Details of the Target
General Information of Target
| Target ID | LDTP13599 | |||||
|---|---|---|---|---|---|---|
| Target Name | Calcium load-activated calcium channel (TMCO1) | |||||
| Gene Name | TMCO1 | |||||
| Gene ID | 54499 | |||||
| Synonyms |
TMCC4; Calcium load-activated calcium channel; CLAC channel; GEL complex subunit TMCO1; Transmembrane and coiled-coil domain-containing protein 1; Transmembrane and coiled-coil domains protein 4; Xenogeneic cross-immune protein PCIA3
|
|||||
| 3D Structure | ||||||
| Sequence |
MRRRAARGPGPPPPGPGLSRLPLPLLLLLALGTRGGCAAPAPAPRAEDLSLGVEWLSRFG
YLPPADPTTGQLQTQEELSKAITAMQQFGGLEATGILDEATLALMKTPRCSLPDLPVLTQ ARRRRQAPAPTKWNKRNLSWRVRTFPRDSPLGHDTVRALMYYALKVWSDIAPLNFHEVAG SAADIQIDFSKADHNDGYPFDGPGGTVAHAFFPGHHHTAGDTHFDDDEAWTFRSSDAHGM DLFAVAVHEFGHAIGLSHVAAAHSIMRPYYQGPVGDPLRYGLPYEDKVRVWQLYGVRESV SPTAQPEEPPLLPEPPDNRSSAPPRKDVPHRCSTHFDAVAQIRGEAFFFKGKYFWRLTRD RHLVSLQPAQMHRFWRGLPLHLDSVDAVYERTSDHKIVFFKGDRYWVFKDNNVEEGYPRP VSDFSLPPGGIDAAFSWAHNDRTYFFKDQLYWRYDDHTRHMDPGYPAQSPLWRGVPSTLD DAMRWSDGASYFFRGQEYWKVLDGELEVAPGYPQSTARDWLVCGDSQADGSVAAGVDAAE GPRAPPGQHDQSRSEDGYEVCSCTSGASSPPGAPGPLVAATMLLLLPPLSPGALWTAAQA LTL |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
TMCO1 family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Calcium-selective channel required to prevent calcium stores from overfilling, thereby playing a key role in calcium homeostasis. In response to endoplasmic reticulum (ER) overloading, assembles into a homotetramer, forming a functional calcium-selective channel, regulating the calcium content in endoplasmic reticulum store. Component of the multi-pass translocon (MPT) complex that mediates insertion of multi-pass membrane proteins into the lipid bilayer of membranes. The MPT complex takes over after the SEC61 complex: following membrane insertion of the first few transmembrane segments of proteins by the SEC61 complex, the MPT complex occludes the lateral gate of the SEC61 complex to promote insertion of subsequent transmembrane regions. Within the MPT complex, the GEL subcomplex may mediate insertion of transmembrane regions into the membrane.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y124(6.19) | LDD0260 | [1] | |
|
1oxF11yne Probe Info |
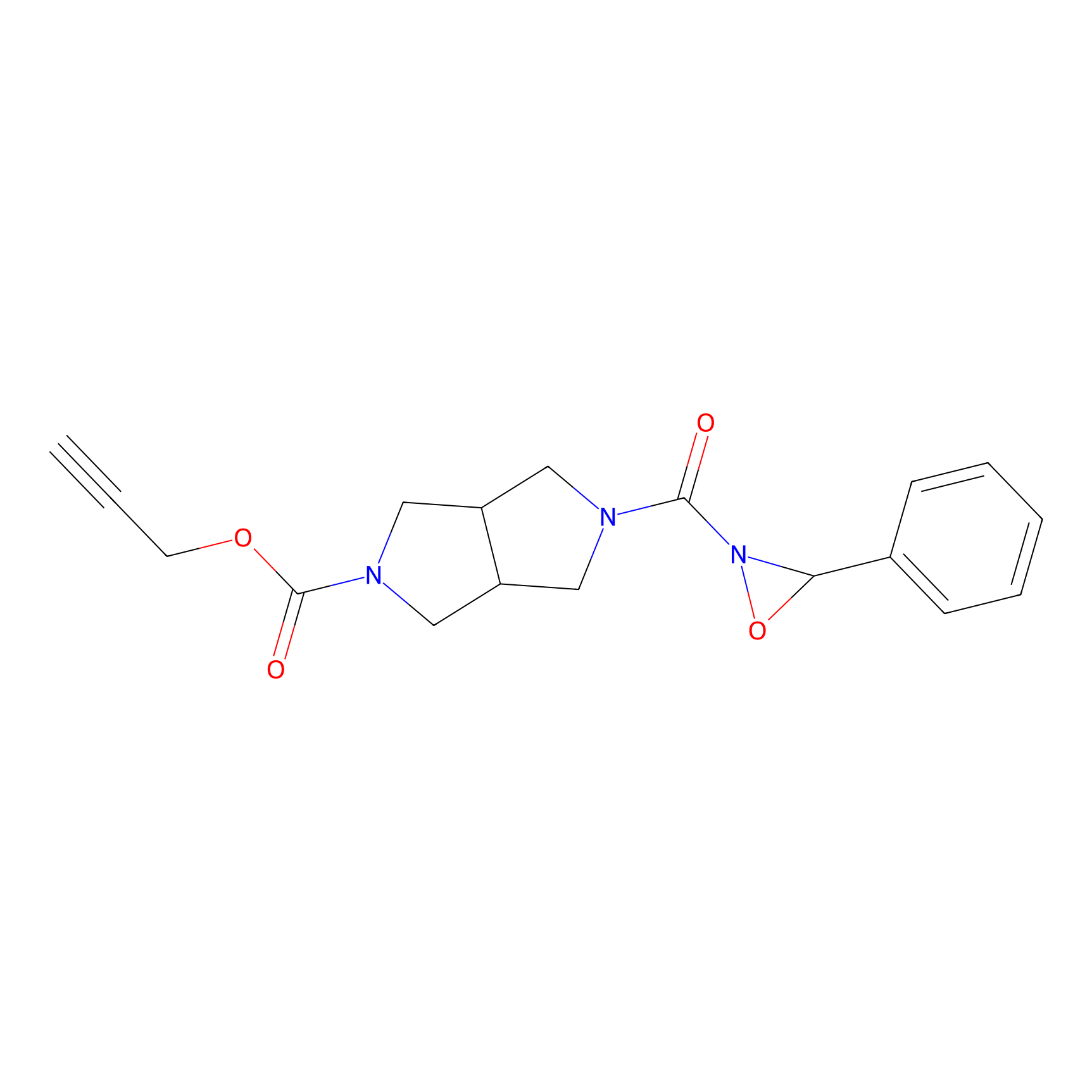 |
N.A. | LDD0193 | [2] | |
|
ONAyne Probe Info |
 |
K91(0.38) | LDD0274 | [3] | |
|
STPyne Probe Info |
 |
K105(6.41); K223(6.59); K91(3.39); K96(5.88) | LDD0277 | [3] | |
|
m-APA Probe Info |
 |
14.33 | LDD0403 | [4] | |
|
Jackson_14 Probe Info |
 |
2.63 | LDD0123 | [5] | |
|
HPAP Probe Info |
 |
3.17 | LDD0063 | [6] | |
|
TER-AC Probe Info |
 |
N.A. | LDD0426 | [7] | |
|
TPP-AC Probe Info |
 |
N.A. | LDD0427 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C040 Probe Info |
 |
8.88 | LDD1740 | [8] | |
|
C056 Probe Info |
 |
17.27 | LDD1753 | [8] | |
|
C094 Probe Info |
 |
34.30 | LDD1785 | [8] | |
|
C235 Probe Info |
 |
20.68 | LDD1908 | [8] | |
|
C289 Probe Info |
 |
29.86 | LDD1959 | [8] | |
|
C349 Probe Info |
 |
8.57 | LDD2010 | [8] | |
|
C350 Probe Info |
 |
22.63 | LDD2011 | [8] | |
|
C362 Probe Info |
 |
27.47 | LDD2023 | [8] | |
|
C388 Probe Info |
 |
42.52 | LDD2047 | [8] | |
|
C390 Probe Info |
 |
20.82 | LDD2049 | [8] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [9] | |
|
FFF probe14 Probe Info |
 |
12.75 | LDD0477 | [9] | |
|
FFF probe2 Probe Info |
 |
20.00 | LDD0463 | [9] | |
|
STS-1 Probe Info |
 |
N.A. | LDD0136 | [10] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [10] | |
|
A-DA Probe Info |
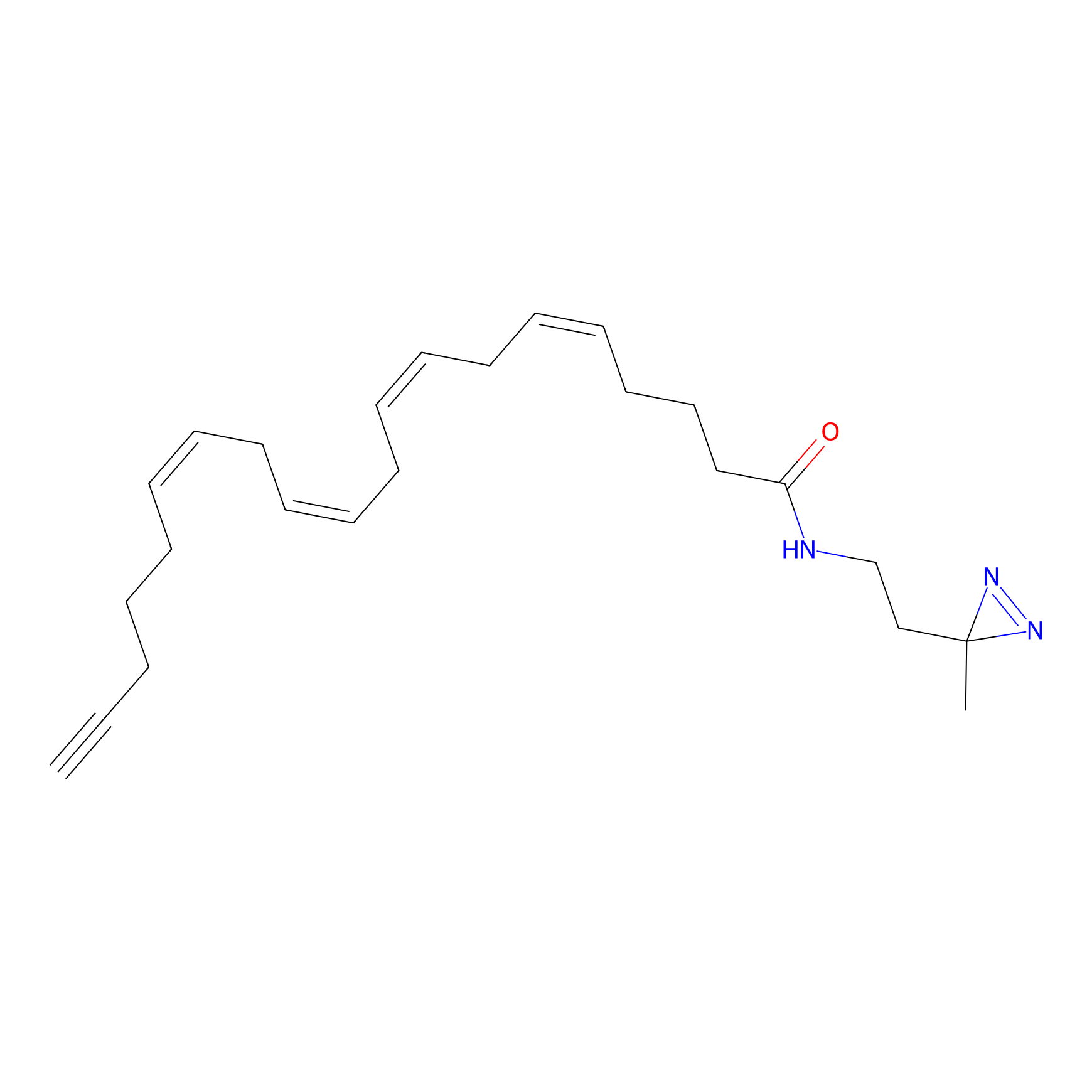 |
2.38 | LDD0145 | [11] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0072 | [12] | |
|
OEA-DA Probe Info |
 |
7.81 | LDD0046 | [13] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References
