0663 | [10] |
| LDCM0347 | AC81 | HCT 116 | C2041(0.66) | LDD0664 | [10] |
| LDCM0348 | AC82 | HCT 116 | C2041(0.86) | LDD0665 | [10] |
| LDCM0349 | AC83 | HCT 116 | C2041(1.74); C2083(2.22); C2088(2.22); C2065(2.81) | LDD0666 | [10] |
| LDCM0350 | AC84 | HCT 116 | C2041(1.49); C2083(1.68); C2088(1.68); C2065(1.92) | LDD0667 | [10] |
| LDCM0351 | AC85 | HCT 116 | C2065(1.28); C2083(1.45); C2088(1.45); C2041(1.60) | LDD0668 | [10] |
| LDCM0352 | AC86 | HCT 116 | C2041(1.33); C2083(1.53); C2088(1.53); C2065(1.54) | LDD0669 | [10] |
| LDCM0353 | AC87 | HCT 116 | C2041(1.08); C2065(1.28); C2083(1.49); C2088(1.49) | LDD0670 | [10] |
| LDCM0354 | AC88 | HCT 116 | C2041(1.21); C2065(1.24); C2083(1.54); C2088(1.54) | LDD0671 | [10] |
| LDCM0355 | AC89 | HCT 116 | C2041(1.45); C2065(1.46); C2083(1.98); C2088(1.98) | LDD0672 | [10] |
| LDCM0357 | AC90 | HCT 116 | C2041(1.00); C2065(1.36); C2083(1.48); C2088(1.48) | LDD0674 | [10] |
| LDCM0358 | AC91 | HCT 116 | C2041(1.71); C2065(2.21); C2083(2.73); C2088(2.73) | LDD0675 | [10] |
| LDCM0359 | AC92 | HCT 116 | C2041(1.59); C2065(2.32); C2083(2.51); C2088(2.51) | LDD0676 | [10] |
| LDCM0360 | AC93 | HCT 116 | C2041(1.30); C2065(1.73); C2083(1.81); C2088(1.81) | LDD0677 | [10] |
| LDCM0361 | AC94 | HCT 116 | C2041(1.58); C2065(1.83); C2083(1.83); C2088(1.83) | LDD0678 | [10] |
| LDCM0362 | AC95 | HCT 116 | C2041(1.27); C2065(1.40); C2083(1.55); C2088(1.55) | LDD0679 | [10] |
| LDCM0363 | AC96 | HCT 116 | C2065(1.35); C2041(1.43); C2083(2.33); C2088(2.33) | LDD0680 | [10] |
| LDCM0364 | AC97 | HCT 116 | C2041(1.69); C2065(1.85); C2083(2.43); C2088(2.43) | LDD0681 | [10] |
| LDCM0365 | AC98 | HCT 116 | C2041(1.89) | LDD0682 | [10] |
| LDCM0366 | AC99 | HCT 116 | C2041(0.84) | LDD0683 | [10] |
| LDCM0248 | AKOS034007472 | HCT 116 | C2041(1.03); C2065(1.26) | LDD0565 | [10] |
| LDCM0356 | AKOS034007680 | HCT 116 | C2065(0.84); C2041(0.95) | LDD0673 | [10] |
| LDCM0275 | AKOS034007705 | HCT 116 | C2065(1.57); C2041(1.62) | LDD0592 | [10] |
| LDCM0156 | Aniline | NCI-H1299 | 12.50 | LDD0403 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C2041(1.18); C2065(1.06); C963(1.01); C382(0.99) | LDD0078 | [10] |
| LDCM0087 | Capsaicin | HEK-293T | 44.47 | LDD0185 | [15] |
| LDCM0630 | CCW28-3 | 231MFP | C1814(1.58) | LDD2214 | [22] |
| LDCM0108 | Chloroacetamide | HeLa | C73(0.00); C963(0.00); C2041(0.00); C1310(0.00) | LDD0222 | [18] |
| LDCM0367 | CL1 | HCT 116 | C2041(1.18) | LDD0684 | [10] |
| LDCM0368 | CL10 | HCT 116 | C2041(1.17) | LDD0685 | [10] |
| LDCM0369 | CL100 | HCT 116 | C2041(1.21) | LDD0686 | [10] |
| LDCM0370 | CL101 | HCT 116 | C2041(1.20); C2065(1.36) | LDD0687 | [10] |
| LDCM0371 | CL102 | HCT 116 | C2065(0.82); C2041(0.95) | LDD0688 | [10] |
| LDCM0372 | CL103 | HCT 116 | C2041(0.92); C2065(0.95) | LDD0689 | [10] |
| LDCM0373 | CL104 | HCT 116 | C2041(0.98); C2065(1.07) | LDD0690 | [10] |
| LDCM0374 | CL105 | HCT 116 | C2041(0.84); C2065(1.23) | LDD0691 | [10] |
| LDCM0375 | CL106 | HCT 116 | C2041(0.76); C2065(1.85) | LDD0692 | [10] |
| LDCM0376 | CL107 | HCT 116 | C2041(0.91); C2065(1.36) | LDD0693 | [10] |
| LDCM0377 | CL108 | HCT 116 | C2041(0.71); C2065(2.01) | LDD0694 | [10] |
| LDCM0378 | CL109 | HCT 116 | C2041(0.84); C2065(1.42) | LDD0695 | [10] |
| LDCM0379 | CL11 | HCT 116 | C2041(0.80) | LDD0696 | [10] |
| LDCM0380 | CL110 | HCT 116 | C2041(0.72); C2065(1.86) | LDD0697 | [10] |
| LDCM0381 | CL111 | HCT 116 | C2041(0.82); C2065(1.67) | LDD0698 | [10] |
| LDCM0382 | CL112 | HCT 116 | C2041(1.08); C2065(1.31) | LDD0699 | [10] |
| LDCM0383 | CL113 | HCT 116 | C2041(1.36); C2065(1.98) | LDD0700 | [10] |
| LDCM0384 | CL114 | HCT 116 | C2041(0.89); C2065(1.25) | LDD0701 | [10] |
| LDCM0385 | CL115 | HCT 116 | C2041(1.29); C2065(1.51) | LDD0702 | [10] |
| LDCM0386 | CL116 | HCT 116 | C2041(1.00); C2065(1.32) | LDD0703 | [10] |
| LDCM0387 | CL117 | HCT 116 | C2083(0.94); C2088(0.94) | LDD0704 | [10] |
| LDCM0388 | CL118 | HCT 116 | C2083(1.27); C2088(1.27) | LDD0705 | [10] |
| LDCM0389 | CL119 | HCT 116 | C2083(1.23); C2088(1.23) | LDD0706 | [10] |
| LDCM0390 | CL12 | HCT 116 | C2041(1.47) | LDD0707 | [10] |
| LDCM0391 | CL120 | HCT 116 | C2083(1.45); C2088(1.45) | LDD0708 | [10] |
| LDCM0392 | CL121 | HCT 116 | C2065(1.26); C2041(1.33) | LDD0709 | [10] |
| LDCM0393 | CL122 | HCT 116 | C2065(1.04); C2041(1.08) | LDD0710 | [10] |
| LDCM0394 | CL123 | HCT 116 | C2041(1.00); C2065(1.61) | LDD0711 | [10] |
| LDCM0395 | CL124 | HCT 116 | C2041(1.18); C2065(1.45) | LDD0712 | [10] |
| LDCM0396 | CL125 | HCT 116 | C2041(1.09) | LDD0713 | [10] |
| LDCM0397 | CL126 | HCT 116 | C2041(1.11) | LDD0714 | [10] |
| LDCM0398 | CL127 | HCT 116 | C2041(1.06) | LDD0715 | [10] |
| LDCM0399 | CL128 | HCT 116 | C2041(1.51) | LDD0716 | [10] |
| LDCM0400 | CL13 | HCT 116 | C2041(1.25) | LDD0717 | [10] |
| LDCM0401 | CL14 | HCT 116 | C2041(1.53) | LDD0718 | [10] |
| LDCM0402 | CL15 | HCT 116 | C2041(1.27) | LDD0719 | [10] |
| LDCM0403 | CL16 | HCT 116 | C2041(1.20) | LDD0720 | [10] |
| LDCM0404 | CL17 | HCT 116 | C2041(1.10) | LDD0721 | [10] |
| LDCM0405 | CL18 | HCT 116 | C2041(1.12) | LDD0722 | [10] |
| LDCM0406 | CL19 | HCT 116 | C2041(1.20) | LDD0723 | [10] |
| LDCM0407 | CL2 | HCT 116 | C2041(1.28) | LDD0724 | [10] |
| LDCM0408 | CL20 | HCT 116 | C2041(1.40) | LDD0725 | [10] |
| LDCM0409 | CL21 | HCT 116 | C2041(1.14) | LDD0726 | [10] |
| LDCM0410 | CL22 | HCT 116 | C2041(1.00) | LDD0727 | [10] |
| LDCM0411 | CL23 | HCT 116 | C2041(1.03) | LDD0728 | [10] |
| LDCM0412 | CL24 | HCT 116 | C2041(1.03) | LDD0729 | [10] |
| LDCM0413 | CL25 | HCT 116 | C2041(0.92) | LDD0730 | [10] |
| LDCM0414 | CL26 | HCT 116 | C2041(1.00) | LDD0731 | [10] |
| LDCM0415 | CL27 | HCT 116 | C2041(1.05) | LDD0732 | [10] |
| LDCM0416 | CL28 | HCT 116 | C2041(1.07) | LDD0733 | [10] |
| LDCM0417 | CL29 | HCT 116 | C2041(1.35) | LDD0734 | [10] |
| LDCM0418 | CL3 | HCT 116 | C2041(1.31) | LDD0735 | [10] |
| LDCM0419 | CL30 | HCT 116 | C2041(0.91) | LDD0736 | [10] |
| LDCM0420 | CL31 | HCT 116 | C2041(0.92) | LDD0737 | [10] |
| LDCM0421 | CL32 | HCT 116 | C2041(1.20) | LDD0738 | [10] |
| LDCM0422 | CL33 | HCT 116 | C2041(1.01) | LDD0739 | [10] |
| LDCM0423 | CL34 | HCT 116 | C2041(0.93) | LDD0740 | [10] |
| LDCM0424 | CL35 | HCT 116 | C2041(1.16) | LDD0741 | [10] |
| LDCM0425 | CL36 | HCT 116 | C2041(1.13) | LDD0742 | [10] |
| LDCM0426 | CL37 | HCT 116 | C2041(1.16) | LDD0743 | [10] |
| LDCM0428 | CL39 | HCT 116 | C2041(1.07) | LDD0745 | [10] |
| LDCM0429 | CL4 | HCT 116 | C2041(1.20) | LDD0746 | [10] |
| LDCM0430 | CL40 | HCT 116 | C2041(1.02) | LDD0747 | [10] |
| LDCM0431 | CL41 | HCT 116 | C2041(0.98) | LDD0748 | [10] |
| LDCM0432 | CL42 | HCT 116 | C2041(1.09) | LDD0749 | [10] |
| LDCM0433 | CL43 | HCT 116 | C2041(1.01) | LDD0750 | [10] |
| LDCM0434 | CL44 | HCT 116 | C2041(1.00) | LDD0751 | [10] |
| LDCM0435 | CL45 | HCT 116 | C2041(0.96) | LDD0752 | [10] |
| LDCM0436 | CL46 | HCT 116 | C2041(1.26) | LDD0753 | [10] |
| LDCM0437 | CL47 | HCT 116 | C2041(1.00) | LDD0754 | [10] |
| LDCM0438 | CL48 | HCT 116 | C2041(0.96) | LDD0755 | [10] |
| LDCM0439 | CL49 | HCT 116 | C2041(0.99) | LDD0756 | [10] |
| LDCM0440 | CL5 | HCT 116 | C2041(1.27) | LDD0757 | [10] |
| LDCM0441 | CL50 | HCT 116 | C2041(0.96) | LDD0758 | [10] |
| LDCM0442 | CL51 | HCT 116 | C2041(1.42) | LDD0759 | [10] |
| LDCM0443 | CL52 | HCT 116 | C2041(1.70) | LDD0760 | [10] |
| LDCM0444 | CL53 | HCT 116 | C2041(1.24) | LDD0761 | [10] |
| LDCM0445 | CL54 | HCT 116 | C2041(1.07) | LDD0762 | [10] |
| LDCM0446 | CL55 | HCT 116 | C2041(1.00) | LDD0763 | [10] |
| LDCM0447 | CL56 | HCT 116 | C2041(0.99) | LDD0764 | [10] |
| LDCM0448 | CL57 | HCT 116 | C2041(0.99) | LDD0765 | [10] |
| LDCM0449 | CL58 | HCT 116 | C2041(1.07) | LDD0766 | [10] |
| LDCM0450 | CL59 | HCT 116 | C2041(1.03) | LDD0767 | [10] |
| LDCM0451 | CL6 | HCT 116 | C2041(1.67) | LDD0768 | [10] |
| LDCM0452 | CL60 | HCT 116 | C2041(0.98) | LDD0769 | [10] |
| LDCM0453 | CL61 | HCT 116 | C2041(1.10); C2083(0.93); C2088(0.93) | LDD0770 | [10] |
| LDCM0454 | CL62 | HCT 116 | C2041(1.22); C2083(0.71); C2088(0.71) | LDD0771 | [10] |
| LDCM0455 | CL63 | HCT 116 | C2041(1.53); C2083(0.87); C2088(0.87) | LDD0772 | [10] |
| LDCM0456 | CL64 | HCT 116 | C2041(1.54); C2083(0.97); C2088(0.97) | LDD0773 | [10] |
| LDCM0457 | CL65 | HCT 116 | C2041(1.08); C2083(0.78); C2088(0.78) | LDD0774 | [10] |
| LDCM0458 | CL66 | HCT 116 | C2041(2.11); C2083(0.58); C2088(0.58) | LDD0775 | [10] |
| LDCM0459 | CL67 | HCT 116 | C2041(1.62); C2083(1.02); C2088(1.02) | LDD0776 | [10] |
| LDCM0460 | CL68 | HCT 116 | C2041(1.53); C2083(0.72); C2088(0.72) | LDD0777 | [10] |
| LDCM0461 | CL69 | HCT 116 | C2041(1.46); C2083(0.69); C2088(0.69) | LDD0778 | [10] |
| LDCM0462 | CL7 | HCT 116 | C2041(1.21) | LDD0779 | [10] |
| LDCM0463 | CL70 | HCT 116 | C2041(1.69); C2083(0.99); C2088(0.99) | LDD0780 | [10] |
| LDCM0464 | CL71 | HCT 116 | C2041(1.58); C2083(0.99); C2088(0.99) | LDD0781 | [10] |
| LDCM0465 | CL72 | HCT 116 | C2041(1.23); C2083(1.35); C2088(1.35) | LDD0782 | [10] |
| LDCM0466 | CL73 | HCT 116 | C2041(1.59); C2083(1.06); C2088(1.06) | LDD0783 | [10] |
| LDCM0467 | CL74 | HCT 116 | C2041(1.74); C2083(1.40); C2088(1.40) | LDD0784 | [10] |
| LDCM0469 | CL76 | HCT 116 | C2041(0.85); C2083(0.43); C2088(0.43) | LDD0786 | [10] |
| LDCM0470 | CL77 | HCT 116 | C2041(0.94); C2083(0.69); C2088(0.69) | LDD0787 | [10] |
| LDCM0471 | CL78 | HCT 116 | C2041(0.90); C2083(1.15); C2088(1.15) | LDD0788 | [10] |
| LDCM0472 | CL79 | HCT 116 | C2041(0.97); C2083(1.27); C2088(1.27) | LDD0789 | [10] |
| LDCM0473 | CL8 | HCT 116 | C2041(1.00) | LDD0790 | [10] |
| LDCM0474 | CL80 | HCT 116 | C2041(0.89); C2083(1.06); C2088(1.06) | LDD0791 | [10] |
| LDCM0475 | CL81 | HCT 116 | C2041(0.80); C2083(0.83); C2088(0.83) | LDD0792 | [10] |
| LDCM0476 | CL82 | HCT 116 | C2041(0.66); C2083(0.70); C2088(0.70) | LDD0793 | [10] |
| LDCM0477 | CL83 | HCT 116 | C2041(0.87); C2083(0.79); C2088(0.79) | LDD0794 | [10] |
| LDCM0478 | CL84 | HCT 116 | C2041(0.80); C2083(0.94); C2088(0.94) | LDD0795 | [10] |
| LDCM0479 | CL85 | HCT 116 | C2041(1.00); C2083(0.90); C2088(0.90) | LDD0796 | [10] |
| LDCM0480 | CL86 | HCT 116 | C2041(0.79); C2083(0.65); C2088(0.65) | LDD0797 | [10] |
| LDCM0481 | CL87 | HCT 116 | C2041(0.78); C2083(0.72); C2088(0.72) | LDD0798 | [10] |
| LDCM0482 | CL88 | HCT 116 | C2041(0.81); C2083(0.91); C2088(0.91) | LDD0799 | [10] |
| LDCM0483 | CL89 | HCT 116 | C2041(0.99); C2083(1.14); C2088(1.14) | LDD0800 | [10] |
| LDCM0484 | CL9 | HCT 116 | C2041(1.13) | LDD0801 | [10] |
| LDCM0485 | CL90 | HCT 116 | C2041(0.73); C2083(0.72); C2088(0.72) | LDD0802 | [10] |
| LDCM0486 | CL91 | HCT 116 | C2041(1.32) | LDD0803 | [10] |
| LDCM0487 | CL92 | HCT 116 | C2041(1.16) | LDD0804 | [10] |
| LDCM0488 | CL93 | HCT 116 | C2041(1.09) | LDD0805 | [10] |
| LDCM0489 | CL94 | HCT 116 | C2041(1.32) | LDD0806 | [10] |
| LDCM0490 | CL95 | HCT 116 | C2041(1.24) | LDD0807 | [10] |
| LDCM0491 | CL96 | HCT 116 | C2041(1.44) | LDD0808 | [10] |
| LDCM0492 | CL97 | HCT 116 | C2041(0.99) | LDD0809 | [10] |
| LDCM0493 | CL98 | HCT 116 | C2041(1.09) | LDD0810 | [10] |
| LDCM0494 | CL99 | HCT 116 | C2041(1.13) | LDD0811 | [10] |
| LDCM0028 | Dobutamine | HEK-293T | 6.71 | LDD0180 | [15] |
| LDCM0027 | Dopamine | HEK-293T | 14.40 | LDD0179 | [15] |
| LDCM0495 | E2913 | HEK-293T | C1228(1.02); C963(0.96); C1814(1.02); C2065(1.08) | LDD1698 | [23] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C1814(0.99) | LDD1702 | [6] |
| LDCM0468 | Fragment33 | HCT 116 | C2041(1.41); C2083(0.66); C2088(0.66) | LDD0785 | [10] |
| LDCM0427 | Fragment51 | HCT 116 | C2041(1.17) | LDD0744 | [10] |
| LDCM0116 | HHS-0101 | DM93 | Y1923(0.06); Y1336(0.36); Y2023(0.62); Y1957(0.62) | LDD0264 | [8] |
| LDCM0117 | HHS-0201 | DM93 | Y1923(0.44); Y2057(0.62); Y2023(0.64); Y2059(0.67) | LDD0265 | [8] |
| LDCM0118 | HHS-0301 | DM93 | Y1906(0.45); Y2057(0.55); Y1974(0.62); Y1955(0.63) | LDD0266 | [8] |
| LDCM0119 | HHS-0401 | DM93 | Y2057(0.56); Y1974(0.63); Y1957(0.66); Y1762(0.73) | LDD0267 | [8] |
| LDCM0120 | HHS-0701 | DM93 | Y1762(0.59); Y1336(0.64); Y1974(0.71); Y2357(0.75) | LDD0268 | [8] |
| LDCM0107 | IAA | HeLa | C73(0.00); H969(0.00); C190(0.00); H204(0.00) | LDD0221 | [18] |
| LDCM0022 | KB02 | HCT 116 | C382(3.93); C2361(2.04); C2083(3.81); C2088(3.81) | LDD0080 | [10] |
| LDCM0023 | KB03 | HCT 116 | C382(6.07); C2361(2.14); C2083(2.28); C2088(2.28) | LDD0081 | [10] |
| LDCM0024 | KB05 | HCT 116 | C382(6.47); C2361(2.43); C2083(2.09); C2088(2.09) | LDD0082 | [10] |
| LDCM0030 | Luteolin | HEK-293T | 68.59 | LDD0182 | [15] |
| LDCM0528 | N-(4-bromophenyl)-2-cyano-N-phenylacetamide | MDA-MB-231 | C1228(0.58) | LDD2121 | [6] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [18] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C1228(0.70) | LDD2089 | [6] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C1228(1.09); C1814(1.32) | LDD2090 | [6] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C1228(0.85) | LDD2098 | [6] |
| LDCM0512 | Nucleophilic fragment 19a | MDA-MB-231 | C1228(1.05) | LDD2105 | [6] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C1228(1.02) | LDD2107 | [6] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C1228(1.00) | LDD2109 | [6] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C1814(0.46) | LDD2118 | [6] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C1228(2.45) | LDD2119 | [6] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C1228(1.04) | LDD2125 | [6] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C1228(1.05) | LDD2129 | [6] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C1228(0.72) | LDD2135 | [6] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C1228(1.29) | LDD2136 | [6] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C1228(1.11) | LDD2140 | [6] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C1228(4.64) | LDD2144 | [6] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C1228(1.08) | LDD2146 | [6] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C1228(0.48) | LDD2150 | [6] |
| LDCM0032 | Oleacein | HEK-293T | 29.94 | LDD0184 | [15] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C382(1.85); C2361(0.45) | LDD2207 | [24] |
| LDCM0029 | Quercetin | HEK-293T | 29.80 | LDD0181 | [15] |
| LDCM0021 | THZ1 | HCT 116 | C2083(1.06); C2088(1.06); C2041(0.98) | LDD2173 | [10] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Leucine-rich repeat serine/threonine-protein kinase 2 (LRRK2) | TKL Ser/Thr protein kinase family | Q5S007 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Cellular tumor antigen p53 (TP53) | P53 family | P04637 | |||
GPCR
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| 5-hydroxytryptamine receptor 6 (HTR6) | G-protein coupled receptor 1 family | P50406 | |||
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Kin of IRRE-like protein 3 (KIRREL3) | Immunoglobulin superfamily | Q8IZU9 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Gigaxonin (GAN) | . | Q9H2C0 | |||
References
